

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.5
(140 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
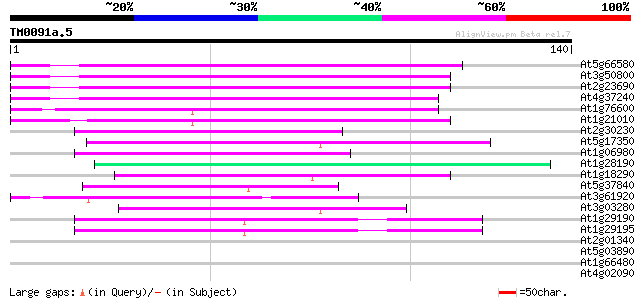
At5g66580 unknown protein 80 4e-16
At3g50800 unknown protein (At3g50800) 73 4e-14
At2g23690 unknown protein 72 1e-13
At4g37240 unknown protein 72 1e-13
At1g76600 unknown protein 64 2e-11
At1g21010 unknown protein 61 2e-10
At2g30230 hypothetical protein 47 3e-06
At5g17350 unknown protein 47 4e-06
At1g06980 unknown protein 46 6e-06
At1g28190 unknown protein 45 2e-05
At1g18290 hypothetical protein 44 3e-05
At5g37840 putative protein 42 1e-04
At3g61920 unknown protein 42 1e-04
At3g03280 unknown protein 41 2e-04
At1g29190 hypothetical protein 40 5e-04
At1g29195 unknown protein 40 5e-04
At2g01340 unknown protein 39 0.001
At5g03890 putative protein 37 0.003
At1g66480 hypothetical protein 36 0.006
At4g02090 unknown protein 33 0.049
>At5g66580 unknown protein
Length = 156
Score = 80.1 bits (196), Expect = 4e-16
Identities = 40/113 (35%), Positives = 62/113 (54%), Gaps = 7/113 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG CAS + + ++ DG LQ+ PVK W +L +NP F+C+S+ M
Sbjct: 1 MGACASRESL-------RSDSAKLILLDGTLQEFSSPVKVWQILQKNPTSFVCNSDEMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSV 113
++ V NEEL+ +YF++P + PL E++ ALA+KA++AL S V
Sbjct: 54 DDAVSAVAGNEELRSGQLYFVLPLTWLNHPLRAEEMAALAVKASSALTKSGGV 106
>At3g50800 unknown protein (At3g50800)
Length = 152
Score = 73.2 bits (178), Expect = 4e-14
Identities = 38/110 (34%), Positives = 61/110 (54%), Gaps = 7/110 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MG CAS + T ++ DG LQ+ PVK W +L +NP F+C+S+ M
Sbjct: 1 MGACASRESR-------RTETAKLILPDGTLQEFSTPVKVWQILQKNPTSFVCNSDDMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHS 110
+ V +E+L+ +YF++P + PL +++ ALA+KA++ALA S
Sbjct: 54 DDAVLAVPGSEDLRPGELYFVLPLTWLNHPLRADEMAALAVKASSALAKS 103
>At2g23690 unknown protein
Length = 278
Score = 72.0 bits (175), Expect = 1e-13
Identities = 37/110 (33%), Positives = 62/110 (55%), Gaps = 7/110 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MGIC+S + +T ++ DG++ + PVK +VL +NP F+C+S+ M
Sbjct: 1 MGICSSYEST-------QVATAKLILHDGRMMEFTSPVKVGYVLQKNPMCFICNSDDMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHS 110
+ ++ + +EE QL +YF +P S L E++ ALA+KA++AL S
Sbjct: 54 DNVVSAISADEEFQLGQLYFALPLSSLHHSLKAEEMAALAVKASSALMRS 103
>At4g37240 unknown protein
Length = 168
Score = 71.6 bits (174), Expect = 1e-13
Identities = 38/107 (35%), Positives = 61/107 (56%), Gaps = 7/107 (6%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYV 60
MGIC+S++ +T ++ DG++ + PVK +VL + P F+C+S+ M
Sbjct: 1 MGICSSSEST-------QVATAKLILQDGRMMEFANPVKVGYVLLKYPMCFICNSDDMDF 53
Query: 61 GSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
+ + +EELQL IYF +P R PL E++ ALA+KA++AL
Sbjct: 54 DDAVAAISADEELQLGQIYFALPLCWLRQPLKAEEMAALAVKASSAL 100
>At1g76600 unknown protein
Length = 216
Score = 64.3 bits (155), Expect = 2e-11
Identities = 40/117 (34%), Positives = 62/117 (52%), Gaps = 13/117 (11%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVL----------SQNPNH 50
MG+C S + +T IV +G L++ PV A VL S + ++
Sbjct: 1 MGLCVSVN---RNEYVSSSTTAKIVTINGDLREYDVPVLASQVLESESTSSSSSSSSSSY 57
Query: 51 FLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
FLC+S+S+Y + + +E LQ N IYF++P SK + LS D+ ALA+KA+ A+
Sbjct: 58 FLCNSDSLYYDDFIPAIESDEILQANQIYFVLPISKRQYRLSASDMAALAVKASVAI 114
>At1g21010 unknown protein
Length = 210
Score = 61.2 bits (147), Expect = 2e-10
Identities = 40/123 (32%), Positives = 63/123 (50%), Gaps = 17/123 (13%)
Query: 1 MGICASTQYAGKGGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVL-------------SQN 47
MGIC S + + TV IV +G L++ PV A VL S+
Sbjct: 1 MGICVSFRREDSNSS----PTVKIVTVNGDLREYNVPVIASQVLEAESAAAYSSSSSSRP 56
Query: 48 PNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAAL 107
++F+C S+S+Y + + E LQ + IYF++P SK + L+ D+ ALA+KA+ A+
Sbjct: 57 SSYFICDSDSLYYDDFIPAIKSEEPLQADQIYFVLPISKRQSRLTASDMAALAVKASVAI 116
Query: 108 AHS 110
+S
Sbjct: 117 QNS 119
>At2g30230 hypothetical protein
Length = 177
Score = 47.4 bits (111), Expect = 3e-06
Identities = 23/67 (34%), Positives = 34/67 (50%)
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLN 76
G + IVH +G + ++ P+ A +L NPNH L S V + + P EL+
Sbjct: 16 GALDLIRIVHLNGHVDEITSPMTAGEILQANPNHVLSKPCSQGVVRKILILSPESELKRG 75
Query: 77 HIYFLVP 83
IYFL+P
Sbjct: 76 SIYFLIP 82
>At5g17350 unknown protein
Length = 183
Score = 46.6 bits (109), Expect = 4e-06
Identities = 24/103 (23%), Positives = 51/103 (49%), Gaps = 2/103 (1%)
Query: 20 STVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLN--H 77
S ++ DG ++ + P+KA ++ + P+ FL ++S+ +G P+ +++LQ+ H
Sbjct: 18 SAAKVILPDGGVRNIHAPMKAAELMMEIPSFFLVDAKSLKIGRKFCPLAADDDLQIKGCH 77
Query: 78 IYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSV 120
+Y P +++ + DL L + A H S++V
Sbjct: 78 VYVAFPMTRATSAANASDLARLFVAAKKQRRHRVGSDHSSAAV 120
>At1g06980 unknown protein
Length = 169
Score = 46.2 bits (108), Expect = 6e-06
Identities = 23/69 (33%), Positives = 35/69 (50%)
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLN 76
G + IVH +G ++++ + A +L NPNH L S V + + P EL+
Sbjct: 16 GALDLIRIVHLNGYVEEITRSITAGEILQANPNHVLSKPCSQGVVRKILILSPESELKRG 75
Query: 77 HIYFLVPRS 85
IYFL+P S
Sbjct: 76 SIYFLIPDS 84
>At1g28190 unknown protein
Length = 266
Score = 44.7 bits (104), Expect = 2e-05
Identities = 29/114 (25%), Positives = 46/114 (39%)
Query: 22 VNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHIYFL 81
+ +V DG LQ PV + P H +C S+ +Y+G + E L+L YFL
Sbjct: 39 IKLVRSDGSLQVYDRPVVVSELTKDFPKHKICRSDLLYIGQKTPVLSETETLKLGLNYFL 98
Query: 82 VPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSSSVPQINFKTHPVHSNVS 135
+P + LS + L N + + +P + Q K + VS
Sbjct: 99 LPSDFFKNDLSFLTIATLKTPQNGGVLVKKTQQQPQPFLIQKGEKGERLRIRVS 152
>At1g18290 hypothetical protein
Length = 176
Score = 43.9 bits (102), Expect = 3e-05
Identities = 25/85 (29%), Positives = 44/85 (51%), Gaps = 1/85 (1%)
Query: 27 FDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQ-LNHIYFLVPRS 85
F G L+ +P+K ++S++ HF+ S + + +T V P+E L+ H+Y L+P
Sbjct: 27 FTGTLEVFSKPIKTSDIVSRHSGHFITDSTLLQISHRVTAVSPDEYLRPRRHLYLLLPTD 86
Query: 86 KSRLPLSLEDLCALAIKANAALAHS 110
L+ E+L ++ KA L S
Sbjct: 87 MLFSVLTQEELSLISNKAAETLNKS 111
>At5g37840 putative protein
Length = 214
Score = 41.6 bits (96), Expect = 1e-04
Identities = 23/65 (35%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 19 QSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNEELQLNH 77
++ V ++ DG + +LK PV A + P L SE++ +G P+ PN+ L+ NH
Sbjct: 9 RNKVKVMKIDGDIFRLKTPVTASDATKEYPGFVLLDSETVKRLGVRAKPLEPNQILKPNH 68
Query: 78 IYFLV 82
YFLV
Sbjct: 69 TYFLV 73
>At3g61920 unknown protein
Length = 187
Score = 41.6 bits (96), Expect = 1e-04
Identities = 28/91 (30%), Positives = 47/91 (50%), Gaps = 9/91 (9%)
Query: 1 MGICASTQYAGKGGNHGM----QSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSE 56
MG C + G GG+ + S + +V +G + +L P+ A + ++ P H + S
Sbjct: 1 MGNCV---FKGNGGSRKLYDKDDSLIKVVTPNGGVMELHPPIFAEFITNEFPGHVIHDSL 57
Query: 57 SMYVGSPMTPVVPNEELQLNHIYFLVPRSKS 87
S+ SP P++ EEL +IY+L+P S S
Sbjct: 58 SLRHSSP--PLLHGEELFPGNIYYLLPLSSS 86
>At3g03280 unknown protein
Length = 166
Score = 41.2 bits (95), Expect = 2e-04
Identities = 19/74 (25%), Positives = 40/74 (53%), Gaps = 2/74 (2%)
Query: 28 DGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLN--HIYFLVPRS 85
DG ++ + P KA ++ + P++FL ++S+ VG P+ +++L L H+Y P +
Sbjct: 24 DGGVRDIHVPTKAAELMMEMPSYFLVDTKSVKVGRKFIPLAADDDLDLGGCHVYVAFPMT 83
Query: 86 KSRLPLSLEDLCAL 99
++ + D+ L
Sbjct: 84 RATSAANASDMARL 97
>At1g29190 hypothetical protein
Length = 330
Score = 39.7 bits (91), Expect = 5e-04
Identities = 28/111 (25%), Positives = 49/111 (43%), Gaps = 16/111 (14%)
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSES---------MYVGSPMTPV 67
G + IVH +G ++++ + A ++ +P H L S + + + V
Sbjct: 16 GALDVIRIVHSNGHVEEISGTITASEIMKAHPKHVLKKPSSPTSDHDERDVISATKIVIV 75
Query: 68 VPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSS 118
P ELQ IYFL+P +KS D CA K +++++V + S
Sbjct: 76 PPEAELQRGKIYFLMPATKS-------DKCAGGGKIRREKSNANAVVKKRS 119
>At1g29195 unknown protein
Length = 193
Score = 39.7 bits (91), Expect = 5e-04
Identities = 28/111 (25%), Positives = 49/111 (43%), Gaps = 16/111 (14%)
Query: 17 GMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSES---------MYVGSPMTPV 67
G + IVH +G ++++ + A ++ +P H L S + + + V
Sbjct: 16 GALDVIRIVHSNGHVEEISGTITASEIMKAHPKHVLKKPSSPTSDHDERDVISATKIVIV 75
Query: 68 VPNEELQLNHIYFLVPRSKSRLPLSLEDLCALAIKANAALAHSSSVFEPSS 118
P ELQ IYFL+P +KS D CA K +++++V + S
Sbjct: 76 PPEAELQRGKIYFLMPATKS-------DKCAGGGKIRREKSNANAVVKKRS 119
>At2g01340 unknown protein
Length = 225
Score = 38.5 bits (88), Expect = 0.001
Identities = 27/89 (30%), Positives = 40/89 (44%), Gaps = 11/89 (12%)
Query: 13 GGNHGMQSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTP----- 66
G + G + T ++ DG+ +LK PV A VL P H L SES+ + G+ P
Sbjct: 2 GNSLGGKKTTKVMKIDGETFKLKTPVTAEEVLKDFPGHVLLDSESVKHYGARAKPLETKR 61
Query: 67 -----VVPNEELQLNHIYFLVPRSKSRLP 90
V + L+ +YF+V K P
Sbjct: 62 LMLFGVQAKQRLEAKRLYFVVEPVKECPP 90
>At5g03890 putative protein
Length = 179
Score = 37.4 bits (85), Expect = 0.003
Identities = 17/70 (24%), Positives = 37/70 (52%), Gaps = 8/70 (11%)
Query: 19 QSTVNIVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESMYVGSPMTPVVPNEELQLNHI 78
+ + IV DGK+ + +EP+ H+L+Q H + + T ++P+ +L +
Sbjct: 9 KKVIKIVRDDGKVLEYREPISVHHILTQFSGHSISHNN--------THLLPDAKLLSGRL 60
Query: 79 YFLVPRSKSR 88
Y+L+P + ++
Sbjct: 61 YYLLPTTMTK 70
>At1g66480 hypothetical protein
Length = 223
Score = 36.2 bits (82), Expect = 0.006
Identities = 20/64 (31%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 24 IVHFDGKLQQLKEPVKAWHVLSQNPNHFLCSSESM-YVGSPMTPVVPNEELQLNHIYFLV 82
++ DG+ ++K PV A V + P + L S+++ + G P+ PN+ L+ YFLV
Sbjct: 14 VMKIDGETFRIKTPVTAREVTADYPGYVLLDSQAVKHFGVRSKPLEPNQTLKPKKTYFLV 73
Query: 83 PRSK 86
K
Sbjct: 74 ELPK 77
>At4g02090 unknown protein
Length = 202
Score = 33.1 bits (74), Expect = 0.049
Identities = 23/109 (21%), Positives = 49/109 (44%), Gaps = 17/109 (15%)
Query: 24 IVHFDGKLQQLKEPVKAWHVLSQNPNHFLC---------SSESMYVGSPMTPVVPNEELQ 74
+V DG++Q L+E ++ +NP H + ++++ V + P+ ++ L+
Sbjct: 20 VVLSDGRVQNLEEETTVAEIMLENPQHVVVEFDPSSISFNNDAKTVKRKLAPLPADKTLE 79
Query: 75 LNHIYFLVPRSK--------SRLPLSLEDLCALAIKANAALAHSSSVFE 115
IY ++P + S L+ E++ + A A + S S +E
Sbjct: 80 PGKIYLVLPAKRSGGRAAKSSSAVLTSEEMRKMLFSATAMVRSSFSYYE 128
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.130 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,311,474
Number of Sequences: 26719
Number of extensions: 130677
Number of successful extensions: 342
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 307
Number of HSP's gapped (non-prelim): 37
length of query: 140
length of database: 11,318,596
effective HSP length: 89
effective length of query: 51
effective length of database: 8,940,605
effective search space: 455970855
effective search space used: 455970855
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0091a.5


