

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.12
(369 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
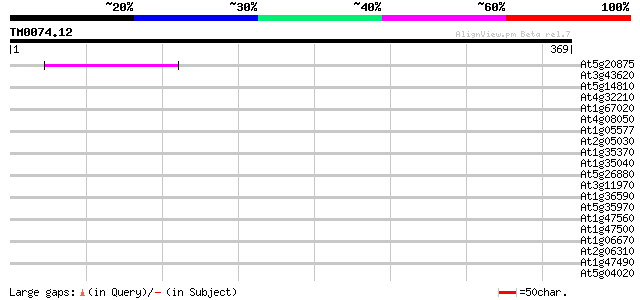
Score E
Sequences producing significant alignments: (bits) Value
At5g20875 unknown protein 44 2e-04
At3g43620 putative protein 37 0.013
At5g14810 putative protein 35 0.050
At4g32210 putative protein 35 0.050
At1g67020 hypothetical protein 33 0.25
At4g08050 32 0.42
At1g05577 hypothetical protein 32 0.42
At2g05030 hypothetical protein 32 0.55
At1g35370 hypothetical protein 32 0.55
At1g35040 hypothetical protein 32 0.55
At5g26880 putative protein 31 1.2
At3g11970 hypothetical protein 31 1.2
At1g36590 hypothetical protein 31 1.2
At5g35970 DNA helicase-like 30 1.6
At1g47560 hypothetical protein 30 1.6
At1g47500 unknown protein 30 2.1
At1g06670 DEIH-box RNA/DNA helicase 30 2.1
At2g06310 putative Athila retroelement ORF1 protein 30 2.7
At1g47490 unknown protein 30 2.7
At5g04020 pathogen-induced calmodulin-binding protein (PICBP) 29 3.6
>At5g20875 unknown protein
Length = 272
Score = 43.5 bits (101), Expect = 2e-04
Identities = 23/89 (25%), Positives = 41/89 (45%), Gaps = 1/89 (1%)
Query: 24 SPPSQPNTSPYYCHNVPSYYPQTHSSLQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQ 83
S ++ N+ + H + Y +++ F G N WL A+ YF + + Q
Sbjct: 88 STAAEDNSIDTHPHRMSRYIQLMKPKIEMPVFEGPNVNSWLTRAERYFEFGSFTDAEKIQ 147
Query: 84 LLAIHLGGKALTWFRLSHQQ-IHSWEEFK 111
L+ + + G+AL WF L ++ W +FK
Sbjct: 148 LVYMSVEGRALCWFNLENRNPFVDWNDFK 176
>At3g43620 putative protein
Length = 221
Score = 37.4 bits (85), Expect = 0.013
Identities = 29/120 (24%), Positives = 47/120 (39%), Gaps = 11/120 (9%)
Query: 10 SILSAQRDPRSIFYSPPSQPNTS-PYYCHNVPSYYPQTH----SSLQLSEFNGENPEYWL 64
S+ R + SP SQ S P + S Y + FNG+ + WL
Sbjct: 24 SMAPHDRSSSMVLGSPESQSGCSDPNHDVRSDSQYHYRRLRRLGKVDFPRFNGDGIKDWL 83
Query: 65 FMADCYFLQNPYPWETRFQLLAIHLGGKALTWFRLSHQQI------HSWEEFKVAFRLYF 118
F + +FL + P E + + +IH TW + Q I H W +K+ ++ +
Sbjct: 84 FQIEQFFLIDHTPEELKVDIASIHFDDIDATWHQSIVQSIMWRHVRHDWWNYKLLLQVRY 143
>At5g14810 putative protein
Length = 716
Score = 35.4 bits (80), Expect = 0.050
Identities = 27/113 (23%), Positives = 46/113 (39%), Gaps = 10/113 (8%)
Query: 16 RDPRSIFYSPPSQPNTSPYYCHNVPSYYPQTHSS----LQLSEFNGENPEYWLFMADCYF 71
R + I +P P+TS Y+P S L+ FNG+ WLF + +F
Sbjct: 68 RSSQLISTAPIVTPDTSKERVGMSNKYFPLEERSDFARLEHLRFNGDRITEWLFQIEQFF 127
Query: 72 LQNPYPWETRFQLLAIHLGGKALTWFRLSHQQI------HSWEEFKVAFRLYF 118
L + P E + ++H A T + Q + H W +K+ ++ +
Sbjct: 128 LIDRTPEELKVGFASLHFDDTAATLHQSIVQSMWWKHVRHDWWSYKLLLQVRY 180
>At4g32210 putative protein
Length = 1086
Score = 35.4 bits (80), Expect = 0.050
Identities = 27/113 (23%), Positives = 46/113 (39%), Gaps = 10/113 (8%)
Query: 16 RDPRSIFYSPPSQPNTSPYYCHNVPSYYPQTHSS----LQLSEFNGENPEYWLFMADCYF 71
R + I +P P+TS Y+P S L+ FNG+ WLF + +F
Sbjct: 68 RSSQLISTAPIVTPDTSKERVGMSNKYFPLEERSDFARLEHLRFNGDRITEWLFQIEQFF 127
Query: 72 LQNPYPWETRFQLLAIHLGGKALTWFRLSHQQI------HSWEEFKVAFRLYF 118
L + P E + ++H A T + Q + H W +K+ ++ +
Sbjct: 128 LIDRTPEELKVGFASLHFDDTAATLHQSIVQSMWWKHVRHDWWSYKLLLQVRY 180
>At1g67020 hypothetical protein
Length = 659
Score = 33.1 bits (74), Expect = 0.25
Identities = 29/124 (23%), Positives = 47/124 (37%), Gaps = 12/124 (9%)
Query: 16 RDPRSIFYS--PPSQPNTSPYYCHNVPSYYPQTHSSLQLSEFNGENPEYWLFMADCYFLQ 73
+ PR F S P SP N S + +++ F+G W + +F
Sbjct: 78 QSPRHSFSSSMPQRGAGESPILLDNRSSLIRR----IEMPVFDGSGVYEWFSKVERFFRV 133
Query: 74 NPYPWETRFQLLAIHLGGKALTWF--RLSHQQIHSWEEFKVAFRLYF----VYNRPNGFS 127
Y + L+A+ L G AL WF +S + W F+ F +++ P
Sbjct: 134 GRYQDSDKLDLVALSLEGVALKWFLREMSTLEFRDWNSFEQRLLARFDPVKIHSSPEVLM 193
Query: 128 TTTV 131
TT+
Sbjct: 194 PTTL 197
>At4g08050
Length = 1428
Score = 32.3 bits (72), Expect = 0.42
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 2/60 (3%)
Query: 81 RFQLLAIHLGGKALTWFR-LSHQQIHSWEEFKVAFRLYFVYNRPNGFSTTTVKYGFEKSS 139
+ +L LG KA W + LSH I +W+++K AF F N + YGF + +
Sbjct: 87 KLRLFPFSLGDKAHIWEKNLSHDSITTWDDYKKAFLSKFFSNARTARLRNEI-YGFSQKT 145
>At1g05577 hypothetical protein
Length = 372
Score = 32.3 bits (72), Expect = 0.42
Identities = 15/51 (29%), Positives = 30/51 (58%)
Query: 223 DAITPDVNKTFVLKTTDLVLRNPRQVFGNLLEGKFVQKNNEAQFEKHFDKL 273
D ITP + +VLK ++++L +P++ + N+ + +V +N E+ KL
Sbjct: 83 DLITPISDNEYVLKGSEILLSSPKEDYPNVEKKAWVTRNGGIDAEEKLQKL 133
>At2g05030 hypothetical protein
Length = 205
Score = 32.0 bits (71), Expect = 0.55
Identities = 14/47 (29%), Positives = 25/47 (52%)
Query: 50 LQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQLLAIHLGGKALTW 96
+ L F+G + WLF + +F + P + + +++AIH A TW
Sbjct: 134 IDLPRFDGTGIKEWLFKVEEFFGIDFTPADLKVKMVAIHFDSHAATW 180
>At1g35370 hypothetical protein
Length = 1447
Score = 32.0 bits (71), Expect = 0.55
Identities = 14/47 (29%), Positives = 23/47 (48%)
Query: 50 LQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQLLAIHLGGKALTW 96
+ F+G WLF + +F + P E + +++AIH A TW
Sbjct: 117 IDFPRFDGSRINEWLFKVEEFFGVDFTPEEMKVKMVAIHFDSHAATW 163
>At1g35040 hypothetical protein
Length = 106
Score = 32.0 bits (71), Expect = 0.55
Identities = 18/62 (29%), Positives = 27/62 (43%), Gaps = 2/62 (3%)
Query: 63 WLFMADCYFLQNPYPWETRFQLLAIHLGGKALTWFRLSHQQI--HSWEEFKVAFRLYFVY 120
W+ M + FL Y + + ++ LG +A WF I +SW + K L F
Sbjct: 42 WITMTEAQFLGGEYSEKDKLACVSGFLGSQAKCWFHKEQSWIPFNSWSQVKDGLLLTFGN 101
Query: 121 NR 122
NR
Sbjct: 102 NR 103
>At5g26880 putative protein
Length = 275
Score = 30.8 bits (68), Expect = 1.2
Identities = 34/127 (26%), Positives = 52/127 (40%), Gaps = 11/127 (8%)
Query: 82 FQLLAIHLGGKALTWFRLSHQQI-----HSWEEFKVAFRLYFVYNRPNGFSTTTVKYGFE 136
FQ+ + AL L H + SW EF+ FRL F+ +P +T T + E
Sbjct: 117 FQVDDARVKRAALLLLNLKHSFVVVKAHSSWAEFQEYFRLQFIVYQPE-IATITSNWVAE 175
Query: 137 KSSLVEIQRSSEQVSPHRVVFSETTLEDGAQLPFEASSDVHLLPEVVSRVELGSAETEIE 196
+ +Q S + + + SET + LP EA SD + P + + ET +
Sbjct: 176 NNGYNLMQDFSYRSGDYLLFGSET-----SGLPPEALSDCNHEPYGGGTLRIPMVETYVR 230
Query: 197 SHESSTS 203
S S
Sbjct: 231 CLNLSVS 237
>At3g11970 hypothetical protein
Length = 1499
Score = 30.8 bits (68), Expect = 1.2
Identities = 13/47 (27%), Positives = 23/47 (48%)
Query: 50 LQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQLLAIHLGGKALTW 96
+ F+G + WLF + +F + P + + ++ AIH A TW
Sbjct: 112 IDFPRFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTW 158
>At1g36590 hypothetical protein
Length = 1499
Score = 30.8 bits (68), Expect = 1.2
Identities = 13/47 (27%), Positives = 23/47 (48%)
Query: 50 LQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQLLAIHLGGKALTW 96
+ F+G + WLF + +F + P + + ++ AIH A TW
Sbjct: 112 IDFPRFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTW 158
>At5g35970 DNA helicase-like
Length = 750
Score = 30.4 bits (67), Expect = 1.6
Identities = 18/48 (37%), Positives = 27/48 (55%), Gaps = 3/48 (6%)
Query: 298 EKMLVTRNTRICVDNTFEKLLETYINLLLDWQRMRPMNLTSEIALSSL 345
E++LVT T VDN EKLL +N++ + P ++S +A SL
Sbjct: 321 ERVLVTAPTNAAVDNMVEKLLHLGLNIV---RVGNPARISSAVASKSL 365
>At1g47560 hypothetical protein
Length = 1564
Score = 30.4 bits (67), Expect = 1.6
Identities = 16/77 (20%), Positives = 32/77 (40%), Gaps = 6/77 (7%)
Query: 48 SSLQLSEFNGENPEYWLFMADCYFLQNPYPWETRFQLLAIHLGGKALTWFRLSHQQ---- 103
S L F+G+ WLF + +F + P E + + + G A W++ +
Sbjct: 966 SKLDFHRFSGDRVRDWLFKLEQFFSLDFTPEELKVSIASTLCDGAAAKWYKSLFESDFGV 1025
Query: 104 --IHSWEEFKVAFRLYF 118
+ +W +K+ +F
Sbjct: 1026 KLLSNWNMYKLLLEEHF 1042
>At1g47500 unknown protein
Length = 434
Score = 30.0 bits (66), Expect = 2.1
Identities = 14/45 (31%), Positives = 24/45 (53%), Gaps = 1/45 (2%)
Query: 21 IFYSPPSQPNTSPYYCH-NVPSYYPQTHSSLQLSEFNGENPEYWL 64
+ Y+PP SPY+ + N ++ Q+ + + FNGEN W+
Sbjct: 63 MMYAPPPPMPFSPYHQYPNHHHFHHQSRGNKHQNAFNGENKTIWV 107
>At1g06670 DEIH-box RNA/DNA helicase
Length = 1576
Score = 30.0 bits (66), Expect = 2.1
Identities = 19/74 (25%), Positives = 36/74 (47%), Gaps = 2/74 (2%)
Query: 181 EVVSRVELGSAETEIESHESSTSEDSSLITTPVDSTPYATRSDAITPDVNKTFVLKTTDL 240
E+ +++ LG+ E + S+ + +E T+PV+ AT+ + + + L D
Sbjct: 1167 ELANKLGLGNMEESLPSNFADGNEQPDPNTSPVEDVSAATKQKKMQSESKRCKSLNNVD- 1225
Query: 241 VLRNPRQVFGNLLE 254
L N + FGN+ E
Sbjct: 1226 -LGNIEENFGNMEE 1238
>At2g06310 putative Athila retroelement ORF1 protein
Length = 733
Score = 29.6 bits (65), Expect = 2.7
Identities = 16/40 (40%), Positives = 21/40 (52%), Gaps = 1/40 (2%)
Query: 83 QLLAIHLGGKALTWFR-LSHQQIHSWEEFKVAFRLYFVYN 121
+L LGGKA W + L H I +W++ K AF F N
Sbjct: 89 RLFPFSLGGKAHIWEKNLPHDSITTWDDCKKAFLSKFFSN 128
>At1g47490 unknown protein
Length = 432
Score = 29.6 bits (65), Expect = 2.7
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 8/50 (16%)
Query: 18 PRSIFYSPPSQPNTSPYYCHNVPSYYPQTHSSL---QLSEFNGENPEYWL 64
P + Y+PP P PY H P+++ H S + FNGEN W+
Sbjct: 61 PHQMMYAPPPFP---PY--HQYPNHHHLHHQSRGNKHQNAFNGENKTIWV 105
>At5g04020 pathogen-induced calmodulin-binding protein (PICBP)
Length = 1495
Score = 29.3 bits (64), Expect = 3.6
Identities = 26/110 (23%), Positives = 49/110 (43%), Gaps = 10/110 (9%)
Query: 159 ETTLEDGAQLPFEASSDVHLLPEVVSRVELGSAETEIESHESSTSEDSSLITTPVDSTPY 218
E T DG ++ + V LL EV+ + L ++ + ++E + + +L + V +
Sbjct: 1343 EDTSVDGEKMELYQTEAVELLGEVIDGISLEESQDQNLNNEETRQKSETLQVSKVRIDRW 1402
Query: 219 ATRSDAITPDVNKTFVLKTTDLVLRNPRQVFGNLLEGKFVQKNNEAQFEK 268
+ AI + + FV ++ NPR E +F+ N E + EK
Sbjct: 1403 SNLKRAI---LLRRFVKALENVRKFNPR-------EPRFLPPNPEVEAEK 1442
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,122,259
Number of Sequences: 26719
Number of extensions: 338913
Number of successful extensions: 1064
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 1047
Number of HSP's gapped (non-prelim): 35
length of query: 369
length of database: 11,318,596
effective HSP length: 101
effective length of query: 268
effective length of database: 8,619,977
effective search space: 2310153836
effective search space used: 2310153836
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0074.12


