

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0087.11
(146 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
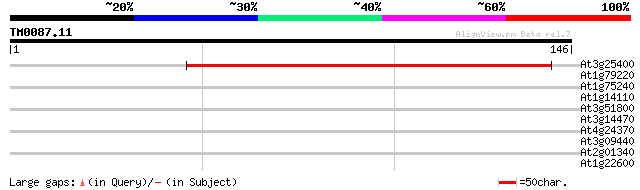
At3g25400 unknown protein 117 2e-27
At1g79220 unknown protein 27 2.9
At1g75240 unknown protein 27 2.9
At1g14110 hypothetical protein 27 2.9
At3g51800 G2p (AtG2) 27 5.0
At3g14470 disease resistance protein, putative 27 5.0
At4g24370 unknown protein 26 6.6
At3g09440 heat-shock protein (At-hsc70-3) 26 8.6
At2g01340 unknown protein 26 8.6
At1g22600 Similar to seed maturation protein PM27 26 8.6
>At3g25400 unknown protein
Length = 99
Score = 117 bits (294), Expect = 2e-27
Identities = 58/95 (61%), Positives = 73/95 (76%)
Query: 47 VGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALA 106
VGEVGELSEIFQWKGEVARG P+W ++K HL EELSDVLLYLVRL+D CG+DLG+AAL
Sbjct: 4 VGEVGELSEIFQWKGEVARGCPDWKEEEKVHLGEELSDVLLYLVRLSDACGVDLGKAALR 63
Query: 107 KIVKNAQKYPVTSAINNGNSKTGEKTTAVVKIDNQ 141
KI NA KYPV ++ GE +++ +K++ +
Sbjct: 64 KIELNAIKYPVPKKTDDHCVGDGEDSSSKIKLNEE 98
>At1g79220 unknown protein
Length = 399
Score = 27.3 bits (59), Expect = 2.9
Identities = 14/39 (35%), Positives = 17/39 (42%)
Query: 38 SPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKE 76
S R VGEVGE ++ + L W CDD E
Sbjct: 58 SSRRKKFDYVGEVGEKEKVRDITSSPLQVLRRWGCDDDE 96
>At1g75240 unknown protein
Length = 309
Score = 27.3 bits (59), Expect = 2.9
Identities = 12/40 (30%), Positives = 19/40 (47%)
Query: 19 DLSKRLAEFAEVRGWDQYHSPRNLLLALVGEVGELSEIFQ 58
D +R+ +FAE GW L GE+G ++F+
Sbjct: 250 DQKERMMDFAEKLGWRMNKQDEEELKRFCGEIGVKRQVFK 289
>At1g14110 hypothetical protein
Length = 435
Score = 27.3 bits (59), Expect = 2.9
Identities = 17/62 (27%), Positives = 29/62 (46%), Gaps = 7/62 (11%)
Query: 33 WDQYHSPRNLLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRL 92
W+ +N+ L GE E++Q GE + D K H ++ L+++ YL+ L
Sbjct: 295 WEYSDHLKNMFLEQASSTGETIEVYQPSGEKIQ-----QTDKKLHDQKALAEI--YLLSL 347
Query: 93 AD 94
D
Sbjct: 348 TD 349
>At3g51800 G2p (AtG2)
Length = 392
Score = 26.6 bits (57), Expect = 5.0
Identities = 11/27 (40%), Positives = 16/27 (58%)
Query: 7 NGFDGARDVSLQDLSKRLAEFAEVRGW 33
NG D +LQ+L K+ E E++GW
Sbjct: 325 NGSDRITSHTLQELPKKTIEDPEIKGW 351
>At3g14470 disease resistance protein, putative
Length = 1054
Score = 26.6 bits (57), Expect = 5.0
Identities = 23/81 (28%), Positives = 39/81 (47%), Gaps = 4/81 (4%)
Query: 42 LLLALVGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVRLADVCGLDLG 101
L L + E + + Q +G ++ G ++ + EHLE L V + L RLA + LG
Sbjct: 92 LRLNIGAESSSSNRLRQLRGRMSLG--DFLDGNSEHLETRLEKVTIRLERLASQRNI-LG 148
Query: 102 QAALAKIVKNAQKYPVTSAIN 122
L ++ Q+ P TS ++
Sbjct: 149 LKELTAMIPK-QRLPTTSLVD 168
>At4g24370 unknown protein
Length = 164
Score = 26.2 bits (56), Expect = 6.6
Identities = 11/30 (36%), Positives = 19/30 (62%)
Query: 47 VGEVGELSEIFQWKGEVARGLPNWSCDDKE 76
+GE E+ ++ QW + AR P+ S DD++
Sbjct: 117 IGEDAEVDKLIQWAIDAARLDPSPSSDDEQ 146
>At3g09440 heat-shock protein (At-hsc70-3)
Length = 649
Score = 25.8 bits (55), Expect = 8.6
Identities = 14/44 (31%), Positives = 24/44 (53%), Gaps = 7/44 (15%)
Query: 69 NWSCDDKEHLEEELSDVLLYLVRLADVCGLDLGQAALAKIVKNA 112
N+ +DKE EE+S ++L +R ++ +A L +KNA
Sbjct: 110 NYKGEDKEFSAEEISSMILIKMR-------EIAEAYLGTTIKNA 146
>At2g01340 unknown protein
Length = 225
Score = 25.8 bits (55), Expect = 8.6
Identities = 11/18 (61%), Positives = 14/18 (77%)
Query: 124 GNSKTGEKTTAVVKIDNQ 141
GNS G+KTT V+KID +
Sbjct: 2 GNSLGGKKTTKVMKIDGE 19
>At1g22600 Similar to seed maturation protein PM27
Length = 385
Score = 25.8 bits (55), Expect = 8.6
Identities = 17/67 (25%), Positives = 28/67 (41%), Gaps = 1/67 (1%)
Query: 44 LALVGEVGELSEIFQWKGEVARGLPNWSCDDKEHLEEELSDVLLYLVR-LADVCGLDLGQ 102
+ + E E+ E WK ARG N K H +++ + + VR + V LG
Sbjct: 130 MTVAREALEVEEKVSWKAREARGKVNERATKKAHRVQKVLEKVQIAVRGIGTVVATALGL 189
Query: 103 AALAKIV 109
+ +V
Sbjct: 190 TKIGSVV 196
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.132 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,003,350
Number of Sequences: 26719
Number of extensions: 112622
Number of successful extensions: 233
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 227
Number of HSP's gapped (non-prelim): 10
length of query: 146
length of database: 11,318,596
effective HSP length: 90
effective length of query: 56
effective length of database: 8,913,886
effective search space: 499177616
effective search space used: 499177616
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0087.11


