

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089a.1
(119 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
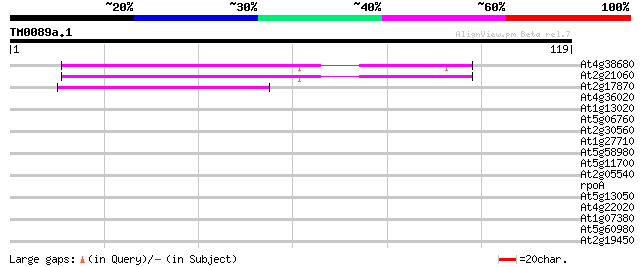
Score E
Sequences producing significant alignments: (bits) Value
At4g38680 glycine-rich protein 2 (GRP2) 56 3e-09
At2g21060 glycine-rich protein (AtGRP2) 54 1e-08
At2g17870 putative glycine-rich, zinc-finger DNA-binding protein 38 7e-04
At4g36020 glycine-rich protein 34 0.013
At1g13020 eukaryotic initiation factor 4B (EIF4B5) 28 0.95
At5g06760 late embryogenesis abundant protein LEA like 27 2.1
At2g30560 putative glycine-rich protein 27 2.1
At1g27710 unknown protein 26 2.8
At5g58980 random slug protein - like 26 3.6
At5g11700 putative protein 26 3.6
At2g05540 putative glycine-rich protein 26 3.6
rpoA -chloroplast genome- RNA polymerase alpha subunit 25 4.7
At5g13050 5-formyltetrahydrofolate cyclo-ligase-like protein 25 4.7
At4g22020 glycine-rich protein 25 4.7
At1g07380 unknown protein 25 4.7
At5g60980 ras-GTPase-activating protein SH3-domain binding prote... 25 8.1
At2g19450 diacylglycerol O-acyltransferase (DAGAT) 25 8.1
>At4g38680 glycine-rich protein 2 (GRP2)
Length = 203
Score = 56.2 bits (134), Expect = 3e-09
Identities = 40/98 (40%), Positives = 51/98 (51%), Gaps = 19/98 (19%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI-------LIPM 64
+K FD QKGF IT DDGG+DLF+ S I+S+GF LA E V+ + I I +
Sbjct: 15 VKWFDTQKGFGFITPDDGGDDLFVHQSSIRSEGFRSLAAEEAVEFEVEIDNNNRPKAIDV 74
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSS----GYGRGGG 98
+ PD AP+ G GG+ G GRGGG
Sbjct: 75 SG--------PDGAPVQGNSGGGSSGGRGGFGGGRGGG 104
>At2g21060 glycine-rich protein (AtGRP2)
Length = 201
Score = 54.3 bits (129), Expect = 1e-08
Identities = 37/94 (39%), Positives = 48/94 (50%), Gaps = 15/94 (15%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI-------LIPM 64
+K FD QKGF IT DGG+DLF+ S I+S+GF LA E V+ + + I +
Sbjct: 19 VKWFDTQKGFGFITPSDGGDDLFVHQSSIRSEGFRSLAAEESVEFDVEVDNSGRPKAIEV 78
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGG 98
+ PD AP+ G GG S G G GG
Sbjct: 79 SG--------PDGAPVQGNSGGGGSSGGRGGFGG 104
>At2g17870 putative glycine-rich, zinc-finger DNA-binding protein
Length = 301
Score = 38.1 bits (87), Expect = 7e-04
Identities = 21/45 (46%), Positives = 26/45 (57%)
Query: 11 KLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVD 55
K+ F D KG+ IT DDGG +LF+ S I S GF L E V+
Sbjct: 14 KVSWFSDGKGYGFITPDDGGEELFVHQSSIVSDGFRSLTLGESVE 58
>At4g36020 glycine-rich protein
Length = 299
Score = 33.9 bits (76), Expect = 0.013
Identities = 18/49 (36%), Positives = 28/49 (56%)
Query: 11 KLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
K+ F+ KG+ IT DDG +LF+ S I S+G+ L + V+ +T
Sbjct: 14 KVNWFNASKGYGFITPDDGSVELFVHQSSIVSEGYRSLTVGDAVEFAIT 62
>At1g13020 eukaryotic initiation factor 4B (EIF4B5)
Length = 549
Score = 27.7 bits (60), Expect = 0.95
Identities = 11/33 (33%), Positives = 18/33 (54%)
Query: 82 GTCCGGNDSSGYGRGGGYQEVVDIRIWRNGRQK 114
G+ GG +G G GGG+ + ++ W G+ K
Sbjct: 201 GSYGGGGAGAGSGGGGGFSKADEVDNWAAGKAK 233
>At5g06760 late embryogenesis abundant protein LEA like
Length = 158
Score = 26.6 bits (57), Expect = 2.1
Identities = 10/17 (58%), Positives = 12/17 (69%)
Query: 83 TCCGGNDSSGYGRGGGY 99
T GG ++GYG GGGY
Sbjct: 140 THVGGGGATGYGTGGGY 156
>At2g30560 putative glycine-rich protein
Length = 171
Score = 26.6 bits (57), Expect = 2.1
Identities = 11/18 (61%), Positives = 11/18 (61%)
Query: 82 GTCCGGNDSSGYGRGGGY 99
G C GG S G G GGGY
Sbjct: 35 GGCGGGGKSGGGGGGGGY 52
>At1g27710 unknown protein
Length = 212
Score = 26.2 bits (56), Expect = 2.8
Identities = 11/18 (61%), Positives = 12/18 (66%)
Query: 82 GTCCGGNDSSGYGRGGGY 99
G CGG+ S G G GGGY
Sbjct: 182 GGGCGGSCSGGGGGGGGY 199
>At5g58980 random slug protein - like
Length = 733
Score = 25.8 bits (55), Expect = 3.6
Identities = 9/17 (52%), Positives = 11/17 (63%)
Query: 85 CGGNDSSGYGRGGGYQE 101
CGG + YGRG GY +
Sbjct: 329 CGGKNEQCYGRGPGYPD 345
>At5g11700 putative protein
Length = 1411
Score = 25.8 bits (55), Expect = 3.6
Identities = 13/23 (56%), Positives = 15/23 (64%), Gaps = 3/23 (13%)
Query: 83 TCCGGNDSSGYGRGGGYQEVVDI 105
+ CGG SGYG GGG + VDI
Sbjct: 288 SACGG---SGYGGGGGGRVSVDI 307
>At2g05540 putative glycine-rich protein
Length = 135
Score = 25.8 bits (55), Expect = 3.6
Identities = 11/16 (68%), Positives = 12/16 (74%), Gaps = 1/16 (6%)
Query: 86 GGNDSSGYG-RGGGYQ 100
GGN GYG RGGGY+
Sbjct: 64 GGNPGGGYGNRGGGYR 79
>rpoA -chloroplast genome- RNA polymerase alpha subunit
Length = 329
Score = 25.4 bits (54), Expect = 4.7
Identities = 9/19 (47%), Positives = 12/19 (62%)
Query: 93 YGRGGGYQEVVDIRIWRNG 111
YG G QE++ + IW NG
Sbjct: 189 YGNGNEKQEILFLEIWTNG 207
>At5g13050 5-formyltetrahydrofolate cyclo-ligase-like protein
Length = 277
Score = 25.4 bits (54), Expect = 4.7
Identities = 10/25 (40%), Positives = 14/25 (56%)
Query: 94 GRGGGYQEVVDIRIWRNGRQKRWRW 118
GRGGGY + R ++K WR+
Sbjct: 206 GRGGGYYDTFLKRYQDRAKEKGWRY 230
>At4g22020 glycine-rich protein
Length = 396
Score = 25.4 bits (54), Expect = 4.7
Identities = 9/14 (64%), Positives = 10/14 (71%)
Query: 86 GGNDSSGYGRGGGY 99
GG + SGYG G GY
Sbjct: 299 GGGNGSGYGSGSGY 312
>At1g07380 unknown protein
Length = 779
Score = 25.4 bits (54), Expect = 4.7
Identities = 9/17 (52%), Positives = 11/17 (63%)
Query: 85 CGGNDSSGYGRGGGYQE 101
CGG + YGRG GY +
Sbjct: 372 CGGKNEMCYGRGPGYPD 388
>At5g60980 ras-GTPase-activating protein SH3-domain binding protein
- like
Length = 460
Score = 24.6 bits (52), Expect = 8.1
Identities = 9/13 (69%), Positives = 9/13 (69%)
Query: 86 GGNDSSGYGRGGG 98
GG GYGRGGG
Sbjct: 402 GGGGRGGYGRGGG 414
>At2g19450 diacylglycerol O-acyltransferase (DAGAT)
Length = 520
Score = 24.6 bits (52), Expect = 8.1
Identities = 9/13 (69%), Positives = 11/13 (84%)
Query: 86 GGNDSSGYGRGGG 98
GG D++G GRGGG
Sbjct: 83 GGGDNNGGGRGGG 95
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.359 0.166 0.627
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,512,036
Number of Sequences: 26719
Number of extensions: 88232
Number of successful extensions: 910
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 845
Number of HSP's gapped (non-prelim): 61
length of query: 119
length of database: 11,318,596
effective HSP length: 95
effective length of query: 24
effective length of database: 8,780,291
effective search space: 210726984
effective search space used: 210726984
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0089a.1


