

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089b.6
(647 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
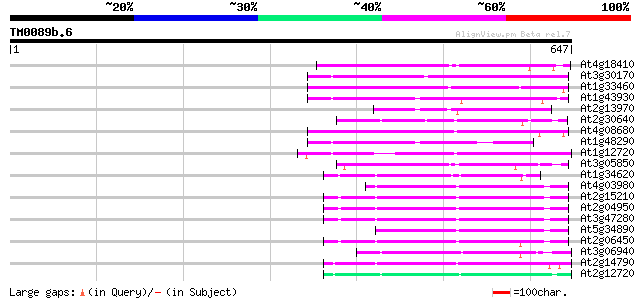
Sequences producing significant alignments: (bits) Value
At4g18410 MuDR transposable element - like protein 114 2e-25
At3g30170 unknown protein 110 2e-24
At1g33460 mutator transposase MUDRA, putative 106 4e-23
At1g43930 hypothetical protein 105 6e-23
At2g13970 Mutator-like transposase 98 1e-20
At2g30640 Mutator-like transposase 97 3e-20
At4g08680 putative MuDR-A-like transposon protein 89 6e-18
At1g48290 hypothetical protein 85 1e-16
At1g12720 hypothetical protein 84 2e-16
At3g05850 unknown protein 84 2e-16
At1g34620 Mutator-like protein 84 2e-16
At4g03980 putative transposon protein 78 2e-14
At2g15210 putaive transposase of transposon FARE2.7 (cds1) 77 2e-14
At2g04950 putative transposase of transposon FARE2.9 (cds1) 77 3e-14
At3g47280 putative protein 77 4e-14
At5g34890 putative protein 72 1e-12
At2g06450 putative transposase of FARE2.11 72 1e-12
At3g06940 putative mudrA protein 70 3e-12
At2g14790 Mutator-like transposase 69 6e-12
At2g12720 putative protein 69 6e-12
>At4g18410 MuDR transposable element - like protein
Length = 633
Score = 114 bits (284), Expect = 2e-25
Identities = 78/304 (25%), Positives = 140/304 (45%), Gaps = 23/304 (7%)
Query: 355 VEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQ 414
+EH ++H+Y N K +G + +I+ L+ + A + + + + +++ + Y +M+
Sbjct: 316 IEHRMCVQHIYGNLKKTYGSKTMIKPLLWNLAWSYNETEYKQHLEKIRCYDTKVYESVMK 375
Query: 415 IPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNE 474
+W + + C+ + N ESFN ++ AR+K + M++ IR M R A +
Sbjct: 376 TNPRSWVRAFQKIGSFCEDVDNNSVESFNGSLNKAREKQFVAMLETIRRMAMVRIAKRSV 435
Query: 475 KMDKY*RGVMPKPLERLKWE--IQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCM 532
+ + P ++ L E + T+ + P G M+EV+H D VDL +TC
Sbjct: 436 ESHTHTGVCTPYVMKFLAGEHKVASTAKVAPGTNG--MYEVRH--GGDTHRVDLAAYTCT 491
Query: 533 CNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTND 592
C W++ GIP H I H +PEDFV ++R A ++ Y + P G K WP +N
Sbjct: 492 CIKWQICGIPCEHAYGVILHKKLQPEDFVCQWFRTAMWKKNYTDGLFPQRGPKFWPESNG 551
Query: 593 EYILSP-----LYRRALGRPKKLRMRDPHEDTTNNHTR-----LHIGVSRNKFSRCGQFG 642
+ + + + + +K R + +E T + +H GV CG
Sbjct: 552 PRVFAAEPPEGEEDKKMTKAEKKRKKGVNESPTKKQPKAKKRIMHCGV-------CGAAD 604
Query: 643 HNTR 646
HN+R
Sbjct: 605 HNSR 608
>At3g30170 unknown protein
Length = 995
Score = 110 bits (276), Expect = 2e-24
Identities = 89/306 (29%), Positives = 136/306 (44%), Gaps = 10/306 (3%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L+ ++ EH +H+ N+K + ++ L A++ + + MMELQ
Sbjct: 481 LVKAIHTILPHAEHRQCSKHIMDNWKRD-SHDLELQRLFWKIARSYTVEEFNNHMMELQQ 539
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
+ +AY L W + F T C+ +N L ESFN TI AR KP+L M++ IR
Sbjct: 540 YNRQAYESLQLTSPVTWSRAFFRIGTCCNDNLNNLSESFNRTIRQARRKPILDMLEDIRR 599
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKH--IVSHDQ 521
M R A +K + + +R EI+ + QC E H V++
Sbjct: 600 QCMVRSAKRFLIAEK----LQTRFTKRAHEEIEKMIVGLRQCERYMARENLHEIYVNNVS 655
Query: 522 FIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPI 581
+ VD+++ TC C ++VGIP H I +K ED+V YY + +R TY + P+
Sbjct: 656 YFVDMEQKTCDCRKCQMVGIPCIHATCVIIGKKEKVEDYVSDYYTKVRWRETYLKGIRPV 715
Query: 582 SGQKLWPTTNDEYILSPLYRRA-LGRPKKLRMRDPHEDTT--NNHTRLHIGVSRNKFSRC 638
G LW TN +L P +RR GRP R + +N ++ S C
Sbjct: 716 QGMPLWCRTNRLPVLPPPWRRGNAGRPSNYARRKGRNEAAAPSNPNKMSREKRIMTCSNC 775
Query: 639 GQFGHN 644
Q GHN
Sbjct: 776 HQEGHN 781
>At1g33460 mutator transposase MUDRA, putative
Length = 826
Score = 106 bits (265), Expect = 4e-23
Identities = 80/312 (25%), Positives = 138/312 (43%), Gaps = 14/312 (4%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
+++ + + EH ++H+ N K G +++ + + A + + + E++A
Sbjct: 444 IISAVKNALPNAEHRPCVKHIVENLKKRHGSLDLLKKFVWNLAWSYSDTQYKANLNEMRA 503
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
Y +M+ + WC+ F + C+ + N ESFNATI+ AR K ++ M++ IR
Sbjct: 504 YIMSLYEDVMKEEPNTWCRSWFRIGSYCEDVDNNATESFNATIVKARAKALVPMMETIRR 563
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLK--WEIQMTSHLIPQCCG*DMFEVKHIVSHDQ 521
M R + K+ K+ + + E LK WE+ + + + FEV + +
Sbjct: 564 QAMARISKRKAKIGKWKKKISEYVSEILKEEWELAVKCEVTKGTH--EKFEVWFDGNSNA 621
Query: 522 FIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPI 581
+ ++ C C W++ GIP H AAI+ G+ EDFV + A+R Y+ P+
Sbjct: 622 VSLKTEEWDCSCCKWQVTGIPCEHAYAAINDVGKDVEDFVIPMFYTIAWREQYDTGPDPV 681
Query: 582 SGQKLWPTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSR---- 637
GQ WP T I +PL KK ++ H N + G R K +
Sbjct: 682 RGQMYWP-TGLGLITAPLQDPVPPGRKKGEKKNYHRIKGPNESPKKKGPQRLKVGKKGVV 740
Query: 638 -----CGQFGHN 644
CG+ GHN
Sbjct: 741 MHCKSCGEAGHN 752
>At1g43930 hypothetical protein
Length = 946
Score = 105 bits (263), Expect = 6e-23
Identities = 88/312 (28%), Positives = 136/312 (43%), Gaps = 19/312 (6%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L+ ++ EH RH+ N+K + ++ L ++ + M L++
Sbjct: 509 LVKAIHSVIPQAEHRQCARHIMENWKRN-SHDMELQRLFWKIVRSYTEGEFGAHMRALKS 567
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
+ A+ L++ W + F + C+ +N L ESFN TI AR KP+L M++ IR
Sbjct: 568 YNASAFELLLKTLPVTWSRAFFKIGSCCNDNLNNLSESFNRTIREARRKPLLDMLEDIRR 627
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSH---D 520
M R NEK + + +R EI+ C KH + D
Sbjct: 628 QCMVR----NEKRYVIADRLRTRFTKRAHAEIEKMIAGSQVCQRWMARHNKHEIKVGPVD 683
Query: 521 QFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSP 580
F VD+ +TC C W++ GIP H + I QK ED+V +Y + ++ TY + P
Sbjct: 684 SFTVDMNNNTCGCMKWQMTGIPCIHAASVIIGKRQKVEDYVSDWYTTSMWKQTYNDGIGP 743
Query: 581 ISGQKLWPTTNDEYILSPLYRRA-LGRPKKLRMR-------DPHEDTTNNHTRLHIGVSR 632
+ G+ LWPT N +L P +RR GRPK R +T +RLH ++
Sbjct: 744 VQGKLLWPTVNKVGVLPPPWRRGNPGRPKNHARRKGVFESSTASSSSTTELSRLHRVMT- 802
Query: 633 NKFSRCGQFGHN 644
S C GHN
Sbjct: 803 --CSNCQGEGHN 812
>At2g13970 Mutator-like transposase
Length = 597
Score = 98.2 bits (243), Expect = 1e-20
Identities = 71/214 (33%), Positives = 101/214 (47%), Gaps = 15/214 (7%)
Query: 420 WCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY 479
W + F + C+ +N L ESFN TI AR KP+L M++ IR M R NEK
Sbjct: 236 WSRAFFRIGSCCNDNLNNLSESFNRTIRNARRKPLLDMLEDIRRQCMVR----NEKRFVI 291
Query: 480 *RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKH------IVSHDQFIVDLKKHTCMC 533
+ + +R EI+ C D + +H + D F VD+ TC C
Sbjct: 292 AYRLKSRFTKRAHMEIEKMIDGAAVC---DRWMARHNRIEVRVGLDDSFSVDMNDRTCGC 348
Query: 534 NFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDE 593
W++ GIP H + I N QK EDFV +Y ++ TY + P+ G+ LWP N
Sbjct: 349 RKWQMTGIPCIHAASVIIGNRQKVEDFVSDWYTTYMWKQTYYDGIMPVQGKMLWPIVNRV 408
Query: 594 YILSPLYRRA-LGRPKKL-RMRDPHEDTTNNHTR 625
+L P +RR GRPK R + +E T + +R
Sbjct: 409 GVLPPPWRRGNPGRPKNHDRRKGIYESATASTSR 442
>At2g30640 Mutator-like transposase
Length = 754
Score = 97.1 bits (240), Expect = 3e-20
Identities = 78/277 (28%), Positives = 122/277 (43%), Gaps = 22/277 (7%)
Query: 377 VIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMN 436
++ NL+ +AAK + KM E+ + PEA +W+ I S W F T+ L
Sbjct: 481 ILVNLVWEAAKCLTDFEFEGKMGEIAQISPEAASWIRNIQHSQWATYCFEG-TRFGHLTA 539
Query: 437 KLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQ 496
+ ES N+ + A P++ M++ IR +M F E ++ G++ ER E
Sbjct: 540 NVSESLNSWVQDASGLPIIQMLESIRRQLMTLFNERRETSMQW-SGMLVPSAERHVLEAI 598
Query: 497 MTSHLIP-QCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQ 555
L P FEV + S ++IVD++ TC C WEL G+P H +AA+ Q
Sbjct: 599 EECRLYPVHKANEAQFEV--MTSEGKWIVDIRCRTCYCRGWELYGLPCSHAVAALLACRQ 656
Query: 556 KPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTT---------NDEYILSPLYRRALGR 606
F +Y+ A YR TY + P+ + W TT +E ++ P R L +
Sbjct: 657 NVYRFTESYFTVANYRRTYAETIHPVPDKTEWKTTEPAGESGEDGEEIVIRP--PRDLRQ 714
Query: 607 PKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGH 643
P + + R + R+ + SRC Q GH
Sbjct: 715 PPRPKKRRSQGEDRGRQKRV------VRCSRCNQAGH 745
>At4g08680 putative MuDR-A-like transposon protein
Length = 761
Score = 89.4 bits (220), Expect = 6e-18
Identities = 70/314 (22%), Positives = 128/314 (40%), Gaps = 15/314 (4%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
LL+ + EH ++H+ N K + +++ L+ A + + + K + L+
Sbjct: 378 LLSAVKQELPNAEHRMCVKHIVENLKKNHAKKDMLKTLVWKLAWSYNEKEYGKNLNNLRC 437
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
Y ++ W + + + C+ + N ESFN+TI AR K ++ M++ IR
Sbjct: 438 YDEALYNDVLNEEPHTWSRCFYKLGSCCEDVDNNATESFNSTITKARAKSLIPMLETIRR 497
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFI 523
M R N+K ++ L+ L E C +FEV + +
Sbjct: 498 QGMTRIVKRNKKSLRHEGRFTKYALKMLALEKTDADRSKVYRCTHGVFEV--YIDENGHR 555
Query: 524 VDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISG 583
VD+ K C C W++ GIP H A+ G E+++ ++ +R +YE P+ G
Sbjct: 556 VDIPKTQCSCGKWQISGIPCEHSYGAMIEAGLDAENYISEFFSTDLWRDSYETATMPLRG 615
Query: 584 QKLWPTTNDEYILSPLYRRALGRPKK-------LRMRDPHEDTTNNHTRLHIGVSRNKFS 636
K W ++ + +P GR K+ R++ +E + + K
Sbjct: 616 PKYWLNSSYGLVTAPPEPILPGRKKEKSKKEKFARIKGKNESPKKKKRKKNEVEKLGKKG 675
Query: 637 R------CGQFGHN 644
+ CG+ GHN
Sbjct: 676 KIIHCKSCGEAGHN 689
>At1g48290 hypothetical protein
Length = 444
Score = 85.1 bits (209), Expect = 1e-16
Identities = 72/263 (27%), Positives = 113/263 (42%), Gaps = 24/263 (9%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L+ + + EH +H+ N+K + ++ L A++ +T + EL+
Sbjct: 165 LVKAIHNRIPAAEHRQCAKHIMDNWKRN-SHDMELQRLFWKIARSYTIGEYTANLEELKT 223
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
+P A LM W + F + C+ +N L ESFN TI AR KP+L M++ IRS
Sbjct: 224 YNPGAAASLMNTKPMEWSRAFFRIGSCCNDNLNNLSESFNRTIRQARRKPLLDMLEDIRS 283
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSH--DQ 521
M R NEK + +R EI+ C KH +SH +
Sbjct: 284 QCMVR----NEKRYIIAGRWKSRFTKRAHEEIEKMIAGSQFCERSMARHNKHEISHFGRK 339
Query: 522 FIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPI 581
+ VD+ +TC C W++ ED+V +Y ++ TY ++P+
Sbjct: 340 YSVDMNDNTCGCRKWQMT-----------------VEDYVSDWYTTRMWQLTYNDGIAPV 382
Query: 582 SGQKLWPTTNDEYILSPLYRRAL 604
GQ LWP N +L P +RR L
Sbjct: 383 QGQLLWPRVNRLGVLPPPWRRGL 405
>At1g12720 hypothetical protein
Length = 585
Score = 84.3 bits (207), Expect = 2e-16
Identities = 78/323 (24%), Positives = 131/323 (40%), Gaps = 35/323 (10%)
Query: 333 ICTEVSLFL-----QDLLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAK 387
+C V+L L L+ +++ EH +H+ N+K + ++ L ++
Sbjct: 175 LCEMVNLALISDKQSGLVKAIHNVLPQAEHRQCSKHIMDNWKRD-SHDMELQRLFWKISR 233
Query: 388 ATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATIL 447
+ + + M L++ +P+AY L W TI
Sbjct: 234 SYTIEEFNTHMANLKSYNPQAYASLQLTSPMTW------------------------TIR 269
Query: 448 LARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG 507
AR KP+L M++ IR M R A ++ P+ ++ I ++
Sbjct: 270 QARRKPLLDMLEDIRRQCMVRTAKRFIIAERLKSRFTPRAHAEIEKMIAGSAGCERHLAR 329
Query: 508 *DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRR 567
++ E+ V+ + VD+ K TC C WE+VGIP H I +K ED+V YY +
Sbjct: 330 NNLHEI--YVNDVGYFVDMDKKTCGCRKWEMVGIPCVHTPCVIIGRKEKVEDYVSDYYTK 387
Query: 568 AAYRATYEHLVSPISGQKLWPTTNDEYILSPLYRRA-LGRPKKLRMRDPHEDT--TNNHT 624
+R TY + P+ G LWP + +L P +RR GR + +T ++N
Sbjct: 388 VRWRETYRDGIRPVQGMPLWPRMSRLPVLPPPWRRGNPGRQSNYARKKGRYETASSSNKN 447
Query: 625 RLHIGVSRNKFSRCGQFGHNTRS 647
++ S C Q GHN S
Sbjct: 448 KMSRANRIMTCSNCKQEGHNKSS 470
>At3g05850 unknown protein
Length = 777
Score = 84.0 bits (206), Expect = 2e-16
Identities = 71/278 (25%), Positives = 121/278 (42%), Gaps = 26/278 (9%)
Query: 378 IRNLMMD----AAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDV 433
I+ L++D AA A + + + ++ L PEAY W++Q A+ + +
Sbjct: 504 IKRLIVDDFYSAAYAPRADSFERHVENIKGLSPEAYDWIVQKSQPDHWANAYFRGARYNH 563
Query: 434 LMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKW 493
+ + E F + A D P+ MVD IR IMG ++ + P +L+
Sbjct: 564 MTSHSGEPFFSWASDANDLPITQMVDVIRGKIMGLIHVRRISANEANGNLTPSMEVKLEK 623
Query: 494 EI--QMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAIS 551
E T H+ P ++F+V+ +V++ + C C W+L G+P H +A I+
Sbjct: 624 ESLRAQTVHVAPSADN-NLFQVR---GETYELVNMAECDCSCKGWQLTGLPCHHAVAVIN 679
Query: 552 HNGQKPEDFVHAYYRRAAYRATYEHLVSPI---SGQKLWPTTNDE--YILSPLYRRALGR 606
+ G+ P D+ Y+ A YR+TY ++P+ G+ ++ + P RR GR
Sbjct: 680 YYGRNPYDYCSKYFTVAYYRSTYAQSINPVPLLEGEMCRESSGGSAVTVTPPPTRRPPGR 739
Query: 607 PKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
P K + P E+ + SRC GHN
Sbjct: 740 PPK--KKTPAEEVMKRQLQC---------SRCKGLGHN 766
>At1g34620 Mutator-like protein
Length = 1034
Score = 84.0 bits (206), Expect = 2e-16
Identities = 68/258 (26%), Positives = 130/258 (50%), Gaps = 14/258 (5%)
Query: 363 HLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCK 422
HL+ N ++ + + + R LM +AA+A + + + +++Q L+P +L+ + S W +
Sbjct: 653 HLWRNIRHLYKPKSLCR-LMSEAAQAFHVTDFNRIFLKIQKLNPGCAAYLVDLGFSEWTR 711
Query: 423 QAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RG 482
+ + +++ + + ES+N I AR+ P++ M++ IR+ +M FAT + +
Sbjct: 712 -VHSKGRRFNIMDSNICESWNNVIREAREYPLICMLEYIRTTLMDWFATRRAQAEDCPTT 770
Query: 483 VMPKPLERLKWEIQMTSHL-IPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGI 541
+ P+ ER++ Q + + C + F+V+ + FIV L + TC C ++ +GI
Sbjct: 771 LAPRVQERVEENYQSAMSMSVKPICNFE-FQVQERTG-ECFIVKLDESTCSCLEFQGLGI 828
Query: 542 PFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPIS--GQKLWP-----TTNDEY 594
P H IAA + G + Y + +YE + PI G ++ P TT +
Sbjct: 829 PCAHAIAAAARIGVPTDSLAANGYFNELVKLSYEEKIYPIHSVGGEVAPEIAAGTTGE-- 886
Query: 595 ILSPLYRRALGRPKKLRM 612
+ P RR GRP+K+R+
Sbjct: 887 LQPPFVRRPPGRPRKIRI 904
>At4g03980 putative transposon protein
Length = 580
Score = 77.8 bits (190), Expect = 2e-14
Identities = 63/238 (26%), Positives = 105/238 (43%), Gaps = 12/238 (5%)
Query: 411 WLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFA 470
+L ++ W + A++ + +++ + L ES NA + R+ P++ + + IRS + F
Sbjct: 343 YLWEVDVRKWSR-AYSPSNRYNIMTSNLAESVNALLKQNREYPIVCLFESIRSIMTRWFN 401
Query: 471 TMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQC-CG*DMFEVKHIVSHDQFIVDLKKH 529
E+ ++ V +++K ++ + C + FEVK +V+L K
Sbjct: 402 ERREESSQHPSAVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKR 459
Query: 530 TCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLW-- 587
TC C +++ P+ H IA+ H FV ++ +R Y + P + W
Sbjct: 460 TCTCCMFDIDKFPYAHGIASAKHINLNKNMFVDEFHSTYRWRQAYSESIHPNGDMEYWEI 519
Query: 588 PTTNDEYI-LSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
P T E I L P R GR KK R+ E H R +K SRCGQ GHN
Sbjct: 520 PDTVSEVICLPPSTRVPSGRRKKKRIPSVWE-----HGRSQPKPKLHKCSRCGQSGHN 572
>At2g15210 putaive transposase of transposon FARE2.7 (cds1)
Length = 580
Score = 77.4 bits (189), Expect = 2e-14
Identities = 73/287 (25%), Positives = 122/287 (42%), Gaps = 15/287 (5%)
Query: 363 HLYANFKNFFGGRVVIRNLMMDAAKATYHQL-WTKKMMELQALHPEAYTWLMQIPTSAWC 421
HL N + F G +I + AA Y + + E+ + +L ++ W
Sbjct: 296 HLQKNLETHFRGSSLIP--VYYAASRVYTKTEFDSLFWEITNSDKKLAQYLWEVHVRKWS 353
Query: 422 KQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*R 481
+ A++ + +++ + L ES NA + R+ P++ + + IRS + F E+ ++
Sbjct: 354 R-AYSPSNRYNIMTSNLAESVNALLKQNREYPIVCLFESIRSIMTRWFNERREESSQHPS 412
Query: 482 GVMPKPLERLKWEIQMTSHLIPQC-CG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVG 540
V +++K ++ + C + FEVK +V+L K TC C +++
Sbjct: 413 AVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKRTCTCCMFDIDK 470
Query: 541 IPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLW--PTTNDEYI-LS 597
P H IA+ +H FV ++ +R Y + P + W P T E I L
Sbjct: 471 FPCAHGIASANHINLNENMFVDEFHSTYRWRQAYSESIHPNGDMEYWEIPETISEVICLP 530
Query: 598 PLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
P R GR KK R+ E H R +K SRCGQ GHN
Sbjct: 531 PSTRVPSGRRKKKRIPSVWE-----HGRSQPKPKLHKCSRCGQSGHN 572
>At2g04950 putative transposase of transposon FARE2.9 (cds1)
Length = 616
Score = 77.0 bits (188), Expect = 3e-14
Identities = 73/287 (25%), Positives = 121/287 (41%), Gaps = 15/287 (5%)
Query: 363 HLYANFKNFFGGRVVIRNLMMDAAKATYHQL-WTKKMMELQALHPEAYTWLMQIPTSAWC 421
HL N + F G +I + AA Y + + E+ + +L ++ W
Sbjct: 332 HLQKNLETHFRGSSLIP--VYYAASRVYTKTEFDSLFWEITNSDKKLAQYLWEVDVRKWS 389
Query: 422 KQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*R 481
+ A++ + +++ + L ES NA + R+ P++ + + IRS + F E+ ++
Sbjct: 390 R-AYSPSNRYNIMTSNLAESVNALLKQNREYPIVCLFESIRSIMTRWFNERREESSQHPS 448
Query: 482 GVMPKPLERLKWEIQMTSHLIPQC-CG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVG 540
V +++K ++ + C + FEVK +V+L K TC C +++
Sbjct: 449 AVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKRTCTCCMFDIDK 506
Query: 541 IPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLW--PTTNDEYI-LS 597
P H IA+ H FV ++ +R Y + P + W P T E I L
Sbjct: 507 FPCAHGIASAKHINLNENMFVDEFHSTYKWRQAYSKSIHPNGDMEYWEIPETISEVICLP 566
Query: 598 PLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
P R GR KK R+ E H R +K SRCGQ GHN
Sbjct: 567 PSTRVPSGRRKKKRIPSVWE-----HGRSQPKPKLHKCSRCGQSGHN 608
>At3g47280 putative protein
Length = 739
Score = 76.6 bits (187), Expect = 4e-14
Identities = 73/287 (25%), Positives = 121/287 (41%), Gaps = 15/287 (5%)
Query: 363 HLYANFKNFFGGRVVIRNLMMDAAKATYHQL-WTKKMMELQALHPEAYTWLMQIPTSAWC 421
HL N + F G +I + AA Y + + E+ + +L ++ W
Sbjct: 455 HLQKNLETHFRGSSLIP--VYYAASRIYTKTEFDSLFWEITNSDKKLAQYLCEVDVRKWS 512
Query: 422 KQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*R 481
+ A++ + +++ + L ES NA + R+ PV+ + + IRS + F E+ ++
Sbjct: 513 R-AYSPSNRYNIMTSNLAESVNALLKQNREYPVVCLFESIRSIMTRWFNERREESSQHPS 571
Query: 482 GVMPKPLERLKWEIQMTSHLIPQC-CG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVG 540
V +++K ++ + C + FEVK +V+L K TC C +++
Sbjct: 572 AVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKRTCTCCMFDIDK 629
Query: 541 IPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLW--PTTNDEYI-LS 597
P H IA+ H FV ++ +R Y + P + W P T E I L
Sbjct: 630 FPCAHGIASAKHINLNENMFVDEFHSTYRWRQAYSESIHPNGDMEYWEIPETISEVICLP 689
Query: 598 PLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
P R G+ KK R+ E H R +K SRCGQ GHN
Sbjct: 690 PSTRVPTGKRKKKRIPSVWE-----HGRSQPKPKLHKCSRCGQSGHN 731
>At5g34890 putative protein
Length = 681
Score = 72.0 bits (175), Expect = 1e-12
Identities = 60/226 (26%), Positives = 97/226 (42%), Gaps = 11/226 (4%)
Query: 423 QAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RG 482
+A++ + +++ + L ES NA + R+ P++ + + IRS + F E+ ++
Sbjct: 455 RAYSPSNRYNIMTSNLAESVNALLKQNREYPIVCLFESIRSIMTRWFNEQREESSQHLSA 514
Query: 483 VMPKPLERLKWEIQMTSHLIPQC-CG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGI 541
V +++K ++ + C + FEVK + +L K TC C +++
Sbjct: 515 VTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLANLDKRTCTCCMFDIDKF 572
Query: 542 PFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLW--PTTNDEYI-LSP 598
P H IA+ H FV ++ +R Y + P K W P T E I L P
Sbjct: 573 PCAHGIASAKHINLNENMFVDEFHSTYRWRQAYSESIHPNGDMKYWEIPETISEVICLPP 632
Query: 599 LYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
R GR K R+ E H R +K SRCGQ GHN
Sbjct: 633 STRVPSGRRNKKRIPSVWE-----HGRSQPKPKLHKCSRCGQSGHN 673
>At2g06450 putative transposase of FARE2.11
Length = 580
Score = 71.6 bits (174), Expect = 1e-12
Identities = 67/287 (23%), Positives = 119/287 (41%), Gaps = 15/287 (5%)
Query: 363 HLYANFKNFFGGRVVIRNLMMDAAKATYHQL-WTKKMMELQALHPEAYTWLMQIPTSAWC 421
HL N + F G +I + AA Y + + E+ + +L ++ W
Sbjct: 296 HLQKNLETHFRGSSLIP--VYYAASRVYTKTEFDSLFWEITNSDKKLAQYLWEVDVRKWS 353
Query: 422 KQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*R 481
+ ++ + +++ + L ES NA + R+ P++ + + IRS + F E+ ++
Sbjct: 354 R-TYSPSNRYNIMTSNLAESVNALLKQNREYPIVCLFESIRSIMTRWFNERREESSQHPS 412
Query: 482 GVMPKPLERLKWEIQMTSHLIPQC-CG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVG 540
V +++K ++ + C + FEVK +V+L K TC C +++
Sbjct: 413 AVTINVGKKMKASYDTSTRWLEVCQVNQEEFEVKG--DTKTHLVNLDKRTCTCCMFDIDK 470
Query: 541 IPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLW---PTTNDEYILS 597
P H IA+ H FV ++ +R Y + P + W T ++ L
Sbjct: 471 FPCAHGIASAKHINLNENMFVDEFHSTYRWRQAYSESIHPNGDMEYWEILETISEVICLP 530
Query: 598 PLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHN 644
P R GR +K R+ E H R +K SRCG+ GHN
Sbjct: 531 PSTRVPSGRREKKRIPSVWE-----HGRSQPKPKLHKCSRCGRSGHN 572
>At3g06940 putative mudrA protein
Length = 749
Score = 70.5 bits (171), Expect = 3e-12
Identities = 57/251 (22%), Positives = 106/251 (41%), Gaps = 17/251 (6%)
Query: 401 LQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD* 460
++++ P+AY W+++ W F + + + + + F + + A + P+ M+D
Sbjct: 508 IKSISPDAYNWVIESEPHHWANALFQG-ERYNKMNSNFGKDFYSWVSEAHEFPITQMIDE 566
Query: 461 IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHD 520
R+ +M T K ++ + P E+L+ EI++ L +F+V S
Sbjct: 567 FRAKMMQSIYTRQVKSREWVTTLTPSNEEKLQKEIELAQLLQVSSPEGSLFQVNGGESSV 626
Query: 521 QFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSP 580
IVD+ + C C W L G+P H IA I + P ++ Y ++R Y + P
Sbjct: 627 S-IVDINQCDCDCKTWRLTGLPCSHAIAVIGCIEKSPYEYCSTYLTVESHRLMYAESIQP 685
Query: 581 ISGQKLW----PTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFS 636
+ P + P RR GRP K++ +P L + + + S
Sbjct: 686 VPNMDRMMLDDPPEGLVCVTPPPTRRTPGRP-KIKKVEP----------LDMMKRQLQCS 734
Query: 637 RCGQFGHNTRS 647
+C GHN ++
Sbjct: 735 KCKGLGHNKKT 745
>At2g14790 Mutator-like transposase
Length = 895
Score = 69.3 bits (168), Expect = 6e-12
Identities = 78/313 (24%), Positives = 131/313 (40%), Gaps = 31/313 (9%)
Query: 363 HLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCK 422
HL N ++G + + L AAKA + + K L+ + +L I W +
Sbjct: 574 HLERNLSTYYG-KFGVSALFFSAAKAYRVRDFEKYFELLREKSGKCAKYLEDIGFEHWTR 632
Query: 423 QAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RG 482
A + +++ + ES N + A+ P++ M++ IR +M FA+ +K+ +
Sbjct: 633 -AHCRGERYNIMSSNNSESMNHVLTKAKTYPIVYMIEFIRDVLMRWFASRRKKVARCKSS 691
Query: 483 VMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIP 542
V P+ ER E+ + + G ++V S + F V L TC C + + IP
Sbjct: 692 VTPEVDERFLQELPASGKYAVKMSGPWSYQVTS-KSGEHFHVVLDHCTCTCLRYTKLRIP 750
Query: 543 FRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQK--LWPTTNDEYILSPLY 600
H +AA +G P+ V +Y + +++ + PI K + P + IL P Y
Sbjct: 751 CEHALAAAIEHGIDPKSCVGWWYGLQTFSDSFQEPILPIPDPKDVVIPQHIVDLILIPPY 810
Query: 601 -RRALGRP--KKLRMRDPHEDTTN-----------------NHTRLHIGVSR------NK 634
RR GRP K++ R + TN N L + R N+
Sbjct: 811 SRRPPGRPLSKRIPSRGENRVCTNIVHSYTCILKYYHAEICNFLSLSLQKKRQKRLLPNQ 870
Query: 635 FSRCGQFGHNTRS 647
SRC Q GHN ++
Sbjct: 871 CSRCRQAGHNKKT 883
>At2g12720 putative protein
Length = 819
Score = 69.3 bits (168), Expect = 6e-12
Identities = 66/286 (23%), Positives = 114/286 (39%), Gaps = 9/286 (3%)
Query: 363 HLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCK 422
HLY N F R + L+ AA+ +T E++ L P + +L + W +
Sbjct: 539 HLYKNILVKFK-RKDLFPLVKKAARCYRLNDFTNAFNEIEELDPLLHAYLQRAGVEMWAR 597
Query: 423 QAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RG 482
F + +++ + ES N + A++ P++ M++ IR + FA + K
Sbjct: 598 AHFPG-DRYNLMTTNIAESMNRALSQAKNLPIVRMLEAIRQMMTRWFAERRDDASKQHTQ 656
Query: 483 VMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIP 542
+ P + L+ + + L Q +V + S +V++ + C C + +P
Sbjct: 657 LTPGVEKLLQTRVTSSRLLDVQTIDASRVQVAYEAS--LHVVNVDEKQCTCRLFNKEKLP 714
Query: 543 FRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWP-TTNDEYILSPLYR 601
H IAA H G YY+ + Y + P + P D+ L P+
Sbjct: 715 CIHAIAAAEHMGVSRISLCSPYYKSSHLVNVYACAIMPSDTEVPVPQIVIDQPCLPPIVA 774
Query: 602 RALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHNTRS 647
GRPKKLRM+ E + SRC + GHN ++
Sbjct: 775 NQPGRPKKLRMKSALEVAVETKRPR----KEHACSRCKETGHNVKT 816
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.341 0.148 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,505,637
Number of Sequences: 26719
Number of extensions: 540173
Number of successful extensions: 1538
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 1428
Number of HSP's gapped (non-prelim): 81
length of query: 647
length of database: 11,318,596
effective HSP length: 106
effective length of query: 541
effective length of database: 8,486,382
effective search space: 4591132662
effective search space used: 4591132662
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0089b.6


