

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.9
(351 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
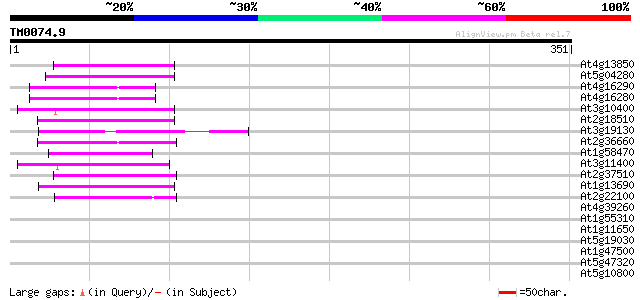
Score E
Sequences producing significant alignments: (bits) Value
At4g13850 glycine-rich RNA-binding protein AtGRP2 - like 45 6e-05
At5g04280 RNA-binding protein-like 44 1e-04
At4g16290 FCA delta protein 43 2e-04
At4g16280 FCA gamma protein 43 2e-04
At3g10400 hypothetical protein 43 3e-04
At2g18510 putative spliceosome associated protein 43 3e-04
At3g19130 nuclear acid binding protein, putative 42 4e-04
At2g36660 putative poly(A) binding protein 42 4e-04
At1g58470 unknown protein 42 4e-04
At3g11400 eukaryotic translation initiation factor 3 subunit lik... 42 7e-04
At2g37510 putative RNA-binding protein 42 7e-04
At1g13690 AtE1 (atE1) 42 7e-04
At2g22100 putative RNA-binding protein 41 9e-04
At4g39260 glycine-rich protein (clone AtGRP8) 40 0.001
At1g55310 Serine/arginine-rich protein 40 0.001
At1g11650 putative DNA binding protein 40 0.001
At5g19030 unknown protein 40 0.002
At1g47500 unknown protein 40 0.002
At5g47320 40S ribosomal protein S19 40 0.002
At5g10800 putative protein 40 0.002
>At4g13850 glycine-rich RNA-binding protein AtGRP2 - like
Length = 158
Score = 45.1 bits (105), Expect = 6e-05
Identities = 26/76 (34%), Positives = 36/76 (47%)
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LF+ GL T + +R F FG VV V + R FGFV F + AA+
Sbjct: 36 KLFIGGLSWGTDDASLRDAFAHFGDVVDAKVIVDRETGRSRGFGFVNFNDEGAATAAISE 95
Query: 88 LHGTRLNSFFLSINPA 103
+ G LN + +NPA
Sbjct: 96 MDGKELNGRHIRVNPA 111
>At5g04280 RNA-binding protein-like
Length = 310
Score = 44.3 bits (103), Expect = 1e-04
Identities = 23/81 (28%), Positives = 38/81 (46%)
Query: 23 AEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGI 82
A+E R+FV GL + + + F RFG ++ + R FGF+ F +R
Sbjct: 3 AKEGSRIFVGGLSPEVTDRDLERAFSRFGDILDCQIMLERDTGRSRGFGFITFADRRAMD 62
Query: 83 AAMRSLHGTRLNSFFLSINPA 103
++R +HG +S+N A
Sbjct: 63 ESIREMHGRDFGDRVISVNRA 83
>At4g16290 FCA delta protein
Length = 533
Score = 43.1 bits (100), Expect = 2e-04
Identities = 25/79 (31%), Positives = 42/79 (52%), Gaps = 1/79 (1%)
Query: 13 RWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGF 72
R++ ER + +LFV L++ +V ++F +FG V V++ + R R GF
Sbjct: 197 RYADGERERIGTLEFKLFVGSLNKQATEKEVEEIFLQFGHVEDVYLMRDEYRQSR-GCGF 255
Query: 73 VLFQSKRDGIAAMRSLHGT 91
V + SK +AA+ L+GT
Sbjct: 256 VKYSSKETAMAAIDGLNGT 274
Score = 37.4 bits (85), Expect = 0.012
Identities = 18/63 (28%), Positives = 35/63 (54%)
Query: 27 IRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMR 86
++LFV + R ++R FE+ G V+ V + K + +++ FV + + +D A+R
Sbjct: 120 VKLFVGSVPRTATEEEIRPYFEQHGNVLEVALIKDKRTGQQQGCCFVKYATSKDADRAIR 179
Query: 87 SLH 89
+LH
Sbjct: 180 ALH 182
>At4g16280 FCA gamma protein
Length = 747
Score = 43.1 bits (100), Expect = 2e-04
Identities = 25/79 (31%), Positives = 42/79 (52%), Gaps = 1/79 (1%)
Query: 13 RWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGF 72
R++ ER + +LFV L++ +V ++F +FG V V++ + R R GF
Sbjct: 197 RYADGERERIGTLEFKLFVGSLNKQATEKEVEEIFLQFGHVEDVYLMRDEYRQSR-GCGF 255
Query: 73 VLFQSKRDGIAAMRSLHGT 91
V + SK +AA+ L+GT
Sbjct: 256 VKYSSKETAMAAIDGLNGT 274
Score = 37.4 bits (85), Expect = 0.012
Identities = 18/63 (28%), Positives = 35/63 (54%)
Query: 27 IRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMR 86
++LFV + R ++R FE+ G V+ V + K + +++ FV + + +D A+R
Sbjct: 120 VKLFVGSVPRTATEEEIRPYFEQHGNVLEVALIKDKRTGQQQGCCFVKYATSKDADRAIR 179
Query: 87 SLH 89
+LH
Sbjct: 180 ALH 182
>At3g10400 hypothetical protein
Length = 261
Score = 42.7 bits (99), Expect = 3e-04
Identities = 33/111 (29%), Positives = 46/111 (40%), Gaps = 13/111 (11%)
Query: 6 SARAPPPRWSCHERSEVAEENI-------------RLFVDGLDRDTKHSQVRKLFERFGT 52
S APPP H+ S A+ + L+V LD +S + LF FG
Sbjct: 23 SVAAPPPSNPKHQPSSSAKSSAPGGGSGGLAPSKSTLYVSNLDFSLTNSDIHTLFSTFGK 82
Query: 53 VVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPA 103
V V V K + FVL+ S+ D A RS+ LN L+++ A
Sbjct: 83 VARVTVLKDRHTRQSRGVAFVLYVSREDAAKAARSMDAKILNGRKLTVSIA 133
>At2g18510 putative spliceosome associated protein
Length = 363
Score = 42.7 bits (99), Expect = 3e-04
Identities = 23/86 (26%), Positives = 46/86 (52%)
Query: 18 ERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
+ S ++ ++V GLD + +LF + G VV+V+VPK + +GF+ ++S
Sbjct: 16 QHSAERNQDATVYVGGLDAQLSEELLWELFVQAGPVVNVYVPKDRVTNLHQNYGFIEYRS 75
Query: 78 KRDGIAAMRSLHGTRLNSFFLSINPA 103
+ D A++ L+ +L+ + +N A
Sbjct: 76 EEDADYAIKVLNMIKLHGKPIRVNKA 101
Score = 30.0 bits (66), Expect = 2.0
Identities = 23/84 (27%), Positives = 38/84 (44%), Gaps = 5/84 (5%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIK---RVRREKFGFVLFQSKRDGIAAM 85
LF+ LD D + F FG + S PK ++ FGF+ + S AA+
Sbjct: 114 LFIGNLDPDVDEKLLYDTFSAFGVIAS--NPKIMRDPDTGNSRGFGFISYDSFEASDAAI 171
Query: 86 RSLHGTRLNSFFLSINPARFVDRK 109
S+ G L++ ++++ A D K
Sbjct: 172 ESMTGQYLSNRQITVSYAYKKDTK 195
>At3g19130 nuclear acid binding protein, putative
Length = 435
Score = 42.4 bits (98), Expect = 4e-04
Identities = 32/131 (24%), Positives = 59/131 (44%), Gaps = 20/131 (15%)
Query: 19 RSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSK 78
+S+ N +FV G+D D +R+ F +FG VVSV +P + GFV F +
Sbjct: 313 QSDGESTNATIFVGGIDPDVIDEDLRQPFSQFGEVVSVKIPV------GKGCGFVQFADR 366
Query: 79 RDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRV 138
+ A+ SL+GT + + ++ R ++++ R Q+W +
Sbjct: 367 KSAEDAIESLNGTVIGKNTVRLSWGRSPNKQW--------------RGDSGQQWNGGYSR 412
Query: 139 GAPFSGTGGFS 149
G ++ GG++
Sbjct: 413 GHGYNNGGGYA 423
>At2g36660 putative poly(A) binding protein
Length = 609
Score = 42.4 bits (98), Expect = 4e-04
Identities = 25/87 (28%), Positives = 44/87 (49%), Gaps = 1/87 (1%)
Query: 18 ERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
+R + E+ L++ LD D +R+ F FG +VS+ + K R+ R + FV F +
Sbjct: 192 DRVKPEEKYTNLYMKNLDADVSEDLLREKFAEFGKIVSLAIAKDENRLCR-GYAFVNFDN 250
Query: 78 KRDGIAAMRSLHGTRLNSFFLSINPAR 104
D A +++GT+ S L + A+
Sbjct: 251 PEDARRAAETVNGTKFGSKCLYVGRAQ 277
Score = 35.4 bits (80), Expect = 0.047
Identities = 18/74 (24%), Positives = 39/74 (52%), Gaps = 1/74 (1%)
Query: 17 HERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQ 76
HE ++ + ++V ++ ++RK F + GT+ S + ++ + + FGFV F
Sbjct: 294 HEEQKMIAKVSNIYVKNVNVAVTEEELRKHFSQCGTITSTKL-MCDEKGKSKGFGFVCFS 352
Query: 77 SKRDGIAAMRSLHG 90
+ + I A+++ HG
Sbjct: 353 TPEEAIDAVKTFHG 366
Score = 34.3 bits (77), Expect = 0.10
Identities = 26/91 (28%), Positives = 47/91 (51%), Gaps = 10/91 (10%)
Query: 1 MRPTGSARAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPK 60
+R S RAP R R+ V +FV L ++ ++ +F++FG +VS V
Sbjct: 95 IRVMWSVRAPDAR-----RNGVGN----VFVKNLPESVTNAVLQDMFKKFGNIVSCKV-A 144
Query: 61 TIKRVRREKFGFVLFQSKRDGIAAMRSLHGT 91
T++ + +GFV F+ + AA+++L+ T
Sbjct: 145 TLEDGKSRGYGFVQFEQEDAAHAAIQTLNST 175
>At1g58470 unknown protein
Length = 360
Score = 42.4 bits (98), Expect = 4e-04
Identities = 20/65 (30%), Positives = 34/65 (51%)
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAA 84
+ +LFV G+ ++T +++ F R+G V+ V K + FGFV F + D + A
Sbjct: 4 DRYKLFVGGIAKETSEEALKQYFSRYGAVLEAVVAKEKVTGKPRGFGFVRFANDCDVVKA 63
Query: 85 MRSLH 89
+R H
Sbjct: 64 LRDTH 68
Score = 36.2 bits (82), Expect = 0.028
Identities = 21/66 (31%), Positives = 29/66 (43%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
V ++FV GL +T + + FERFG V V R FGFV + S+
Sbjct: 115 VTSRTKKIFVGGLSSNTTEEEFKSYFERFGRTTDVVVMHDGVTNRPRGFGFVTYDSEDSV 174
Query: 82 IAAMRS 87
M+S
Sbjct: 175 EVVMQS 180
>At3g11400 eukaryotic translation initiation factor 3 subunit like
protein
Length = 294
Score = 41.6 bits (96), Expect = 7e-04
Identities = 30/101 (29%), Positives = 45/101 (43%), Gaps = 6/101 (5%)
Query: 6 SARAPPPRWSCHERSEVAEENIR------LFVDGLDRDTKHSQVRKLFERFGTVVSVFVP 59
+A PP + +RS V + R + V L DT+ + +LF FG V V+V
Sbjct: 186 AAYVPPSMRAGADRSAVGSDMRRRNDENSVRVTNLSEDTREPDLMELFHPFGAVTRVYVA 245
Query: 60 KTIKRVRREKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSI 100
K FGFV F S+ D A+ L+G ++ L +
Sbjct: 246 IDQKTGVSRGFGFVNFVSREDAQRAINKLNGYGYDNLILRV 286
>At2g37510 putative RNA-binding protein
Length = 142
Score = 41.6 bits (96), Expect = 7e-04
Identities = 24/77 (31%), Positives = 39/77 (50%)
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
RLFV GL R T + +++ F FG +V V R + FGFV + + D A
Sbjct: 35 RLFVSGLSRLTTNEKLQDAFASFGQLVDARVITDRDSGRSKGFGFVTYATIEDAEKAKAE 94
Query: 88 LHGTRLNSFFLSINPAR 104
++ L+ + + ++PAR
Sbjct: 95 MNAKFLDGWVIFVDPAR 111
>At1g13690 AtE1 (atE1)
Length = 177
Score = 41.6 bits (96), Expect = 7e-04
Identities = 25/85 (29%), Positives = 38/85 (44%)
Query: 19 RSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSK 78
+ + A + L+V GL + S + F FG + V P + FGFV F +
Sbjct: 5 QQQQAMQKNTLYVGGLADEVNESILHAAFIPFGDIKDVKTPLDQANQKHRSFGFVTFLER 64
Query: 79 RDGIAAMRSLHGTRLNSFFLSINPA 103
D AAM ++ G L L++N A
Sbjct: 65 EDASAAMDNMDGAELYGRVLTVNYA 89
>At2g22100 putative RNA-binding protein
Length = 382
Score = 41.2 bits (95), Expect = 9e-04
Identities = 25/76 (32%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
+FV GL DT H ++ FE +G + V R + FGFVLF++++ AA+++
Sbjct: 165 IFVRGLGWDTTHENLKAAFEVYGEITECSVVMDKDTGRAKGFGFVLFKTRKGARAALKNP 224
Query: 89 HGTRLNSFFLSINPAR 104
R+ + +S PAR
Sbjct: 225 E-KRMYNRTVSCLPAR 239
>At4g39260 glycine-rich protein (clone AtGRP8)
Length = 169
Score = 40.4 bits (93), Expect = 0.001
Identities = 22/83 (26%), Positives = 42/83 (50%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
++E R FV GL T +++ F +FG V+ + + R FGFV F+ ++
Sbjct: 1 MSEVEYRCFVGGLAWATNDEDLQRTFSQFGDVIDSKIINDRESGRSRGFGFVTFKDEKAM 60
Query: 82 IAAMRSLHGTRLNSFFLSINPAR 104
A+ ++G L+ +++N A+
Sbjct: 61 RDAIEEMNGKELDGRVITVNEAQ 83
>At1g55310 Serine/arginine-rich protein
Length = 220
Score = 40.4 bits (93), Expect = 0.001
Identities = 26/83 (31%), Positives = 36/83 (43%), Gaps = 2/83 (2%)
Query: 8 RAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRR 67
R+P PR RS + L V L D + +RK FE+FG V +++P+
Sbjct: 19 RSPSPRGRYGGRSRDLPTS--LLVRNLRHDCRQEDLRKSFEQFGPVKDIYLPRDYYTGDP 76
Query: 68 EKFGFVLFQSKRDGIAAMRSLHG 90
FGFV F D A + G
Sbjct: 77 RGFGFVQFMDPADAADAKHHMDG 99
>At1g11650 putative DNA binding protein
Length = 405
Score = 40.4 bits (93), Expect = 0.001
Identities = 24/89 (26%), Positives = 41/89 (45%), Gaps = 6/89 (6%)
Query: 26 NIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAM 85
N +FV GLD ++ +F ++G +V V +P ++ GFV F K A+
Sbjct: 260 NTTVFVGGLDASVTDDHLKNVFSQYGEIVHVKIP------AGKRCGFVQFSEKSCAEEAL 313
Query: 86 RSLHGTRLNSFFLSINPARFVDRKYKWNP 114
R L+G +L + ++ R K +P
Sbjct: 314 RMLNGVQLGGTTVRLSWGRSPSNKQSGDP 342
>At5g19030 unknown protein
Length = 172
Score = 40.0 bits (92), Expect = 0.002
Identities = 21/83 (25%), Positives = 40/83 (47%)
Query: 18 ERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQS 77
E S + LFV G +++K+F FG V +V + + + +G+V F S
Sbjct: 68 EASSLPSSISSLFVKGFSDSVSEGRLKKVFSEFGQVTNVKIIANERTRQSLGYGYVWFNS 127
Query: 78 KRDGIAAMRSLHGTRLNSFFLSI 100
K D +A+ +++G + F+ +
Sbjct: 128 KEDAQSAVEAMNGKFFDGRFILV 150
>At1g47500 unknown protein
Length = 434
Score = 40.0 bits (92), Expect = 0.002
Identities = 35/128 (27%), Positives = 55/128 (42%), Gaps = 19/128 (14%)
Query: 19 RSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSK 78
RSE N +FV GLD +++ F FG +VSV +P + GFV F ++
Sbjct: 298 RSEGDTINTTIFVGGLDSSVTDEDLKQPFSEFGEIVSVKIPV------GKGCGFVQFVNR 351
Query: 79 RDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRV 138
+ A+ L+GT + + ++ R NP KQ R K +W +
Sbjct: 352 PNAEEALEKLNGTVIGKQTVRLSWGR--------NPANKQ-----PRDKYGNQWVDPYYG 398
Query: 139 GAPFSGTG 146
G ++G G
Sbjct: 399 GQFYNGYG 406
Score = 33.1 bits (74), Expect = 0.23
Identities = 24/95 (25%), Positives = 43/95 (45%), Gaps = 5/95 (5%)
Query: 14 WSCHERSEVAEEN----IRLFVDGLDRDTKHSQVRKLF-ERFGTVVSVFVPKTIKRVRRE 68
W+ E EN + +FV L D + + + F E++ +V + V R +
Sbjct: 182 WASFSTGEKRLENNGPDLSIFVGDLAPDVSDALLHETFSEKYPSVKAAKVVLDANTGRSK 241
Query: 69 KFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPA 103
+GFV F + + AM ++G + +S + I PA
Sbjct: 242 GYGFVRFGDENERTKAMTEMNGVKCSSRAMRIGPA 276
>At5g47320 40S ribosomal protein S19
Length = 212
Score = 39.7 bits (91), Expect = 0.002
Identities = 22/77 (28%), Positives = 37/77 (47%)
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+L++ GL T ++ F F V V R +GFV F S+ +A+ +
Sbjct: 32 KLYIGGLSPGTDEHSLKDAFSSFNGVTEARVMTNKVTGRSRGYGFVNFISEDSANSAISA 91
Query: 88 LHGTRLNSFFLSINPAR 104
++G LN F +S+N A+
Sbjct: 92 MNGQELNGFNISVNVAK 108
>At5g10800 putative protein
Length = 957
Score = 39.7 bits (91), Expect = 0.002
Identities = 26/85 (30%), Positives = 42/85 (48%), Gaps = 3/85 (3%)
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFV--PKTIKRVRREKF-GFVLFQSKRDG 81
+ L+V L + + + F RFG + SV + P+T + RRE+ GFV F ++ DG
Sbjct: 169 QTTNLYVVNLSSKVDENFLLRTFGRFGPIASVKIMWPRTEEEKRRERHCGFVAFMNRADG 228
Query: 82 IAAMRSLHGTRLNSFFLSINPARFV 106
AA + G + + L I + V
Sbjct: 229 EAAKEKMQGIIVYEYELKIGWGKVV 253
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,999,178
Number of Sequences: 26719
Number of extensions: 329953
Number of successful extensions: 1051
Number of sequences better than 10.0: 140
Number of HSP's better than 10.0 without gapping: 97
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 877
Number of HSP's gapped (non-prelim): 200
length of query: 351
length of database: 11,318,596
effective HSP length: 100
effective length of query: 251
effective length of database: 8,646,696
effective search space: 2170320696
effective search space used: 2170320696
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0074.9


