

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
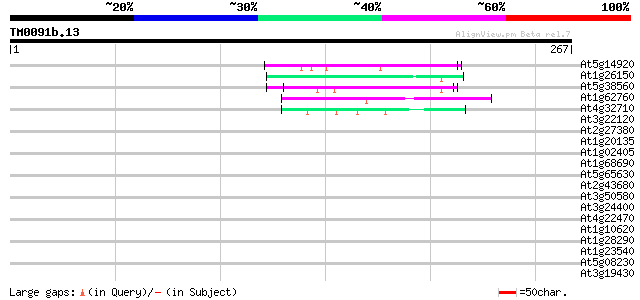
Query= TM0091b.13
(267 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g14920 unknown protein 46 2e-05
At1g26150 Pto kinase interactor, putative 45 4e-05
At5g38560 putative protein 44 7e-05
At1g62760 hypothetical protein 42 3e-04
At4g32710 putative protein kinase 41 6e-04
At3g22120 unknown protein 40 0.001
At2g27380 hypothetical protein 40 0.001
At1g20135 anter-specific proline-rich protein APG precursor 40 0.001
At1g02405 putative protein 40 0.001
At1g68690 protein kinase, putative 40 0.002
At5g65630 unknown protein 39 0.002
At2g43680 SF16 protein {Helianthus annuus} like protein 39 0.002
At3g50580 proline-rich protein 39 0.003
At3g24400 protein kinase, putative 39 0.003
At4g22470 extensin - like protein 39 0.004
At1g10620 putative serine/threonine protein kinase emb|CAA18823.1 39 0.004
At1g28290 proline-rich protein, putative 38 0.005
At1g23540 putative serine/threonine protein kinase 38 0.005
At5g08230 putative protein 38 0.006
At3g19430 putative late embryogenesis abundant protein 38 0.006
>At5g14920 unknown protein
Length = 275
Score = 45.8 bits (107), Expect = 2e-05
Identities = 30/102 (29%), Positives = 42/102 (40%), Gaps = 11/102 (10%)
Query: 122 TPITMIRYNPPFKKPS-PSMPAQPKTDSP-------KPPQIKGCQPKNVLPKADPPKSSP 173
TPI PP K P+ P P +P +P +PP K P P PP P
Sbjct: 67 TPIKPPTTKPPVKPPTIPVTPVKPPVSTPPIKLPPVQPPTYKPPTPTVKPPSVQPPTYKP 126
Query: 174 SNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLR 215
PP T+P+K + V +P P+ +P T P++
Sbjct: 127 PTPTVK---PPTTSPVKPPTTPPVQSPPVQPPTYKPPTSPVK 165
Score = 45.4 bits (106), Expect = 3e-05
Identities = 29/95 (30%), Positives = 39/95 (40%), Gaps = 3/95 (3%)
Query: 122 TPITMIRYNPPFKKPSPSMPA-QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSN--AAE 178
TP+ PP K P P +P T + KPP ++ K P PP +SP
Sbjct: 86 TPVKPPVSTPPIKLPPVQPPTYKPPTPTVKPPSVQPPTYKPPTPTVKPPTTSPVKPPTTP 145
Query: 179 AIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+++PP P K + V P P K P T P
Sbjct: 146 PVQSPPVQPPTYKPPTSPVKPPTTTPPVKPPTTTP 180
Score = 40.0 bits (92), Expect = 0.001
Identities = 32/108 (29%), Positives = 47/108 (42%), Gaps = 5/108 (4%)
Query: 106 VFVEHDDEELPFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPK 165
VF ++E L TP T+ +P K PSP++ +P T S KPP + K P
Sbjct: 19 VFAASNEESNALVSLPTP-TLPSPSPATKPPSPAL--KPPTPSYKPPTLPTTPIKP--PT 73
Query: 166 ADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP P+ ++ P +T P+K Q + K P+ QP
Sbjct: 74 TKPPVKPPTIPVTPVKPPVSTPPIKLPPVQPPTYKPPTPTVKPPSVQP 121
Score = 30.8 bits (68), Expect = 0.79
Identities = 15/48 (31%), Positives = 21/48 (43%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPP 169
+P+ PP K P+ + P QP T +P +K V P PP
Sbjct: 162 SPVKPPTTTPPVKPPTTTPPVQPPTYNPPTTPVKPPTAPPVKPPTPPP 209
>At1g26150 Pto kinase interactor, putative
Length = 760
Score = 45.1 bits (105), Expect = 4e-05
Identities = 33/96 (34%), Positives = 38/96 (39%), Gaps = 3/96 (3%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P T I P P P + P +P PP PK V P PP+ PS A I
Sbjct: 134 PTTPITSPSPPTNPPPPPESPPSLPAPDPPSNPLPPPKLVPPSHSPPRHLPSPPASEIPP 193
Query: 183 PPNTAPMKKALSQHVSAPKKVT--PSKRPATQPLRR 216
PP P A S+ S P + PS P P RR
Sbjct: 194 PPRHLPSPPA-SERPSTPPSDSEHPSPPPPGHPKRR 228
Score = 39.3 bits (90), Expect = 0.002
Identities = 28/93 (30%), Positives = 42/93 (45%), Gaps = 7/93 (7%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
T T + +PP + P S P +P SP P + G P +P + PP+ SP
Sbjct: 53 TTNTPAQSSPPPETPLSSPPPEP---SPPSPSLTG-PPPTTIPVSPPPEPSPPPPLPT-E 107
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQPL 214
APP P+ + S+P P++ P T P+
Sbjct: 108 APPPANPVSSPPPE--SSPPPPPPTEAPPTTPI 138
Score = 36.2 bits (82), Expect = 0.019
Identities = 27/97 (27%), Positives = 34/97 (34%), Gaps = 13/97 (13%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P T I +PP P PS P T++P P P P PP +P +
Sbjct: 86 PPTTIPVSPP---PEPSPPPPLPTEAPPPANPVSSPPPESSPPPPPPTEAPPTTPITSPS 142
Query: 183 PPN----------TAPMKKALSQHVSAPKKVTPSKRP 209
PP + P S + PK V PS P
Sbjct: 143 PPTNPPPPPESPPSLPAPDPPSNPLPPPKLVPPSHSP 179
Score = 35.8 bits (81), Expect = 0.024
Identities = 25/92 (27%), Positives = 38/92 (41%), Gaps = 2/92 (2%)
Query: 123 PITMIRYNPPFKKP-SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
P T + PP P SPS+ P T P P + P LP PP ++P ++
Sbjct: 64 PETPLSSPPPEPSPPSPSLTGPPPTTIPVSPPPEP-SPPPPLPTEAPPPANPVSSPPPES 122
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP P + + +++P T P P
Sbjct: 123 SPPPPPPTEAPPTTPITSPSPPTNPPPPPESP 154
Score = 34.3 bits (77), Expect = 0.071
Identities = 23/80 (28%), Positives = 32/80 (39%), Gaps = 4/80 (5%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P K+P+PS P+ + P P P+ LP PPK SP +PP P
Sbjct: 234 PGSKRPTPSPPSPSDSKRPVHPS-PPSPPEETLP---PPKPSPDPLPSNSSSPPTLLPPS 289
Query: 191 KALSQHVSAPKKVTPSKRPA 210
+S K V+ P+
Sbjct: 290 SVVSPPSPPRKSVSGPDNPS 309
Score = 33.5 bits (75), Expect = 0.12
Identities = 23/110 (20%), Positives = 41/110 (36%), Gaps = 18/110 (16%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSS---------- 172
P+ + PP P +P+ P ++ P PP+ P + P P S
Sbjct: 166 PLPPPKLVPPSHSPPRHLPSPPASEIPPPPRHLPSPPASERPSTPPSDSEHPSPPPPGHP 225
Query: 173 -------PSNAAEAIRAPPNTAPMKKALSQHV-SAPKKVTPSKRPATQPL 214
P + +PP+ + K+ + S P++ P +P+ PL
Sbjct: 226 KRREQPPPPGSKRPTPSPPSPSDSKRPVHPSPPSPPEETLPPPKPSPDPL 275
>At5g38560 putative protein
Length = 681
Score = 44.3 bits (103), Expect = 7e-05
Identities = 30/90 (33%), Positives = 37/90 (40%), Gaps = 2/90 (2%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P ++ +PP P+ + PA P+T SP PP P PPK SPS E +
Sbjct: 89 PPPVVIASPPPSTPATTPPAPPQTVSPPPPPDASPSPPAPTTTNPPPKPSPSPPGET-PS 147
Query: 183 PPNTAPMKKALSQHVSAPKKVT-PSKRPAT 211
PP P S P T P PAT
Sbjct: 148 PPGETPSPPKPSPSTPTPTTTTSPPPPPAT 177
Score = 40.8 bits (94), Expect = 8e-04
Identities = 25/89 (28%), Positives = 37/89 (41%), Gaps = 6/89 (6%)
Query: 131 PPFKKPSPSMPAQPK--TDSPKPPQ----IKGCQPKNVLPKADPPKSSPSNAAEAIRAPP 184
PP + P+ P+ P T P PPQ + P + + PP SSP + I +PP
Sbjct: 22 PPPLQTQPTTPSAPPPVTPPPSPPQSPPPVVSSSPPPPVVSSPPPSSSPPPSPPVITSPP 81
Query: 185 NTAPMKKALSQHVSAPKKVTPSKRPATQP 213
T +++P TP+ P P
Sbjct: 82 PTVASSPPPPVVIASPPPSTPATTPPAPP 110
Score = 35.4 bits (80), Expect = 0.032
Identities = 29/105 (27%), Positives = 37/105 (34%), Gaps = 17/105 (16%)
Query: 123 PITMIRYNPPFKKPSPSMPA---QPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEA 179
P T+ PP PSP P P SP PP G P PPK SPS
Sbjct: 110 PQTVSPPPPPDASPSPPAPTTTNPPPKPSPSPP---GETPSPPGETPSPPKPSPSTPTPT 166
Query: 180 I-----------RAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP++ P + P V P ++P +P
Sbjct: 167 TTTSPPPPPATSASPPSSNPTDPSTLAPPPTPLPVVPREKPIAKP 211
Score = 33.5 bits (75), Expect = 0.12
Identities = 25/93 (26%), Positives = 37/93 (38%), Gaps = 14/93 (15%)
Query: 131 PPFKKPSPSMPAQPK---TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEA-------I 180
PP P PS P P + SP PP + P + P + P +SP + I
Sbjct: 35 PPPVTPPPSPPQSPPPVVSSSPPPPVVSSPPPSSSPPPSPPVITSPPPTVASSPPPPVVI 94
Query: 181 RAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP + P + + P+ V+P P P
Sbjct: 95 ASPPPSTP----ATTPPAPPQTVSPPPPPDASP 123
>At1g62760 hypothetical protein
Length = 312
Score = 42.4 bits (98), Expect = 3e-04
Identities = 31/112 (27%), Positives = 46/112 (40%), Gaps = 16/112 (14%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADP------------PKSSPSNAA 177
+PP PS + P+ SP P + P P + P P SSPS+A
Sbjct: 39 SPPSSSPSSAPPSSLSPSSPPPLSLSPSSPPPPPPSSSPLSSLSPSLSPSPPSSSPSSAP 98
Query: 178 EAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKNLE 229
+ +P + P LS S+P PS P + S+ +SN+ NL+
Sbjct: 99 PSSLSPSSPPP----LSLSPSSPPPPPPSSSPLSSLSPSSSSSTYSNQTNLD 146
Score = 32.0 bits (71), Expect = 0.35
Identities = 21/74 (28%), Positives = 33/74 (44%), Gaps = 9/74 (12%)
Query: 137 SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKS-SPSNAAEAIRAPPNTAPMKKALSQ 195
SPS+ P + SP P ++ P + PP S SPS+ PP+++P+
Sbjct: 33 SPSLSPSPPSSSPS-----SAPPSSLSPSSPPPLSLSPSSPPP---PPPSSSPLSSLSPS 84
Query: 196 HVSAPKKVTPSKRP 209
+P +PS P
Sbjct: 85 LSPSPPSSSPSSAP 98
>At4g32710 putative protein kinase
Length = 731
Score = 41.2 bits (95), Expect = 6e-04
Identities = 33/101 (32%), Positives = 41/101 (39%), Gaps = 20/101 (19%)
Query: 130 NPPFKKPSPSM---PAQPKTDSPKPPQI-----KGCQPKNVLP-KADPPKSSPSNAA--- 177
+PP + P P + P P +SP PP G P LP K PP SSP +
Sbjct: 73 SPPIESPPPPLLESPPPPPLESPSPPSPHVSAPSGSPPLPFLPAKPSPPPSSPPSETVPP 132
Query: 178 -EAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRS 217
I PP + P + S P T S P + P RRS
Sbjct: 133 GNTISPPPRSLPSE-------STPPVNTASPPPPSPPRRRS 166
Score = 35.8 bits (81), Expect = 0.024
Identities = 24/77 (31%), Positives = 32/77 (41%), Gaps = 2/77 (2%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP P+ S P P P PP I+ P + PP SPS + + AP + P+
Sbjct: 54 PPLSAPTASPPPLPVESPPSPP-IESPPPPLLESPPPPPLESPSPPSPHVSAPSGSPPL- 111
Query: 191 KALSQHVSAPKKVTPSK 207
L S P PS+
Sbjct: 112 PFLPAKPSPPPSSPPSE 128
Score = 33.9 bits (76), Expect = 0.093
Identities = 32/111 (28%), Positives = 42/111 (37%), Gaps = 27/111 (24%)
Query: 127 IRYNPPFKKPSPSMPAQPKTD-SPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAP-- 183
I +PP PSP+ P+ P+T PKPP P + P PPK SP+ P
Sbjct: 176 INSSPP--NPSPNTPSLPETSPPPKPPLSTTPFPSSSTP---PPKKSPAAVTLPFFGPAG 230
Query: 184 ----------------PNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSN 218
P T+P A S P+ + P P T RS+
Sbjct: 231 QLPDGTVAPPIGPVIEPKTSP---AESISPGTPQPLVPKSLPVTTSYHRSS 278
Score = 32.3 bits (72), Expect = 0.27
Identities = 27/88 (30%), Positives = 34/88 (37%), Gaps = 10/88 (11%)
Query: 131 PPFKKPSPSMPAQPKTDSPKP-PQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP PS S P P P P + PK P P SSP N + + P T+P
Sbjct: 139 PPRSLPSESTPPVNTASPPPPSPPRRRSGPKPSFPP--PINSSPPNPSPNTPSLPETSPP 196
Query: 190 KKALSQHVSAPKKVTPSKRPATQPLRRS 217
K P TP +T P ++S
Sbjct: 197 PKP-------PLSTTPFPSSSTPPPKKS 217
Score = 31.2 bits (69), Expect = 0.60
Identities = 23/78 (29%), Positives = 30/78 (37%), Gaps = 5/78 (6%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNV---LPKADPPKSSPSNAAEAIRAPPNTA 187
PP P + + P SP P P + P + P +S P E+ PP +
Sbjct: 35 PPLSPLPPPLSSPPPLPSPPPLSAPTASPPPLPVESPPSPPIESPPPPLLESPPPPPLES 94
Query: 188 PMKKALSQHVSAPKKVTP 205
P S HVSAP P
Sbjct: 95 PSPP--SPHVSAPSGSPP 110
>At3g22120 unknown protein
Length = 351
Score = 40.4 bits (93), Expect = 0.001
Identities = 25/85 (29%), Positives = 33/85 (38%), Gaps = 7/85 (8%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP P+P P + PKPP +K +P P PP P + PP P
Sbjct: 46 PPKPSPAPHKPPKHPVKPPKPPAVKPPKP----PAVKPPTPKPPTVKPHPK-PPTVKPHP 100
Query: 191 K--ALSQHVSAPKKVTPSKRPATQP 213
K + H P P +P T+P
Sbjct: 101 KPPTVKPHPKPPTVKPPHPKPPTKP 125
Score = 35.4 bits (80), Expect = 0.032
Identities = 28/85 (32%), Positives = 32/85 (36%), Gaps = 9/85 (10%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKN--VLPKADPPKSSPSNAAEAIRAPPNTAP 188
PP KP P P K PKPP PK V P PP S+P + PP + P
Sbjct: 102 PPTVKPHPKPPTV-KPPHPKPPTKPHPHPKPPIVKPPTKPPPSTPKPPTK----PPPSTP 156
Query: 189 MKKALSQHVSAPKKVTPSKRPATQP 213
S PK P +P P
Sbjct: 157 KPPTTKPPPSTPK--PPHHKPPPTP 179
Score = 34.3 bits (77), Expect = 0.071
Identities = 27/86 (31%), Positives = 35/86 (40%), Gaps = 6/86 (6%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGC-QPKNVLPKADPPKSSPSNAAEAIRAPPNTAP- 188
PP KP P K PKPP +K +P V P PP P + + P+ P
Sbjct: 74 PPAVKPPTPKPPTVKPH-PKPPTVKPHPKPPTVKPHPKPPTVKPPHPKPPTKPHPHPKPP 132
Query: 189 -MKKALSQHVSAPKKVTPSKRPATQP 213
+K S PK P+K P + P
Sbjct: 133 IVKPPTKPPPSTPK--PPTKPPPSTP 156
Score = 32.3 bits (72), Expect = 0.27
Identities = 23/86 (26%), Positives = 32/86 (36%), Gaps = 6/86 (6%)
Query: 123 PITMIRYNPPFKKPSPSM---PAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEA 179
PI PP P P P+ PK + KPP P + P P +P+
Sbjct: 132 PIVKPPTKPPPSTPKPPTKPPPSTPKPPTTKPPPSTPKPPHHKPPPTPCPPPTPTPTPPV 191
Query: 180 IRAPPNTAPMKKALSQHVSAPKKVTP 205
+ P T P+ ++ P VTP
Sbjct: 192 VTPPTPTPPV---ITPPTPTPPVVTP 214
Score = 31.2 bits (69), Expect = 0.60
Identities = 23/82 (28%), Positives = 29/82 (35%), Gaps = 15/82 (18%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT---- 186
PP KP P +P +PKPP +P PK K PS PP T
Sbjct: 131 PPIVKP----PTKPPPSTPKPP----TKPPPSTPKPPTTKPPPSTPKPPHHKPPPTPCPP 182
Query: 187 ---APMKKALSQHVSAPKKVTP 205
P ++ P +TP
Sbjct: 183 PTPTPTPPVVTPPTPTPPVITP 204
Score = 28.1 bits (61), Expect = 5.1
Identities = 22/80 (27%), Positives = 30/80 (37%), Gaps = 10/80 (12%)
Query: 131 PPFKKPSPSM--PAQPKTDSPKPPQIKGCQPKNVL---PKADPPKSSPSNAAEAIRAPPN 185
PP KP P+ P P +P PP + P + P PP +P + PP
Sbjct: 170 PPHHKPPPTPCPPPTP---TPTPPVVTPPTPTPPVITPPTPTPPVVTPPTPTPPVITPPT 226
Query: 186 TAPMKKALSQHVSAPKKVTP 205
P ++ P VTP
Sbjct: 227 PTP--PVITPPTPTPPVVTP 244
>At2g27380 hypothetical protein
Length = 761
Score = 40.4 bits (93), Expect = 0.001
Identities = 30/97 (30%), Positives = 38/97 (38%), Gaps = 9/97 (9%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPPKSSPSNA--AE 178
TPI Y+PP K P P P P KPP +K P P PP P +
Sbjct: 446 TPI----YSPPVKPPPVHKPPTPTYSPPIKPPPVKPPTPTYSPPVQPPPVQKPPTPTYSP 501
Query: 179 AIRAPPNTAPMKKALSQHVSAP--KKVTPSKRPATQP 213
++ PP P S + P K TP+ P +P
Sbjct: 502 PVKPPPIQKPPTPTYSPPIKPPPVKPPTPTYSPPIKP 538
Score = 39.3 bits (90), Expect = 0.002
Identities = 28/85 (32%), Positives = 35/85 (40%), Gaps = 5/85 (5%)
Query: 129 YNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPP-NT 186
Y+PP K P P P P KPP +K P P PP P I +PP
Sbjct: 315 YSPPIKPPPVQKPPTPTYSPPIKPPPVKPPTPIYSPPVKPPPVHKPPT---PIYSPPVKP 371
Query: 187 APMKKALSQHVSAPKKVTPSKRPAT 211
P+ K + S P K P ++P T
Sbjct: 372 PPVHKPPTPIYSPPVKPPPIQKPPT 396
Score = 38.5 bits (88), Expect = 0.004
Identities = 27/91 (29%), Positives = 34/91 (36%), Gaps = 6/91 (6%)
Query: 129 YNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPPKSSPSNA--AEAIRAPPN 185
Y+PP K P MP P P KPP + P PP P + I+ PP
Sbjct: 146 YSPPVKPPPVQMPPTPTYSPPIKPPPVHKPPTPTYSPPIKPPVHKPPTPIYSPPIKPPPV 205
Query: 186 TAPMKKALSQHVSAP---KKVTPSKRPATQP 213
P S + P K TP+ P +P
Sbjct: 206 HKPPTPIYSPPIKPPPVHKPPTPTYSPPVKP 236
Score = 37.0 bits (84), Expect = 0.011
Identities = 27/90 (30%), Positives = 34/90 (37%), Gaps = 5/90 (5%)
Query: 129 YNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPPKSSPSNA--AEAIRAPPN 185
Y+PP K P P P P K P +K P P PP P + ++ PP
Sbjct: 399 YSPPIKPPPLQKPPTPTYSPPIKLPPVKPPTPIYSPPVKPPPVHKPPTPIYSPPVKPPPV 458
Query: 186 TAPMKKALSQHVSAP--KKVTPSKRPATQP 213
P S + P K TP+ P QP
Sbjct: 459 HKPPTPTYSPPIKPPPVKPPTPTYSPPVQP 488
Score = 35.4 bits (80), Expect = 0.032
Identities = 31/98 (31%), Positives = 37/98 (37%), Gaps = 10/98 (10%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSP-KPPQI-KGCQPKNVLPKADPPKSSPSNA--A 177
TPI Y+PP K P P P P KPP + K P P PP P +
Sbjct: 261 TPI----YSPPVKPPPVQTPPTPIYSPPVKPPPVHKPPTPTYSPPVKSPPVQKPPTPTYS 316
Query: 178 EAIRAPPNTAPMKKALSQHVSAP--KKVTPSKRPATQP 213
I+ PP P S + P K TP P +P
Sbjct: 317 PPIKPPPVQKPPTPTYSPPIKPPPVKPPTPIYSPPVKP 354
Score = 35.0 bits (79), Expect = 0.042
Identities = 28/99 (28%), Positives = 38/99 (38%), Gaps = 11/99 (11%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPP---KSSPSNAA 177
TPI Y+PP K P P P P KPP ++ P PP K +
Sbjct: 244 TPI----YSPPIKPPPVHKPPTPIYSPPVKPPPVQTPPTPIYSPPVKPPPVHKPPTPTYS 299
Query: 178 EAIRAPPNTAPMKKALSQHVSAP---KKVTPSKRPATQP 213
+++PP P S + P K TP+ P +P
Sbjct: 300 PPVKSPPVQKPPTPTYSPPIKPPPVQKPPTPTYSPPIKP 338
Score = 34.7 bits (78), Expect = 0.054
Identities = 28/92 (30%), Positives = 37/92 (39%), Gaps = 7/92 (7%)
Query: 129 YNPPFKKPSPSMPAQPKTDSP-KPPQI-KGCQPKNVLPKADPPKSSPSNA--AEAIRAPP 184
Y+PP K P P P P KPP + K P P PP +P + ++ PP
Sbjct: 230 YSPPVKPPPVHKPPTPIYSPPIKPPPVHKPPTPIYSPPVKPPPVQTPPTPIYSPPVKPPP 289
Query: 185 NTAPMKKALSQHVSAP---KKVTPSKRPATQP 213
P S V +P K TP+ P +P
Sbjct: 290 VHKPPTPTYSPPVKSPPVQKPPTPTYSPPIKP 321
Score = 33.5 bits (75), Expect = 0.12
Identities = 27/92 (29%), Positives = 34/92 (36%), Gaps = 8/92 (8%)
Query: 129 YNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPP---KSSPSNAAEAIRAPP 184
Y+PP K P P P P P KPP + P PP K + I+ PP
Sbjct: 516 YSPPIK-PPPVKPPTPTYSPPIKPPPVHKPPTPTYSPPIKPPPIHKPPTPTYSPPIKPPP 574
Query: 185 NTAPMKKALSQHVSAP---KKVTPSKRPATQP 213
P S + P K TP+ P +P
Sbjct: 575 VHKPPTPTYSPPIKPPPVHKPPTPTYSPPIKP 606
Score = 33.5 bits (75), Expect = 0.12
Identities = 26/85 (30%), Positives = 33/85 (38%), Gaps = 4/85 (4%)
Query: 129 YNPPFKKPSPSMPAQPKTDSP-KPPQI-KGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT 186
Y+PP K P P P P KPP + K P P PP P + P
Sbjct: 600 YSPPIKPPPVHKPPTPTYSPPIKPPPVHKPPTPTYSPPIKPPPVHKPPTPTYS--PPIKP 657
Query: 187 APMKKALSQHVSAPKKVTPSKRPAT 211
P++K + S P K P + P T
Sbjct: 658 PPVQKPPTPTYSPPVKPPPVQLPPT 682
Score = 33.1 bits (74), Expect = 0.16
Identities = 26/85 (30%), Positives = 35/85 (40%), Gaps = 5/85 (5%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQI-KGCQPKNVLPKADPPKSSPSNAAEAIRAPP-NT 186
Y+PP K P P + KPP + K P P PP P I +PP
Sbjct: 332 YSPPIKPPPVKPPTPIYSPPVKPPPVHKPPTPIYSPPVKPPPVHKPPT---PIYSPPVKP 388
Query: 187 APMKKALSQHVSAPKKVTPSKRPAT 211
P++K + S P K P ++P T
Sbjct: 389 PPIQKPPTPTYSPPIKPPPLQKPPT 413
Score = 32.7 bits (73), Expect = 0.21
Identities = 25/85 (29%), Positives = 31/85 (36%), Gaps = 6/85 (7%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSP--SNAAEAIRAPPNT 186
Y+PP K P P +P T + PP K P PP P +PP
Sbjct: 466 YSPPIKPP----PVKPPTPTYSPPVQPPPVQKPPTPTYSPPVKPPPIQKPPTPTYSPPIK 521
Query: 187 APMKKALSQHVSAPKKVTPSKRPAT 211
P K + S P K P +P T
Sbjct: 522 PPPVKPPTPTYSPPIKPPPVHKPPT 546
Score = 31.2 bits (69), Expect = 0.60
Identities = 26/88 (29%), Positives = 33/88 (36%), Gaps = 13/88 (14%)
Query: 129 YNPPFKKPSPSMPAQPKTDSP-KPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN-- 185
Y+PP K P P P P KPP ++ LP P SP ++ PP
Sbjct: 651 YSPPIKPPPVQKPPTPTYSPPVKPPPVQ-------LPPT--PTYSPPVKPPPVQVPPTPT 701
Query: 186 -TAPMKKALSQHVSAPKKVTPSKRPATQ 212
+ P+K Q P P K P Q
Sbjct: 702 YSPPVKPPPVQVPPTPTYSPPIKPPPVQ 729
Score = 31.2 bits (69), Expect = 0.60
Identities = 24/84 (28%), Positives = 32/84 (37%), Gaps = 4/84 (4%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQI-KGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
Y+PP K P P + KPP + K P P PP P +PP
Sbjct: 416 YSPPIKLPPVKPPTPIYSPPVKPPPVHKPPTPIYSPPVKPPPVHKPPTPT---YSPPIKP 472
Query: 188 PMKKALSQHVSAPKKVTPSKRPAT 211
P K + S P + P ++P T
Sbjct: 473 PPVKPPTPTYSPPVQPPPVQKPPT 496
Score = 28.9 bits (63), Expect = 3.0
Identities = 25/89 (28%), Positives = 33/89 (36%), Gaps = 6/89 (6%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQP----KNVLPKADPP-KSSPSNAAEAIRAP 183
+ PP SP + + P P P +P K P PP K P I +P
Sbjct: 291 HKPPTPTYSPPVKSPPVQKPPTPTYSPPIKPPPVQKPPTPTYSPPIKPPPVKPPTPIYSP 350
Query: 184 P-NTAPMKKALSQHVSAPKKVTPSKRPAT 211
P P+ K + S P K P +P T
Sbjct: 351 PVKPPPVHKPPTPIYSPPVKPPPVHKPPT 379
Score = 28.9 bits (63), Expect = 3.0
Identities = 26/94 (27%), Positives = 34/94 (35%), Gaps = 12/94 (12%)
Query: 129 YNPPFKKP---SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN 185
Y PP +KP SP + P P P P + K P SP I+ PP
Sbjct: 68 YPPPIQKPPTYSPPIYPPPIQKPPTPTYSPPIYPPPI-QKPPTPTYSPPIYPPPIQKPPT 126
Query: 186 TA--------PMKKALSQHVSAPKKVTPSKRPAT 211
P++K + S P K P + P T
Sbjct: 127 PTYSPPIYPPPIQKPPTPSYSPPVKPPPVQMPPT 160
Score = 28.1 bits (61), Expect = 5.1
Identities = 27/104 (25%), Positives = 34/104 (31%), Gaps = 13/104 (12%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSP--------KPPQIKGCQPKNVLPKADPPKSSPS 174
PI PP SP + P P KPP I+ P PP P
Sbjct: 469 PIKPPPVKPPTPTYSPPVQPPPVQKPPTPTYSPPVKPPPIQKPPTPTYSPPIKPPPVKPP 528
Query: 175 NA--AEAIRAPPNTAPMKKALSQHVSAP---KKVTPSKRPATQP 213
+ I+ PP P S + P K TP+ P +P
Sbjct: 529 TPTYSPPIKPPPVHKPPTPTYSPPIKPPPIHKPPTPTYSPPIKP 572
Score = 27.7 bits (60), Expect = 6.6
Identities = 12/25 (48%), Positives = 14/25 (56%), Gaps = 2/25 (8%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQ 153
Y+PP K P +P P T P PPQ
Sbjct: 719 YSPPIKPPPVQVPPTPTT--PSPPQ 741
Score = 27.3 bits (59), Expect = 8.7
Identities = 23/88 (26%), Positives = 30/88 (33%), Gaps = 9/88 (10%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
+ PP SP + P P P +P V P S P I+ PP P
Sbjct: 593 HKPPTPTYSPPIKPPPVHKPPTPTYSPPIKPPPVHKPPTPTYSPP------IKPPPVHKP 646
Query: 189 MKKALSQHVSAP---KKVTPSKRPATQP 213
S + P K TP+ P +P
Sbjct: 647 PTPTYSPPIKPPPVQKPPTPTYSPPVKP 674
>At1g20135 anter-specific proline-rich protein APG precursor
Length = 534
Score = 40.4 bits (93), Expect = 0.001
Identities = 31/103 (30%), Positives = 41/103 (39%), Gaps = 16/103 (15%)
Query: 132 PFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADP-----PKSSPSNAAEAI---RAP 183
P P PS P +P+ PKP C P P+ P PK +P A + + P
Sbjct: 103 PSPSPCPSPPPKPQ---PKPVPPPACPPTPPKPQPKPAPPPEPKPAPPPAPKPVPCPSPP 159
Query: 184 PNTAPMKKALSQHVSAPKKV-----TPSKRPATQPLRRSNRLI 221
AP K + H PK PS +PA P + N+ I
Sbjct: 160 KPPAPTPKPVPPHGPPPKPAPAPTPAPSPKPAPSPPKPENKTI 202
Score = 33.1 bits (74), Expect = 0.16
Identities = 25/86 (29%), Positives = 31/86 (35%), Gaps = 6/86 (6%)
Query: 130 NPPFKKPSPS--MPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
NPP PSP P P + PP G P P PK P+ + +PP
Sbjct: 59 NPPTPDPSPKPVAPPGPSSKPVAPP---GPSPCPSPPPKPQPKPPPAPSPSPCPSPP-PK 114
Query: 188 PMKKALSQHVSAPKKVTPSKRPATQP 213
P K + P P +PA P
Sbjct: 115 PQPKPVPPPACPPTPPKPQPKPAPPP 140
Score = 31.6 bits (70), Expect = 0.46
Identities = 24/86 (27%), Positives = 30/86 (33%), Gaps = 8/86 (9%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLP-KADPPKSSPSNAAEAIRAPPNTA 187
Y P+ P+ PK +P P K P P + PPK P + PP +
Sbjct: 52 YPQPWPMNPPTPDPSPKPVAPPGPSSKPVAPPGPSPCPSPPPKPQP-------KPPPAPS 104
Query: 188 PMKKALSQHVSAPKKVTPSKRPATQP 213
P PK V P P T P
Sbjct: 105 PSPCPSPPPKPQPKPVPPPACPPTPP 130
Score = 30.0 bits (66), Expect = 1.3
Identities = 23/76 (30%), Positives = 28/76 (36%), Gaps = 18/76 (23%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP KP P P+ PK +P PK V P PPK +P+ P AP
Sbjct: 147 PPAPKPVPC-PSPPKPPAP--------TPKPVPPHGPPPKPAPA---------PTPAPSP 188
Query: 191 KALSQHVSAPKKVTPS 206
K K P+
Sbjct: 189 KPAPSPPKPENKTIPA 204
>At1g02405 putative protein
Length = 134
Score = 40.4 bits (93), Expect = 0.001
Identities = 19/64 (29%), Positives = 25/64 (38%)
Query: 106 VFVEHDDEELPFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPK 165
+ + +D V C P + PP P PS P SP PP+ C P + P
Sbjct: 26 IIINAEDSSAEVDVNCIPCLQNQPPPPPSPPPPSCTPSPPPPSPPPPKKSSCPPSPLPPP 85
Query: 166 ADPP 169
PP
Sbjct: 86 PPPP 89
>At1g68690 protein kinase, putative
Length = 708
Score = 39.7 bits (91), Expect = 0.002
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 6/82 (7%)
Query: 130 NPPFKKP--SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
+PP ++P SP P+ P S +P Q P++ P+ PP+ SP++ +PP +
Sbjct: 186 SPPSERPTQSPPPPSPPSPPSDRPSQSPPPPPEDTKPQ--PPRRSPNSPPPTFSSPPRSP 243
Query: 188 PMKKALSQHVSAPKKVTPSKRP 209
P + L + P + P+ RP
Sbjct: 244 P--EILVPGSNNPSQNNPTLRP 263
Score = 37.4 bits (85), Expect = 0.008
Identities = 26/89 (29%), Positives = 38/89 (42%), Gaps = 6/89 (6%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKAD--PPKSSPSNAAEAIRAPPNTA 187
+PP P P +P+ P + SP PP + P + P A PP+ SPS PP+
Sbjct: 95 SPP---PQPVIPSPPPSASP-PPALVPPLPSSPPPPASVPPPRPSPSPPILVRSPPPSVR 150
Query: 188 PMKKALSQHVSAPKKVTPSKRPATQPLRR 216
P++ P + P P + P R
Sbjct: 151 PIQSPPPPPSDRPTQSPPPPSPPSPPSER 179
Score = 37.4 bits (85), Expect = 0.008
Identities = 26/90 (28%), Positives = 33/90 (35%), Gaps = 12/90 (13%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA-- 187
+PP P P P T PP + P P A PP +P +PP A
Sbjct: 13 SPPVTSPPP--PLNNATSPATPPPVTSPLP----PSAPPPNRAPPPPPPVTTSPPPVANG 66
Query: 188 ----PMKKALSQHVSAPKKVTPSKRPATQP 213
P+ K P+ V PS P+T P
Sbjct: 67 APPPPLPKPPESSSPPPQPVIPSPPPSTSP 96
Score = 37.0 bits (84), Expect = 0.011
Identities = 24/84 (28%), Positives = 33/84 (38%), Gaps = 5/84 (5%)
Query: 131 PPF-KKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
PP P P P SP PP + P +V P PP P + ++PP +P
Sbjct: 117 PPLPSSPPPPASVPPPRPSPSPPILVRSPPPSVRPIQSPP---PPPSDRPTQSPPPPSPP 173
Query: 190 KKALSQHVSAPKKVTPSKRPATQP 213
+ +P PS+RP P
Sbjct: 174 SPPSERPTQSPPS-PPSERPTQSP 196
Score = 37.0 bits (84), Expect = 0.011
Identities = 24/85 (28%), Positives = 34/85 (39%), Gaps = 3/85 (3%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P P P P PKPP+ P+ V+P + PP +SP PP+ +P
Sbjct: 55 PVTTSPPPVANGAPPPPLPKPPESSSPPPQPVIP-SPPPSTSPPPQPVIPSPPPSASPPP 113
Query: 191 KALSQHVSA--PKKVTPSKRPATQP 213
+ S+ P P RP+ P
Sbjct: 114 ALVPPLPSSPPPPASVPPPRPSPSP 138
Score = 36.6 bits (83), Expect = 0.014
Identities = 29/96 (30%), Positives = 37/96 (38%), Gaps = 13/96 (13%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSP----------SNAAEAI 180
PP KP S P+ P PP P+ V+P + PP +SP S A
Sbjct: 70 PPLPKPPESSSPPPQPVIPSPPPSTSPPPQPVIP-SPPPSASPPPALVPPLPSSPPPPAS 128
Query: 181 RAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRR 216
PP +P L + S P V P + P P R
Sbjct: 129 VPPPRPSPSPPILVR--SPPPSVRPIQSPPPPPSDR 162
Score = 35.4 bits (80), Expect = 0.032
Identities = 22/83 (26%), Positives = 33/83 (39%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP PSP + + S +P Q P + ++ PP S PS +E P + P +
Sbjct: 131 PPRPSPSPPILVRSPPPSVRPIQSPPPPPSDRPTQSPPPPSPPSPPSERPTQSPPSPPSE 190
Query: 191 KALSQHVSAPKKVTPSKRPATQP 213
+ PS RP+ P
Sbjct: 191 RPTQSPPPPSPPSPPSDRPSQSP 213
Score = 35.0 bits (79), Expect = 0.042
Identities = 25/95 (26%), Positives = 35/95 (36%), Gaps = 5/95 (5%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPK---TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAE 178
+P ++R PP +P S P P T SP PP P + P PP +
Sbjct: 137 SPPILVRSPPPSVRPIQSPPPPPSDRPTQSPPPPSPPS--PPSERPTQSPPSPPSERPTQ 194
Query: 179 AIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+ P +P SQ P + T + P P
Sbjct: 195 SPPPPSPPSPPSDRPSQSPPPPPEDTKPQPPRRSP 229
Score = 33.5 bits (75), Expect = 0.12
Identities = 26/92 (28%), Positives = 33/92 (35%), Gaps = 11/92 (11%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR 181
TP + PP P P P + PP + P LPK SSP
Sbjct: 31 TPPPVTSPLPPSAPPPNRAPPPPPPVTTSPPPVANGAPPPPLPKPPE-SSSPPPQPVIPS 89
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP+T+P P+ V PS P+ P
Sbjct: 90 PPPSTSP----------PPQPVIPSPPPSASP 111
Score = 32.7 bits (73), Expect = 0.21
Identities = 24/86 (27%), Positives = 37/86 (42%), Gaps = 7/86 (8%)
Query: 131 PPFKKP--SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAE-AIRAPP--- 184
PP +P SP P+ P S +P Q P ++ PP S PS ++ ++PP
Sbjct: 158 PPSDRPTQSPPPPSPPSPPSERPTQSPPSPPSERPTQSPPPPSPPSPPSDRPSQSPPPPP 217
Query: 185 -NTAPMKKALSQHVSAPKKVTPSKRP 209
+T P S + P +P + P
Sbjct: 218 EDTKPQPPRRSPNSPPPTFSSPPRSP 243
>At5g65630 unknown protein
Length = 590
Score = 39.3 bits (90), Expect = 0.002
Identities = 23/82 (28%), Positives = 35/82 (42%), Gaps = 8/82 (9%)
Query: 134 KKPSPSMPAQ----PKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
K P P++P Q + SP PP P + P+ P+ P + APP+ + +
Sbjct: 328 KPPQPTLPPQLVEPSRVQSPSPPP----PPPVIQPELPQPQPPPPQLEIEVEAPPDVSEV 383
Query: 190 KKALSQHVSAPKKVTPSKRPAT 211
K + PK P+KR T
Sbjct: 384 SKGRKGKLPKPKAKDPNKRLMT 405
>At2g43680 SF16 protein {Helianthus annuus} like protein
Length = 668
Score = 39.3 bits (90), Expect = 0.002
Identities = 27/103 (26%), Positives = 46/103 (44%), Gaps = 4/103 (3%)
Query: 112 DEELPFGVLCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKS 171
+ E ++ P T R NP P P PA P+ SP+P + P+ P+A+ P++
Sbjct: 67 EAERDHNLVFRPPTPDRPNPYSASPPPR-PASPRVASPRPTSPRVASPRVPSPRAEVPRT 125
Query: 172 -SPSNAAEAIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
SP + P + +P K S P+ ++P ++P
Sbjct: 126 LSPKPPSPRAEVPRSLSP--KPPSPRADLPRSLSPKPFDRSKP 166
Score = 38.5 bits (88), Expect = 0.004
Identities = 22/75 (29%), Positives = 36/75 (47%), Gaps = 4/75 (5%)
Query: 134 KKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPK---ADPPKSSPSNAAEAIRAPPNTAPMK 190
K PSP P++ SPKPP + P+++ PK P S+ +NA +R P +
Sbjct: 129 KPPSPRAEV-PRSLSPKPPSPRADLPRSLSPKPFDRSKPSSASANAPPTLRPASTRVPSQ 187
Query: 191 KALSQHVSAPKKVTP 205
+ V +P+ +P
Sbjct: 188 RITPHSVPSPRPSSP 202
Score = 32.3 bits (72), Expect = 0.27
Identities = 21/97 (21%), Positives = 42/97 (42%), Gaps = 4/97 (4%)
Query: 127 IRYNPP---FKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPK-SSPSNAAEAIRA 182
+R +PP +P+ P P D+P+ + PK P++DPP+ +P +
Sbjct: 237 LRADPPRLDAARPTTPRPPSPLADAPRLDAPRPTTPKPPSPRSDPPRLDAPRPTTPKPPS 296
Query: 183 PPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSNR 219
P + +P + V P+ P + + ++ + R
Sbjct: 297 PRSVSPRAVQRREIVYRPEPTLPVQHASATKIQGAFR 333
Score = 32.3 bits (72), Expect = 0.27
Identities = 24/92 (26%), Positives = 38/92 (41%), Gaps = 5/92 (5%)
Query: 129 YNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNA----AEAIRAPP 184
++ P +PS A P+ S KPP + P P+ P+++ A +A R
Sbjct: 192 HSVPSPRPSSPRGASPQAISSKPPSPRAEPPTLDTPRPPSPRAASLRADPPRLDAARPTT 251
Query: 185 NTAPMKKALSQHVSAPKKVTPS-KRPATQPLR 215
P A + + AP+ TP P + P R
Sbjct: 252 PRPPSPLADAPRLDAPRPTTPKPPSPRSDPPR 283
Score = 31.2 bits (69), Expect = 0.60
Identities = 26/79 (32%), Positives = 39/79 (48%), Gaps = 13/79 (16%)
Query: 137 SPSMPAQ----PKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKA 192
SP +P+ P+T SPKPP P+ +P++ PK PS A+ P + +P
Sbjct: 112 SPRVPSPRAEVPRTLSPKPPS-----PRAEVPRSLSPK-PPSPRAD---LPRSLSPKPFD 162
Query: 193 LSQHVSAPKKVTPSKRPAT 211
S+ SA P+ RPA+
Sbjct: 163 RSKPSSASANAPPTLRPAS 181
>At3g50580 proline-rich protein
Length = 265
Score = 38.9 bits (89), Expect = 0.003
Identities = 36/122 (29%), Positives = 47/122 (38%), Gaps = 21/122 (17%)
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADP-------------PKSSPSNAAEAIRA 182
PS +P+ P T SP PP K P PK P PK SPS
Sbjct: 77 PSTPIPSTPSTPSPPPPAPKKSPPPPT-PKKSPSPPSLTPFVPHPTPKKSPSPPPTPSLP 135
Query: 183 PPNTAPMKKALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNRKNLEQSQVVAVQKIQAN 242
PP AP K + + P TP K P P S+ SN + +Q+ +++ N
Sbjct: 136 PP--APKKSPSTPSLPPP---TPKKSPPPPPSHHSSSP--SNPPHHQQNPWEHIERCMIN 188
Query: 243 AG 244
G
Sbjct: 189 MG 190
>At3g24400 protein kinase, putative
Length = 694
Score = 38.9 bits (89), Expect = 0.003
Identities = 24/79 (30%), Positives = 30/79 (37%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP PSP P P+P + P LP A PP P+ A+ PP +
Sbjct: 6 PPGGTPSPPPQPLPIPPPPQPLPVTPPPPPTALPPALPPPPPPTALPPALPPPPPPTTVP 65
Query: 191 KALSQHVSAPKKVTPSKRP 209
S P +TPS P
Sbjct: 66 PIPPSTPSPPPPLTPSPLP 84
Score = 37.7 bits (86), Expect = 0.006
Identities = 29/91 (31%), Positives = 41/91 (44%), Gaps = 12/91 (13%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLP------KADPPKSSPSNAAEAIR-A 182
+PP PSP+ P+ P T SP PP I P P PP SPS + + +
Sbjct: 91 SPPLT-PSPTTPSPPLTPSP-PPAITPSPPLTPSPLPPSPTTPSPPPPSPSIPSPPLTPS 148
Query: 183 PPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
PP ++P++ + P TPS P + P
Sbjct: 149 PPPSSPLRPS---SPPPPSPATPSTPPRSPP 176
Score = 37.7 bits (86), Expect = 0.006
Identities = 31/110 (28%), Positives = 49/110 (44%), Gaps = 13/110 (11%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA-PPNTAP 188
+PP PSPS+P+ P T P PP P + L + PP SP+ + R+ PP + P
Sbjct: 132 SPP--PPSPSIPSPPLT--PSPP------PSSPLRPSSPPPPSPATPSTPPRSPPPPSTP 181
Query: 189 MKKALSQHVSAPKKVTPS--KRPATQPLRRSNRLIWSNRKNLEQSQVVAV 236
+S P +PS + P+T + + K L + +V +
Sbjct: 182 TPPPRVGSLSPPPPASPSGGRSPSTPSTTPGSSPPAQSSKELSKGAMVGI 231
Score = 33.9 bits (76), Expect = 0.093
Identities = 22/80 (27%), Positives = 30/80 (37%), Gaps = 1/80 (1%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIR-APPNTAPM 189
PP P+ P P T SP PP P + + P SP+ + + +PP
Sbjct: 56 PPPPPPTTVPPIPPSTPSPPPPLTPSPLPPSPTTPSPPLTPSPTTPSPPLTPSPPPAITP 115
Query: 190 KKALSQHVSAPKKVTPSKRP 209
L+ P TPS P
Sbjct: 116 SPPLTPSPLPPSPTTPSPPP 135
Score = 32.3 bits (72), Expect = 0.27
Identities = 25/95 (26%), Positives = 32/95 (33%), Gaps = 12/95 (12%)
Query: 131 PPFKKPSPSMPAQPKTDSPK-----------PPQIKGCQPKNVLPKADPPKSSPSNAAEA 179
PP +P P P P T P PP + P +P P SP
Sbjct: 21 PPPPQPLPVTPPPPPTALPPALPPPPPPTALPPALPPPPPPTTVPPIPPSTPSPPPPLTP 80
Query: 180 IRAPPNTAPMKKALSQHVSAPKK-VTPSKRPATQP 213
PP+ L+ + P +TPS PA P
Sbjct: 81 SPLPPSPTTPSPPLTPSPTTPSPPLTPSPPPAITP 115
>At4g22470 extensin - like protein
Length = 375
Score = 38.5 bits (88), Expect = 0.004
Identities = 26/86 (30%), Positives = 33/86 (38%), Gaps = 4/86 (4%)
Query: 131 PPFKKPSPSMP--AQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
PP P P P + P+T P PP I P + P PP + + PP P
Sbjct: 85 PPALPPKPLPPPLSPPQTTPPPPPAITPPPPPAITPPLSPPPPAITPPPPLATTPPALPP 144
Query: 189 MKKALSQHVSAPKKVTPSKRPATQPL 214
K L +S P+ P T PL
Sbjct: 145 --KPLPPPLSPPQTTPPPPPAITPPL 168
Score = 35.8 bits (81), Expect = 0.024
Identities = 26/80 (32%), Positives = 30/80 (37%), Gaps = 3/80 (3%)
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA-PPNTAPMKKALS 194
P P P QP P PP + P N P P +SP A A PP P +
Sbjct: 42 PGPP-PPQPDPQPPTPPTFQPAPPANDQPPPPPQSTSPPPVATTPPALPPKPLPPPLSPP 100
Query: 195 QHV-SAPKKVTPSKRPATQP 213
Q P +TP PA P
Sbjct: 101 QTTPPPPPAITPPPPPAITP 120
Score = 32.7 bits (73), Expect = 0.21
Identities = 32/118 (27%), Positives = 42/118 (35%), Gaps = 29/118 (24%)
Query: 119 VLCTP----ITMIRYNPPFKKPSPS-------MPAQPKTDSPKPPQIK-----------G 156
V C P I + PP +P P PA P D P PP
Sbjct: 28 VYCPPPPPCICICNPGPPPPQPDPQPPTPPTFQPAPPANDQPPPPPQSTSPPPVATTPPA 87
Query: 157 CQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQHVS-APKKVTPSKRPATQP 213
PK + P PP+++P P T P A++ +S P +TP AT P
Sbjct: 88 LPPKPLPPPLSPPQTTPP------PPPAITPPPPPAITPPLSPPPPAITPPPPLATTP 139
Score = 32.0 bits (71), Expect = 0.35
Identities = 24/70 (34%), Positives = 28/70 (39%), Gaps = 5/70 (7%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP P P PA SP PP I P P A PPK P + P T P
Sbjct: 107 PPAITPPPP-PAITPPLSPPPPAITPPPPLATTPPALPPKPLP----PPLSPPQTTPPPP 161
Query: 191 KALSQHVSAP 200
A++ +S P
Sbjct: 162 PAITPPLSPP 171
Score = 29.3 bits (64), Expect = 2.3
Identities = 15/50 (30%), Positives = 19/50 (38%)
Query: 120 LCTPITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPP 169
L P I PP P++P +P PPQ P + P PP
Sbjct: 122 LSPPPPAITPPPPLATTPPALPPKPLPPPLSPPQTTPPPPPAITPPLSPP 171
Score = 28.5 bits (62), Expect = 3.9
Identities = 14/37 (37%), Positives = 18/37 (47%), Gaps = 3/37 (8%)
Query: 136 PSPSM---PAQPKTDSPKPPQIKGCQPKNVLPKADPP 169
PSP++ P P+T P PPQ P + P PP
Sbjct: 256 PSPTISPPPLPPQTLKPPPPQTTPPPPPAITPPLSPP 292
Score = 27.3 bits (59), Expect = 8.7
Identities = 21/75 (28%), Positives = 31/75 (41%), Gaps = 6/75 (8%)
Query: 85 LNDVDAM-----HLGAYAEYKKQNVEVFVEHDDEELPFGVLCTPITMIRYNPPFKKPSPS 139
++D+DA+ +GA N +F + +P G C P +PP P
Sbjct: 212 VSDLDAVTCFCKSVGAPRFSLSPNFGIFFKVCGRRIPQGFSC-PGPSPTISPPPLPPQTL 270
Query: 140 MPAQPKTDSPKPPQI 154
P P+T P PP I
Sbjct: 271 KPPPPQTTPPPPPAI 285
>At1g10620 putative serine/threonine protein kinase emb|CAA18823.1
Length = 718
Score = 38.5 bits (88), Expect = 0.004
Identities = 23/78 (29%), Positives = 29/78 (36%), Gaps = 1/78 (1%)
Query: 137 SPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN-TAPMKKALSQ 195
SP P P T + PP P N P PP + PS+ +I PP+ P S
Sbjct: 47 SPPSPPSPDTQTSPPPATAAQPPPNQPPNTTPPPTPPSSPPPSITPPPSPPQPQPPPQST 106
Query: 196 HVSAPKKVTPSKRPATQP 213
V P +P P
Sbjct: 107 PTGDSPVVIPFPKPQLPP 124
Score = 28.1 bits (61), Expect = 5.1
Identities = 16/62 (25%), Positives = 23/62 (36%), Gaps = 1/62 (1%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+PP + P QP +P PP P ++ P PP+ P + P P
Sbjct: 59 SPPPATAAQPPPNQPPNTTP-PPTPPSSPPPSITPPPSPPQPQPPPQSTPTGDSPVVIPF 117
Query: 190 KK 191
K
Sbjct: 118 PK 119
Score = 27.3 bits (59), Expect = 8.7
Identities = 20/68 (29%), Positives = 27/68 (39%), Gaps = 7/68 (10%)
Query: 131 PPFKKPSPSMPAQPK---TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTA 187
PP P P+ P+ P T P PPQ + P P D P P + PP+
Sbjct: 73 PPNTTPPPTPPSSPPPSITPPPSPPQPQ--PPPQSTPTGDSPVVIPFPKPQL--PPPSLF 128
Query: 188 PMKKALSQ 195
P ++Q
Sbjct: 129 PPPSLVNQ 136
>At1g28290 proline-rich protein, putative
Length = 359
Score = 38.1 bits (87), Expect = 0.005
Identities = 30/95 (31%), Positives = 36/95 (37%), Gaps = 7/95 (7%)
Query: 123 PITMIRYNPPFK---KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEA 179
P T PP K KP S PA+P P P K P PP P+ A
Sbjct: 94 PPTKAPVKPPTKPPVKPPVSPPAKPPVKPPVYPPTKAPVKPPTKPPVKPPVYPPTKAPV- 152
Query: 180 IRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQPL 214
PP P+K + AP K P+K P P+
Sbjct: 153 --KPPTKPPVKPPVYPPTKAPVK-PPTKPPVKPPV 184
Score = 36.6 bits (83), Expect = 0.014
Identities = 27/84 (32%), Positives = 33/84 (39%), Gaps = 2/84 (2%)
Query: 132 PFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNA-AEAIRAPPNTAPMK 190
P KP P P T +P P K V P A PP P +A PP P+K
Sbjct: 82 PPAKPPVKPPVYPPTKAPVKPPTKPPVKPPVSPPAKPPVKPPVYPPTKAPVKPPTKPPVK 141
Query: 191 KALSQHVSAPKKVTPSKRPATQPL 214
+ AP K P+K P P+
Sbjct: 142 PPVYPPTKAPVK-PPTKPPVKPPV 164
Score = 35.0 bits (79), Expect = 0.042
Identities = 27/86 (31%), Positives = 33/86 (37%), Gaps = 3/86 (3%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSN-AAEAIRAPPNTAPM 189
PP K P S PA+P P P K P PP S P+ + PP AP+
Sbjct: 74 PPVKAPV-SPPAKPPVKPPVYPPTKAPVKPPTKPPVKPPVSPPAKPPVKPPVYPPTKAPV 132
Query: 190 KKALSQHVSAPKKVTPSKRPATQPLR 215
K V P P+K P P +
Sbjct: 133 KPPTKPPVK-PPVYPPTKAPVKPPTK 157
Score = 33.9 bits (76), Expect = 0.093
Identities = 27/95 (28%), Positives = 36/95 (37%), Gaps = 11/95 (11%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P T PP P+ + P +P T P P + V P PP P +
Sbjct: 134 PPTKPPVKPPVYPPTKA-PVKPPTKPPVKPPVYPPTKAPVKPPTKPPVKPPV-------S 185
Query: 183 PPNTAPMKKALSQHVSAPKK---VTPSKRPATQPL 214
PP P+K + AP K P+K P T P+
Sbjct: 186 PPAKPPVKPPVYPPTKAPVKPPVSPPTKPPVTPPV 220
Score = 33.9 bits (76), Expect = 0.093
Identities = 24/88 (27%), Positives = 30/88 (33%), Gaps = 6/88 (6%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P P P P SP P +K P PP P+ A PP P+K
Sbjct: 53 PHHHHPHPHPHPHPPAKSPVKPPVKAPVSPPAKPPVKPPVYPPTKAPVK---PPTKPPVK 109
Query: 191 KALSQHVSAPKK---VTPSKRPATQPLR 215
+S P K P+K P P +
Sbjct: 110 PPVSPPAKPPVKPPVYPPTKAPVKPPTK 137
Score = 33.1 bits (74), Expect = 0.16
Identities = 26/80 (32%), Positives = 31/80 (38%), Gaps = 2/80 (2%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNA-AEAIR 181
P T PP KP P P T +P P K V P A PP P +A
Sbjct: 146 PPTKAPVKPP-TKPPVKPPVYPPTKAPVKPPTKPPVKPPVSPPAKPPVKPPVYPPTKAPV 204
Query: 182 APPNTAPMKKALSQHVSAPK 201
PP + P K ++ V PK
Sbjct: 205 KPPVSPPTKPPVTPPVYPPK 224
Score = 32.7 bits (73), Expect = 0.21
Identities = 27/93 (29%), Positives = 33/93 (35%), Gaps = 5/93 (5%)
Query: 123 PITMIRYNPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRA 182
P T PP KP P P T +P P K V P P P+
Sbjct: 126 PPTKAPVKPP-TKPPVKPPVYPPTKAPVKPPTKPPVKPPVYPPTKAPVKPPTKPPVK--- 181
Query: 183 PPNTAPMKKALSQHVSAPKKVTPSKRPATQPLR 215
PP + P K + V P K P K P + P +
Sbjct: 182 PPVSPPAKPPVKPPVYPPTKA-PVKPPVSPPTK 213
Score = 32.3 bits (72), Expect = 0.27
Identities = 25/88 (28%), Positives = 32/88 (35%), Gaps = 11/88 (12%)
Query: 130 NPPFK---KPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNT 186
+PP K KP P P P P + V P PP P +PP
Sbjct: 65 HPPAKSPVKPPVKAPVSPPAKPPVKPPVYPPTKAPVKPPTKPPVKPPV-------SPPAK 117
Query: 187 APMKKALSQHVSAPKKVTPSKRPATQPL 214
P+K + AP K P+K P P+
Sbjct: 118 PPVKPPVYPPTKAPVK-PPTKPPVKPPV 144
>At1g23540 putative serine/threonine protein kinase
Length = 731
Score = 38.1 bits (87), Expect = 0.005
Identities = 30/96 (31%), Positives = 38/96 (39%), Gaps = 14/96 (14%)
Query: 130 NPPFKKPSPSM----PAQPKTDSPKPPQIK---GCQPKNVLPKADPPKSSPSNAAEAIRA 182
+PP PSP++ P P SP P P + L +PP PS A + A
Sbjct: 132 SPPPSSPSPNVGPTNPESPPLQSPPAPPASDPTNSPPASPLDPTNPPPIQPSGPATSPPA 191
Query: 183 PPN-------TAPMKKALSQHVSAPKKVTPSKRPAT 211
PN T P K S V +P +PSK T
Sbjct: 192 NPNAPPSPFPTVPPKTPSSGPVVSPSLTSPSKGTPT 227
Score = 37.0 bits (84), Expect = 0.011
Identities = 32/105 (30%), Positives = 42/105 (39%), Gaps = 21/105 (20%)
Query: 131 PPFKKPS--------PSMPA--QPKTDSPKPPQIKGCQPKNVLPKADPPKS----SPSNA 176
PP +PS P +P+ P TDSP PP P + P PP S SP
Sbjct: 50 PPLSEPSTPPPDSQLPPLPSILPPLTDSPPPPS-DSSPPVDSTPSPPPPTSNESPSPPED 108
Query: 177 AEAIRAPPNTA------PMKKALSQHVSAPKKVTPSKRPATQPLR 215
+E APPN + P + S S+P P + PL+
Sbjct: 109 SETPPAPPNESNDNNPPPSQDLQSPPPSSPSPNVGPTNPESPPLQ 153
Score = 33.5 bits (75), Expect = 0.12
Identities = 26/90 (28%), Positives = 31/90 (33%), Gaps = 11/90 (12%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM- 189
PP SPS P +T P + P PP SSPS PN P
Sbjct: 96 PPTSNESPSPPEDSETPPAPPNESNDNNPPPSQDLQSPPPSSPS---------PNVGPTN 146
Query: 190 -KKALSQHVSAPKKVTPSKRPATQPLRRSN 218
+ Q AP P+ P PL +N
Sbjct: 147 PESPPLQSPPAPPASDPTNSPPASPLDPTN 176
Score = 33.1 bits (74), Expect = 0.16
Identities = 22/85 (25%), Positives = 39/85 (45%), Gaps = 10/85 (11%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSN--AAEAIRAPPNTAP 188
P PSP P +SP PP+ + P A P +S+ +N ++ +++PP ++P
Sbjct: 87 PVDSTPSP--PPPTSNESPSPPE------DSETPPAPPNESNDNNPPPSQDLQSPPPSSP 138
Query: 189 MKKALSQHVSAPKKVTPSKRPATQP 213
+ +P +P PA+ P
Sbjct: 139 SPNVGPTNPESPPLQSPPAPPASDP 163
Score = 27.7 bits (60), Expect = 6.6
Identities = 19/55 (34%), Positives = 26/55 (46%), Gaps = 5/55 (9%)
Query: 134 KKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
+ PS S PA P D+ PP+ + LP D S PS A++ PP + P
Sbjct: 6 ESPSSSPPA-PPADTAPPPETP--SENSALPPVD--SSPPSPPADSSSTPPLSEP 55
Score = 27.7 bits (60), Expect = 6.6
Identities = 22/81 (27%), Positives = 29/81 (35%), Gaps = 3/81 (3%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
PP + PS + P SP P + + PP S +I P +P
Sbjct: 21 PPPETPSENSALPPVDSSPPSPPADSSSTPPLSEPSTPPPDSQLPPLPSILPPLTDSPPP 80
Query: 191 KALSQHVSAPKKVTPSKRPAT 211
+ S S P TPS P T
Sbjct: 81 PSDS---SPPVDSTPSPPPPT 98
>At5g08230 putative protein
Length = 1445
Score = 37.7 bits (86), Expect = 0.006
Identities = 34/125 (27%), Positives = 55/125 (43%), Gaps = 11/125 (8%)
Query: 130 NPPFKKPS-PSMPAQPKTDSP---KPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPN 185
+PP S PS P QP + P PPQ+ P + PP ++P A++I PP+
Sbjct: 1125 SPPLPHESPPSPPPQPPSSPPPPSSPPQLAPAPPPS--DHCLPPPTAPLAPAQSIALPPS 1182
Query: 186 TAPMKKALSQHVSAPKKVTPSKRPATQPLRRSNRLIWSNR--KNLEQSQVVAVQKIQANA 243
+ + ++ H S P + P P PL + I R ++ S +A + A
Sbjct: 1183 SI-TRPSMPSHPSLP--LQPGFAPPAYPLLQHEYQISMQRDHSSIATSNQIAPVPVNAAH 1239
Query: 244 GDNAE 248
G +A+
Sbjct: 1240 GRHAD 1244
Score = 32.7 bits (73), Expect = 0.21
Identities = 22/80 (27%), Positives = 33/80 (40%), Gaps = 7/80 (8%)
Query: 131 PPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMK 190
P F SP +P + P P P + P + PP+ +P+ PP TAP+
Sbjct: 1119 PSFPAGSPPLPHESPPSPPPQP------PSSPPPPSSPPQLAPAPPPSDHCLPPPTAPLA 1172
Query: 191 KALSQHVSAPKKVTPSKRPA 210
A S + P +T P+
Sbjct: 1173 PAQSIAL-PPSSITRPSMPS 1191
>At3g19430 putative late embryogenesis abundant protein
Length = 550
Score = 37.7 bits (86), Expect = 0.006
Identities = 26/92 (28%), Positives = 40/92 (43%), Gaps = 10/92 (10%)
Query: 130 NPPFKKPSPSMPA-------QPKTDSPKPPQIKGCQPKNVLPKADPPKS-SPSNAAEAIR 181
+PP P+PS+P+ P T +P P P + +P PP S P ++
Sbjct: 134 SPPPPTPTPSVPSPTPPVSPPPPTPTPSVPSPTPPVPTDPMPSPPPPVSPPPPTPTPSVP 193
Query: 182 APPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
+PP+ P S V +P VTP+ + P
Sbjct: 194 SPPDVTPTPPTPS--VPSPPDVTPTPPTPSVP 223
Score = 36.2 bits (82), Expect = 0.019
Identities = 26/95 (27%), Positives = 37/95 (38%), Gaps = 7/95 (7%)
Query: 122 TPITMIRYNPPFKKPSPSMPAQPKTDSP---KPPQIKGCQPKNVLPKADPPKSSPSNAAE 178
+P + PP PS P P P PP + P PP +P+
Sbjct: 146 SPTPPVSPPPPTPTPSVPSPTPPVPTDPMPSPPPPVSPPPPTPTPSVPSPPDVTPTPPTP 205
Query: 179 AIRAPPNTAPMKKALSQHVSAPKKVTPSKRPATQP 213
++ +PP+ P S V +P VTP+ P T P
Sbjct: 206 SVPSPPDVTPTPPTPS--VPSPPDVTPT--PPTPP 236
Score = 35.8 bits (81), Expect = 0.024
Identities = 27/86 (31%), Positives = 35/86 (40%), Gaps = 15/86 (17%)
Query: 130 NPPFKKPSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPM 189
+PP P+PS+P+ SP PP P +P PP S P PP P
Sbjct: 98 SPPPPTPTPSVPSPTPPVSPPPP-----TPTPSVPSPTPPVSPP---------PPTPTPS 143
Query: 190 KKALSQHVSAPKKV-TPSKRPATQPL 214
+ + VS P TPS T P+
Sbjct: 144 VPSPTPPVSPPPPTPTPSVPSPTPPV 169
Score = 34.3 bits (77), Expect = 0.071
Identities = 20/70 (28%), Positives = 33/70 (46%), Gaps = 1/70 (1%)
Query: 136 PSPSMPAQPKTDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAPMKKALSQ 195
P+PS+P+ P +P PP P +V P P S P+ + PP + + A ++
Sbjct: 203 PTPSVPSPPDV-TPTPPTPSVPSPPDVTPTPPTPPSVPTPSGSPPYVPPPSDEEEAAGAK 261
Query: 196 HVSAPKKVTP 205
V K+ +P
Sbjct: 262 RVRCKKQRSP 271
Score = 32.7 bits (73), Expect = 0.21
Identities = 22/85 (25%), Positives = 35/85 (40%), Gaps = 18/85 (21%)
Query: 130 NPPFKKPSPSMPAQPK-TDSPKPPQIKGCQPKNVLPKADPPKSSPSNAAEAIRAPPNTAP 188
+PP P+PS+P+ P T +P P + PP +P+ ++ +PP+ P
Sbjct: 182 SPPPPTPTPSVPSPPDVTPTPPTPSV-----------PSPPDVTPTPPTPSVPSPPDVTP 230
Query: 189 MKKALSQHVSAPKKVTPSKRPATQP 213
+ P TPS P P
Sbjct: 231 TPP------TPPSVPTPSGSPPYVP 249
Score = 28.9 bits (63), Expect = 3.0
Identities = 22/68 (32%), Positives = 27/68 (39%), Gaps = 2/68 (2%)
Query: 116 PFGVLCTPITMIRYNPPFKKPSPSMPAQPKTD--SPKPPQIKGCQPKNVLPKADPPKSSP 173
P V TP T +PP P+P P+ P +P PP + P PP S
Sbjct: 195 PPDVTPTPPTPSVPSPPDVTPTPPTPSVPSPPDVTPTPPTPPSVPTPSGSPPYVPPPSDE 254
Query: 174 SNAAEAIR 181
AA A R
Sbjct: 255 EEAAGAKR 262
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.133 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,932,839
Number of Sequences: 26719
Number of extensions: 349370
Number of successful extensions: 3027
Number of sequences better than 10.0: 197
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 126
Number of HSP's that attempted gapping in prelim test: 1431
Number of HSP's gapped (non-prelim): 976
length of query: 267
length of database: 11,318,596
effective HSP length: 98
effective length of query: 169
effective length of database: 8,700,134
effective search space: 1470322646
effective search space used: 1470322646
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0091b.13


