

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.6
(242 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
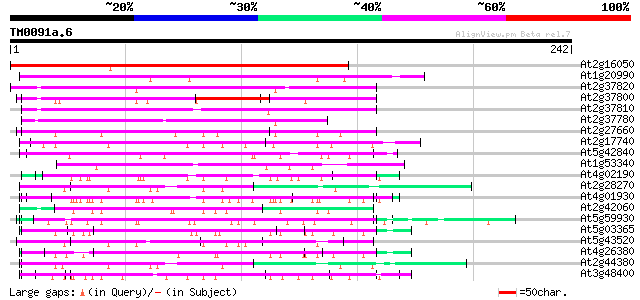
Score E
Sequences producing significant alignments: (bits) Value
At2g16050 hypothetical protein 259 9e-70
At1g20990 hypothetical protein 106 1e-23
At2g37820 hypothetical protein 100 5e-22
At2g37800 hypothetical protein 95 4e-20
At2g37810 hypothetical protein 89 2e-18
At2g37780 hypothetical protein 75 2e-14
At2g27660 hypothetical protein 75 2e-14
At2g17740 unknown protein 69 2e-12
At5g42840 CHP-rich zinc finger protein-like 69 3e-12
At1g53340 hypothetical protein 69 3e-12
At4g02190 CHP-rich hypothetical protein similar to T10M13.17.1 68 4e-12
At2g28270 pseudogene 67 7e-12
At4g01930 putative CHP-rich zinc finger protein 67 9e-12
At2g42060 unknown protein 66 1e-11
At5g59930 putative protein 66 2e-11
At5g03365 putative protein 65 3e-11
At5g43520 putative protein 65 3e-11
At4g26380 hypothetical protein 65 3e-11
At2g44380 unknown protein 65 4e-11
At3g48400 putative protein 64 7e-11
>At2g16050 hypothetical protein
Length = 167
Score = 259 bits (662), Expect = 9e-70
Identities = 109/151 (72%), Positives = 128/151 (84%), Gaps = 5/151 (3%)
Query: 1 MKYSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCS-----ICDYDLHMQCAIP 55
MKY++ISHFSHPQH L+F++ PFKCDGC E+GIGSRY+CS CD+DLH CA+P
Sbjct: 1 MKYADISHFSHPQHNLKFDYTEKPFKCDGCNEVGIGSRYRCSGDNHGSCDFDLHAHCALP 60
Query: 56 SPTLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVL 115
S ++ HPFY KC+FQF++ PPGN RYCNACEKDV+GF+YHCK+CGFDLHPCCA LPMVL
Sbjct: 61 SASITHPFYKKCTFQFVARPPGNERRYCNACEKDVSGFLYHCKACGFDLHPCCAMLPMVL 120
Query: 116 DDGEVKLYLYRKVSSPCHRCGRKGRSWSYRS 146
DDGE L+LYRKVSS CHRCG+KGRSWSYRS
Sbjct: 121 DDGETNLFLYRKVSSSCHRCGKKGRSWSYRS 151
>At1g20990 hypothetical protein
Length = 319
Score = 106 bits (264), Expect = 1e-23
Identities = 64/193 (33%), Positives = 86/193 (44%), Gaps = 21/193 (10%)
Query: 5 EISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL-VHPF 63
E+ HF HPQH L + C GCKE G G RY C CDY LH CA+ P L HPF
Sbjct: 67 EMVHFGHPQHVLVKVELPDIYTCAGCKEEGAGVRYVCQECDYQLHEFCALAPPQLKSHPF 126
Query: 64 YTKCSFQFMSSPP--GNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVK 121
+ + F + P G + C+ C + G+ + CK+C F +HP CA L L +
Sbjct: 127 HYQHQLLFFAKPAKGGIVKSKCDVCGRSPKGYTFRCKACSFQMHPGCAMLSPSLSSSSLH 186
Query: 122 LYLYRKVSSP--------------CHRCGRKGRSWS-YRSKCKSYNLHVACVREMVVENW 166
+ R + S C C R R+ YR Y+LH C ++ V
Sbjct: 187 HHPLRLLPSSSAGSTTGGDSGGFLCGECKRGKRTGRVYRCTVCDYHLHAVCAKDAAVNG- 245
Query: 167 DHVYVAGGKGRNK 179
+ G KGR+K
Sbjct: 246 --LRANGHKGRDK 256
>At2g37820 hypothetical protein
Length = 290
Score = 100 bits (250), Expect = 5e-22
Identities = 55/165 (33%), Positives = 78/165 (46%), Gaps = 9/165 (5%)
Query: 1 MKYSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCA---IPSP 57
+K + HF+H H L ++ F CDGCK G G Y+C C+YDLH CA + P
Sbjct: 6 LKRHTVQHFTHI-HPLTEFNSIGDFICDGCKTYGSGKTYRCEPCNYDLHEYCATCPLTLP 64
Query: 58 TLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPM---- 113
T +HP + R C+ C++ V G Y CK C FD+HP C +LP
Sbjct: 65 TFIHPQHELSLVVRKQQSTRQNDRACDICDESVEGLFYRCKICEFDVHPLCTQLPQHVRH 124
Query: 114 VLDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
VL L +S C C +SW YR + +++H+ C+
Sbjct: 125 VLHPAH-HLEFRPSGASTCMVCHGPCQSWRYRCELCRFDIHMECI 168
Score = 37.7 bits (86), Expect = 0.006
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 1/49 (2%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAI 54
+ H HP H L F + C C RY+C +C +D+HM+C +
Sbjct: 122 VRHVLHPAHHLEFRPSGAS-TCMVCHGPCQSWRYRCELCRFDIHMECIL 169
>At2g37800 hypothetical protein
Length = 396
Score = 94.7 bits (234), Expect = 4e-20
Identities = 51/160 (31%), Positives = 74/160 (45%), Gaps = 8/160 (5%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYT 65
+ HF+H H L F CDGCK G G Y+C+ CDY+LH CA TL +
Sbjct: 145 VQHFTHI-HPLTKVDGYGEFTCDGCKTYGFGKTYRCTRCDYNLHDHCATCPSTLATFMHP 203
Query: 66 KCSFQFMSSPPGNI---PRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKL 122
+ + + P + R C+ C++ G Y C+ CGFD+HP C +LP +
Sbjct: 204 QHELRLVFRGPEHTHQNKRMCDICDESAEGLYYQCEPCGFDVHPLCTQLPQHVRHVPHPA 263
Query: 123 YLYR----KVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
+L SS C C RSW Y+ ++H+ C+
Sbjct: 264 HLLELSQWGASSICMVCRGAIRSWRYKCGPCGLDVHMECI 303
Score = 51.6 bits (122), Expect = 4e-07
Identities = 35/114 (30%), Positives = 50/114 (43%), Gaps = 15/114 (13%)
Query: 4 SEISHFSHPQHKLRF-----EH-NAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCA-IPS 56
S ++ F HPQH+LR EH + CD C E G Y+C C +D+H C +P
Sbjct: 195 STLATFMHPQHELRLVFRGPEHTHQNKRMCDICDESAEGLYYQCEPCGFDVHPLCTQLPQ 254
Query: 57 PT--LVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCC 108
+ HP + Q+ +S C C + + Y C CG D+H C
Sbjct: 255 HVRHVPHPAHLLELSQWGAS------SICMVCRGAIRSWRYKCGPCGLDVHMEC 302
Score = 48.1 bits (113), Expect = 4e-06
Identities = 16/32 (50%), Positives = 21/32 (65%)
Query: 81 RYCNACEKDVNGFVYHCKSCGFDLHPCCAKLP 112
R C+ C++ G Y CK CGFD+HP C +LP
Sbjct: 34 RMCDICDESAEGLYYQCKPCGFDVHPLCTQLP 65
Score = 36.6 bits (83), Expect = 0.012
Identities = 17/54 (31%), Positives = 23/54 (42%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL 59
+ H HP H L C C+ RYKC C D+HM+C S ++
Sbjct: 256 VRHVPHPAHLLELSQWGASSICMVCRGAIRSWRYKCGPCGLDVHMECISSSASV 309
Score = 32.3 bits (72), Expect = 0.23
Identities = 30/128 (23%), Positives = 39/128 (30%), Gaps = 50/128 (39%)
Query: 38 RYKCSICDYDLHMQCAIPS-------------------PTLVHPFY-------------T 65
RYKC C D+HM+C S P P+Y
Sbjct: 70 RYKCGQCRVDVHMECVDSSTSVAETGIQQRGFGQQFEFPQYHQPYYNHGYNYGYINQENN 129
Query: 66 KCSFQFMSSPP------------------GNIPRYCNACEKDVNGFVYHCKSCGFDLHPC 107
K + + SS P G C+ C+ G Y C C ++LH
Sbjct: 130 KKTTKMSSSRPEQLVVQHFTHIHPLTKVDGYGEFTCDGCKTYGFGKTYRCTRCDYNLHDH 189
Query: 108 CAKLPMVL 115
CA P L
Sbjct: 190 CATCPSTL 197
Score = 30.0 bits (66), Expect = 1.2
Identities = 11/26 (42%), Positives = 14/26 (53%)
Query: 27 CDGCKEIGIGSRYKCSICDYDLHMQC 52
CD C E G Y+C C +D+H C
Sbjct: 36 CDICDESAEGLYYQCKPCGFDVHPLC 61
>At2g37810 hypothetical protein
Length = 233
Score = 89.0 bits (219), Expect = 2e-18
Identities = 51/157 (32%), Positives = 71/157 (44%), Gaps = 8/157 (5%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYT 65
+ HFSH H L F C GCK G G Y+C+ CDY LH CA TL+ +
Sbjct: 11 VQHFSHT-HPLTKVDEYGEFTCGGCKTYGFGKTYRCAPCDYVLHDHCATCPFTLISFMHP 69
Query: 66 KCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAK----LPMVLDDGEVK 121
+ + + N+ CN C + V G YHC++CGFD+HP C + + V +
Sbjct: 70 QHELRLFVNGSENM---CNICHRLVEGVYYHCETCGFDVHPLCTQHHQHVSYVPHPAHLL 126
Query: 122 LYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
S+ C C SW Y+ ++HV CV
Sbjct: 127 ELSQCGASNTCMVCRGAILSWRYKCGPCMLDVHVECV 163
Score = 37.4 bits (85), Expect = 0.007
Identities = 18/61 (29%), Positives = 26/61 (42%), Gaps = 1/61 (1%)
Query: 3 YSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPS-PTLVH 61
+ +S+ HP H L C C+ + RYKC C D+H++C S P
Sbjct: 113 HQHVSYVPHPAHLLELSQCGASNTCMVCRGAILSWRYKCGPCMLDVHVECVNSSAPAATE 172
Query: 62 P 62
P
Sbjct: 173 P 173
>At2g37780 hypothetical protein
Length = 286
Score = 75.5 bits (184), Expect = 2e-14
Identities = 44/138 (31%), Positives = 61/138 (43%), Gaps = 8/138 (5%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYT 65
I HF+H H L + + CDGCK G G Y+CS CDYDLH CA L++ +
Sbjct: 7 IQHFTH-NHLLTQVNGIGTYTCDGCKLYGEGRTYRCSDCDYDLHEYCATCPSILLNSCHG 65
Query: 66 KCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPM------VLDDGE 119
+ R C C + G Y C+ C F+ HP C PM +L +
Sbjct: 66 P-DHELSLFNGHMTERSCYVCRVSIQGMFYKCRQCSFEAHPLCTYAPMHASSPDLLVTKQ 124
Query: 120 VKLYLYRKVSSPCHRCGR 137
L+ + SP H+ G+
Sbjct: 125 RSLHGHAGQPSPPHQYGQ 142
>At2g27660 hypothetical protein
Length = 718
Score = 75.5 bits (184), Expect = 2e-14
Identities = 48/167 (28%), Positives = 73/167 (42%), Gaps = 13/167 (7%)
Query: 4 SEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSR-YKCSICDYDLHMQCAIPSPTLVHP 62
S I+HFSHP H+L+ C CK G R Y C C++ LH C+ + HP
Sbjct: 10 SFINHFSHP-HRLQLTPATSSPPCSACKLTGGNGRVYSCRPCNFSLHESCSKMKQVITHP 68
Query: 63 FYTKCSFQFMSSPPGNIPRY-CNACEKDVNGFVYHCKSCGFDLHPCCAKLPM-VLDDGEV 120
+ + + +P + + C+ C GF Y C C FD+H CA P+ ++
Sbjct: 69 SHPSHTLSLLVAPVYDGGYFNCDGCGIHGTGFSYQCSVCDFDIHALCAYKPLSIIHKSHP 128
Query: 121 KLYLYRKVSSP--------CHRCGRKGRS-WSYRSKCKSYNLHVACV 158
+ L SP C C + G++ W YR ++ HV C+
Sbjct: 129 QHNLKLAFQSPYGANKGFSCDICRKIGKNQWLYRCIPCEFDAHVGCI 175
Score = 70.9 bits (172), Expect = 6e-13
Identities = 39/114 (34%), Positives = 56/114 (48%), Gaps = 7/114 (6%)
Query: 6 ISHFSHPQHKLRF----EHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVH 61
I+H SHP H L ++ F CDGC G G Y+CS+CD+D+H CA +++H
Sbjct: 65 ITHPSHPSHTLSLLVAPVYDGGYFNCDGCGIHGTGFSYQCSVCDFDIHALCAYKPLSIIH 124
Query: 62 PFYTKCSFQ--FMSSPPGNIPRYCNACEK-DVNGFVYHCKSCGFDLHPCCAKLP 112
+ + + + F S N C+ C K N ++Y C C FD H C P
Sbjct: 125 KSHPQHNLKLAFQSPYGANKGFSCDICRKIGKNQWLYRCIPCEFDAHVGCITGP 178
>At2g17740 unknown protein
Length = 248
Score = 68.9 bits (167), Expect = 2e-12
Identities = 39/115 (33%), Positives = 57/115 (48%), Gaps = 9/115 (7%)
Query: 5 EISHFSHPQHKLRF----EHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLV 60
EI H SHP H L ++N + CD C E G G Y CSIC YD+H+ C T+
Sbjct: 62 EIRHKSHPDHPLILLYSPQNNNSTYTCDACGEYGSGFTYNCSICQYDVHVGCVSVPETMK 121
Query: 61 HPFYTKCSFQFMSSP-PGNIPRYCNACEKDV--NGFVYHCKSC--GFDLHPCCAK 110
H + +P P + C+ C++ + N ++Y+C+ C G LH C A+
Sbjct: 122 HDEHVHPLALIYKAPCPKDHIFTCDVCDETMPHNLWLYYCQKCDYGAHLHSCVAE 176
Score = 61.2 bits (147), Expect = 5e-10
Identities = 54/184 (29%), Positives = 75/184 (40%), Gaps = 24/184 (13%)
Query: 10 SHPQ--HKLRFEHNAFPFKCDGCKEIGIGSRYKC--SICDYDLHMQCAIPSPTLVHPFYT 65
+HP HK + E C GC +G+ +KC S CDY LH C + H +
Sbjct: 13 NHPLRGHKAQVEDEII---CSGCDLDLLGAYFKCTKSECDYFLHKSCFDLPREIRHKSHP 69
Query: 66 KCSFQFMSSPPGNIPRY-CNACEKDVNGFVYHCKSCGFDLHPCCAKLP--MVLDDGEVKL 122
+ SP N Y C+AC + +GF Y+C C +D+H C +P M D+ L
Sbjct: 70 DHPLILLYSPQNNNSTYTCDACGEYGSGFTYNCSICQYDVHVGCVSVPETMKHDEHVHPL 129
Query: 123 YLYRKVSSP------CHRCGR--KGRSWSYRSKCKSYNLHV-ACVREMVVENWDHVYVAG 173
L K P C C W Y + Y H+ +CV E + G
Sbjct: 130 ALIYKAPCPKDHIFTCDVCDETMPHNLWLYYCQKCDYGAHLHSCVAEE-----EEKSKKG 184
Query: 174 GKGR 177
G+GR
Sbjct: 185 GRGR 188
>At5g42840 CHP-rich zinc finger protein-like
Length = 671
Score = 68.6 bits (166), Expect = 3e-12
Identities = 58/179 (32%), Positives = 76/179 (42%), Gaps = 21/179 (11%)
Query: 5 EISHFSHPQHKLRFEHNAFP----FKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLV 60
EI+H SH +H L+ N P C C + Y C IC ++L+M+CAI PT V
Sbjct: 169 EITHPSHVRHPLKLRSNGAPDYTNLNCHICGDATGNLLYHCDICKFNLNMRCAIREPTPV 228
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDL-HPCCAKLPMVL---- 115
K ++ P I C+AC + Y C C F + H CAKLP V+
Sbjct: 229 ALSNMKVHEHTLTLMPRLISFVCDACGMKGDRAPYVCVQCDFMIFHQECAKLPRVIHVNH 288
Query: 116 DDGEVKLYLYRKVSSPCH-RCG--RKGRSWSYR----SKCKSYNLHVACVREMVVENWD 167
D V Y+ P RCG + WSY S C +Y +H C V WD
Sbjct: 289 HDHRVS---YKYPLGPGEWRCGVCWEEIDWSYGAYSCSLCPNYAMHSLCATRKDV--WD 342
Score = 43.5 bits (101), Expect = 1e-04
Identities = 35/163 (21%), Positives = 61/163 (36%), Gaps = 18/163 (11%)
Query: 8 HFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIP---SPTLVHPFY 64
+F +H + + +CD C G C C + +H +C HP +
Sbjct: 4 YFEGHEHHVSIIKHRDGLECDACDR-SFGDVISCGECKFTVHRKCVFMFDIQEIFDHPSH 62
Query: 65 T-KCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFD------LHPCCAKLPMVL-- 115
C + P + + C+ C K +YHC C + + CA+ P+ L
Sbjct: 63 DGHCLKLLTTGAPDDTDQKCHLCGKKTKRLLYHCSDCKLNVDIDCIIDYICARSPLKLPW 122
Query: 116 -DDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVAC 157
D +K+ L + PC C G + +C+ + +H C
Sbjct: 123 HDHPLIKVDL--GANMPCDFCNESGIDYCC-PRCR-FMIHERC 161
Score = 39.7 bits (91), Expect = 0.001
Identities = 32/109 (29%), Positives = 44/109 (40%), Gaps = 12/109 (11%)
Query: 8 HFSHPQHKLRFEHNAFPFK-CDGC----KEIGIGSRYKCSICDYDLHMQCAIPSPTLVHP 62
H SH H L + K C+ C KE IG C+ C+Y L +CA T+ P
Sbjct: 485 HGSHDHHLLFLKLGRGNVKTCECCGIVQKEYAIG----CTKCNYFLDFRCATLPLTVRLP 540
Query: 63 FYTKCSFQFM-SSPPGNIPRYCNACEKDVN--GFVYHCKSCGFDLHPCC 108
Y + +C+ CE++ N + Y CK CG H C
Sbjct: 541 RYDDHPLTLCYGDEKASGKCWCDICERETNPKSWFYTCKDCGVTFHIFC 589
Score = 36.2 bits (82), Expect = 0.016
Identities = 41/168 (24%), Positives = 60/168 (35%), Gaps = 17/168 (10%)
Query: 8 HFSHPQHKLRFEHNAFPF----KCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPF 63
H SH H L+ P KC C + Y CS C ++ + C I P
Sbjct: 59 HPSHDGHCLKLLTTGAPDDTDQKCHLCGKKTKRLLYHCSDCKLNVDIDCIIDYICARSPL 118
Query: 64 YTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKL---PMVLDDGEV 120
+ C+ C + +G Y C C F +H CA + P + V
Sbjct: 119 KLPWHDHPLIKVDLGANMPCDFCNE--SGIDYCCPRCRFMIHERCASVFDSPEITHPSHV 176
Query: 121 KLYL-YRKVSSP------CHRCGRKGRSWSYRSKCKSYNLHVAC-VRE 160
+ L R +P CH CG + Y +NL++ C +RE
Sbjct: 177 RHPLKLRSNGAPDYTNLNCHICGDATGNLLYHCDICKFNLNMRCAIRE 224
Score = 32.7 bits (73), Expect = 0.18
Identities = 28/98 (28%), Positives = 39/98 (39%), Gaps = 11/98 (11%)
Query: 71 FMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVL-----DDGEVKL-YL 124
F+ GN+ + C C + C C + L CA LP+ + DD + L Y
Sbjct: 494 FLKLGRGNV-KTCECCGIVQKEYAIGCTKCNYFLDFRCATLPLTVRLPRYDDHPLTLCYG 552
Query: 125 YRKVSSP--CHRCGRK--GRSWSYRSKCKSYNLHVACV 158
K S C C R+ +SW Y K H+ CV
Sbjct: 553 DEKASGKCWCDICERETNPKSWFYTCKDCGVTFHIFCV 590
Score = 31.6 bits (70), Expect = 0.40
Identities = 22/91 (24%), Positives = 35/91 (38%), Gaps = 12/91 (13%)
Query: 83 CNACEKDVNGFVYHCKSCGFDLHPCCA---KLPMVLD----DGEVKLYLYR----KVSSP 131
C+AC++ G V C C F +H C + + D DG L
Sbjct: 23 CDACDRSF-GDVISCGECKFTVHRKCVFMFDIQEIFDHPSHDGHCLKLLTTGAPDDTDQK 81
Query: 132 CHRCGRKGRSWSYRSKCKSYNLHVACVREMV 162
CH CG+K + Y N+ + C+ + +
Sbjct: 82 CHLCGKKTKRLLYHCSDCKLNVDIDCIIDYI 112
>At1g53340 hypothetical protein
Length = 667
Score = 68.6 bits (166), Expect = 3e-12
Identities = 49/157 (31%), Positives = 65/157 (41%), Gaps = 19/157 (12%)
Query: 21 NAFPFKCDGCKEIGIG-SRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSPPGNI 79
N + F C C I G S Y C C H +C HPFY S Q S P G +
Sbjct: 121 NIYDFICRACTTIEQGTSYYVCVTCGDQFHKECVGAPLEFKHPFYPSLSLQLYSPPSGRV 180
Query: 80 PRYCNACEKDVNGFVYHCKSCGFDLHPCCA--KLPMVLDDGE---VKLYLYRKVS-SPCH 133
C+ C+K + G Y+C + F LH CA +P V+D + L + K S PCH
Sbjct: 181 --LCSCCQKPIYGMNYYCPTSNFTLHLFCAFKPIPFVIDHPKRHPHPLTFFPKQSFLPCH 238
Query: 134 RCGRKGRSWSYRSKCKSYNLHVACVREMVVENWDHVY 170
C S K + C+R V + D +Y
Sbjct: 239 VC----------SLIKKFIPTYICIRCAFVVHQDCIY 265
Score = 38.1 bits (87), Expect = 0.004
Identities = 43/167 (25%), Positives = 66/167 (38%), Gaps = 34/167 (20%)
Query: 27 CDGCKEIGIGSRYKCSICDYDLHMQCAI-PSPTLV-------HP--FYTKCSFQFMSSPP 76
C C++ G Y C ++ LH+ CA P P ++ HP F+ K SF
Sbjct: 182 CSCCQKPIYGMNYYCPTSNFTLHLFCAFKPIPFVIDHPKRHPHPLTFFPKQSF------- 234
Query: 77 GNIPRYCNACEKDVNGFV--YHCKSCGFDLHPCCAKLPMV--LDDGEVKLYLYRKVSSPC 132
+P C+ C + F+ Y C C F +H C P V + ++ +SS
Sbjct: 235 --LP--CHVCSL-IKKFIPTYICIRCAFVVHQDCIYFPYVIKISRHHHRISYTSSLSSGK 289
Query: 133 HRCG--RKGRSWSYR----SKCKSYNLHVACVREMVVENWDHVYVAG 173
CG R+ Y +KC Y +H C + WD + + G
Sbjct: 290 WSCGVCRQEVDNDYGAYSCNKCDDYFVHSRCALRRDI--WDGIELEG 334
Score = 30.0 bits (66), Expect = 1.2
Identities = 48/170 (28%), Positives = 61/170 (35%), Gaps = 27/170 (15%)
Query: 6 ISHFSHPQHKLRFEHNAFPFK--CDGCK-EIGIGSRYK-CSICDYDLHMQCAIPSPTLVH 61
I HFSH H A+ C C I G Y CD+ LH CA H
Sbjct: 354 ILHFSHGHHLKLDTSKAYDENKLCQACTLPIYEGGYYSGVDECDFILHEACANAPCKKYH 413
Query: 62 PF--YTKCSFQFMSSPPGNIPRY-CNACEKDVNGFVY---------HCKSCGFDLHPCCA 109
Y + N R+ C+AC+++ GFVY K F + CA
Sbjct: 414 ALNPYPLTLKVVTNEYHDNKGRFRCDACQRESCGFVYVDDFRGCYADTKDYKFKIDIRCA 473
Query: 110 KLPMVLD--DGEVKLYL----YRKVSSPCHRCGR-KGRSWSYRSKCKSYN 152
+ D E LYL + S+ CH C K S+S CK N
Sbjct: 474 SVSEPFDYLGHEHPLYLALNPEEEESAICHICQESKDESFS----CKKLN 519
Score = 27.3 bits (59), Expect = 7.5
Identities = 18/67 (26%), Positives = 27/67 (39%), Gaps = 8/67 (11%)
Query: 7 SHFSHPQHKLRF----EHNAFPFKCDGCKEIGIGSR----YKCSICDYDLHMQCAIPSPT 58
+ + H +H L+F E N CD C+ R Y C C LH+ C +
Sbjct: 538 ARYQHDKHFLKFYEAKEANDHSEWCDVCERRIADLRKKGFYSCDDCCTTLHIDCLLGEDM 597
Query: 59 LVHPFYT 65
+ P +T
Sbjct: 598 YMKPGHT 604
>At4g02190 CHP-rich hypothetical protein similar to T10M13.17.1
Length = 659
Score = 68.2 bits (165), Expect = 4e-12
Identities = 55/183 (30%), Positives = 73/183 (39%), Gaps = 26/183 (14%)
Query: 12 PQHKLRFEHNAFPFKCDG-CKEIGIGSRYKCSICDYDLHMQCAI-PS-----PTLVHPFY 64
P HK F CD C+EIG G YKC C H+ C + PS P ++HP
Sbjct: 118 PCHKRTLLKEKIEFHCDAKCEEIGYGFPYKCLECGLTFHIDCVLHPSGVKHLPEVIHPLD 177
Query: 65 TKCSFQFMSSP-----PGNIPRY----CNACEKDVNG-FVYHCKSCGFDLHPCCAKLPMV 114
K P G +P Y C CEK ++ YHC C F L CA P
Sbjct: 178 VKNHSCHPLHPLKLVLTGQLPDYSDGKCRLCEKKIDSPLFYHCSPCNFTLDMRCALNPPS 237
Query: 115 LDDGEVKLY------LYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV---REMVVEN 165
+ + K + L R S C+ CG KG Y ++ +H C+ R + +
Sbjct: 238 ISFEDSKTHDHQLTLLPRLDSFTCNACGLKGDRSPYICVQCNFIIHQECLTLPRLININR 297
Query: 166 WDH 168
DH
Sbjct: 298 HDH 300
Score = 55.1 bits (131), Expect = 3e-08
Identities = 44/157 (28%), Positives = 66/157 (42%), Gaps = 18/157 (11%)
Query: 15 KLRFEHNAFPF----KCDGCKEI-GIGSRYKCSICDYDLHMQCAIP-SPTLVHPFYTKCS 68
+L EH+ P K D C I Y C CD+ +H +C S ++ HP + +
Sbjct: 9 RLIHEHSMIPCNHLRKGDCCGRFEAISDGYYCKTCDFFVHKKCVDEASESIEHPSHPGHT 68
Query: 69 FQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKL--PMVLDDGEV----KL 122
Q +S RYCN C +D+ YH + DL+ CAK P V++ E +
Sbjct: 69 LQLLSKQKY---RYCNLCGRDIKDLCYHFGNFDVDLY--CAKYPPPEVIESSETPCHKRT 123
Query: 123 YLYRKVSSPCH-RCGRKGRSWSYRSKCKSYNLHVACV 158
L K+ C +C G + Y+ H+ CV
Sbjct: 124 LLKEKIEFHCDAKCEEIGYGFPYKCLECGLTFHIDCV 160
Score = 48.9 bits (115), Expect = 2e-06
Identities = 36/141 (25%), Positives = 54/141 (37%), Gaps = 10/141 (7%)
Query: 6 ISHFSHPQHKLRFEHNAFPF----KCDGCKE-IGIGSRYKCSICDYDLHMQCAIPSPTLV 60
I HFSH +H L+ + N +C C + + S Y C CD+ LH CA
Sbjct: 376 IQHFSHEEHYLKLDDNGVLCDENKRCSACTHSVCLESFYGCMDCDFILHQNCAKFPKRKR 435
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMV--LDDG 118
H + + + G+ +CN C + NGF Y D+ C P V
Sbjct: 436 HVLHNE-RLTLFTREAGHF--WCNVCGRISNGFSYQYGDMKLDVICCSVLEPFVHPSHPD 492
Query: 119 EVKLYLYRKVSSPCHRCGRKG 139
Y+ ++ C+ C G
Sbjct: 493 HPLYYISPEMEEVCNGCNMSG 513
Score = 36.6 bits (83), Expect = 0.012
Identities = 24/69 (34%), Positives = 31/69 (44%), Gaps = 7/69 (10%)
Query: 83 CNACEKDV--NGFVYHCKSCGFDLHPCCAKLPM----VLDDGEVKLYLYRKVSSPCHRCG 136
C+AC V F Y C C F LH CAK P VL + + L+ C+ CG
Sbjct: 401 CSACTHSVCLESF-YGCMDCDFILHQNCAKFPKRKRHVLHNERLTLFTREAGHFWCNVCG 459
Query: 137 RKGRSWSYR 145
R +SY+
Sbjct: 460 RISNGFSYQ 468
>At2g28270 pseudogene
Length = 248
Score = 67.4 bits (163), Expect = 7e-12
Identities = 52/191 (27%), Positives = 75/191 (39%), Gaps = 24/191 (12%)
Query: 27 CDGCKEIGIGSRYKC--SICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSPPGNIPRY-C 83
C GC+ IG +KC S CDY LH C + H +T + SPP + Y C
Sbjct: 33 CSGCEHDLIGQAFKCTKSECDYFLHKSCFDLPGEIHHKSHTNHPLTLLHSPPNGLSTYTC 92
Query: 84 NACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKLYLYRKVSSPCHRCGRKG---- 139
+AC + + F YHC C + +H CA +P + + + L ++PC GRK
Sbjct: 93 DACGEYGSAFTYHCSECKYHVHVGCAFVPENVKREDHEHPLTLLYNTPCK--GRKDGVVF 150
Query: 140 -----------RSWSYRSKCKSYNLHVACVREMVVENWDHVYVAGGKGRNKVVETSVPGL 188
W Y K Y HV N D+ GG+ + S +
Sbjct: 151 ICDVCEVDVSENLWVYYCKECDYGTHV----HSCTTNEDNGPKKGGEEEGESSSLSTSRI 206
Query: 189 KNTLYAAHNSR 199
K+ + A R
Sbjct: 207 KSLMEAEKEMR 217
Score = 57.4 bits (137), Expect = 7e-09
Identities = 37/116 (31%), Positives = 48/116 (40%), Gaps = 20/116 (17%)
Query: 5 EISHFSHPQHKLRFEHNA----FPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL- 59
EI H SH H L H+ + CD C E G Y CS C Y +H+ CA +
Sbjct: 66 EIHHKSHTNHPLTLLHSPPNGLSTYTCDACGEYGSAFTYHCSECKYHVHVGCAFVPENVK 125
Query: 60 ----VHP----FYTKCSFQFMSSPPGNIPRYCNACEKDV--NGFVYHCKSCGFDLH 105
HP + T C + C+ CE DV N +VY+CK C + H
Sbjct: 126 REDHEHPLTLLYNTPC-----KGRKDGVVFICDVCEVDVSENLWVYYCKECDYGTH 176
>At4g01930 putative CHP-rich zinc finger protein
Length = 652
Score = 67.0 bits (162), Expect = 9e-12
Identities = 46/152 (30%), Positives = 69/152 (45%), Gaps = 15/152 (9%)
Query: 19 EHNAFPF----KCDGCKEI-GIGSRYKCSICDYDLHMQCAIPSPTLV-HPFYTKCSFQFM 72
+H+ P+ K D C+ I Y C CD+ +H C S + HP + + Q +
Sbjct: 13 KHHMLPWNDMRKGDCCRRFEAISDGYYCKTCDFFVHKSCGDRSSEYIEHPSHPSHALQLL 72
Query: 73 SSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKL--PMVLDDGEV---KLYLYRK 127
P R CN C + Y+C+ C F++H CAK P V+D E KL LY+K
Sbjct: 73 RKPGHT--RNCNLCGISIKSLFYNCEICNFNVHMYCAKYPPPQVIDISETHHHKLNLYKK 130
Query: 128 VS-SPC-HRCGRKGRSWSYRSKCKSYNLHVAC 157
+ C +CG+ +SY+ + HV C
Sbjct: 131 PNYIDCGDKCGKLSYEFSYKCQECDLAFHVDC 162
Score = 57.0 bits (136), Expect = 9e-09
Identities = 51/177 (28%), Positives = 73/177 (40%), Gaps = 20/177 (11%)
Query: 5 EISHFSHPQHKLRFEHNAFPFKCDG-CKEIG--IGSR--YKCSICDYDLHMQCAI--PSP 57
E++HF H H L+ P D C+ G IG R Y CS C++ L M+C + P
Sbjct: 174 EVNHFYHSLHPLKLVRGKLPDYSDRKCRLCGRYIGDRLFYHCSSCNFTLDMRCVLNPPQQ 233
Query: 58 TLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVL-- 115
+L++ K ++ P CNAC + Y C CGF +H C LP V+
Sbjct: 234 SLLN---LKAHDHQLTLLPRLDSFTCNACGLKGDRSPYVCFQCGFMIHQDCLDLPRVINI 290
Query: 116 ---DDGEVKLYLYRKVSSPCHRCGRKGRSWSYR----SKCKSYNLHVACVREMVVEN 165
D + + V S C C +K W+ +C Y +H C V N
Sbjct: 291 NRHDHRVSRTSVLGVVKSVCGVCHQK-VDWTCGGFSCQRCHGYVVHSKCATRKDVWN 346
Score = 52.4 bits (124), Expect = 2e-07
Identities = 54/185 (29%), Positives = 73/185 (39%), Gaps = 26/185 (14%)
Query: 5 EISHFSHPQHKLRFEHNAFPFKC-DGCKEIGIGSRYKCSICDYDLHMQCA-IPS----PT 58
+IS H HKL C D C ++ YKC CD H+ CA PS P
Sbjct: 116 DISETHH--HKLNLYKKPNYIDCGDKCGKLSYEFSYKCQECDLAFHVDCAWRPSESNLPL 173
Query: 59 LVHPFYTKCSFQFMSSPPGNIPRY----CNACEKDV-NGFVYHCKSCGFDLHPCCAKLP- 112
V+ FY S + G +P Y C C + + + YHC SC F L C P
Sbjct: 174 EVNHFYH--SLHPLKLVRGKLPDYSDRKCRLCGRYIGDRLFYHCSSCNFTLDMRCVLNPP 231
Query: 113 ------MVLDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV---REMVV 163
+ D ++ L L R S C+ CG KG Y + +H C+ R + +
Sbjct: 232 QQSLLNLKAHDHQLTL-LPRLDSFTCNACGLKGDRSPYVCFQCGFMIHQDCLDLPRVINI 290
Query: 164 ENWDH 168
DH
Sbjct: 291 NRHDH 295
Score = 48.9 bits (115), Expect = 2e-06
Identities = 41/134 (30%), Positives = 54/134 (39%), Gaps = 20/134 (14%)
Query: 6 ISHFSHPQHKLRFEHNAFPFK----CDGCKE-IGIGSRYKCSICDYDLHMQCAIPSPTLV 60
I HFSH +H LR N C C I + S Y C CD+ LH CA
Sbjct: 371 IQHFSHKEHYLRLHVNGVLCDDNKWCSACTHPICLQSFYGCMDCDFILHQNCAGFRRMKW 430
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFD----------LHPCCAK 110
H + + ++ P C AC++ NGF YH + FD LHP
Sbjct: 431 HVLHNERLTLVINEAE---PFRCYACDRWSNGFRYHQGNITFDVRCGSIAEPFLHPSHPD 487
Query: 111 LPM--VLDDGEVKL 122
P+ L DG +K+
Sbjct: 488 HPLYHTLPDGGIKI 501
Score = 45.4 bits (106), Expect = 3e-05
Identities = 45/163 (27%), Positives = 66/163 (39%), Gaps = 19/163 (11%)
Query: 8 HFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKC 67
H H + + A PF+C C G RY +D+ +C + +HP +
Sbjct: 431 HVLHNERLTLVINEAEPFRCYACDRWSNGFRYHQGNITFDV--RCGSIAEPFLHPSHPDH 488
Query: 68 SFQFMSSPPGNIPRYCNACEKDVNGF--VYHC--KSCGFDLHPCCAKLPMVL----DDGE 119
+ + P G I + CN C+ NGF V C C F + CA P V+ DD
Sbjct: 489 PL-YHTLPDGGI-KICNGCK---NGFYDVLGCIEDDCRFAICFKCATFPQVVKHRVDDHP 543
Query: 120 VKLYLYRKVSSP--CHRCGRKGRSWSYRSKCKSY--NLHVACV 158
+ L K S C C ++ + CK Y +LH+ CV
Sbjct: 544 LSLCYGEKASGKYWCDICEKETNPEIWFYTCKDYQASLHIECV 586
Score = 31.6 bits (70), Expect = 0.40
Identities = 19/71 (26%), Positives = 31/71 (42%), Gaps = 6/71 (8%)
Query: 95 YHCKSCGFDLHPCCAKLPMVLDDGEVK----LYLYRKVSSP--CHRCGRKGRSWSYRSKC 148
Y+CK+C F +H C + L L RK C+ CG +S Y +
Sbjct: 38 YYCKTCDFFVHKSCGDRSSEYIEHPSHPSHALQLLRKPGHTRNCNLCGISIKSLFYNCEI 97
Query: 149 KSYNLHVACVR 159
++N+H+ C +
Sbjct: 98 CNFNVHMYCAK 108
>At2g42060 unknown protein
Length = 268
Score = 66.2 bits (160), Expect = 1e-11
Identities = 57/185 (30%), Positives = 74/185 (39%), Gaps = 36/185 (19%)
Query: 5 EISHFSHPQHKLRFEHNAFPFK-------CDGCKEIGIGSR--YKCSICDYDLHMQCAIP 55
E HFSHP H L+ + P K C GC+ S Y CS CD++LH QC
Sbjct: 2 EYKHFSHP-HTLKLQQIQ-PHKSSDSSVICSGCESAISESETAYICSTCDFNLHEQCGNA 59
Query: 56 SPTLVHPFYTKCSF----QFMSSPPGNIPRYCNACE-KDVNGFVYHCKSCGFDLHPCCAK 110
+ HP + + + G C AC GF Y C C FDLH CA
Sbjct: 60 VRGMQHPSHAGLHHLTLVPYTTYSAGTF--LCRACGCTGGKGFSYCCPLCDFDLHVQCAH 117
Query: 111 LPMVL--DDGEVKLYLYRKVSSP--------------CHRCG--RKGRSWSYRSKCKSYN 152
LP VL + + L S+P C+ C GR WSY +Y+
Sbjct: 118 LPQVLVHESHPMHSLLLVYNSTPPMSFTQFGFGNQLVCNLCNMTMDGRFWSYNCYACNYH 177
Query: 153 LHVAC 157
+H +C
Sbjct: 178 IHASC 182
Score = 57.0 bits (136), Expect = 9e-09
Identities = 33/102 (32%), Positives = 51/102 (49%), Gaps = 12/102 (11%)
Query: 20 HNAFPFKCDGCKEIG-IGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKCS--FQFMSSPP 76
++A F C C G G Y C +CD+DLH+QCA LVH + S + S+PP
Sbjct: 82 YSAGTFLCRACGCTGGKGFSYCCPLCDFDLHVQCAHLPQVLVHESHPMHSLLLVYNSTPP 141
Query: 77 GNIPRY-------CNACEKDVNG--FVYHCKSCGFDLHPCCA 109
+ ++ CN C ++G + Y+C +C + +H CA
Sbjct: 142 MSFTQFGFGNQLVCNLCNMTMDGRFWSYNCYACNYHIHASCA 183
Score = 35.0 bits (79), Expect = 0.036
Identities = 21/74 (28%), Positives = 27/74 (36%), Gaps = 14/74 (18%)
Query: 8 HFSHPQHKLRFEHNAFP------------FKCDGCKEIGIGS--RYKCSICDYDLHMQCA 53
H SHP H L +N+ P C+ C G Y C C+Y +H CA
Sbjct: 124 HESHPMHSLLLVYNSTPPMSFTQFGFGNQLVCNLCNMTMDGRFWSYNCYACNYHIHASCA 183
Query: 54 IPSPTLVHPFYTKC 67
+ P V C
Sbjct: 184 VNKPNPVAASAENC 197
>At5g59930 putative protein
Length = 656
Score = 65.9 bits (159), Expect = 2e-11
Identities = 46/156 (29%), Positives = 65/156 (41%), Gaps = 10/156 (6%)
Query: 11 HPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQ 70
HP L+ P C+ CK +GS Y C C+ H+ C S + HP ++
Sbjct: 125 HPLVFLKKRQEKTP--CEVCKNRILGSSYSCLGCELYFHVGCIHLSKEVNHPCHSDHPLM 182
Query: 71 FMSSPP--GNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKLYLY--- 125
+ S N + C C + +G +YHC C F L C K P L VK + +
Sbjct: 183 LVESESLTDNDKKTCFLCGQQPHGILYHCSICNFTLCIGCIKSPPPLVVENVKTHKHPLT 242
Query: 126 ---RKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
R V C+ CG KG SY + +H +CV
Sbjct: 243 FFPRGVYCTCNVCGVKGLILSYMCLQCGFAVHCSCV 278
Score = 60.1 bits (144), Expect = 1e-09
Identities = 47/182 (25%), Positives = 69/182 (37%), Gaps = 25/182 (13%)
Query: 4 SEISHFSHPQHK--LRFEHNAFPFKCDGCKEIGIGSR-YKCSICDYDLHMQCAI--PSPT 58
SEI+H HPQH L +E + C C E Y CS C + + C + P P
Sbjct: 57 SEINHLFHPQHPLLLTYEEGWWSCFCALCGETKPAKYFYSCSTCKIKVDLTCGMNPPPPA 116
Query: 59 LVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDL------------HP 106
+ HP F+ P C C+ + G Y C C HP
Sbjct: 117 IEHPICHDHPLVFLKKRQEKTP--CEVCKNRILGSSYSCLGCELYFHVGCIHLSKEVNHP 174
Query: 107 CCAKLPMVLDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACVRE---MVV 163
C + P++L + E L C CG++ Y ++ L + C++ +VV
Sbjct: 175 CHSDHPLMLVESE---SLTDNDKKTCFLCGQQPHGILYHCSICNFTLCIGCIKSPPPLVV 231
Query: 164 EN 165
EN
Sbjct: 232 EN 233
Score = 50.1 bits (118), Expect = 1e-06
Identities = 60/240 (25%), Positives = 93/240 (38%), Gaps = 34/240 (14%)
Query: 5 EISHFSHPQHKLRFEHNAF-----PFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL 59
E++H H H L + C C + G Y CSIC++ L + C P L
Sbjct: 170 EVNHPCHSDHPLMLVESESLTDNDKKTCFLCGQQPHGILYHCSICNFTLCIGCIKSPPPL 229
Query: 60 VHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFV--YHCKSCGFDLHPCCAKLPMVLDD 117
V K ++ P + CN C V G + Y C CGF +H C LP V++
Sbjct: 230 VVE-NVKTHKHPLTFFPRGVYCTCNVC--GVKGLILSYMCLQCGFAVHCSCVDLPQVINI 286
Query: 118 G--EVKLYLYRKVSSPCHRCGRKGRSWSYRS------KCKSYNLHVACVREMVVENWDHV 169
+ ++ + + CG RS + C +Y +H C + WD V
Sbjct: 287 NRHDHRISFTQHLGPGYLNCGVCRRSVCQFNGAYSCLVCPNYAVHSRCATRYDI--WDGV 344
Query: 170 YVAGGKGRN-------KVVETSVPGLKNTLYAAHNSRRSKGKV----KKCCEIAGLAVQF 218
+ GK N KVV + +++ ++ H+ R K + CE G V+F
Sbjct: 345 ELE-GKAENIEDIAPFKVVGDDL--IRHFSHSKHSLRLHKNNIIYDDSTRCEACGHPVEF 401
Score = 46.2 bits (108), Expect = 2e-05
Identities = 44/165 (26%), Positives = 63/165 (37%), Gaps = 17/165 (10%)
Query: 6 ISHFSHPQHKLRFEHNAFPF----KCDGCKE-IGIGSRYKCSICDYDLHMQCA-IPSPTL 59
I HFSH +H LR N + +C+ C + G Y C C + LH +CA +P
Sbjct: 367 IRHFSHSKHSLRLHKNNIIYDDSTRCEACGHPVEFGRIYNCEECCFILHEKCANLPMKKR 426
Query: 60 VHPFYTKCSFQFMSSPPGN---IPRYCNACEKDVNGFVYHC---KSCGFDLHPCCAKLPM 113
+ + F+ PGN C C GF Y + D+H P
Sbjct: 427 L----VFSTLPFLLLGPGNDVCKVLDCKLCGMFFTGFEYKSQERRQSDIDVHCGSLTEPF 482
Query: 114 VLDDGEVKLYLYRKVSS-PCHRCGRKGRSWSYRSKCKSYNLHVAC 157
V D LY +K PC+ C + R + +NL + C
Sbjct: 483 VHDGHLHPLYFDQKQKELPCNACQKTIRDYMLCCADCDFNLCLWC 527
Score = 36.2 bits (82), Expect = 0.016
Identities = 24/97 (24%), Positives = 38/97 (38%), Gaps = 3/97 (3%)
Query: 14 HKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMS 73
H L F+ C+ C++ C+ CD++L + CA P + +
Sbjct: 489 HPLYFDQKQKELPCNACQKTIRDYMLCCADCDFNLCLWCA-SLPKKIRHSNDEHILTLCC 547
Query: 74 SPPGNIPRYCNACEK--DVNGFVYHCKSCGFDLHPCC 108
N +C+ CE D + + Y C CG LH C
Sbjct: 548 GEEANGKYWCDICEAELDPSKWFYTCFGCGGTLHVEC 584
>At5g03365 putative protein
Length = 1022
Score = 65.5 bits (158), Expect = 3e-11
Identities = 44/131 (33%), Positives = 57/131 (42%), Gaps = 9/131 (6%)
Query: 6 ISHFSHPQHKLRFEHNAFPFK-----CDGC-KEIGIGSRYKCSICDYDLHMQCAIPSPTL 59
I HFSHPQH L+ + N C C I G+ Y C CDY LH +CA +
Sbjct: 731 IQHFSHPQHHLKLDENTSKDYDEDKLCQACIMPIYFGNFYSCVQCDYILHDECASFPRKM 790
Query: 60 VHPFYTKCSFQFMSSPPGNIPRYCNACE-KDVNGFVYHCKSCGFDLHPCCAKL--PMVLD 116
HP + S N+ + C+AC GF Y C F LH CA + P+V +
Sbjct: 791 HHPIHPHLLTLVNSDGVINVRKKCSACPWMCTTGFFYECSKEDFQLHVQCANVSEPLVHE 850
Query: 117 DGEVKLYLYRK 127
L+L K
Sbjct: 851 SHMHPLFLTTK 861
Score = 48.9 bits (115), Expect = 2e-06
Identities = 23/80 (28%), Positives = 40/80 (49%), Gaps = 1/80 (1%)
Query: 37 SRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSPPGNI-PRYCNACEKDVNGFVY 95
+ ++C C+ + H + HP + K S Q +SS P ++ R C C++D+ Y
Sbjct: 508 THFRCRGCNGENHKGYEQAPVDVKHPLHPKHSLQLVSSQPSSVRTRECYCCDEDLAHIFY 567
Query: 96 HCKSCGFDLHPCCAKLPMVL 115
C +C + ++ C K P VL
Sbjct: 568 CCLACNYAMNVACVKKPSVL 587
Score = 44.7 bits (104), Expect = 5e-05
Identities = 46/189 (24%), Positives = 76/189 (39%), Gaps = 27/189 (14%)
Query: 5 EISHFSHPQHKLRF----EHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLV 60
++ H HP+H L+ + +C C E Y C C+Y +++ C + P+++
Sbjct: 529 DVKHPLHPKHSLQLVSSQPSSVRTRECYCCDEDLAHIFYCCLACNYAMNVAC-VKKPSVL 587
Query: 61 HPFYTKCSFQFMSSPPG--NIPRYCNACE-KDVNGFVYHCKSCGFDLHPCCAKLPMVLDD 117
+ YT+ ++ P CN C D + +Y C C F +H C LP V+
Sbjct: 588 YRDYTRWHEHKLALFPRLQASSLTCNLCALADSSSPIYMCPPCDFVVHQRCTGLPRVI-- 645
Query: 118 GEVKLYLYR--------KVSSPCHRCGRK-----GRSWSYRSKCKSYNLHVACVREMVVE 164
+ +L+R + C C RK G + C SY H C + V
Sbjct: 646 -RISRHLHRISFTYSFDQGDLCCSVCRRKIDNDYGGYSCIKDGC-SYAAHSRCATQRNV- 702
Query: 165 NWDHVYVAG 173
WD + + G
Sbjct: 703 -WDGIELEG 710
Score = 43.9 bits (102), Expect = 8e-05
Identities = 44/147 (29%), Positives = 61/147 (40%), Gaps = 17/147 (11%)
Query: 26 KCDGCKEI-GIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSPPGNIPRYCN 84
KC C + G Y+CS D+ LH+QCA S LVH + F+++ P R C+
Sbjct: 813 KCSACPWMCTTGFFYECSKEDFQLHVQCANVSEPLVHESHMHP--LFLTTKPEEW-RVCS 869
Query: 85 ACEKD---VNGFVYHCKSCGFDLHPCCAKLPMVL----DDGEVKLYLYRKVS---SPCHR 134
AC++ V ++ C F L CA LP L D + L + S S C
Sbjct: 870 ACKESDCTVTNETFNSIECDFALCFRCATLPQKLRYKHDKHTLTLSYGEETSTMPSWCEI 929
Query: 135 CGRKGRSWSYRSKCKSY---NLHVACV 158
C R C Y LH+ C+
Sbjct: 930 CERNINPQERFYMCDEYCCVTLHIECM 956
>At5g43520 putative protein
Length = 250
Score = 65.1 bits (157), Expect = 3e-11
Identities = 42/122 (34%), Positives = 61/122 (49%), Gaps = 10/122 (8%)
Query: 27 CDGCKEIGIGSRYKC--SICDYDLHMQCA-IPSPTLVHPFYTKCSFQFMSSPPGNIPRY- 82
C GC+ G +KC S CDY LH C +PS T + + + SPP + Y
Sbjct: 38 CSGCELELTGQAFKCIKSECDYFLHKSCFDLPSETN-NKSHQDHPLTLLHSPPDDRSVYT 96
Query: 83 CNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKLYLYRKVSSPCHRCGRKGRSW 142
C+AC++ +GF YHC +C + LH CA +P +D + + L +PC KGR
Sbjct: 97 CDACDQYGSGFSYHCSNCNYSLHVGCAFIPETVDREDHEHPLTLLYCTPC-----KGRED 151
Query: 143 SY 144
+Y
Sbjct: 152 TY 153
Score = 58.9 bits (141), Expect = 2e-09
Identities = 37/117 (31%), Positives = 55/117 (46%), Gaps = 11/117 (9%)
Query: 4 SEISHFSHPQHKLRFEHNA----FPFKCDGCKEIGIGSRYKCSICDYDLHMQCA-IPSPT 58
SE ++ SH H L H+ + CD C + G G Y CS C+Y LH+ CA IP
Sbjct: 70 SETNNKSHQDHPLTLLHSPPDDRSVYTCDACDQYGSGFSYHCSNCNYSLHVGCAFIPETV 129
Query: 59 LVHPFYTKCSFQFMSSPPGNIPRY--CNACEKDVNG--FVYHCKSC--GFDLHPCCA 109
+ + + G Y C+AC++ ++ ++Y+CK C G LH C A
Sbjct: 130 DREDHEHPLTLLYCTPCKGREDTYFTCSACDETISEDLWMYYCKDCDYGTHLHSCAA 186
Score = 41.6 bits (96), Expect = 4e-04
Identities = 23/85 (27%), Positives = 37/85 (43%), Gaps = 10/85 (11%)
Query: 83 CNACEKDVNGFVYHC--KSCGFDLHPCCAKLPMVLDDGEVKLYLYRKVSSP--------C 132
C+ CE ++ G + C C + LH C LP ++ + + + SP C
Sbjct: 38 CSGCELELTGQAFKCIKSECDYFLHKSCFDLPSETNNKSHQDHPLTLLHSPPDDRSVYTC 97
Query: 133 HRCGRKGRSWSYRSKCKSYNLHVAC 157
C + G +SY +Y+LHV C
Sbjct: 98 DACDQYGSGFSYHCSNCNYSLHVGC 122
>At4g26380 hypothetical protein
Length = 1016
Score = 65.1 bits (157), Expect = 3e-11
Identities = 45/131 (34%), Positives = 56/131 (42%), Gaps = 9/131 (6%)
Query: 6 ISHFSHPQHKLRFEHNAF-----PFKCDGC-KEIGIGSRYKCSICDYDLHMQCAIPSPTL 59
I HF HPQH LR + N C C I G+ Y C+ C+Y LH CA S +
Sbjct: 740 IQHFGHPQHHLRLDENTSRDYDEDKLCQACVMPIYFGNFYSCTQCNYILHEACANFSRKM 799
Query: 60 VHPFYTKCSFQFMSSPPGNIPRYCNACE-KDVNGFVYHCKSCGFDLHPCCAKL--PMVLD 116
HP + S N+ + C AC GF Y C F LH CA L P+V +
Sbjct: 800 HHPVHPHVLTLVNSDGAINVRKECAACPWMCTTGFFYECSKEDFQLHVQCANLSEPLVHE 859
Query: 117 DGEVKLYLYRK 127
L+L K
Sbjct: 860 SHMHPLFLTSK 870
Score = 51.2 bits (121), Expect = 5e-07
Identities = 48/156 (30%), Positives = 67/156 (42%), Gaps = 22/156 (14%)
Query: 16 LRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSP 75
+R E A P+ C G Y+CS D+ LH+QCA S LVH + F++S
Sbjct: 819 VRKECAACPWMCT------TGFFYECSKEDFQLHVQCANLSEPLVHESHMHP--LFLTSK 870
Query: 76 PGNIPRYCNACEK---DVNGFVYHCKSCGFDLHPCCAKLPMVL---DDGEVKLYLYRKVS 129
PG R C+AC++ V ++C C F L CA LP + D + Y + +
Sbjct: 871 PGE-RRLCSACKESYFSVTRETFNCIECDFALCFRCATLPQKVRYKHDKHMLTLSYGEET 929
Query: 130 SP----CHRCGRKGRSWSYRSKCKSY---NLHVACV 158
S C C R KC Y LH+ C+
Sbjct: 930 STMTYWCEVCERNLNPKERFYKCDEYCSITLHIECM 965
Score = 47.8 bits (112), Expect = 5e-06
Identities = 44/187 (23%), Positives = 74/187 (39%), Gaps = 25/187 (13%)
Query: 5 EISHFSHPQHKLRF----EHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLV 60
E+ H HP+H L+ +C C E + Y C C++ +++ C P ++
Sbjct: 540 EVKHPLHPKHPLQLVLSQSSGGRTRECYCCDEDLLHIFYCCLACNFSMNVACT-KKPAVL 598
Query: 61 HPFYTKCSFQFMSSPPGNIPRYCNACE-KDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGE 119
+ + + ++ P CN C D + +Y C C F +H C LP V+ +
Sbjct: 599 YRNHPRWHEHTLALFPRQASLTCNLCALADSSSPIYMCPPCDFVVHLRCTGLPRVI---K 655
Query: 120 VKLYLYR--------KVSSPCHRCGRK-----GRSWSYRSKCKSYNLHVACVREMVVENW 166
+ +L+R + C C RK G + C SY H C + V W
Sbjct: 656 ISRHLHRISFTFSFDRGDLSCSVCRRKIDEDYGGYTCIKDGC-SYAAHSKCATQKNV--W 712
Query: 167 DHVYVAG 173
D + + G
Sbjct: 713 DGIELEG 719
Score = 46.6 bits (109), Expect = 1e-05
Identities = 27/106 (25%), Positives = 46/106 (42%), Gaps = 7/106 (6%)
Query: 37 SRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQF-MSSPPGNIPRYCNACEKDVNGFVY 95
+ ++C C+ D H + + HP + K Q +S G R C C++D+ Y
Sbjct: 519 THFRCRGCNGDNHKENEKAPLEVKHPLHPKHPLQLVLSQSSGGRTRECYCCDEDLLHIFY 578
Query: 96 HCKSCGFDLHPCCAKLPMVLDDGEVKLYLY------RKVSSPCHRC 135
C +C F ++ C K P VL + + + R+ S C+ C
Sbjct: 579 CCLACNFSMNVACTKKPAVLYRNHPRWHEHTLALFPRQASLTCNLC 624
>At2g44380 unknown protein
Length = 247
Score = 64.7 bits (156), Expect = 4e-11
Identities = 60/215 (27%), Positives = 84/215 (38%), Gaps = 34/215 (15%)
Query: 6 ISHFSHPQHKLRF--EHNAFPFKCDGCKEIGIGSRYKC--SICDYDLHMQCAIPSPTLVH 61
+ H SH H LR + C GC+ G +KC S CDY LH C H
Sbjct: 12 VRHASH-NHPLRVFKARDEDEVVCSGCELELTGQAFKCMKSDCDYFLHKSCFDLPRETNH 70
Query: 62 PFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVK 121
+ S + SPP C+AC + +GF Y+C C +D+H CA +P ++ + +
Sbjct: 71 KSHPNHSLTLLHSPPYGQSYTCDACGEYGSGFTYNCSECQYDVHVGCAFIPETVEREDHE 130
Query: 122 LYLYRKVSSPCHRCGRKGRS------------------WSYRSKCKSYNLHV-ACVREMV 162
L ++PC KGR W Y K Y HV +C V
Sbjct: 131 HPLTLLYNTPC-----KGREDGAKFICDVCEEKMSENLWVYYCKECDYGTHVHSCA---V 182
Query: 163 VENWDHVYVAGGKGRNKVVETSVPGLKNTLYAAHN 197
E DH G GR + +S +L A +
Sbjct: 183 YE--DHESEKRGGGREEGEASSAVSRMKSLMKAQD 215
Score = 63.5 bits (153), Expect = 1e-10
Identities = 38/115 (33%), Positives = 51/115 (44%), Gaps = 19/115 (16%)
Query: 5 EISHFSHPQHKLRFEHN---AFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL-- 59
E +H SHP H L H+ + CD C E G G Y CS C YD+H+ CA T+
Sbjct: 67 ETNHKSHPNHSLTLLHSPPYGQSYTCDACGEYGSGFTYNCSECQYDVHVGCAFIPETVER 126
Query: 60 ---VHP----FYTKCSFQFMSSPPGNIPRYCNACEKDV--NGFVYHCKSCGFDLH 105
HP + T C C+ CE+ + N +VY+CK C + H
Sbjct: 127 EDHEHPLTLLYNTPC-----KGREDGAKFICDVCEEKMSENLWVYYCKECDYGTH 176
>At3g48400 putative protein
Length = 669
Score = 63.9 bits (154), Expect = 7e-11
Identities = 51/182 (28%), Positives = 78/182 (42%), Gaps = 16/182 (8%)
Query: 5 EISHFSHPQHKLR-FEHNAFPFK----CDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL 59
E+ H SH H L F + P C C ++ Y CS+C++ + + CAI P L
Sbjct: 172 EVHHSSHTNHTLELFGSESLPDSAQKTCLLCDDVSDHLIYHCSVCNFSICVYCAINPPPL 231
Query: 60 VHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLD--- 116
+ K ++ P + CNAC +G Y C CG+ +H C LP V++
Sbjct: 232 AIE-HLKTHEHTLTLLPRQVKFICNACGMKGDGCPYFCLGCGYLIHRKCIDLPRVININR 290
Query: 117 -DGEVKLYLYRKVS-SPCHRCGR--KGRSWSYR-SKCKSYNLHVACVREMVVENWDHVYV 171
D + L + + S C C + G Y S C +Y +H C M + WD + +
Sbjct: 291 HDHRISLTHHLGLQYSECGVCRQSVNGYYGGYSCSLCPNYVVHPQCA--MKDDVWDGIEL 348
Query: 172 AG 173
G
Sbjct: 349 EG 350
Score = 58.9 bits (141), Expect = 2e-09
Identities = 42/154 (27%), Positives = 63/154 (40%), Gaps = 12/154 (7%)
Query: 27 CDGCKEIGIGSR-YKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSP--PGNIPRYC 83
C CK+ G Y C CD H++C S + H +T + + S P + + C
Sbjct: 140 CGVCKKTVFGMYPYVCLECDVYFHVECINISREVHHSSHTNHTLELFGSESLPDSAQKTC 199
Query: 84 NACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKLY------LYRKVSSPCHRCGR 137
C+ + +YHC C F + CA P L +K + L R+V C+ CG
Sbjct: 200 LLCDDVSDHLIYHCSVCNFSICVYCAINPPPLAIEHLKTHEHTLTLLPRQVKFICNACGM 259
Query: 138 KGRSWSYRSKCKSYNLHVACV---REMVVENWDH 168
KG Y Y +H C+ R + + DH
Sbjct: 260 KGDGCPYFCLGCGYLIHRKCIDLPRVININRHDH 293
Score = 55.1 bits (131), Expect = 3e-08
Identities = 41/142 (28%), Positives = 52/142 (35%), Gaps = 10/142 (7%)
Query: 6 ISHFSHPQHKLRFEHNAFPF------KCDGCK-EIGIGSRYKCSICDYDLHMQCAIPSPT 58
I HFSH +H L N +C C + Y C CDY LH CA
Sbjct: 370 IKHFSHEKHNLMLNKNGIITQHDEITRCQACVLPLSSDPSYSCGQCDYVLHETCANLPRK 429
Query: 59 LVHPFYTKCSFQFMSSPPGNIPRY--CNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLD 116
H F K F + + + C+AC NGF Y D+H P++ D
Sbjct: 430 KRHVFNNK-PFTLHTGGKDSSRNWFECDACRTISNGFRYVSGDWILDVHCGSMSEPLLHD 488
Query: 117 DGEVKLYLYRKVSSPCHRCGRK 138
LY YR+ C C K
Sbjct: 489 GHSHPLYYYRRQGKSCRSCHNK 510
Score = 43.5 bits (101), Expect = 1e-04
Identities = 39/146 (26%), Positives = 54/146 (36%), Gaps = 19/146 (13%)
Query: 25 FKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYTKCSFQFMSSPPGNIPRYCN 84
F+CD C+ I G RY D+ L + C S L+H ++ + + + C
Sbjct: 453 FECDACRTISNGFRYVSG--DWILDVHCGSMSEPLLHDGHSHPLYYYRRQG-----KSCR 505
Query: 85 ACE-KDVNGFVYHCKSCGFDLHPCCAKL-----------PMVLDDGEVKLYLYRKVSSPC 132
+C K V V C +C FDL CA L P+ L GE + K
Sbjct: 506 SCHNKVVKPRVLCCDTCDFDLDFYCANLPKTVKHCYDEHPLSLCHGEDDVIKDNKYWCDI 565
Query: 133 HRCGRKGRSWSYRSKCKSYNLHVACV 158
+ W Y LHV CV
Sbjct: 566 CETETDPKKWFYTCYDCGVTLHVKCV 591
Score = 41.2 bits (95), Expect = 5e-04
Identities = 30/108 (27%), Positives = 46/108 (41%), Gaps = 15/108 (13%)
Query: 12 PQHKLRFEHNA--FPFKCDGCKEIG-IGSRYKCSICDYD---LHMQCAIPSPTLVHPFYT 65
P HK H++ +C GCK+ I S Y+C+ D + H +C P + HP +
Sbjct: 8 PIHKHPLYHSSRFINGRCRGCKQTSPIYSGYRCNDSDCNHVWYHKECGESLPEINHPSHL 67
Query: 66 K---CSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAK 110
+ C + S C C + G YHC C F++ CA+
Sbjct: 68 EHPLCLIDYRGSGT------CAFCGAFLFGPTYHCWICAFNIDIACAR 109
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.138 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,118,228
Number of Sequences: 26719
Number of extensions: 280770
Number of successful extensions: 2283
Number of sequences better than 10.0: 173
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 789
Number of HSP's gapped (non-prelim): 801
length of query: 242
length of database: 11,318,596
effective HSP length: 96
effective length of query: 146
effective length of database: 8,753,572
effective search space: 1278021512
effective search space used: 1278021512
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0091a.6


