

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.15
(489 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
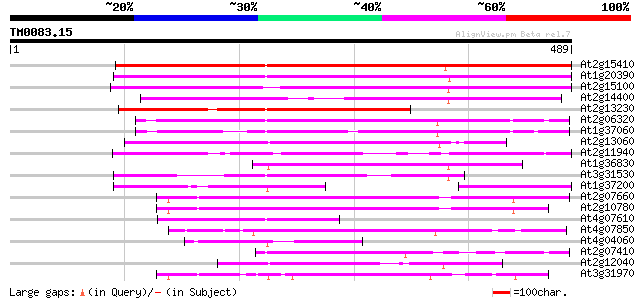
Score E
Sequences producing significant alignments: (bits) Value
At2g15410 putative retroelement pol polyprotein 337 1e-92
At1g20390 hypothetical protein 321 7e-88
At2g15100 putative retroelement pol polyprotein 300 1e-81
At2g14400 putative retroelement pol polyprotein 254 1e-67
At2g13230 pseudogene 216 2e-56
At2g06320 putative retroelement pol polyprotein 194 7e-50
At1g37060 Athila retroelment ORF 1, putative 179 4e-45
At2g13060 pseudogene 174 1e-43
At2g11940 putative retroelement gag/pol polyprotein 172 3e-43
At1g36830 hypothetical protein 171 8e-43
At3g31530 hypothetical protein 165 6e-41
At1g37200 hypothetical protein 133 3e-31
At2g07660 putative retroelement pol polyprotein 114 9e-26
At2g10780 pseudogene 112 5e-25
At4g07610 105 6e-23
At4g07850 putative polyprotein 100 2e-21
At4g04060 putative transposon protein 97 2e-20
At2g07410 putative retroelement pol polyprotein 97 2e-20
At2g12040 T10J7.2 96 3e-20
At3g31970 hypothetical protein 96 4e-20
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 337 bits (863), Expect = 1e-92
Identities = 175/399 (43%), Positives = 244/399 (60%), Gaps = 3/399 (0%)
Query: 93 LPEEKAEAKKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLG 152
+P++K A++L R++ YT+ ++ L + + LL CL + A V+ EIHEG+ G+H G
Sbjct: 1389 VPKDKWAARRLRARSAQYTLPHEHLLRWSATGVLLSCLDDEEAQQVMREIHEGAGGNHSG 1448
Query: 153 GRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGM 212
GR+LA K+ + G YWPT+ D V +C+ CQ HA H P L + + P+PF +W M
Sbjct: 1449 GRALALKIRKHGQYWPTMNADCEKFVARCEKCQGHAPFIHIPSEVLQTAMPPYPFMRWAM 1508
Query: 213 DLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLIT 272
D++ PF + KY++V DY+TKW EAE A I S VQ F +KN+I G+P ++T
Sbjct: 1509 DIIRPFPASRQ-KKYILVMTDYFTKWFEAESYARIQSKEVQNFVWKNIICHHGLPYEIVT 1567
Query: 273 DNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAE 332
DNGTQFTS F I+ ++ +PQ NGQAE+ N+ IL GL+KRL + KG WA+
Sbjct: 1568 DNGTQFTSLQFEGFCAKWKIRLSKSTPRYPQCNGQAEATNKTILDGLKKRLDEKKGAWAD 1627
Query: 333 QLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGF--DLNQNDQLLGED 390
+LD VLW+YRTTP +T +P+ LT+G EA+ P E P+ R + D NDQ+L ++
Sbjct: 1628 ELDGVLWSYRTTPRRSTDRTPFSLTYGMEALAPCEAGLPTVRRSMLINDPTLNDQMLLDN 1687
Query: 391 LDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAK 450
LD L E+R A R +Q AA YNKKV R FA GDLVL+ + GKL A
Sbjct: 1688 LDTLEEQRDQALLRIQNYQQAAAKFYNKKVKNRLFAEGDLVLRKVFENTVEQDAGKLGAH 1747
Query: 451 WEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNLRRYY 489
WEGPY + + G Y+L T+ G IPR+WN+ NL+RYY
Sbjct: 1748 WEGPYLISKVVKPGVYELLTMDGTPIPRSWNSANLKRYY 1786
>At1g20390 hypothetical protein
Length = 1791
Score = 321 bits (822), Expect = 7e-88
Identities = 167/401 (41%), Positives = 241/401 (59%), Gaps = 3/401 (0%)
Query: 91 GWLPEEKAEAKKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHH 150
G LP +K A++L +A+ YT++ + L K +L CL ++ E HEG+ G+H
Sbjct: 1391 GTLPSDKWTARRLRIKAAKYTLMKEHLLKVSAFGAMLNCLHGTEINEIMKETHEGAAGNH 1450
Query: 151 LGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQW 210
GGR+LA K+ + G+YWPT+ D +C+ CQRHA H P L + V+P+PF +W
Sbjct: 1451 SGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIHQPTELLRAGVAPYPFMRW 1510
Query: 211 GMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVL 270
MD++GP + ++++V DY+TKW+EAE ATI + VQ F +K +I R G+P +
Sbjct: 1511 AMDIVGPMPASRQ-KRFILVMTDYFTKWVEAESYATIRANDVQNFVWKFIICRHGLPYEI 1569
Query: 271 ITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDW 330
ITDNG+QF S F + I+ ++ +PQ NGQAE+ N+ IL GL+KRL + KG W
Sbjct: 1570 ITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAEATNKTILSGLKKRLDEKKGAW 1629
Query: 331 AEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLNQ--NDQLLG 388
A++LD VLW+YRTTP S T ++P+ +G EA+ PAE+ S R + N ND+++
Sbjct: 1630 ADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVKNPELNDRMML 1689
Query: 389 EDLDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLA 448
+ LD L E R A CR + AA YN+KV R F GDLVL+ GKL
Sbjct: 1690 DRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDVGDLVLRKVFENTAEINAGKLG 1749
Query: 449 AKWEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNLRRYY 489
A WEG Y+V + G Y+L T++G +PRTWN+ +L+RYY
Sbjct: 1750 ANWEGSYQVSKIVRPGDYELLTMSGTAVPRTWNSMHLKRYY 1790
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 300 bits (768), Expect = 1e-81
Identities = 157/403 (38%), Positives = 232/403 (56%), Gaps = 16/403 (3%)
Query: 89 LSGWLPEEKAEAKKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCG 148
L+G LP EK A+KL + + I ND L++R FS P C+ ++ V+ E+H+G+CG
Sbjct: 940 LNGTLPAEKWAARKLKATCARFCIANDILYRRIFSAPDAVCIFGEQTRTVMKEVHDGTCG 999
Query: 149 HHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFH 208
+H GGRSLA KV + GYYWPTL D + ++C+ CQ+HA L P L+++ +P+PF
Sbjct: 1000 NHTGGRSLAFKVRKYGYYWPTLVADCEAYARKCEQCQKHAPLILQPAELLTTVSAPYPFM 1059
Query: 209 QWGMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPG 268
+W MD++GP + T+ +EA + IT +V F +K++I R G+P
Sbjct: 1060 KWLMDIVGPLHVS--------------TRGVEAAAYSNITHVQVWNFIWKDIICRHGLPY 1105
Query: 269 VLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKG 328
++TDNG+QF S+ F + I+ ++ +PQ NGQAE+ N+ I+ L+K+L KG
Sbjct: 1106 EIVTDNGSQFISEQFEVFCEEWQIRLSHSTPRYPQGNGQAEAMNKTIISNLKKKLNAYKG 1165
Query: 329 DWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLN--QNDQL 386
W +L +VLWA RTTP T E+P+ L +G EAVIPAEI+ PSAR N +N+++
Sbjct: 1166 AWFGELQNVLWAVRTTPRRATDETPFSLIYGMEAVIPAEIKVPSARRIRNPQNETENNEM 1225
Query: 387 LGEDLDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGK 446
+ + +D + ERR A R A YN V RSF G LVL+ GK
Sbjct: 1226 IIDVIDTIDERRNRALARMQNYHNAGARYYNSNVRNRSFEVGTLVLRRVQQNKAEKGAGK 1285
Query: 447 LAAKWEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNLRRYY 489
L WEGPY++ G Y+L + GK + R WN+ +L+R+Y
Sbjct: 1286 LGISWEGPYKITHVVRNGVYRLINMEGKTVRRAWNSMHLKRFY 1328
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 254 bits (648), Expect = 1e-67
Identities = 136/369 (36%), Positives = 195/369 (51%), Gaps = 44/369 (11%)
Query: 115 DDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDA 174
+ L++RG S P L + V++E+HEG CG H GR++A K+ + YYWPT+ D
Sbjct: 951 NSLYRRGVSDPYLLSIFGPVVEIVMSEVHEGLCGSHSSGRAMAFKIKKMCYYWPTMITDC 1010
Query: 175 ADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDTAPG*LKYLVVAVDY 234
+ ++C CQ HA L H P SS+ +P+PF +W MD++GP + ++YL+V DY
Sbjct: 1011 VKYAQRCKRCQLHAPLIHQPSELFSSISAPYPFMRWSMDIIGPLHRSTRGVQYLLVLTDY 1070
Query: 235 YTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQ 294
++KWIEAE ++ +G IK
Sbjct: 1071 FSKWIEAEA----------------ILLNWG--------------------------IKV 1088
Query: 295 RFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPY 354
++++ +PQ NGQAE+AN+ IL L+KRL KG W ++L VLWAYRTTP +TGE+P+
Sbjct: 1089 SYSTLRYPQGNGQAEAANKTILSNLKKRLNHLKGGWYDELQPVLWAYRTTPRRSTGETPF 1148
Query: 355 RLTFGTEAVIPAEIREPSARTAGFDLN--QNDQLLGEDLDLLAERRAIANCRELIAKQEA 412
L +G +AV+PAE+ P R LN +N +L + LD + ERR A + + A
Sbjct: 1149 SLVYGMKAVVPAELNVPGLRRTEAPLNEKENSAMLDDSLDTINERRDQALIQIQNYQHAA 1208
Query: 413 AIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAKWEGPYRVVEATGTGAYKLETLA 472
A YN KV R F GD VL+ + GKL WEGPY V E G Y+L+ L
Sbjct: 1209 ARYYNSKVKSRPFFVGDYVLKRVFDNKKEEGAGKLGINWEGPYIVTEVVRNGVYRLKDLE 1268
Query: 473 GKEIPRTWN 481
+ + R WN
Sbjct: 1269 DRPVQRPWN 1277
>At2g13230 pseudogene
Length = 353
Score = 216 bits (551), Expect = 2e-56
Identities = 109/254 (42%), Positives = 162/254 (62%), Gaps = 8/254 (3%)
Query: 96 EKAEAKKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRS 155
+K EA+KL A+ + +V + LFK S PL+ C+ K+ A V+ E+H GSCG++ GGR+
Sbjct: 108 DKWEARKLKAEAARFVLVEEKLFKWRLSGPLMTCVEKEAARKVMKEVHGGSCGNNSGGRA 167
Query: 156 LARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLL 215
LA K+ R GY+WPT+ KD C+ CQR+A H P LSS+ SP+P +W MD++
Sbjct: 168 LAIKIKRLGYFWPTMIKD-------CEKCQRNAPTIHLPAELLSSIASPYPLMRWSMDII 220
Query: 216 GPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDNG 275
GP + K ++V DY++K EAE A+I A+V+ F +K+++ R GVP ++TDNG
Sbjct: 221 GPMHASKQ-KKLVLVLTDYFSKLREAESYASIKEAQVESFMWKHILCRHGVPYEIVTDNG 279
Query: 276 TQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLD 335
+QF S F+ D I+ ++ + Q NGQA++AN+ IL GL+KRL KG W ++L+
Sbjct: 280 SQFISTRFQGFCDKWGIRLSKSTPRYLQGNGQADAANKTILDGLKKRLTAKKGSWVDELE 339
Query: 336 HVLWAYRTTPHSTT 349
VLW++RTTP T
Sbjct: 340 GVLWSHRTTPRRAT 353
>At2g06320 putative retroelement pol polyprotein
Length = 466
Score = 194 bits (494), Expect = 7e-50
Identities = 120/382 (31%), Positives = 190/382 (49%), Gaps = 17/382 (4%)
Query: 110 YTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAGYYWPT 169
YT+ D +++ +++D +L H+ + G H K+L+AG++WPT
Sbjct: 79 YTLCKDKIYR--------SYVSEDEVEGILLHCHDSAYGGHFATFKTVSKILQAGFWWPT 130
Query: 170 LEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDTAPG*LKYLV 229
+ KDA + V +CD CQR ++ + ++ F WG+D +GPF ++ G KY++
Sbjct: 131 MFKDAQEFVSKCDSCQRKGNISRRNEMPQNPILEVDIFDVWGIDFMGPFPSSYG-NKYIL 189
Query: 230 VAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDG 289
V VDY +KW+EA T + V + F + RFGV V+I+D G F +K F +LL
Sbjct: 190 VVVDYVSKWVEAIASPTNDAKVVLKLFKTIIFPRFGVSWVVISDGGKHFINKVFENLLKK 249
Query: 290 LHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTT 349
+K + + HPQT+GQ E +NR I L K +G + DW+ +LD LWAY+T +
Sbjct: 250 HGVKHKVATPYHPQTSGQVEISNREIKTILEKTVGITRKDWSTKLDDALWAYKTAFKTPI 309
Query: 350 GESPYRLTFGTEAVIPAEIREP---SARTAGFDLNQNDQLLGEDLDLLAERRAIANCREL 406
G +P+ L + E+ + + FD+ ++ L L E R A
Sbjct: 310 GTTPFNLLCVKSCHLHVELEYKAMWAVKLLNFDIKTAEEKRLIQLSELDEIRLEAYESSK 369
Query: 407 IAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAKWEGPYRVVEATGTGAY 466
I K+ + ++KK++ + F GD VL S GKL ++W GP+ + E GA
Sbjct: 370 IYKERTKLFHDKKIITKDFQVGDQVLLFNS--RLKIFPGKLKSRWSGPFCITEVRPYGAV 427
Query: 467 KLETLAGKEIPRTWNATNLRRY 488
TLAGK T N L++Y
Sbjct: 428 ---TLAGKSGVFTVNGQRLKKY 446
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 179 bits (453), Expect = 4e-45
Identities = 117/382 (30%), Positives = 184/382 (47%), Gaps = 45/382 (11%)
Query: 110 YTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAGYYWPT 169
YT+ D +++ C+++D +L H + G H K+L+AG++WP+
Sbjct: 1375 YTLCKDKIYRT--------CVSEDEIEGILLHCHGFAYGGHFATFKTMSKILQAGFWWPS 1426
Query: 170 LEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDTAPG*LKYLV 229
+ KDA + + +CD + F WG+D +GPF ++ G KY++
Sbjct: 1427 MFKDAQEFISKCDSFEN--------------------FDVWGIDFMGPFPSSYG-NKYIL 1465
Query: 230 VAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDG 289
VA+DY +KW+EA T + V + F + RFGVP ++I+D G F +KGF +LL
Sbjct: 1466 VAIDYVSKWVEAIASHTNDARVVLKLFKTIIFPRFGVPRIVISDGGKHFINKGFENLLKK 1525
Query: 290 LHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTT 349
+K + T+GQ E +NR I L K +G + DW+ +L+ LWAYRT +
Sbjct: 1526 HGVKHK--------TSGQVEISNREIKAILEKTVGSTRKDWSAKLNDTLWAYRTAFKTPI 1577
Query: 350 GESPYRLTFGTEAVIPAEIREP---SARTAGFDLNQNDQLLGEDLDLLAERRAIANCREL 406
G +P+ L +G +P E+ + + FD+ ++ L+ L + R A
Sbjct: 1578 GTTPFNLLYGKSCHLPVELEYKAMWAVKLLNFDIKTAEEKRLIQLNDLNKIRLEAYESSK 1637
Query: 407 IAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAKWEGPYRVVEATGTGAY 466
I K+ ++KK+V R F GD VL S GKL ++W GP+ V T Y
Sbjct: 1638 IYKERTKSFHDKKIVSRDFKVGDQVLLFNS--RLRLFPGKLKSRWSGPFSV---TAVRPY 1692
Query: 467 KLETLAGKEIPRTWNATNLRRY 488
TLAGK T N L++Y
Sbjct: 1693 GAITLAGKNGDFTVNGQRLKKY 1714
>At2g13060 pseudogene
Length = 930
Score = 174 bits (440), Expect = 1e-43
Identities = 103/336 (30%), Positives = 164/336 (48%), Gaps = 10/336 (2%)
Query: 101 KKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHHLGGRSLARKV 160
K+ +R Y L+K + +C+A +L+ H S G H KV
Sbjct: 161 KRFLREIRRYQWDEPYLYKHSYDGIYRRCIAATEVPSILSHCHSSSYGGHFATFKTVSKV 220
Query: 161 LRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDT 220
L+A ++WPT+ +D + QCDPCQR + + + F +WG+D +GPF
Sbjct: 221 LQADFWWPTMFRDTQKFISQCDPCQRRGKISKRNEMPPNFRLEVEVFDRWGIDFMGPFPP 280
Query: 221 APG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTS 280
+ L Y++V VDY +KW+EA SA V + F + RFGVP ++I+D F +
Sbjct: 281 SNKNL-YILVDVDYVSKWVEAIASLKNDSAVVMKLFKSIIFPRFGVPRIVISDGDKHFIN 339
Query: 281 KGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWA 340
K LL ++ R + HPQT+GQ E +NR I L K +G AK +W+ +L LWA
Sbjct: 340 KILEKLLLQYGVQHRVATPYHPQTSGQVEVSNRQIKEILEKTVGKAKKEWSYKLYDALWA 399
Query: 341 YRTTPHSTTGESPYRLTFGTEAVIPAEIREPSA---RTAGFDLNQNDQLLGEDLDLLAER 397
Y+T + G +P+ L +G +P E+ +A + FD+ + +LD E
Sbjct: 400 YKTAFKTPLGTTPFHLLYGKACHLPVELEHKAAWAVKMMNFDIKSAGE---SELD---EI 453
Query: 398 RAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQ 433
R A + K+ ++KK++ R+F D +L+
Sbjct: 454 RIHAYDNSKLYKERTKAYHDKKILTRTFEPNDQILR 489
>At2g11940 putative retroelement gag/pol polyprotein
Length = 1212
Score = 172 bits (437), Expect = 3e-43
Identities = 118/400 (29%), Positives = 184/400 (45%), Gaps = 72/400 (18%)
Query: 90 SGWLPEEKAEAKKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGH 149
+G LP +K EA+KL + + + +D L++R S P C+ ++ V+ EIH+G+CG+
Sbjct: 884 NGVLPTDKWEARKLKATCARFCLADDILYRRIISAPDAICIFGEQTRTVIKEIHDGTCGN 943
Query: 150 HLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQ 209
H GGRSLA K+ + GY+WPT+ D C+ +A H +SP PF +
Sbjct: 944 HSGGRSLAFKLKKYGYFWPTMVAD----------CEAYA---HRSRRTFHFGISPLPFMK 990
Query: 210 WGMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGV 269
W MD++GP + +++L++ +W+EA + IT +V+ F +K +I R G+P
Sbjct: 991 WSMDIVGPLHVSTRGVRFLLL------EWVEAAAYSYITQVQVRHFIWKEIICRHGLPYE 1044
Query: 270 LITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGD 329
++TDNG QF S+ F+ I+ ++ +PQ N +A
Sbjct: 1045 IVTDNGPQFISEQFKAFYAEWQIRLNRSTPRYPQGNVEA--------------------- 1083
Query: 330 WAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLNQNDQLLGE 389
V+ A P + IR P A N++++ +
Sbjct: 1084 -------VIPAEIKVPRT------------------RRIRNPKNEAA------NNKMMVD 1112
Query: 390 DLDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAA 449
+D + ERR A R AA YN V RSF G LVL+ GKL
Sbjct: 1113 VIDTIDERRNRALVRMQNYHNAAAQYYNSNVRSRSFDVGTLVLRRIHQNTAEKGAGKLGI 1172
Query: 450 KWEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNLRRYY 489
WEGPY++ G L + GK + R WNA +L+R+Y
Sbjct: 1173 SWEGPYKITHIVRNGDC-LINMEGKTVRRAWNAAHLKRFY 1211
>At1g36830 hypothetical protein
Length = 243
Score = 171 bits (433), Expect = 8e-43
Identities = 94/240 (39%), Positives = 136/240 (56%), Gaps = 4/240 (1%)
Query: 212 MDLLGPFDTAPG*--LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGV 269
MD++GP + + G LK L+V DY TKWIEA+ +T +V+ F ++N++ + G+P
Sbjct: 1 MDVVGPMEASGGKKKLKNLLVLTDYSTKWIEAKAFQQVTEKQVEDFLWENIVCQHGIPYE 60
Query: 270 LITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGD 329
+ITDNGT TS+ + D I+ ++ +PQ NGQAE+AN+ IL ++K L K
Sbjct: 61 IITDNGTNLTSRKIKAFCDKWKIRLTTSTPHYPQGNGQAEAANKAILSNIKKILDSKKSM 120
Query: 330 WAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLN--QNDQLL 387
W++ L VLWAYRTTP +T E+P+ L +G EAVIP PS R N N Q+L
Sbjct: 121 WSDVLHGVLWAYRTTPRKSTQETPFSLAYGLEAVIPIVTIIPSVRRTASHANSDMNTQML 180
Query: 388 GEDLDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKL 447
+++D + ERR A R +Q YN + +R F G+LVL+ R GKL
Sbjct: 181 RDNMDFIDERRDQAMIRVQNYQQAVTRYYNSNIKIRRFEVGELVLRKVFSNTRELNAGKL 240
>At3g31530 hypothetical protein
Length = 831
Score = 165 bits (417), Expect = 6e-41
Identities = 103/308 (33%), Positives = 154/308 (49%), Gaps = 63/308 (20%)
Query: 91 GWLPEEKAEAKKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHH 150
G LP +K +A+KL +A+ + +V+ L+K S PL+ C+ + ++ EIH GS
Sbjct: 585 GKLPSDKWKARKLKAQAARFVLVDTKLYKWRLSGPLMTCVEAEAICKIMKEIHSGS---- 640
Query: 151 LGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQW 210
HA H P LSS+ SP+PF +W
Sbjct: 641 ------------------------------------HAPTIHQPAELLSSIASPYPFMRW 664
Query: 211 GMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVL 270
MD++GP + K ++V DY++KWIEAE A+I A+V+ F K+++ R G+P +
Sbjct: 665 SMDIIGPMHPSKQ-KKLVLVLTDYFSKWIEAESYASIKDAQVENFVLKHILCRHGIPYEI 723
Query: 271 ITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDW 330
+TDNG+QF S F+ D I+ ++ +PQ NGQAE+AN
Sbjct: 724 VTDNGSQFISTRFQGFCDKWGIRLSKSTPRYPQGNGQAEAAN------------------ 765
Query: 331 AEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIREPSARTAGFDLN--QNDQLLG 388
+L+ VLW +RTTP T E+P+ +GTE VIPAE+ PS R + N N Q
Sbjct: 766 --ELEGVLWLHRTTPRRATRETPFASIYGTECVIPAEMIVPSLRRSLSPENDPDNTQTPL 823
Query: 389 EDLDLLAE 396
++LDL+ E
Sbjct: 824 DELDLIDE 831
>At1g37200 hypothetical protein
Length = 1564
Score = 133 bits (334), Expect = 3e-31
Identities = 69/187 (36%), Positives = 104/187 (54%), Gaps = 12/187 (6%)
Query: 91 GWLPEEKAEAKKLVRRASWYTIVNDDLFKRGFSTPLLKCLAKDRAAYVLAEIHEGSCGHH 150
G P +K E +KL +++ Y I+ L KR + P + C + ++ +HEG CG H
Sbjct: 1259 GITPPDKWETRKLKAQSARYCIMEGRLMKRSVAGPYMVCTYGQQTKDLMKSMHEGQCGSH 1318
Query: 151 LGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQW 210
GR+L R ++ A D H +CD CQRHA H PP +SS+ SP+PF +W
Sbjct: 1319 YSGRTL-RCIMLA---------DCIAHSLRCDKCQRHAPTLHQPPEEMSSISSPYPFMKW 1368
Query: 211 GMDLLGPFDTAPG--*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPG 268
MD++ P + + G LK L+V DY TKWIE + +T +V+ F +N++ R G+P
Sbjct: 1369 SMDVVRPMEASGGKKRLKNLLVLTDYSTKWIEGKAFQQVTEKQVEVFLLENIVYRHGIPY 1428
Query: 269 VLITDNG 275
++TDNG
Sbjct: 1429 EIVTDNG 1435
Score = 68.6 bits (166), Expect = 8e-12
Identities = 37/99 (37%), Positives = 50/99 (50%), Gaps = 1/99 (1%)
Query: 392 DLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAKW 451
D + ERR A + +Q A YN + +R F G+LVL+ R GKL W
Sbjct: 1465 DFIDERRDHAMIQVQNYQQPAVRYYNSNIKIRRFEVGELVLRKVFSNTRELNAGKLGTNW 1524
Query: 452 EGPYRVVEATGTGAYKL-ETLAGKEIPRTWNATNLRRYY 489
EGPYR+ E G YKL + G R WNA +L++Y+
Sbjct: 1525 EGPYRITEVVRDGIYKLVKVFNGVPELRPWNAMHLKKYH 1563
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 114 bits (286), Expect = 9e-26
Identities = 100/370 (27%), Positives = 156/370 (42%), Gaps = 21/370 (5%)
Query: 129 CLAKDRAAY--VLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQR 186
C+ DRA +L E H+ H G + R + R Y+W + KD A V +C CQ
Sbjct: 544 CVPNDRALKEEILREAHQSKFSIHPGSNKMYRDLKRY-YHWVGMRKDVARWVAKCPTCQL 602
Query: 187 HADLHHAPPARLSSL-VSPWPFHQWGMDLLGPFDTAPG*LKYLV-VAVDYYTKWIEAEPL 244
H P L +L +S W + MD + T V V VD TK +
Sbjct: 603 VKAEHQVPSGLLQNLPISEWKWDHITMDFVTRLPTGIKSKHNAVWVVVDRLTKSAHFMAI 662
Query: 245 ATITSARV-QRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQ 303
+ A + + ++R G+P +++D T+FTSK + L + ++ HPQ
Sbjct: 663 SDKDGAEIIAEKYIDEIMRLHGIPVSIVSDRDTRFTSKFWNAFQKALGTRVNLSTAYHPQ 722
Query: 304 TNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAV 363
T+GQ+E + + LR + D G+W + L + +AY + ++ G SPY +G
Sbjct: 723 TDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLIEFAYNNSFQASIGMSPYEALYGRACR 782
Query: 364 IPAEIREPSARTAGFDLNQNDQLLGEDL-DLLAERRAIANCRELIAKQEAAIRYNKKVVL 422
P R +L G + D ER + A+ NK+
Sbjct: 783 TPLCWTPVGER----------RLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKE 832
Query: 423 RSFAAGDLVLQNASI---GARSTERGKLAAKWEGPYRVVEATGTGAYKLETLAGKEI-PR 478
F GDLV A R T R KL+ ++ GPY+V+E G AYKL+ +
Sbjct: 833 LEFQVGDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLDLPPKLNVFHN 892
Query: 479 TWNATNLRRY 488
++ + LR+Y
Sbjct: 893 VFHVSQLRKY 902
>At2g10780 pseudogene
Length = 1611
Score = 112 bits (280), Expect = 5e-25
Identities = 97/350 (27%), Positives = 148/350 (41%), Gaps = 20/350 (5%)
Query: 129 CLAKDRAAY--VLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQR 186
C+ DRA +L E H+ H G + R + R Y+W ++KD A V +C CQ
Sbjct: 1152 CVPNDRALKEEILREAHQSKFSIHPGSNKMYRDLKRY-YHWVGMKKDVARWVAKCPTCQL 1210
Query: 187 HADLHHAPPARLSSLVSP-WPFHQWGMDLLGPFDTAPG*LKYLV-VAVDYYTKWIEAEPL 244
H P L +L P W + MD + T V V VD TK +
Sbjct: 1211 VKAEHQVPSGLLQNLPIPEWKWDHITMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHFMAI 1270
Query: 245 ATITSARV-QRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQ 303
+ A + + ++R G+P +++D T+FTSK ++ L + ++ HPQ
Sbjct: 1271 SDKDGAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKAFQKALGTRVNLSTAYHPQ 1330
Query: 304 TNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAV 363
T+ Q+E + + LR + D G+W + L V +AY + ++ G SPY +G
Sbjct: 1331 TDEQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGRACR 1390
Query: 364 IPAEIREPSARTAGFDLNQNDQLLGEDL-DLLAERRAIANCRELIAKQEAAIRYNKKVVL 422
P R +L G + D ER + A+ NK+
Sbjct: 1391 TPLCWTPVGER----------RLFGPTIVDETTERMKFLKIKLKEAQDRQKSYANKRRKE 1440
Query: 423 RSFAAGDLVLQNASI---GARSTERGKLAAKWEGPYRVVEATGTGAYKLE 469
F GDLV A R T R KL+ ++ GPY+V+E G AYKL+
Sbjct: 1441 LEFQVGDLVYLKAMTYKGAGRFTSRKKLSPRYVGPYKVIERVGAVAYKLD 1490
>At4g07610
Length = 276
Score = 105 bits (262), Expect = 6e-23
Identities = 55/158 (34%), Positives = 88/158 (54%), Gaps = 1/158 (0%)
Query: 130 LAKDRAAYVLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHAD 189
+++D +L H + G + K+L+AG++WPT+ K+A + V +CD C R +
Sbjct: 76 VSEDEVEGILLHCHCSTYGGNFTTFKTVSKILQAGFWWPTMFKEAQEFVLKCDSCHRKGN 135
Query: 190 LHHAPPARLSSLVSPWPFHQWGMDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITS 249
+ + ++ F WG+D +GPF ++ G KY++VAVDY +KWIEA T +
Sbjct: 136 ISKRNEMPQNPILEVELFDVWGIDFIGPFPSSYG-NKYILVAVDYVSKWIEATASPTNDA 194
Query: 250 ARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLL 287
V + F + RFGVP V+I+D G F +K F +LL
Sbjct: 195 KVVLKLFKTIIFSRFGVPRVVISDGGKHFINKVFENLL 232
>At4g07850 putative polyprotein
Length = 1138
Score = 100 bits (248), Expect = 2e-21
Identities = 92/357 (25%), Positives = 152/357 (41%), Gaps = 22/357 (6%)
Query: 139 LAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARL 198
+ E H G H G S KV++ ++WP +++D ++C C++ A P
Sbjct: 763 IREAHGGGLMGHFGV-SKTLKVMQDHFHWPHMKRDVERMCERCTTCKQ-AKAKSQPHGLC 820
Query: 199 SSLVSPWPFHQWG---MDLLGPFDTAPG*LKYLVVAVDYYTKWIEAEPL-ATITSARVQR 254
+ L P P H W MD + + V VD ++K P T + +
Sbjct: 821 TPL--PIPLHPWNDISMDFVVGLPRTRTGKDSIFVVVDRFSKMAHFIPCHKTDDAMHIAN 878
Query: 255 FFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRV 314
F++ V+R G+P +++D T+F S ++ L L K F++ HPQT+GQ E NR
Sbjct: 879 LFFREVVRLHGMPKTIVSDRDTKFLSYFWKTLWSKLGTKLLFSTTCHPQTDGQTEVVNRT 938
Query: 315 ILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTEAVIPAEIR----- 369
+ LR + W + L HV +AY + HS T SP+++ +G + P ++
Sbjct: 939 LSTLLRALIKKNLKTWEDCLPHVEFAYNHSVHSATKFSPFQIVYGFNPITPLDLMPLPLI 998
Query: 370 EPSARTAGFDLNQNDQLLGE-DLDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAG 428
R F + +L+ D +R N L + E +R N V
Sbjct: 999 SIHLRKERFPNKRKSKLMPRMDGSFKVLKRINDNAYSLDLQDETDLRSNPFQV----GED 1054
Query: 429 DLVLQNASIGARSTERGKLAAKWEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNL 485
D+++ + GA+ +L + G + +E GA K E L E P T + T L
Sbjct: 1055 DVIMGSLDHGAKE----QLEPEEFGERKQLEDGELGAKKDEALHVLEGPITRSKTKL 1107
>At4g04060 putative transposon protein
Length = 375
Score = 97.4 bits (241), Expect = 2e-20
Identities = 57/158 (36%), Positives = 83/158 (52%), Gaps = 22/158 (13%)
Query: 153 GRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQRHADLHHAPPARLSSLVSPWPFHQWGM 212
GR + R V+ GY+WPT+ + H +CD CQRHA H P +SS+ SP+PF +W M
Sbjct: 186 GRLMKRSVV--GYFWPTMLANCIAHSLRCDKCQRHAPTLHQLPKEMSSISSPYPFMKWSM 243
Query: 213 DLLGPFDTAPG--*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVL 270
D++GP + + G LK L+V T +V+ F ++N++ R G+P +
Sbjct: 244 DVVGPMEASGGKKKLKNLLV-----------------TEKQVEDFLWENIVCRHGIPYEI 286
Query: 271 ITDNGTQFTSKGFRDLLDGLHIKQRF-TSVEHPQTNGQ 307
ITDNGT TS + D I+ T V H T+ Q
Sbjct: 287 ITDNGTNLTSGKIKAFCDKCKIRLTISTPVTHKVTDRQ 324
>At2g07410 putative retroelement pol polyprotein
Length = 411
Score = 97.1 bits (240), Expect = 2e-20
Identities = 83/278 (29%), Positives = 126/278 (44%), Gaps = 38/278 (13%)
Query: 215 LGPFDTAPG*LKYLVVAVDYYTKWIEAEPLATITSARVQRFFYKNVIRRFGVPGVLITDN 274
+G F ++ G KY++VAVDY +KW+EA T + V + F + RFGV V+I+D
Sbjct: 1 MGSFPSSYG-NKYILVAVDYVSKWVEAIVSPTNDAKVVLKLFKTIIFTRFGVHMVVISDG 59
Query: 275 GTQFTSKGFRDLLDGLHIKQRFTSVEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQL 334
F +K F +LL +K + + HPQT+GQ E +NR I L K +G + D + +L
Sbjct: 60 RKHFINKAFENLLKKHGVKHKVATSYHPQTSGQVEISNREIKAILEKTVGVTRKDLSAKL 119
Query: 335 DHVLWAYRT---TPHST-TGESPYRLTFGTEAVIPAEIREPSARTAGFDLNQNDQLLGED 390
D LWAY+T T T G +P+ L + +P E+ F +L+ D
Sbjct: 120 DDALWAYKTAFKTAFKTPIGTTPFNLLYAKSGHLPIELE--------FKAMWAVKLMNFD 171
Query: 391 LDLLAERRAIANCRELIAKQEAAIRYNKKVVLRSFAAGDLVLQNASIGARSTERGKLAAK 450
+ A++E I+ N +R L+ + KL ++
Sbjct: 172 IK--------------TAEEERLIQLNDLDEIR--------LEAYESSKNYKGKRKLKSR 209
Query: 451 WEGPYRVVEATGTGAYKLETLAGKEIPRTWNATNLRRY 488
GP+ V E GA TLA K T N L++Y
Sbjct: 210 LFGPFCVTEIRPYGAV---TLAVKSGDFTVNGQRLKKY 244
>At2g12040 T10J7.2
Length = 976
Score = 96.3 bits (238), Expect = 3e-20
Identities = 70/251 (27%), Positives = 113/251 (44%), Gaps = 24/251 (9%)
Query: 182 DPCQRHADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDTAPG*LKYLVVAVDYYTKWIEA 241
DPCQR + + ++ F WG++ +GP + L Y++V VDY +KW+EA
Sbjct: 544 DPCQRRGKISNCNEMPQKFILEVKVFDCWGINFMGPIPPSNKNL-YILVVVDYVSKWVEA 602
Query: 242 EPLATITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTSVEH 301
SA V + F + FGVP ++I+D G F +K LL ++ R + H
Sbjct: 603 IASPKNDSAVVMKLFKCIIFPHFGVPRIVISDGGKHFINKILTKLLLQYGVQHRVATPYH 662
Query: 302 PQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTFGTE 361
PQT+GQ E +NR I L K + A++T G +P+ L +G
Sbjct: 663 PQTSGQVEVSNRQIKEILEKTV----------------AFKT----PLGTTPFHLLYGKA 702
Query: 362 AVIPAEIREPSARTA---GFDLNQNDQLLGEDLDLLAERRAIANCRELIAKQEAAIRYNK 418
+P E+ +A T FD+ + L+ L E + A + K+ ++K
Sbjct: 703 CHLPVELEHKAAWTVKMLNFDIKSAGERRLIQLNELDEIQIHAYDNSKLYKERTKAYHDK 762
Query: 419 KVVLRSFAAGD 429
K++ R+F D
Sbjct: 763 KILTRTFEPKD 773
>At3g31970 hypothetical protein
Length = 1329
Score = 95.9 bits (237), Expect = 4e-20
Identities = 94/357 (26%), Positives = 155/357 (43%), Gaps = 36/357 (10%)
Query: 129 CLAKDRAAY--VLAEIHEGSCGHHLGGRSLARKVLRAGYYWPTLEKDAADHVKQCDPCQR 186
C+ KD +L+E H H + R + R Y W +++D A+ V +CD CQ
Sbjct: 895 CVPKDEELRREILSEAHASMFSIHPRATKMYRDLKRY-YQWVGMKRDVANWVTECDVCQL 953
Query: 187 HADLHHAPPARLSSLVSPWPFHQWGMDLLGPFDTAPG*-----LKYLVVAVDYYTKWIEA 241
H P L SL P +W D + D G + V VD TK A
Sbjct: 954 VKAEHQVPGGLLQSL----PILEWKWDFI-TMDFVVGLPVSRTKDAIWVIVDRLTK--SA 1006
Query: 242 EPLA---TITSARVQRFFYKNVIRRFGVPGVLITDNGTQFTSKGFRDLLDGLHIKQRFTS 298
LA T + + + + ++ GVP +++D ++FTS +R + K + ++
Sbjct: 1007 HFLAIRKTDGAVLLAKKYVSEIVELHGVPVSIVSDRDSKFTSAFWRAFQGEMGTKVQMST 1066
Query: 299 VEHPQTNGQAESANRVILRGLRKRLGDAKGDWAEQLDHVLWAYRTTPHSTTGESPYRLTF 358
HPQT+GQ+E + + LR + D G WA+ L V +AY + ++ +P+ +
Sbjct: 1067 AYHPQTDGQSERTIQTLEDMLRMCVLDRGGHWADHLSLVEFAYNNSYQASIRMAPFEALY 1126
Query: 359 GTEAVIP---AEIREPSARTAGFDLNQNDQLLGEDLDLLAERRAIANCRELIAKQEAAIR 415
G P ++ E S A + L +++ R N +E +Q +
Sbjct: 1127 GRPCRTPLCWTQVGERSIYGADYVLETTERI----------RVLKLNMKEAQDRQRSYAD 1176
Query: 416 YNKKVVLRSFAAGDLV-LQNASIGA--RSTERGKLAAKWEGPYRVVEATGTGAYKLE 469
++ + F GD V L+ A + RS KL ++ GP+R+VE G AY+LE
Sbjct: 1177 KRRREL--EFEVGDRVYLKMAMLRGPNRSISETKLTPRYMGPFRIVERVGPVAYRLE 1231
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.137 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,917,451
Number of Sequences: 26719
Number of extensions: 461460
Number of successful extensions: 1114
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 950
Number of HSP's gapped (non-prelim): 125
length of query: 489
length of database: 11,318,596
effective HSP length: 103
effective length of query: 386
effective length of database: 8,566,539
effective search space: 3306684054
effective search space used: 3306684054
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0083.15


