

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079a.1
(139 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
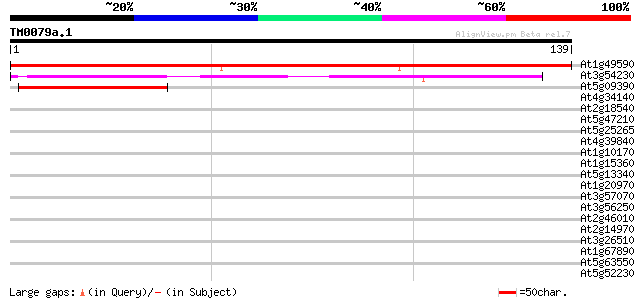
Score E
Sequences producing significant alignments: (bits) Value
At1g49590 unknown protein 164 1e-41
At3g54230 unknown protein 49 8e-07
At5g09390 unknown protein 47 4e-06
At4g34140 hypothetical protein 32 0.082
At2g18540 putative vicilin storage protein (globulin-like) 32 0.14
At5g47210 unknown protein 31 0.18
At5g25265 Unknown protein 31 0.18
At4g39840 unknown protein 31 0.24
At1g10170 hypothetical protein 31 0.24
At1g15360 ethylene responsive element like protein 30 0.31
At5g13340 putative protein 30 0.41
At1g20970 hypothetical protein 30 0.41
At3g57070 unknown protein 29 0.69
At3g56250 putative protein 29 0.69
At2g46010 hypothetical protein 29 0.69
At2g14970 En/Spm-like transposon protein 29 0.69
At3g26510 unknown protein 29 0.91
At1g67890 putative protein kinase (At1g67890) 29 0.91
At5g63550 unknown protein 28 1.2
At5g52230 unknown protein 28 1.5
>At1g49590 unknown protein
Length = 242
Score = 164 bits (415), Expect = 1e-41
Identities = 87/142 (61%), Positives = 105/142 (73%), Gaps = 3/142 (2%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTK-KASSTSQS 59
W +S+SGYYY++TNG +YD +SGFYYSD+IG WVTQ+EAYA+ +S TK S
Sbjct: 100 WMLDSASGYYYNQTNGLHYDSQSGFYYSDSIGHWVTQDEAYAAVKTSSGTKVPLVKKPVS 159
Query: 60 NKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLG--KRKRPGEKPKVVSEEEKSALKA 117
+ G P G PGR+VT SLNP R VK A SS+ LG KRKR EKPK VS EEK+ALKA
Sbjct: 160 SSGAGPSVGKPPGRLVTASLNPKRAVKGAASSVDLGNNKRKRQDEKPKKVSAEEKAALKA 219
Query: 118 REAARKRVQEREKSLLGLYSKP 139
REAARKRV++REK LLGLY++P
Sbjct: 220 REAARKRVEDREKPLLGLYNRP 241
>At3g54230 unknown protein
Length = 1007
Score = 48.9 bits (115), Expect = 8e-07
Identities = 34/133 (25%), Positives = 56/133 (41%), Gaps = 21/133 (15%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASPHFASTTKKASSTSQSN 60
WD +SGYYY +G+YYD SG YY G W + ++ T + Q+N
Sbjct: 586 WD--EASGYYYDAASGYYYDGNSGLYYDSNSGLWYSYDQ--------QTQQYVPCPDQNN 635
Query: 61 KGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPG-EKPKVVSEEEKSALKARE 119
+ N P + + K++ + + P EK + + ++A A
Sbjct: 636 ESKVTENQP----------DSAKKEKSSQQKVIISAATTPNVEKVLSLPDAVQAAAAAAI 685
Query: 120 AARKRVQEREKSL 132
A+ KR +ER K +
Sbjct: 686 ASEKREKERVKEI 698
>At5g09390 unknown protein
Length = 351
Score = 46.6 bits (109), Expect = 4e-06
Identities = 18/37 (48%), Positives = 23/37 (61%)
Query: 3 FESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEE 39
F+ SSGYYY + G+YYDP +G Y GKW +E
Sbjct: 298 FDESSGYYYSSSLGYYYDPNTGLYCYATTGKWYRYDE 334
>At4g34140 hypothetical protein
Length = 392
Score = 32.3 bits (72), Expect = 0.082
Identities = 14/34 (41%), Positives = 20/34 (58%)
Query: 5 SSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQE 38
S+SG+Y+ G+YY K G YY G++V E
Sbjct: 36 SNSGFYHDPNAGWYYCSKDGRYYKHENGEYVPLE 69
Score = 30.4 bits (67), Expect = 0.31
Identities = 15/41 (36%), Positives = 21/41 (50%), Gaps = 5/41 (12%)
Query: 4 ESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYASP 44
ESSS +GFY+DP +G+YY G++ E P
Sbjct: 32 ESSSS-----NSGFYHDPNAGWYYCSKDGRYYKHENGEYVP 67
>At2g18540 putative vicilin storage protein (globulin-like)
Length = 699
Score = 31.6 bits (70), Expect = 0.14
Identities = 17/35 (48%), Positives = 21/35 (59%)
Query: 96 KRKRPGEKPKVVSEEEKSALKAREAARKRVQEREK 130
+RKR E+ K EE K + E ARKR +EREK
Sbjct: 483 ERKREEEEAKRREEERKKREEEAEQARKREEEREK 517
>At5g47210 unknown protein
Length = 357
Score = 31.2 bits (69), Expect = 0.18
Identities = 22/75 (29%), Positives = 35/75 (46%), Gaps = 2/75 (2%)
Query: 60 NKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKARE 119
N G + + G + TS PT V+ +P + G E K ++ EEK+ +A E
Sbjct: 166 NGGGRGNWGTTEDDIPPTSEEPTTEVEKSPVAEKQGGEDETPEAKKELTAEEKAQKEAEE 225
Query: 120 AARKR--VQEREKSL 132
A + ++E EK L
Sbjct: 226 AEAREMTLEEYEKIL 240
>At5g25265 Unknown protein
Length = 366
Score = 31.2 bits (69), Expect = 0.18
Identities = 21/60 (35%), Positives = 25/60 (41%), Gaps = 2/60 (3%)
Query: 1 WDFESSSGYYYHKTNGFYYDPKSGFYYSDAIGKWVTQEEAYAS-PHFASTTKKASSTSQS 59
WD E Y H T G YD K Y IG+W + +Y S P + T SQS
Sbjct: 288 WDIEVGDKYIIHYTYGCDYDMKGKLTYG-KIGEWRFDKRSYDSKPPPRNLTMPPPGVSQS 346
>At4g39840 unknown protein
Length = 451
Score = 30.8 bits (68), Expect = 0.24
Identities = 19/85 (22%), Positives = 40/85 (46%)
Query: 45 HFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKP 104
H ++TK +S+T++++ K N T+S+ + ++ + SS + K P K
Sbjct: 123 HKLNSTKSSSNTTKTSSELKKLNSGTKSTNSTSSIKKSADLSKSSSSKNKTTIKPPSSKL 182
Query: 105 KVVSEEEKSALKAREAARKRVQERE 129
E+KS ++ + + E+E
Sbjct: 183 SSPPSEKKSQPSSKPVTKSKQSEKE 207
>At1g10170 hypothetical protein
Length = 1188
Score = 30.8 bits (68), Expect = 0.24
Identities = 22/77 (28%), Positives = 36/77 (46%), Gaps = 7/77 (9%)
Query: 43 SPHFASTTKKASSTSQSNKGNKPHNGPL-------PGRVVTTSLNPTRNVKAAPSSLSLG 95
SP S+T +S +S K N P + P + T++L P ++ + SS + G
Sbjct: 1109 SPIGGSSTDAQASALRSAKSNSPIVTSVNRWSVLEPKKASTSTLEPIAQIEESSSSKTTG 1168
Query: 96 KRKRPGEKPKVVSEEEK 112
K+ G +VV + EK
Sbjct: 1169 KQPVEGSGEEVVDDWEK 1185
>At1g15360 ethylene responsive element like protein
Length = 199
Score = 30.4 bits (67), Expect = 0.31
Identities = 24/78 (30%), Positives = 34/78 (42%), Gaps = 5/78 (6%)
Query: 42 ASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPG 101
AS H K+A S S S+ GP TTS + A + + +G G
Sbjct: 121 ASSHIGVWQKRAGSKSDSSWVMTVELGPASSSQETTSKASQDAILAPTTEVEIG-----G 175
Query: 102 EKPKVVSEEEKSALKARE 119
+ +V+ EEEK AL+ E
Sbjct: 176 SREEVLDEEEKVALQMIE 193
>At5g13340 putative protein
Length = 242
Score = 30.0 bits (66), Expect = 0.41
Identities = 23/85 (27%), Positives = 40/85 (47%), Gaps = 3/85 (3%)
Query: 50 TKKASSTSQSNKGNKPHNGPLPG---RVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKV 106
T++ S S +K P P R ++SL+P+ + A + R + ++
Sbjct: 30 TRRDRSRSPYTSRHKKSRSPAPRQHQRDRSSSLSPSEHRIAIEVKKEQEDKARLQHEAEL 89
Query: 107 VSEEEKSALKAREAARKRVQEREKS 131
EE++A + EA RK V+ER K+
Sbjct: 90 KRLEEETAQRIEEAVRKNVEERMKT 114
>At1g20970 hypothetical protein
Length = 1498
Score = 30.0 bits (66), Expect = 0.41
Identities = 21/82 (25%), Positives = 41/82 (49%), Gaps = 4/82 (4%)
Query: 47 ASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKV 106
A T++ S T++S K KP P +VT ++ ++ + + K E+ ++
Sbjct: 1162 AKPTEQKSQTTKSKKAVKPDQPP---SIVTELVSGKEEIEKSATPEEEEPPKLTKEEEEL 1218
Query: 107 VSEEEKSALKAREAARKRVQER 128
+ +EE+ K +EAA+ + Q R
Sbjct: 1219 IKKEEEKR-KQKEAAKMKEQHR 1239
>At3g57070 unknown protein
Length = 417
Score = 29.3 bits (64), Expect = 0.69
Identities = 27/84 (32%), Positives = 38/84 (45%), Gaps = 7/84 (8%)
Query: 38 EEAYAS---PHFASTTKKA-SSTSQSNKGN--KPHNGPLPGRVVTTSLNPTRNVKAAPSS 91
EE++ S P S+ KKA SS SN N PH P + S NP+ + P S
Sbjct: 190 EESFLSDLDPSIVSSYKKALSSKLLSNHSNTRNPHR-PTKSLSCSPSSNPSILISEEPKS 248
Query: 92 LSLGKRKRPGEKPKVVSEEEKSAL 115
+S + KP++ E+K L
Sbjct: 249 VSSSQLISSPAKPRLPGTEDKIVL 272
>At3g56250 putative protein
Length = 222
Score = 29.3 bits (64), Expect = 0.69
Identities = 27/95 (28%), Positives = 39/95 (40%), Gaps = 7/95 (7%)
Query: 42 ASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKR-- 99
ASP A + + +S S +K P V + + K A SS S R R
Sbjct: 91 ASPASAPS-RPQTSRSDMGVSDKDFTEPWSVYTVRRKASGQKRSKDASSSTSAAARIRLD 149
Query: 100 ----PGEKPKVVSEEEKSALKAREAARKRVQEREK 130
G+K + SE +K LK R+R ++EK
Sbjct: 150 LSSSSGKKTRQASENKKKTLKVAPKKRQRTVKQEK 184
>At2g46010 hypothetical protein
Length = 942
Score = 29.3 bits (64), Expect = 0.69
Identities = 22/87 (25%), Positives = 43/87 (49%), Gaps = 7/87 (8%)
Query: 48 STTKKASSTSQSNKGNKPH--NGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPK 105
++TKK+ ++ + +P N P P V + P N SSL G+ + ++
Sbjct: 393 ASTKKSLGPAEHLQMQQPRQLNTPTPNLVAPSDTGPLSN-----SSLQSGQGTQQAQQRS 447
Query: 106 VVSEEEKSALKAREAARKRVQEREKSL 132
++++ LKA+ A +R+++ E SL
Sbjct: 448 GFTKQQLHVLKAQILAFRRLKKGEGSL 474
>At2g14970 En/Spm-like transposon protein
Length = 771
Score = 29.3 bits (64), Expect = 0.69
Identities = 36/116 (31%), Positives = 53/116 (45%), Gaps = 15/116 (12%)
Query: 30 AIGKWVTQEEAYASPHFASTTKKASSTSQSNK------GNKPHNGPLPGRVVTTSLNPTR 83
A+ K Q P +++ K+++ SQ N G K NG G+ + N +
Sbjct: 577 AVQKAPLQSSQSTYPTTSASKGKSAAASQQNGATEKSLGEKQQNGATKGK---SGHNSSI 633
Query: 84 NVKAAPSSLSLGK-RKRPGEK-PKVV--SEEEKSALKAREAARKRVQEREKSLLGL 135
V AA S S G K GEK P VV S +K+ L ++ A R+ R+K+ GL
Sbjct: 634 QVSAAKGSESAGTIAKSLGEKQPTVVPKSHVKKTQLASQTALRR--SPRQKTSEGL 687
>At3g26510 unknown protein
Length = 196
Score = 28.9 bits (63), Expect = 0.91
Identities = 14/35 (40%), Positives = 20/35 (57%)
Query: 47 ASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNP 81
+STT SS+ +S +KP P P R+ T + NP
Sbjct: 139 SSTTSSTSSSPRSPSLSKPPLPPSPPRITTVTKNP 173
Score = 27.3 bits (59), Expect = 2.6
Identities = 16/53 (30%), Positives = 26/53 (48%), Gaps = 2/53 (3%)
Query: 58 QSNKGNKPHNGPL--PGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVVS 108
+S G+K PL P T+S + T + ++P S SL K P P++ +
Sbjct: 116 KSAAGSKKSPPPLALPSSTTTSSSSTTSSTSSSPRSPSLSKPPLPPSPPRITT 168
>At1g67890 putative protein kinase (At1g67890)
Length = 765
Score = 28.9 bits (63), Expect = 0.91
Identities = 15/63 (23%), Positives = 28/63 (43%)
Query: 30 AIGKWVTQEEAYASPHFASTTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAP 89
++ WV E+ H +++ S S +++ NKP N G V S + + +
Sbjct: 405 SVWPWVHNEQQKEEAHHSNSYNSVKSESLASESNKPANNENMGSVNVNSASSASSCGSTS 464
Query: 90 SSL 92
SS+
Sbjct: 465 SSV 467
>At5g63550 unknown protein
Length = 530
Score = 28.5 bits (62), Expect = 1.2
Identities = 20/91 (21%), Positives = 37/91 (39%), Gaps = 2/91 (2%)
Query: 48 STTKKASSTSQSNKGNKPHNGPLPGRVVTTSLNPTRNVKAAPSSLSLGKRKRPGEKPKVV 107
S+ K S KG+ ++ ++ +P + K S K K+ KP+
Sbjct: 350 SSGSKGKDKQPSAKGSARSGEKSSKQIAKSTSSPAKKQKVDHVESSKEKSKKQPSKPQAK 409
Query: 108 SEEEKSALKAREAARKRVQEREKSLLGLYSK 138
+EK KA + + + + K +L + SK
Sbjct: 410 GSKEKG--KATKKGKAKAEPTRKEMLEVVSK 438
>At5g52230 unknown protein
Length = 746
Score = 28.1 bits (61), Expect = 1.5
Identities = 13/49 (26%), Positives = 25/49 (50%)
Query: 84 NVKAAPSSLSLGKRKRPGEKPKVVSEEEKSALKAREAARKRVQEREKSL 132
N K+ S++ R + +K V+ EEE+ + R +V+E++ L
Sbjct: 213 NAKSEKDSVNSSVRSQKPKKEAVMKEEEEQDSSEKRITRSKVEEKKNEL 261
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.306 0.125 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,342,271
Number of Sequences: 26719
Number of extensions: 136180
Number of successful extensions: 662
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 600
Number of HSP's gapped (non-prelim): 93
length of query: 139
length of database: 11,318,596
effective HSP length: 89
effective length of query: 50
effective length of database: 8,940,605
effective search space: 447030250
effective search space used: 447030250
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0079a.1


