

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.5
(176 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
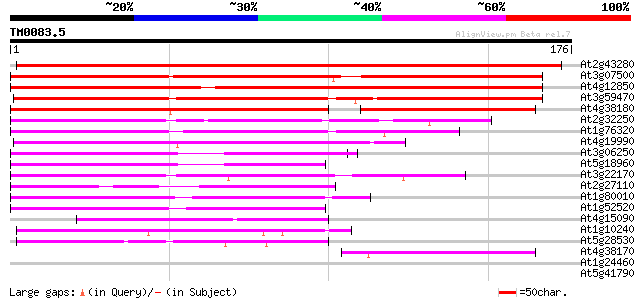
Score E
Sequences producing significant alignments: (bits) Value
At2g43280 unknown protein 286 5e-78
At3g07500 hypothetical protein 166 5e-42
At4g12850 putative protein 159 6e-40
At3g59470 unknown protein 142 7e-35
At4g38180 unknown protein (At4g38180) 111 2e-25
At2g32250 Mutator-like transposase 87 6e-18
At1g76320 putative phytochrome A signaling protein 79 1e-15
At4g19990 putative protein 72 2e-13
At3g06250 unknown protein 72 2e-13
At5g18960 FAR1 - like protein 71 3e-13
At3g22170 far-red impaired response protein, putative 71 3e-13
At2g27110 Mutator-like transposase 69 1e-12
At1g80010 hypothetical protein 62 2e-10
At1g52520 F6D8.26 60 6e-10
At4g15090 unknown protein 52 1e-07
At1g10240 unknown protein 47 5e-06
At5g28530 far-red impaired response protein (FAR1) - like 41 4e-04
At4g38170 hypothetical protein 40 5e-04
At1g24460 unknown protein 33 0.10
At5g41790 myosin heavy chain-like protein 31 0.30
>At2g43280 unknown protein
Length = 206
Score = 286 bits (731), Expect = 5e-78
Identities = 131/171 (76%), Positives = 156/171 (90%)
Query: 3 FESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIR 62
FESEDAAK+FYD+Y+R +GFVMRVMSCRRSE+DGRILARR GCNKEGHC++IRGK GS+R
Sbjct: 28 FESEDAAKMFYDDYSRRLGFVMRVMSCRRSEKDGRILARRFGCNKEGHCVSIRGKFGSVR 87
Query: 63 RPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQELTA 122
+PR STREGCKAMIH+KYD+SGKWVITKFVK+HNHPLVVSPREAR T+DEKDK+IQELT
Sbjct: 88 KPRPSTREGCKAMIHVKYDRSGKWVITKFVKEHNHPLVVSPREARHTLDEKDKRIQELTI 147
Query: 123 EIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFEPIEELL 173
E+R KKRLCA Y+EQL +F KIVEEHS++++ K+ ++VNNLKEFE +E L
Sbjct: 148 ELRNKKRLCAAYKEQLDAFAKIVEEHSNQIAKKVENVVNNLKEFEHLEHAL 198
>At3g07500 hypothetical protein
Length = 214
Score = 166 bits (421), Expect = 5e-42
Identities = 86/168 (51%), Positives = 119/168 (70%), Gaps = 8/168 (4%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEFESE+AAK FYD YA +GFVMRV + RRS RDG ++ RRL CNKEG R +
Sbjct: 45 MEFESEEAAKSFYDNYATCMGFVMRVDAFRRSMRDGTVVWRRLVCNKEGF-RRSRPRRSE 103
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLV-VSPREARQTMDEKDKKIQE 119
R+PRA TREGCKA+I +K +KSG W++TKF K+HNHPL+ +SP DEKD KI+E
Sbjct: 104 SRKPRAITREGCKALIVVKREKSGTWLVTKFEKEHNHPLLPLSPN------DEKDAKIRE 157
Query: 120 LTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFE 167
L+AE+ ++R C Q+QL +K +EEHS+ L++ I+ ++ ++++ E
Sbjct: 158 LSAELSRERRRCTALQQQLDMVLKEMEEHSNHLTININSVIQSVRDIE 205
>At4g12850 putative protein
Length = 183
Score = 159 bits (403), Expect = 6e-40
Identities = 80/167 (47%), Positives = 117/167 (69%), Gaps = 4/167 (2%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
++FESE+ AK FY EY++ +GFV+R+M RRS DGR LARRLGCNK+G N + S
Sbjct: 14 LKFESEEEAKDFYVEYSKRLGFVVRMMQRRRSGIDGRTLARRLGCNKQGFGPNNQRSSSS 73
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQEL 120
+S+REGCKA I +K +KSGKWV+T+F+K+HNH L + + +K++KI+EL
Sbjct: 74 ----SSSSREGCKATILVKMEKSGKWVVTRFIKEHNHSLQFIGSSSYDSFADKERKIKEL 129
Query: 121 TAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFE 167
T EI + RLC Y+++L SF+ VE ++++LS+K+ IV N+K+ E
Sbjct: 130 TEEIECQDRLCDVYRDRLVSFIDNVEHYTEELSLKVRDIVENVKKLE 176
>At3g59470 unknown protein
Length = 251
Score = 142 bits (359), Expect = 7e-35
Identities = 81/172 (47%), Positives = 107/172 (62%), Gaps = 11/172 (6%)
Query: 2 EFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSI 61
EFESE AA FY+ YA VGFV+RV RS DG + R+L CNKEG+ + K +
Sbjct: 75 EFESEAAAHGFYNAYATKVGFVIRVSKLSRSRHDGSPIGRQLVCNKEGY--RLPSKRDKV 132
Query: 62 RRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREAR------QTMDEKDK 115
R RA TR GCKAMI I+ + SGKWVITKFVK+HNH L+ P R Q +E D
Sbjct: 133 IRQRAETRVGCKAMILIRKENSGKWVITKFVKEHNHSLM--PGRVRRGCIYDQYPNEHD- 189
Query: 116 KIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKEFE 167
KIQEL ++ +K+ ATY+ L + +E+H++ LS +I HIV+N++ E
Sbjct: 190 KIQELMQQLAAEKKRAATYKRHLEMLFEQIEQHNESLSKRIQHIVDNVRNLE 241
>At4g38180 unknown protein (At4g38180)
Length = 788
Score = 111 bits (277), Expect = 2e-25
Identities = 56/102 (54%), Positives = 69/102 (66%), Gaps = 2/102 (1%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEG--HCLNIRGKL 58
+EFESE+AAK FY+ YAR +GF RV S RRS RDG I+ R+ C KEG + R K
Sbjct: 77 LEFESEEAAKAFYNSYARRIGFSTRVSSSRRSRRDGAIIQRQFVCAKEGFRNMNEKRTKD 136
Query: 59 GSIRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLV 100
I+RPR TR GCKA + +K SGKW+++ FVKDHNH LV
Sbjct: 137 REIKRPRTITRVGCKASLSVKMQDSGKWLVSGFVKDHNHELV 178
Score = 43.5 bits (101), Expect = 6e-05
Identities = 20/55 (36%), Positives = 34/55 (61%)
Query: 111 DEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLKE 165
DE DKKI +L E+ + R C Y+ L S +K +E+ ++S+K+ +I +LK+
Sbjct: 732 DEMDKKINQLRNELELANRKCEAYRTNLLSVLKEMEDQKLQVSIKVQNIKISLKD 786
>At2g32250 Mutator-like transposase
Length = 684
Score = 86.7 bits (213), Expect = 6e-18
Identities = 56/152 (36%), Positives = 85/152 (55%), Gaps = 11/152 (7%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
M+FES++AA FY EYAR VGF + + + RRS+R G+ + ++ C++ G R K +
Sbjct: 42 MDFESKEAAYYFYREYARSVGFGITIKASRRSKRSGKFIDVKIACSRFG---TKREKATA 98
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQEL 120
I PR+ + GCKA +H+K + KWVI FVK+HNH + P + ++ K+K
Sbjct: 99 I-NPRSCPKTGCKAGLHMKRKEDEKWVIYNFVKEHNHE--ICPDDFYVSVRGKNKP---- 151
Query: 121 TAEIRIKKRL-CATYQEQLSSFMKIVEEHSDK 151
+ IKK L A +E L ++ E DK
Sbjct: 152 AGALAIKKGLQLALEEEDLKLLLEHFMEMQDK 183
>At1g76320 putative phytochrome A signaling protein
Length = 670
Score = 79.3 bits (194), Expect = 1e-15
Identities = 46/142 (32%), Positives = 74/142 (51%), Gaps = 7/142 (4%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEFE+ + A +FY +YA+ VGF +S RRS + + C + G + +
Sbjct: 1 MEFETHEDAYLFYKDYAKSVGFGTAKLSSRRSRASKEFIDAKFSCIRYGS----KQQSDD 56
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKK-IQE 119
PRAS + GCKA +H+K GKW + FVK+HNH L+ P +A ++ + ++
Sbjct: 57 AINPRASPKIGCKASMHVKRRPDGKWYVYSFVKEHNHDLL--PEQAHYFRSHRNTELVKS 114
Query: 120 LTAEIRIKKRLCATYQEQLSSF 141
+ +R KK T + LS++
Sbjct: 115 NDSRLRRKKNTPLTDCKHLSAY 136
>At4g19990 putative protein
Length = 672
Score = 71.6 bits (174), Expect = 2e-13
Identities = 40/135 (29%), Positives = 67/135 (49%), Gaps = 13/135 (9%)
Query: 2 EFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHC---------- 51
EFES++ A FY EYA VGF + + RRS G+ + + C + G
Sbjct: 26 EFESKEEAFEFYKEYANSVGFTTIIKASRRSRMTGKFIDAKFVCTRYGSKKEDIDTGLGT 85
Query: 52 --LNIRGKLGSIRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQT 109
NI R R+S++ CKA +H+K + G+WV+ VK+HNH + ++ +
Sbjct: 86 DGFNIPQARKRGRINRSSSKTDCKAFLHVKRRQDGRWVVRSLVKEHNHEIFTGQADSLRE 145
Query: 110 MDEKDKKIQELTAEI 124
+ + +K+++L I
Sbjct: 146 LSGR-RKLEKLNGAI 159
>At3g06250 unknown protein
Length = 764
Score = 71.6 bits (174), Expect = 2e-13
Identities = 38/109 (34%), Positives = 57/109 (51%), Gaps = 14/109 (12%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
+EF++ + A+ +Y+ YA GF +R RS DG + +RR C+KEG LN
Sbjct: 32 LEFDTAEEARDYYNSYATRTGFKVRTGQLYRSRTDGTVSSRRFVCSKEGFQLN------- 84
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQT 109
+R GC A I ++ +GKWV+ + K+HNH L EA+ T
Sbjct: 85 -------SRTGCPAFIRVQRRDTGKWVLDQIQKEHNHDLGGHIEEAQTT 126
Score = 67.8 bits (164), Expect = 3e-12
Identities = 39/106 (36%), Positives = 53/106 (49%), Gaps = 14/106 (13%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
+EF S + A FY YA VGF +R+ RS+ DG I +RR C+KEG
Sbjct: 194 LEFNSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSKEGF---------- 243
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREA 106
+ +R GC A + IK SG W++ + KDHNH L + A
Sbjct: 244 ----QHPSRMGCGAYMRIKRQDSGGWIVDRLNKDHNHDLEPGKKNA 285
>At5g18960 FAR1 - like protein
Length = 788
Score = 71.2 bits (173), Expect = 3e-13
Identities = 36/99 (36%), Positives = 53/99 (53%), Gaps = 14/99 (14%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
+EF++ + A+ FY+ YA GF +R RS DG + +RR C+KEG LN
Sbjct: 47 LEFDTAEEAREFYNAYAARTGFKVRTGQLYRSRTDGTVSSRRFVCSKEGFQLN------- 99
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPL 99
+R GC A I ++ +GKWV+ + K+HNH L
Sbjct: 100 -------SRTGCTAFIRVQRRDTGKWVLDQIQKEHNHEL 131
Score = 65.5 bits (158), Expect = 1e-11
Identities = 37/99 (37%), Positives = 51/99 (51%), Gaps = 14/99 (14%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
+EF S + A FY YA VGF +R+ RS+ DG I +RR C++EG
Sbjct: 215 LEFGSANEACQFYQAYAEVVGFRVRIGQLFRSKVDGSITSRRFVCSREGF---------- 264
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPL 99
+ +R GC A + IK SG W++ + KDHNH L
Sbjct: 265 ----QHPSRMGCGAYMRIKRQDSGGWIVDRLNKDHNHDL 299
>At3g22170 far-red impaired response protein, putative
Length = 814
Score = 70.9 bits (172), Expect = 3e-13
Identities = 50/159 (31%), Positives = 73/159 (45%), Gaps = 24/159 (15%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEFES A FY EY+R +GF + + RRS+ + + C++ G R S
Sbjct: 74 MEFESHGEAYSFYQEYSRAMGFNTAIQNSRRSKTTREFIDAKFACSRYG---TKREYDKS 130
Query: 61 IRRPRAS---------------TREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPRE 105
RPRA + CKA +H+K GKWVI FV++HNH L+ +
Sbjct: 131 FNRPRARQSKQDPENMAGRRTCAKTDCKASMHVKRRPDGKWVIHSFVREHNHELLPA--- 187
Query: 106 ARQTMDEKDKKIQELTA-EIRIKKRLCATYQEQLSSFMK 143
Q + E+ +KI A + K + + + SSF K
Sbjct: 188 --QAVSEQTRKIYAAMAKQFAEYKTVISLKSDSKSSFEK 224
>At2g27110 Mutator-like transposase
Length = 851
Score = 69.3 bits (168), Expect = 1e-12
Identities = 38/102 (37%), Positives = 51/102 (49%), Gaps = 16/102 (15%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEF SE AK FYDEY+R +GF +++ DG + R C+ S
Sbjct: 53 MEFNSEKEAKSFYDEYSRQLGFTSKLLP----RTDGSVSVREFVCSS------------S 96
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVS 102
+R + E C AM+ I+ KWV+TKFVK+H H L S
Sbjct: 97 SKRSKRRLSESCDAMVRIELQGHEKWVVTKFVKEHTHGLASS 138
>At1g80010 hypothetical protein
Length = 696
Score = 61.6 bits (148), Expect = 2e-10
Identities = 39/113 (34%), Positives = 51/113 (44%), Gaps = 7/113 (6%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEFES D A FY+ YAR +GF +RV S L CN +G L L
Sbjct: 70 MEFESYDDAYSFYNSYARELGFAIRVKSSWTKRNSKEKRGAVLCCNCQGFKL-----LKD 124
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEK 113
R TR GC+AMI ++ +W + + DHNH P+ A + K
Sbjct: 125 AHSRRKETRTGCQAMIRLRLIHFDRWKVDQVKLDHNHSF--DPQRAHNSKSHK 175
>At1g52520 F6D8.26
Length = 703
Score = 60.1 bits (144), Expect = 6e-10
Identities = 35/99 (35%), Positives = 49/99 (49%), Gaps = 5/99 (5%)
Query: 1 MEFESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGS 60
MEFES D A +Y+ YA VGF +RV + R L C+ +G ++
Sbjct: 89 MEFESYDDAYNYYNCYASEVGFRVRVKNSWFKRRSKEKYGAVLCCSSQGF-----KRIND 143
Query: 61 IRRPRASTREGCKAMIHIKYDKSGKWVITKFVKDHNHPL 99
+ R R TR GC AMI ++ S +W + + DHNH L
Sbjct: 144 VNRVRKETRTGCPAMIRMRQVDSKRWRVVEVTLDHNHLL 182
>At4g15090 unknown protein
Length = 768
Score = 52.4 bits (124), Expect = 1e-07
Identities = 26/79 (32%), Positives = 40/79 (49%), Gaps = 1/79 (1%)
Query: 22 FVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIRRPRASTREGCKAMIHIKYD 81
F + + RRS++ + + C++ G S RR + CKA +H+K
Sbjct: 17 FTTSIKNSRRSKKTKDFIDAKFACSRYGVTPESESSGSSSRRSTVKKTD-CKASMHVKRR 75
Query: 82 KSGKWVITKFVKDHNHPLV 100
GKW+I +FVKDHNH L+
Sbjct: 76 PDGKWIIHEFVKDHNHELL 94
>At1g10240 unknown protein
Length = 680
Score = 47.0 bits (110), Expect = 5e-06
Identities = 35/109 (32%), Positives = 53/109 (48%), Gaps = 5/109 (4%)
Query: 3 FESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARR-LGCNKEGHCLNIRGKLGSI 61
F + D A FY +A+ GF +R + G+ L RR C++ G+ G
Sbjct: 54 FLTHDTAYEFYSTFAKRCGFSIRRHRTEGKDGVGKGLTRRYFVCHRAGNTPIKTLSEGKP 113
Query: 62 RRPRASTREGCKAMIHI-KYDKSG--KWVITKFVKDHNHPLVVSPREAR 107
+R R S+R GC+A + I K + G +W +T F HNH L + P + R
Sbjct: 114 QRNRRSSRCGCQAYLRISKLTELGSTEWRVTGFANHHNHEL-LEPNQVR 161
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 40.8 bits (94), Expect = 4e-04
Identities = 29/103 (28%), Positives = 46/103 (44%), Gaps = 8/103 (7%)
Query: 3 FESEDAAKVFYDEYARHVGFVMRVMSCRRSERDGRILARRLGCNKEGHCLNIRGKLGSIR 62
F ++D A +Y +AR GF +R S+ G + R C + G N K ++
Sbjct: 76 FTTDDEAFEYYSTFARKSGFSIRKARSTESQNLG-VYRRDFVCYRSG--FNQPRKKANVE 132
Query: 63 RPRA--STREGCKAMIHIK---YDKSGKWVITKFVKDHNHPLV 100
PR S R GC +++ D W +++F HNH L+
Sbjct: 133 HPRERKSVRCGCDGKLYLTKEVVDGVSHWYVSQFSNVHNHELL 175
>At4g38170 hypothetical protein
Length = 531
Score = 40.4 bits (93), Expect = 5e-04
Identities = 24/68 (35%), Positives = 39/68 (57%), Gaps = 7/68 (10%)
Query: 105 EARQTMD-------EKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIH 157
EAR+T + EK++ I ELTAE+ + C Y+ L S ++ +EE +LS+K+
Sbjct: 464 EARETANATNHPGGEKERTILELTAELERTGQRCEVYRANLLSILRDMEEQKFQLSLKVQ 523
Query: 158 HIVNNLKE 165
+ +LKE
Sbjct: 524 NARLSLKE 531
>At1g24460 unknown protein
Length = 1791
Score = 32.7 bits (73), Expect = 0.10
Identities = 23/112 (20%), Positives = 49/112 (43%), Gaps = 7/112 (6%)
Query: 72 CKAMIHIKYDKSGKWVITKFVKDHNHPLVVSPREARQTMDEKDKKIQELTAEIRIKKRLC 131
C + +K ++ + ++D N V ++ + + ++L AE+ ++K C
Sbjct: 259 CSELFELKQKEAAFFERLSHLEDENRNFVEQVNREKEMCESMRTEFEKLKAELELEKTKC 318
Query: 132 ATYQEQLSSFM---KIVEEHSD----KLSVKIHHIVNNLKEFEPIEELLHQT 176
+E+LS + K + ++ D +LS K + N L E + E L +
Sbjct: 319 TNTKEKLSMAVTKGKALVQNRDALKHQLSEKTTELANRLTELQEKEIALESS 370
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 31.2 bits (69), Expect = 0.30
Identities = 19/87 (21%), Positives = 39/87 (43%)
Query: 90 KFVKDHNHPLVVSPREARQTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHS 149
K ++ N + E +++ IQEL AE+ K + +LSS +++ E H
Sbjct: 178 KAAEEENKAISSKNVETMNKLEQTQNTIQELMAELGKLKDSHREKESELSSLVEVHETHQ 237
Query: 150 DKLSVKIHHIVNNLKEFEPIEELLHQT 176
S+ + + ++ + + L+QT
Sbjct: 238 RDSSIHVKELEEQVESSKKLVAELNQT 264
Score = 30.8 bits (68), Expect = 0.39
Identities = 17/71 (23%), Positives = 33/71 (45%)
Query: 105 EARQTMDEKDKKIQELTAEIRIKKRLCATYQEQLSSFMKIVEEHSDKLSVKIHHIVNNLK 164
E + + K+QEL E+ K + +LSSF+++ E H S ++ + ++
Sbjct: 524 EITDELKQAQSKVQELVTELAESKDTLTQKENELSSFVEVHEAHKRDSSSQVKELEARVE 583
Query: 165 EFEPIEELLHQ 175
E + L+Q
Sbjct: 584 SAEEQVKELNQ 594
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,858,870
Number of Sequences: 26719
Number of extensions: 151419
Number of successful extensions: 643
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 586
Number of HSP's gapped (non-prelim): 60
length of query: 176
length of database: 11,318,596
effective HSP length: 93
effective length of query: 83
effective length of database: 8,833,729
effective search space: 733199507
effective search space used: 733199507
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0083.5


