

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.13
(195 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
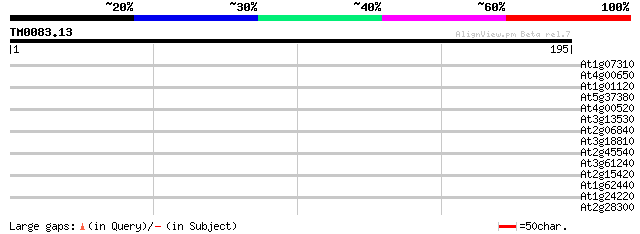
Score E
Sequences producing significant alignments: (bits) Value
At1g07310 unknown protein 33 0.096
At4g00650 Flowering Time gene FRIGIDA (FRI) 29 1.8
At1g01120 putative fatty acid elongase 3-ketoacyl-CoA synthase 1... 28 2.4
At5g37380 unknown protein 28 3.1
At4g00520 acetyl CoA thioesterase like protein 28 3.1
At3g13530 MAP3K epsilon protein kinase 28 3.1
At2g06840 putative retroelement pol polyprotein 28 4.0
At3g18810 protein kinase, putative 27 5.3
At2g45540 unknown protein 27 5.3
At3g61240 DEAD box RNA helicase RH12 27 6.9
At2g15420 hypothetical protein 27 6.9
At1g62440 putative extensin-like protein (gnl|PID|e1310400 27 6.9
At1g24220 hypothetical protein 27 6.9
At2g28300 unknown protein 27 9.0
>At1g07310 unknown protein
Length = 352
Score = 33.1 bits (74), Expect = 0.096
Identities = 14/50 (28%), Positives = 23/50 (46%)
Query: 17 VRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSP 66
++ + +H PP+ + + L P GN Y P PPQ +T+P
Sbjct: 135 IKDRPIPPPQHPPPRPQSQPLDYYSAPQGNHYYSPSPPPPPPPQAPITAP 184
>At4g00650 Flowering Time gene FRIGIDA (FRI)
Length = 609
Score = 28.9 bits (63), Expect = 1.8
Identities = 12/27 (44%), Positives = 16/27 (58%)
Query: 26 RHTPPKARGRGLAKQRDPPGNDTYHPK 52
RH+P + R GL+ QR P N + PK
Sbjct: 583 RHSPSEERYLGLSNQRSPRSNSSLDPK 609
>At1g01120 putative fatty acid elongase 3-ketoacyl-CoA synthase 1
(At1g01120)
Length = 528
Score = 28.5 bits (62), Expect = 2.4
Identities = 19/82 (23%), Positives = 37/82 (44%), Gaps = 4/82 (4%)
Query: 45 GNDTYHPKSQASHPPQDKMTSPEAYLKALRAEALDCNLYYSYQTQVTAPGRIRSLVSGLL 104
G++TY P+ S PP+ M+ A +A+ ALD ++ P + L+
Sbjct: 171 GDETYLPRGITSTPPKLNMSEARAEAEAVMFGALDS----LFEKTGIKPAEVGILIVNCS 226
Query: 105 TEDLDPQLKAGLLTFLELTEEM 126
+ P L A ++ ++ E++
Sbjct: 227 LFNPTPSLSAMIVNHYKMREDI 248
>At5g37380 unknown protein
Length = 431
Score = 28.1 bits (61), Expect = 3.1
Identities = 13/40 (32%), Positives = 22/40 (54%), Gaps = 1/40 (2%)
Query: 23 TAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDK 62
T R+ PP + A++ +PP T +P +Q ++PP K
Sbjct: 163 TTPRNNPPAQKTNPPAQKTNPPAQKT-NPPAQKNNPPTQK 201
>At4g00520 acetyl CoA thioesterase like protein
Length = 425
Score = 28.1 bits (61), Expect = 3.1
Identities = 23/76 (30%), Positives = 34/76 (44%), Gaps = 5/76 (6%)
Query: 105 TEDLDPQLKAGL-LTFLELTEEMWELHRRRLEVNMLIWGLEVSATEHKGEETGTM----G 159
T L+P + G+ + L L MW R + +L + +ATE +G TG M G
Sbjct: 340 TISLNPHRREGMSVAALSLDHSMWFHRPVRADDWLLFVIVSPTATESRGFATGKMFNRKG 399
Query: 160 EIEAELTSLLATREGL 175
E+ LT RE +
Sbjct: 400 ELVVSLTQEAVLREAV 415
>At3g13530 MAP3K epsilon protein kinase
Length = 1368
Score = 28.1 bits (61), Expect = 3.1
Identities = 16/43 (37%), Positives = 21/43 (48%), Gaps = 5/43 (11%)
Query: 28 TPPKARGRGLAKQRDPPG----NDTYHPKSQASH-PPQDKMTS 65
TP G LA+ DPPG +D +HP + S P + TS
Sbjct: 450 TPSSVSGNELARFSDPPGDASLHDLFHPLDKVSEGKPNEASTS 492
>At2g06840 putative retroelement pol polyprotein
Length = 1102
Score = 27.7 bits (60), Expect = 4.0
Identities = 15/44 (34%), Positives = 18/44 (40%)
Query: 7 KGHRPIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYH 50
KGHRP P K+ V K H+ P GL+ P H
Sbjct: 563 KGHRPKHPPVYLKDYVAHKVHSSPHTSSPGLSDSNVSPTVSANH 606
>At3g18810 protein kinase, putative
Length = 700
Score = 27.3 bits (59), Expect = 5.3
Identities = 12/34 (35%), Positives = 15/34 (43%)
Query: 28 TPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
TPP +Q PP +D+ P A PP D
Sbjct: 15 TPPSPSSSDNQQQSSPPPSDSSSPSPPAPPPPDD 48
>At2g45540 unknown protein
Length = 2946
Score = 27.3 bits (59), Expect = 5.3
Identities = 18/64 (28%), Positives = 27/64 (42%), Gaps = 7/64 (10%)
Query: 11 PIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSPEAYL 70
P P H+R+++ +R T K +PP N+ A P+DK + A L
Sbjct: 1769 PPTPSHLRRDSSMLERKTAKLQTFSSFQKPLEPPNNN-------APPRPRDKAAAKAAAL 1821
Query: 71 KALR 74
A R
Sbjct: 1822 AAAR 1825
>At3g61240 DEAD box RNA helicase RH12
Length = 498
Score = 26.9 bits (58), Expect = 6.9
Identities = 14/40 (35%), Positives = 17/40 (42%), Gaps = 3/40 (7%)
Query: 33 RGR---GLAKQRDPPGNDTYHPKSQASHPPQDKMTSPEAY 69
RGR G+ R P N YH + PPQD+ Y
Sbjct: 5 RGRYPPGVGTGRGAPPNPDYHQSYRQQQPPQDQQYVQRGY 44
>At2g15420 hypothetical protein
Length = 957
Score = 26.9 bits (58), Expect = 6.9
Identities = 19/54 (35%), Positives = 25/54 (46%)
Query: 140 IWGLEVSATEHKGEETGTMGEIEAELTSLLATREGLDRDLGAVHLRYNAMRVTP 193
I G E+ + GE +G GEI E L REGL G + +R + V P
Sbjct: 144 IVGEELGVCSNGGELSGGDGEINGEREELGGEREGLVGCEGEIGVRRVDVAVRP 197
>At1g62440 putative extensin-like protein (gnl|PID|e1310400
Length = 786
Score = 26.9 bits (58), Expect = 6.9
Identities = 21/70 (30%), Positives = 25/70 (35%), Gaps = 6/70 (8%)
Query: 24 AKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSPEAYLKALRAEALDCNLY 83
A + PP A Q PP Y+P AS PP P Y + +Y
Sbjct: 574 ATQSPPPPPPPTYYAVQSPPPPPPVYYPPVTASPPP------PPVYYTPVIQSPPPPPVY 627
Query: 84 YSYQTQVTAP 93
YS TQ P
Sbjct: 628 YSPVTQSPPP 637
>At1g24220 hypothetical protein
Length = 744
Score = 26.9 bits (58), Expect = 6.9
Identities = 17/54 (31%), Positives = 23/54 (42%), Gaps = 7/54 (12%)
Query: 24 AKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSPEAYLKALRAEA 77
A R PPKA P + T PK+ + PP+ K T P + + EA
Sbjct: 580 ASRTIPPKAT-------IPPKSSRTISPKANRTIPPKSKKTFPREAKRTIPREA 626
>At2g28300 unknown protein
Length = 2218
Score = 26.6 bits (57), Expect = 9.0
Identities = 14/33 (42%), Positives = 19/33 (57%), Gaps = 2/33 (6%)
Query: 10 RPIRPCHVRKEAVTAKRHTPPKARGRGLAKQRD 42
+P+ P VR ++ T K T P RGRG K+ D
Sbjct: 97 QPMEP--VRPQSHTLKEETQPIKRGRGRPKRTD 127
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,449,624
Number of Sequences: 26719
Number of extensions: 180695
Number of successful extensions: 493
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 482
Number of HSP's gapped (non-prelim): 19
length of query: 195
length of database: 11,318,596
effective HSP length: 94
effective length of query: 101
effective length of database: 8,807,010
effective search space: 889508010
effective search space used: 889508010
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0083.13


