

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091b.10
(353 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
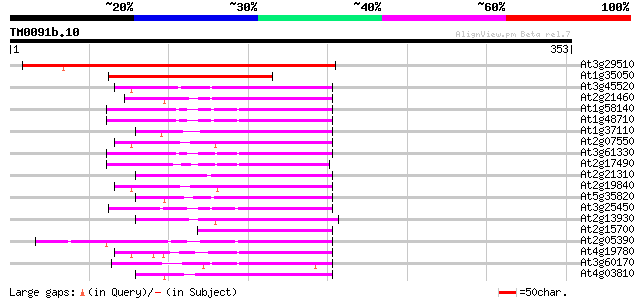
Score E
Sequences producing significant alignments: (bits) Value
At3g29510 hypothetical protein 165 4e-41
At1g35050 putative protein 114 6e-26
At3g45520 copia-like polyprotein 75 7e-14
At2g21460 putative retroelement pol polyprotein 74 1e-13
At1g58140 hypothetical protein 70 2e-12
At1g48710 hypothetical protein 70 2e-12
At1g37110 69 4e-12
At2g07550 putative retroelement pol polyprotein 69 5e-12
At3g61330 copia-type polyprotein 68 7e-12
At2g17490 putative retroelement pol polyprotein 67 1e-11
At2g21310 putative retroelement pol polyprotein 67 1e-11
At2g19840 copia-like retroelement pol polyprotein 67 1e-11
At5g35820 copia-like retrotransposable element 64 1e-10
At3g25450 hypothetical protein 64 2e-10
At2g13930 putative retroelement pol polyprotein 63 2e-10
At2g15700 copia-like retroelement pol polyprotein 63 3e-10
At2g05390 putative retroelement pol polyprotein 63 3e-10
At4g19780 putative LTR retrotransposon (fragment) 62 6e-10
At3g60170 putative protein 60 1e-09
At4g03810 putative retrotransposon protein 60 2e-09
>At3g29510 hypothetical protein
Length = 1158
Score = 165 bits (417), Expect = 4e-41
Identities = 92/202 (45%), Positives = 132/202 (64%), Gaps = 5/202 (2%)
Query: 9 QNDPTNSCIFMMRYEIV-YVIINVY--DNNLLRL*KSSQKL*IC*RKNLR*KS*-E*NFC 64
+NDP + CIF+ ++ +VII VY D N+L + +K K + FC
Sbjct: 804 KNDPISPCIFIKKFASKGFVIIAVYVDDLNILGTSGEIAQTVEYLKKEFEMKDLGKTKFC 863
Query: 65 LALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEEL 124
L L +EY+ + ++Q+ Y VLK F MDK+ PL + + +RSL ++ DPF P + +EE+
Sbjct: 864 LGLQLEYIDKGILVHQKAYTETVLKRFNMDKAHPLTSPMQVRSLGLDSDPFGPKKDDEEI 923
Query: 125 LDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDM 183
L PE+PYLSAI AL+YL+SHTR DI FA+NLL+R+SS PT+RHW G K + RYL+G +D
Sbjct: 924 LGPEMPYLSAIGALMYLSSHTRPDICFAVNLLSRFSSCPTKRHWEGIKHLLRYLQGTIDF 983
Query: 184 SLFFPFMSKLNLIGYADAGYLS 205
L++ +K L+G+ADAGYLS
Sbjct: 984 GLYYTNHNKEGLVGFADAGYLS 1005
>At1g35050 putative protein
Length = 450
Score = 114 bits (286), Expect = 6e-26
Identities = 58/103 (56%), Positives = 74/103 (71%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
FCL L IE+ +D +F++Q Y K+LK F MDK+ PL T +V RSLNV DPFRP E NE
Sbjct: 254 FCLGLQIEHFQDGIFVHQSNYTKKILKRFNMDKANPLSTPMVNRSLNVENDPFRPCEDNE 313
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQ 165
+ L P+VPY+SAI L+YLA+ TR DI+FA NLLAR ++ Q
Sbjct: 314 DFLGPKVPYMSAIGGLMYLANCTRPDIAFATNLLARSMTNHIQ 356
>At3g45520 copia-like polyprotein
Length = 1363
Score = 74.7 bits (182), Expect = 7e-14
Identities = 48/142 (33%), Positives = 76/142 (52%), Gaps = 7/142 (4%)
Query: 67 LSIEYLKDE----VFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
L IE + D +++ Q +Y+ KVLK F M +S P T L L + E
Sbjct: 1079 LGIEIIIDREAGVLWLSQESYLNKVLKTFNMLESKPALTPLGAH-LKMKSATEEKLSTEE 1137
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
E ++ VPY SA+ +++Y TR D+++ + +++R+ S P + HW G K V RY+KG +
Sbjct: 1138 EYMN-SVPYSSAVGSIMYAMIGTRPDLAYPVGVVSRFMSQPAKEHWLGVKWVLRYIKGTV 1196
Query: 182 DMSLFFPFMSKLNLIGYADAGY 203
D L + S ++ GY DA Y
Sbjct: 1197 DTRLCYKRNSDFSICGYCDADY 1218
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 73.9 bits (180), Expect = 1e-13
Identities = 42/136 (30%), Positives = 77/136 (55%), Gaps = 11/136 (8%)
Query: 73 KDEVFMYQRTYIAKVLKGFYMDKS----CPLCTLLVLRSLNVNKDPFRP*EKNEELLDPE 128
++ +++ Q Y+ K+L+ + M +S PL L +R+ V K E++E+ +
Sbjct: 1059 ENTLWLSQNGYLNKILETYNMAESKHVVTPLGAHLKMRAATVEKQ-----EQDEDYMK-S 1112
Query: 129 VPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFF 187
+PY SA+ +++Y TR D+++ + +++RY S P + HW G K V RY+KG+L L +
Sbjct: 1113 IPYSSAVGSIMYAMIGTRPDLAYPVGIISRYMSQPAREHWLGVKWVLRYIKGSLGTKLQY 1172
Query: 188 PFMSKLNLIGYADAGY 203
S ++GY DA +
Sbjct: 1173 KRSSDFKVVGYCDADH 1188
>At1g58140 hypothetical protein
Length = 1320
Score = 69.7 bits (169), Expect = 2e-12
Identities = 43/143 (30%), Positives = 79/143 (55%), Gaps = 10/143 (6%)
Query: 62 NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKN 121
++ L + ++ + +F+ Q Y +VLK F MD S P+CT + + ++K ++
Sbjct: 1033 SYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMEC-GIKLSK------KEE 1085
Query: 122 EELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWN-GKQVFRYLKGA 180
E +DP + S + +L YL TR DI +A+ +++RY PT H+ K++ RY+KG
Sbjct: 1086 GEGVDP-TTFKSLVGSLRYLTC-TRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGT 1143
Query: 181 LDMSLFFPFMSKLNLIGYADAGY 203
++ L + S L+GY+D+ +
Sbjct: 1144 VNFGLHYSTTSDYKLVGYSDSDW 1166
>At1g48710 hypothetical protein
Length = 1352
Score = 69.7 bits (169), Expect = 2e-12
Identities = 43/143 (30%), Positives = 79/143 (55%), Gaps = 10/143 (6%)
Query: 62 NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKN 121
++ L + ++ + +F+ Q Y +VLK F MD S P+CT + + ++K ++
Sbjct: 1065 SYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMEC-GIKLSK------KEE 1117
Query: 122 EELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWN-GKQVFRYLKGA 180
E +DP + S + +L YL TR DI +A+ +++RY PT H+ K++ RY+KG
Sbjct: 1118 GEGVDP-TTFKSLVGSLRYLTC-TRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGT 1175
Query: 181 LDMSLFFPFMSKLNLIGYADAGY 203
++ L + S L+GY+D+ +
Sbjct: 1176 VNFGLHYSTTSDYKLVGYSDSDW 1198
>At1g37110
Length = 1356
Score = 68.9 bits (167), Expect = 4e-12
Identities = 45/130 (34%), Positives = 66/130 (50%), Gaps = 16/130 (12%)
Query: 80 QRTYIAKVLKGFYMD----KSCPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEV-PYLSA 134
Q YI KVL F M + P+ L ++ + +E +D +V PY SA
Sbjct: 1091 QEIYIRKVLDRFNMSGAKMTNAPVGAHFKLAAVR----------EEDECVDTDVVPYSSA 1140
Query: 135 IKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKL 193
+ +++Y TR D+++AI L++RY S P HW K V RYLKGA D++L F
Sbjct: 1141 VGSIMYAMLGTRPDLAYAICLISRYMSKPGSMHWEAVKWVMRYLKGAQDLNLVFTKEKDF 1200
Query: 194 NLIGYADAGY 203
+ GY D+ Y
Sbjct: 1201 TVTGYCDSNY 1210
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 68.6 bits (166), Expect = 5e-12
Identities = 42/146 (28%), Positives = 77/146 (51%), Gaps = 15/146 (10%)
Query: 67 LSIEYLKDE----VFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
L +E ++D +++ Q Y+ K+L+ + M ++ P T L F+ + +
Sbjct: 1072 LGMEIIRDRTLGVLWLSQEGYLNKILETYNMAEAKPAMTPLGAHF------KFQAATEQK 1125
Query: 123 ELLDPE----VPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYL 177
+ D + VPY SA+ +++Y TR D+++ + +++R+ S P + HW G K V RY+
Sbjct: 1126 LIRDEDFMKSVPYSSAVGSIMYAMLGTRPDLAYPVGIISRFMSQPIKEHWLGVKWVLRYI 1185
Query: 178 KGALDMSLFFPFMSKLNLIGYADAGY 203
KG L L + S +++GY DA Y
Sbjct: 1186 KGTLKTRLCYKKSSSFSIVGYCDADY 1211
>At3g61330 copia-type polyprotein
Length = 1352
Score = 68.2 bits (165), Expect = 7e-12
Identities = 42/143 (29%), Positives = 79/143 (54%), Gaps = 10/143 (6%)
Query: 62 NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKN 121
++ L + ++ + +F+ Q Y +VLK F +D S P+CT + + ++K ++
Sbjct: 1065 SYYLGIEVKQEDNGIFITQEGYAKEVLKKFKIDDSNPVCTPMEC-GIKLSK------KEE 1117
Query: 122 EELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGA 180
E +DP + S + +L YL TR DI +A+ +++RY PT H+ K++ RY+KG
Sbjct: 1118 GEGVDPTT-FKSLVGSLRYLTC-TRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGT 1175
Query: 181 LDMSLFFPFMSKLNLIGYADAGY 203
++ L + S L+GY+D+ +
Sbjct: 1176 VNFGLHYSTTSDYKLVGYSDSDW 1198
>At2g17490 putative retroelement pol polyprotein
Length = 822
Score = 67.4 bits (163), Expect = 1e-11
Identities = 49/141 (34%), Positives = 75/141 (52%), Gaps = 11/141 (7%)
Query: 62 NFCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKN 121
N+ L + I + +F+ Q Y K+LK F M+ S + T L L KD E++
Sbjct: 691 NYFLGMEIVQSLEGIFLSQECYARKLLKKFNMEDSKIMSTPL----LPQRKDQ----EED 742
Query: 122 EELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWN-GKQVFRYLKGA 180
E L D +V Y S + LLYL S TR ++ F + L+RY P +H+ K+V RY+ G
Sbjct: 743 ESLADSKV-YRSLVGGLLYLTS-TRPNLMFFASYLSRYLKEPKIKHFKEAKRVLRYINGT 800
Query: 181 LDMSLFFPFMSKLNLIGYADA 201
+DM + F + LIG++D+
Sbjct: 801 IDMGMRFTSTMEPKLIGHSDS 821
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 67.0 bits (162), Expect = 1e-11
Identities = 38/125 (30%), Positives = 67/125 (53%), Gaps = 3/125 (2%)
Query: 80 QRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPYLSAIKALL 139
QR+Y+ KVLK F MD+ P+ T L V E+ +++ +PY +A+ +++
Sbjct: 571 QRSYLQKVLKTFRMDECKPVKTPLAPHMKFVAATETEAEEQADQM--KSIPYANAVGSIM 628
Query: 140 YLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKLNLIGY 198
Y +R D++ + +++R+ S P HW G K V RY+KG++D L + + GY
Sbjct: 629 YSMIGSRPDLAHPVGVISRFMSKPLMNHWLGVKWVLRYIKGSIDSRLQYKREGDFVVTGY 688
Query: 199 ADAGY 203
D+ +
Sbjct: 689 CDSDH 693
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 67.0 bits (162), Expect = 1e-11
Identities = 44/146 (30%), Positives = 74/146 (50%), Gaps = 15/146 (10%)
Query: 67 LSIEYLKDE----VFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
L +E ++D + + Q Y+ KVL F MD++ P+ T + +P E
Sbjct: 869 LGMEIVRDRKAGTMSISQEGYLLKVLGNFGMDQAKPVFTPMGAHF------KLKPATDEE 922
Query: 123 ELLDPEV----PYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYL 177
+ EV PY SA+ +L+Y TR D++ ++ L+ R+ S P + HW K + RY+
Sbjct: 923 VMRQSEVMRAVPYQSAVGSLMYSMIGTRPDLAHSVGLVCRFMSKPLKEHWQAVKWILRYI 982
Query: 178 KGALDMSLFFPFMSKLNLIGYADAGY 203
+G++D L + +L L GY D+ Y
Sbjct: 983 RGSIDRKLCYKNEGELILEGYCDSDY 1008
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 63.9 bits (154), Expect = 1e-10
Identities = 42/129 (32%), Positives = 68/129 (52%), Gaps = 11/129 (8%)
Query: 80 QRTYIAKVLKGFYMDKS----CPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPYLSAI 135
Q Y+A VL+ F MD+S PL LR+ N + ++ E + VPY +AI
Sbjct: 1075 QSEYVAGVLRAFGMDQSKVSQTPLGAHFKLRAANE-----KTLARDAEYMKL-VPYPNAI 1128
Query: 136 KALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKLN 194
+++Y +R D+++ + +++R+ S P++ HW K V RY+KG D L F K
Sbjct: 1129 GSIMYSMIGSRPDLAYPVGVVSRFMSKPSKEHWQAVKWVMRYMKGTQDTCLRFKKDDKFE 1188
Query: 195 LIGYADAGY 203
+ GY D+ Y
Sbjct: 1189 IRGYCDSDY 1197
>At3g25450 hypothetical protein
Length = 1343
Score = 63.5 bits (153), Expect = 2e-10
Identities = 44/142 (30%), Positives = 71/142 (49%), Gaps = 10/142 (7%)
Query: 63 FCLALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNE 122
+ L + + KD + + Q Y K+L+ M K C ++ SL ++K ++E
Sbjct: 1048 YYLGIEVLQSKDGITLKQERYAKKILEEAGMSK-CNTVNTPMIASLELSK------AQDE 1100
Query: 123 ELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGAL 181
+ +D E Y I L YL HTR D+S+ + +L+RY P + H KQ+ RYL+G
Sbjct: 1101 KRID-ETDYRRNIGCLRYLL-HTRPDLSYNVGILSRYLQEPRESHGAALKQILRYLQGTT 1158
Query: 182 DMSLFFPFMSKLNLIGYADAGY 203
L+F LIGY+D+ +
Sbjct: 1159 SHGLYFKKGENAGLIGYSDSSH 1180
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 63.2 bits (152), Expect = 2e-10
Identities = 41/132 (31%), Positives = 67/132 (50%), Gaps = 9/132 (6%)
Query: 80 QRTYIAKVLKGFYMDKSCPLCTLLVLR-SLNVNKDPFRP*EKNEELLDPE--VPYLSAIK 136
Q Y+ KVL+ F MD + P+ T L + L D ++ EE + VPY + I
Sbjct: 1068 QEGYVKKVLRSFQMDNAKPVSTPLGIHFKLKAATD-----KEYEEQFERMKIVPYANTIG 1122
Query: 137 ALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFMSKLNL 195
+++Y TR D+++++ +++R+ S P + HW K V RY++G L F L
Sbjct: 1123 SIMYSMIGTRPDLAYSLGVISRFMSKPLKDHWQAVKWVLRYMRGTEKKKLCFRKQEDFLL 1182
Query: 196 IGYADAGYLSKY 207
GY D+ Y S +
Sbjct: 1183 RGYCDSDYGSNF 1194
>At2g15700 copia-like retroelement pol polyprotein
Length = 1166
Score = 62.8 bits (151), Expect = 3e-10
Identities = 30/86 (34%), Positives = 51/86 (58%), Gaps = 1/86 (1%)
Query: 119 EKNEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYL 177
+K E + +PY + + +L+Y +R D++F + ++R+ SSP + HW+ K V RYL
Sbjct: 975 KKTEAVHMERIPYANVVGSLMYAMVGSRPDLAFVVGFISRFMSSPGREHWSAVKWVLRYL 1034
Query: 178 KGALDMSLFFPFMSKLNLIGYADAGY 203
KGA +L F SK + G++D+ Y
Sbjct: 1035 KGAYTQNLIFKKDSKFCIEGFSDSDY 1060
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 62.8 bits (151), Expect = 3e-10
Identities = 52/193 (26%), Positives = 93/193 (47%), Gaps = 16/193 (8%)
Query: 17 IFMMRYEIVYVIINVYDNNLLRL*KSSQKL*IC*RKNLR*KS*-----E*NFCLALSIEY 71
++ + E +I+ +Y ++LL + SS L +C +K++ K + + L + + +
Sbjct: 959 VYRRQEEKKLLIVAIYVDDLL-VTGSSLDLILCFKKDMAGKFEMSDLGQLTYYLGIEVLH 1017
Query: 72 LKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPY 131
K+ + + Q Y K+++ M P+ + + L + K EE E Y
Sbjct: 1018 RKNGIILRQERYAMKIIEEAGMSNCNPVL-IPMAAGLELCK-------AQEEKCITERDY 1069
Query: 132 LSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKGALDMSLFFPFM 190
I L Y+ HTR D+S+ + +L+RY P + H N KQV RYLKG + L+
Sbjct: 1070 RRMIGCLRYIV-HTRPDLSYCVGVLSRYLQQPRESHGNALKQVLRYLKGTMSHGLYLKRG 1128
Query: 191 SKLNLIGYADAGY 203
K L+GY+D+ +
Sbjct: 1129 FKSGLVGYSDSSH 1141
>At4g19780 putative LTR retrotransposon (fragment)
Length = 306
Score = 61.6 bits (148), Expect = 6e-10
Identities = 50/144 (34%), Positives = 72/144 (49%), Gaps = 20/144 (13%)
Query: 67 LSIEYLKDE--VFMYQRTYIAKVLK--GFYMDK--SCPLCTLLVLRSLNVNKDPFRP*EK 120
L IE + E + + QR YI ++L GF K S PL S+ +NK+ P
Sbjct: 24 LGIEIARSEKGISICQRKYILELLSTTGFLGSKPSSIPLDP-----SVKLNKEDGVP--- 75
Query: 121 NEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHWNG-KQVFRYLKG 179
L Y + L+YL TR DI++A+N L ++S +PT H N +V RYLKG
Sbjct: 76 ----LTDSTSYRKLVGKLMYLQI-TRPDIAYAVNTLCQFSHAPTSVHLNAVHKVLRYLKG 130
Query: 180 ALDMSLFFPFMSKLNLIGYADAGY 203
+ LF+P K +L GY D+ +
Sbjct: 131 TVGQGLFYPADDKFDLRGYTDSDF 154
>At3g60170 putative protein
Length = 1339
Score = 60.5 bits (145), Expect = 1e-09
Identities = 43/147 (29%), Positives = 73/147 (49%), Gaps = 22/147 (14%)
Query: 65 LALSIEYLKDEVFMYQRTYIAKVLKGFYMDKSCPLCTLLVLRSLNVNKDPFRP*EK---- 120
L + ++ +F+ QR Y +VL F MD+S N K+P P K
Sbjct: 1057 LGIEVKQSDGGIFICQRRYAREVLARFGMDES------------NAVKNPIVPGTKLTKD 1104
Query: 121 -NEELLDPEVPYLSAIKALLYLASHTRLDISFAINLLARYSSSPTQRHW-NGKQVFRYLK 178
N E +D E + + +L+YL TR D+ + + L++R+ S+P HW K++ RYLK
Sbjct: 1105 ENGEKVD-ETMFKQLVGSLMYLTV-TRPDLMYGVCLISRFMSNPRMSHWLAAKRILRYLK 1162
Query: 179 GALDMSLFFPFMS--KLNLIGYADAGY 203
G +++ +F+ L L+ + D+ Y
Sbjct: 1163 GTVELGIFYRRRKNRSLKLMAFTDSDY 1189
>At4g03810 putative retrotransposon protein
Length = 964
Score = 60.1 bits (144), Expect = 2e-09
Identities = 40/129 (31%), Positives = 64/129 (49%), Gaps = 11/129 (8%)
Query: 80 QRTYIAKVLKGFYMDKS----CPLCTLLVLRSLNVNKDPFRP*EKNEELLDPEVPYLSAI 135
Q TYI KVL F M S P+ + L P +E ++PY SAI
Sbjct: 692 QDTYIDKVLHRFNMHDSKKGFIPMSHGITLSKTQC------PSTHDERERMSKIPYASAI 745
Query: 136 KALLYLASHTRLDISFAINLLARYSSSPTQRHW-NGKQVFRYLKGALDMSLFFPFMSKLN 194
+++Y +TR D++ A+++ +RY S P + HW + +F+YL+ D L + +L
Sbjct: 746 GSIMYAMLYTRPDVACALSMTSRYQSDPGESHWIVVRNIFKYLRRTKDKFLVYGGSEELV 805
Query: 195 LIGYADAGY 203
+ GY DA +
Sbjct: 806 VSGYTDASF 814
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.357 0.160 0.555
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,725,301
Number of Sequences: 26719
Number of extensions: 247682
Number of successful extensions: 1278
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 56
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 1134
Number of HSP's gapped (non-prelim): 101
length of query: 353
length of database: 11,318,596
effective HSP length: 100
effective length of query: 253
effective length of database: 8,646,696
effective search space: 2187614088
effective search space used: 2187614088
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0091b.10


