

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0055.14
(267 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
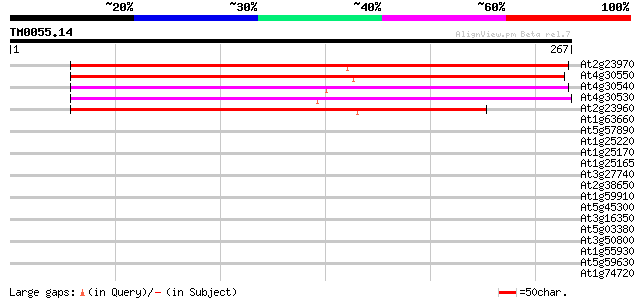
At2g23970 unknown protein 232 2e-61
At4g30550 unknown protein 218 2e-57
At4g30540 unknown protein (At4g30540) 208 2e-54
At4g30530 unknown protein 204 3e-53
At2g23960 unknown protein 182 2e-46
At1g63660 GMP synthase 40 0.001
At5g57890 anthranilate synthase beta chain 35 0.042
At1g25220 unknown protein 35 0.054
At1g25170 35 0.054
At1g25165 putative protein 35 0.054
At3g27740 carbamoyl phosphate synthetase small subunit (carA) 34 0.093
At2g38650 unknown protein 33 0.12
At1g59910 hypothetical protein 33 0.12
At5g45300 beta-amylase-like 33 0.16
At3g16350 putative MYB family transcription factor 33 0.21
At5g03380 farnesylated protein - like 32 0.35
At3g50800 unknown protein (At3g50800) 32 0.35
At1g55930 unknown protein 32 0.35
At5g59630 glycine-rich protein - like 32 0.46
At1g74720 hypothetical protein 32 0.46
>At2g23970 unknown protein
Length = 251
Score = 232 bits (591), Expect = 2e-61
Identities = 110/242 (45%), Positives = 156/242 (64%), Gaps = 5/242 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KR+AL + DS ++ K +GG F F E+GE+WDL++V++GEFP +L YDGFV
Sbjct: 6 KRFALFLATSDSTFVKKAYGGYFNVFVSTFGEDGEQWDLFRVIDGEFPDDKDLDKYDGFV 65
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS +DA + WI L SL LD + KK+LGICFGHQIL R+ GGKVGR+S G D+G
Sbjct: 66 ISGSLNDAFGDDDWIVKLCSLCQKLDDMKKKVLGICFGHQILSRIKGGKVGRASRGLDMG 125
Query: 150 VTTMKLASSS-----SIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
+ ++ + + + +P L+I KCH+DEV LP A ++ +S+K +EM YG+
Sbjct: 126 LRSITMVTDAVKPGGYFGSQIPKSLAIIKCHQDEVLELPESATLLAYSDKYNVEMCSYGN 185
Query: 205 HMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLK 264
H+LGIQGHPE+ +ILF IDR+ + L+++ FA K EP+ + W+ LC NFLK
Sbjct: 186 HLLGIQGHPEYNKEILFEIIDRVVNLKLMEQDFADKAKATMENAEPDRKQWQTLCKNFLK 245
Query: 265 DR 266
R
Sbjct: 246 GR 247
>At4g30550 unknown protein
Length = 249
Score = 218 bits (556), Expect = 2e-57
Identities = 105/241 (43%), Positives = 152/241 (62%), Gaps = 6/241 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KR+AL + DSE++ K +GG F F EEGE+WDL++V++G+FP ++L YDGFV
Sbjct: 8 KRFALFLATCDSEFVKKTYGGYFNVFVSTFGEEGEQWDLFRVIDGQFPDENDLDKYDGFV 67
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA + WI L + LD + KK+LGICFGHQI+ RV GGK+GR+ G D+G
Sbjct: 68 ISGSPHDAFGDADWIVKLCEVCQKLDHMKKKVLGICFGHQIITRVKGGKIGRALKGADMG 127
Query: 150 VTTMKLASSSSIA------LNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYG 203
+ ++ +A + + +P+ L+I KCH+DEV LP A ++ SE +EMF G
Sbjct: 128 LRSITIAKDNEKLRGYFGDVEVPASLAIIKCHQDEVLELPESATLLASSEVCNVEMFSIG 187
Query: 204 DHMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFL 263
DH IQGHPE+ +ILF +DR+ + L+++ FA K +P+ W++LC NFL
Sbjct: 188 DHFFCIQGHPEYNKEILFEIVDRVLNMKLMEQEFADKAKSTMETAQPDRILWQKLCKNFL 247
Query: 264 K 264
K
Sbjct: 248 K 248
>At4g30540 unknown protein (At4g30540)
Length = 248
Score = 208 bits (530), Expect = 2e-54
Identities = 107/240 (44%), Positives = 145/240 (59%), Gaps = 3/240 (1%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
+RYAL DSE++ + +GG F F +EGE+WDL++V++GEFPR ++L Y+GFV
Sbjct: 6 RRYALFQATPDSEFVKEMYGGYFNVFVSAFGDEGEQWDLFRVIDGEFPRDEDLEKYEGFV 65
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA WI +L S+ LD + KKILGICFGHQI+ RV GGKVGR+ G DIG
Sbjct: 66 ISGSLHDAFTEEDWIIELCSVCKKLDVMKKKILGICFGHQIICRVRGGKVGRARKGPDIG 125
Query: 150 ---VTTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDHM 206
+T ++ + LSI +CHRDEV P A +IG+S+K +E+F DH+
Sbjct: 126 LGNITIVQDVIKPGDYFDQIESLSIIQCHRDEVLEPPESARVIGFSDKCDVEIFSVEDHL 185
Query: 207 LGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLKDR 266
L QGHPE+ +IL IDR+ ++E K EP+T+ LC NFLK R
Sbjct: 186 LCFQGHPEYNKEILLEIIDRVHKIKFVEEEILEKAKDSIKKFEPDTQRLHMLCKNFLKGR 245
>At4g30530 unknown protein
Length = 250
Score = 204 bits (520), Expect = 3e-53
Identities = 101/243 (41%), Positives = 144/243 (58%), Gaps = 5/243 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
KRYAL + DSE++ K +GG F +EGE WD ++VV+GEFP +L YDGFV
Sbjct: 5 KRYALFLATLDSEFVKKTYGGYHNVFVTTFGDEGEHWDSFRVVSGEFPDEKDLEKYDGFV 64
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTG---- 145
I+GS HDA N WI L +V +D + KKILGICFGHQI+ RV GG VGR+ G
Sbjct: 65 ISGSSHDAFENDDWILKLCDIVKKIDEMKKKILGICFGHQIIARVRGGTVGRAKKGPELK 124
Query: 146 -WDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
DI + + S +P ++I KCH+DEV LP A+++ +S+ +EM+ D
Sbjct: 125 LGDITIVKDAITPGSYFGNEIPDSIAIIKCHQDEVLVLPETAKVLAYSKNYEVEMYSIED 184
Query: 205 HMLGIQGHPEFTIDILFHFIDRLTSRNLLQECFAVDVKVKAALQEPNTEAWKRLCVNFLK 264
H+ IQGHPE+ +ILF +DR+ + +++ FA K + + + W+ +C NFLK
Sbjct: 185 HLFCIQGHPEYNKEILFEIVDRVLALGYVKQEFADAAKATMENRGADRKLWETICKNFLK 244
Query: 265 DRL 267
R+
Sbjct: 245 GRV 247
>At2g23960 unknown protein
Length = 217
Score = 182 bits (462), Expect = 2e-46
Identities = 87/203 (42%), Positives = 127/203 (61%), Gaps = 5/203 (2%)
Query: 30 KRYALLMCGEDSEYLLKKHGGCFGFFTRMLAEEGERWDLYKVVNGEFPRHDELPLYDGFV 89
K+Y L + DSE+ K +GG F +L +EGE+WD ++VV+GEFP +L Y+GFV
Sbjct: 5 KKYLLFLATPDSEFAKKTYGGYHNVFVSLLGDEGEQWDSFRVVDGEFPEEKDLEKYEGFV 64
Query: 90 ITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWDIG 149
I+GS HDA + WI L ++ LD +NKK+LGICFGHQ++ R GGKV R+ G ++
Sbjct: 65 ISGSSHDAFQDTDWILKLCDIIKKLDDMNKKVLGICFGHQLIARAKGGKVARARKGPELC 124
Query: 150 VTTMKLASSSSIALN-----LPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGD 204
+ + + + + N +P+ L I KCH+DEV LP A+++ +S +EM+ D
Sbjct: 125 LGNITIVKEAVMPENYFGEEVPANLRIIKCHQDEVLELPENAKLLAYSSMYEVEMYSIKD 184
Query: 205 HMLGIQGHPEFTIDILFHFIDRL 227
+ L IQGHPE+ DILF IDR+
Sbjct: 185 NFLCIQGHPEYNRDILFDIIDRV 207
>At1g63660 GMP synthase
Length = 534
Score = 40.0 bits (92), Expect = 0.001
Identities = 35/145 (24%), Positives = 66/145 (45%), Gaps = 12/145 (8%)
Query: 89 VITGSCHDAHA-NHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRSSTGWD 147
+++G H HA + P + + +S +LGIC+G Q++ + LGG V + +
Sbjct: 56 ILSGGPHSVHALDAPSFPE--GFIEWAESNGVSVLGICYGLQLIVQKLGGVVVEGESK-E 112
Query: 148 IGVTTMKLASSSSI--ALNLPSELSIFKCHRDEVRNLPPKAEIIGWSEKTGIEMFRYGDH 205
G +++ S I + + + ++ H DE LP E++ S + +
Sbjct: 113 YGKMEIEVKGKSEIFGSESGGEKQMVWMSHGDEAVKLPEGFEVVAQSAQGAVAALESRKK 172
Query: 206 ML-GIQGHPEFT-----IDILFHFI 224
+ G+Q HPE T ++ L HF+
Sbjct: 173 KIYGLQYHPEVTHSPKGMETLRHFL 197
>At5g57890 anthranilate synthase beta chain
Length = 273
Score = 35.0 bits (79), Expect = 0.042
Identities = 31/106 (29%), Positives = 47/106 (44%), Gaps = 10/106 (9%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTM-----KLASSSSIALNLPSELSIFKCH 175
+ G+C G Q +G GGK+ RS G G ++M K L+ P + +
Sbjct: 145 LFGVCMGLQCIGEAFGGKIVRSPFGVMHGKSSMVHYDEKGEEGLFSGLSNPFLVGRYHSL 204
Query: 176 RDEVRNLP-PKAEIIGWSEKTGIEM---FRYGDHMLGIQGHPEFTI 217
E + P + E+ W+E G+ M R H+ G+Q HPE I
Sbjct: 205 VIEKDSFPSDELEVTAWTE-DGLVMAARHRKYKHIQGVQFHPESII 249
>At1g25220 unknown protein
Length = 276
Score = 34.7 bits (78), Expect = 0.054
Identities = 31/106 (29%), Positives = 46/106 (43%), Gaps = 10/106 (9%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTM-----KLASSSSIALNLPSELSIFKCH 175
+ G+C G Q +G GGK+ RS G G ++M K L+ P + +
Sbjct: 148 LFGVCMGLQCIGEAFGGKIVRSPFGVMHGKSSMVHYDEKGEEGLFSGLSNPFIVGRYHSL 207
Query: 176 RDEVRNLP-PKAEIIGWSEKTGIEM---FRYGDHMLGIQGHPEFTI 217
E P + E+ W+E G+ M R H+ G+Q HPE I
Sbjct: 208 VIEKDTFPSDELEVTAWTE-DGLVMAARHRKYKHIQGVQFHPESII 252
>At1g25170
Length = 469
Score = 34.7 bits (78), Expect = 0.054
Identities = 31/106 (29%), Positives = 46/106 (43%), Gaps = 10/106 (9%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTM-----KLASSSSIALNLPSELSIFKCH 175
+ G+C G Q +G GGK+ RS G G ++M K L+ P + +
Sbjct: 341 LFGVCMGLQCIGEAFGGKIVRSPFGVMHGKSSMVHYDEKGEEGLFSGLSNPFIVGRYHSL 400
Query: 176 RDEVRNLP-PKAEIIGWSEKTGIEM---FRYGDHMLGIQGHPEFTI 217
E P + E+ W+E G+ M R H+ G+Q HPE I
Sbjct: 401 VIEKDTFPSDELEVTAWTE-DGLVMAARHRKYKHIQGVQFHPESII 445
>At1g25165 putative protein
Length = 315
Score = 34.7 bits (78), Expect = 0.054
Identities = 31/106 (29%), Positives = 46/106 (43%), Gaps = 10/106 (9%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTGWDIGVTTM-----KLASSSSIALNLPSELSIFKCH 175
+ G+C G Q +G GGK+ RS G G ++M K L+ P + +
Sbjct: 187 LFGVCMGLQCIGEAFGGKIVRSPFGVMHGKSSMVHYDEKGEEGLFSGLSNPFIVGRYHSL 246
Query: 176 RDEVRNLP-PKAEIIGWSEKTGIEM---FRYGDHMLGIQGHPEFTI 217
E P + E+ W+E G+ M R H+ G+Q HPE I
Sbjct: 247 VIEKDTFPSDELEVTAWTE-DGLVMAARHRKYKHIQGVQFHPESII 291
>At3g27740 carbamoyl phosphate synthetase small subunit (carA)
Length = 430
Score = 33.9 bits (76), Expect = 0.093
Identities = 13/25 (52%), Positives = 17/25 (68%)
Query: 121 ILGICFGHQILGRVLGGKVGRSSTG 145
+ GIC GHQ+LG+ LGGK + G
Sbjct: 314 VYGICMGHQLLGQALGGKTFKMKFG 338
>At2g38650 unknown protein
Length = 619
Score = 33.5 bits (75), Expect = 0.12
Identities = 15/25 (60%), Positives = 20/25 (80%), Gaps = 1/25 (4%)
Query: 14 MESGCGGGGGGGGGKGKRYALLMCG 38
M+ G GGGGGGGGGK +R+ +L+ G
Sbjct: 1 MKGGGGGGGGGGGGK-RRWKVLVIG 24
>At1g59910 hypothetical protein
Length = 929
Score = 33.5 bits (75), Expect = 0.12
Identities = 14/22 (63%), Positives = 16/22 (72%)
Query: 11 ELEMESGCGGGGGGGGGKGKRY 32
E+ +S GGGGGGGGG G RY
Sbjct: 152 EIRDQSNNGGGGGGGGGGGGRY 173
Score = 29.6 bits (65), Expect = 1.7
Identities = 11/14 (78%), Positives = 12/14 (85%)
Query: 19 GGGGGGGGGKGKRY 32
GGGGGGGGG G+ Y
Sbjct: 161 GGGGGGGGGGGRYY 174
Score = 28.5 bits (62), Expect = 3.9
Identities = 11/19 (57%), Positives = 13/19 (67%)
Query: 10 RELEMESGCGGGGGGGGGK 28
R+ G GGGGGGGGG+
Sbjct: 154 RDQSNNGGGGGGGGGGGGR 172
>At5g45300 beta-amylase-like
Length = 689
Score = 33.1 bits (74), Expect = 0.16
Identities = 13/15 (86%), Positives = 13/15 (86%)
Query: 17 GCGGGGGGGGGKGKR 31
G GGGGGGGGKGKR
Sbjct: 73 GSSGGGGGGGGKGKR 87
>At3g16350 putative MYB family transcription factor
Length = 387
Score = 32.7 bits (73), Expect = 0.21
Identities = 14/23 (60%), Positives = 16/23 (68%)
Query: 16 SGCGGGGGGGGGKGKRYALLMCG 38
SG GGGGGGGGG G A+ + G
Sbjct: 29 SGGGGGGGGGGGSGSSSAVKLFG 51
Score = 28.9 bits (63), Expect = 3.0
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 28 GSGGGGGGGGGGG 40
Score = 28.1 bits (61), Expect = 5.1
Identities = 10/12 (83%), Positives = 10/12 (83%)
Query: 18 CGGGGGGGGGKG 29
CGG GGGGGG G
Sbjct: 26 CGGSGGGGGGGG 37
>At5g03380 farnesylated protein - like
Length = 392
Score = 32.0 bits (71), Expect = 0.35
Identities = 13/21 (61%), Positives = 16/21 (75%)
Query: 9 ERELEMESGCGGGGGGGGGKG 29
E++ E+ G GGGGGGGGG G
Sbjct: 265 EKKKEVAVGGGGGGGGGGGDG 285
>At3g50800 unknown protein (At3g50800)
Length = 152
Score = 32.0 bits (71), Expect = 0.35
Identities = 12/16 (75%), Positives = 14/16 (87%)
Query: 16 SGCGGGGGGGGGKGKR 31
+GCGG G GGGGKG+R
Sbjct: 126 NGCGGRGCGGGGKGRR 141
>At1g55930 unknown protein
Length = 846
Score = 32.0 bits (71), Expect = 0.35
Identities = 28/99 (28%), Positives = 38/99 (38%)
Query: 83 PLYDGFVITGSCHDAHANHPWIHDLLSLVHTLDSINKKILGICFGHQILGRVLGGKVGRS 142
PL D I GS ++ W+ S V + I+GI + +L V GK+ S
Sbjct: 549 PLVDVVAIDGSGSLVDFHNFWVTHQYSRVPVFEQRIDNIVGIAYAMDLLDYVPKGKLLES 608
Query: 143 STGWDIGVTTMKLASSSSIALNLPSELSIFKCHRDEVRN 181
+T D+ S NL E I K H V N
Sbjct: 609 TTVVDMAHKPAFFVPDSMSVWNLLREFRIRKVHMAVVLN 647
>At5g59630 glycine-rich protein - like
Length = 253
Score = 31.6 bits (70), Expect = 0.46
Identities = 16/31 (51%), Positives = 19/31 (60%)
Query: 17 GCGGGGGGGGGKGKRYALLMCGEDSEYLLKK 47
G GGGGGGGGG G + LL S+ + KK
Sbjct: 107 GGGGGGGGGGGGGGKLQLLGLKPYSKKMKKK 137
Score = 29.6 bits (65), Expect = 1.7
Identities = 12/14 (85%), Positives = 12/14 (85%)
Query: 16 SGCGGGGGGGGGKG 29
SG GGGGGGGGG G
Sbjct: 71 SGGGGGGGGGGGGG 84
Score = 28.5 bits (62), Expect = 3.9
Identities = 13/22 (59%), Positives = 14/22 (63%)
Query: 17 GCGGGGGGGGGKGKRYALLMCG 38
G GGGGGGGGG G L + G
Sbjct: 105 GGGGGGGGGGGGGGGGKLQLLG 126
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 81 GGGGGGGGGGGGG 93
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 82 GGGGGGGGGGGGG 94
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 87 GGGGGGGGGGGGG 99
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 101 GGGGGGGGGGGGG 113
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 92 GGGGGGGGGGGGG 104
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 94 GGGGGGGGGGGGG 106
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 95 GGGGGGGGGGGGG 107
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 80 GGGGGGGGGGGGG 92
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 88 GGGGGGGGGGGGG 100
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 75 GGGGGGGGGGGGG 87
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 77 GGGGGGGGGGGGG 89
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 74 GGGGGGGGGGGGG 86
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 76 GGGGGGGGGGGGG 88
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 86 GGGGGGGGGGGGG 98
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 79 GGGGGGGGGGGGG 91
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 96 GGGGGGGGGGGGG 108
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 93 GGGGGGGGGGGGG 105
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 103 GGGGGGGGGGGGG 115
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 90 GGGGGGGGGGGGG 102
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 78 GGGGGGGGGGGGG 90
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 83 GGGGGGGGGGGGG 95
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 99 GGGGGGGGGGGGG 111
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 73 GGGGGGGGGGGGG 85
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 85 GGGGGGGGGGGGG 97
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 102 GGGGGGGGGGGGG 114
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 97 GGGGGGGGGGGGG 109
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 104 GGGGGGGGGGGGG 116
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 84 GGGGGGGGGGGGG 96
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 91 GGGGGGGGGGGGG 103
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 89 GGGGGGGGGGGGG 101
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 98 GGGGGGGGGGGGG 110
Score = 28.1 bits (61), Expect = 5.1
Identities = 11/13 (84%), Positives = 11/13 (84%)
Query: 17 GCGGGGGGGGGKG 29
G GGGGGGGGG G
Sbjct: 100 GGGGGGGGGGGGG 112
Score = 27.3 bits (59), Expect = 8.7
Identities = 11/14 (78%), Positives = 11/14 (78%)
Query: 16 SGCGGGGGGGGGKG 29
S GGGGGGGGG G
Sbjct: 70 SSGGGGGGGGGGGG 83
>At1g74720 hypothetical protein
Length = 1081
Score = 31.6 bits (70), Expect = 0.46
Identities = 18/42 (42%), Positives = 24/42 (56%), Gaps = 6/42 (14%)
Query: 12 LEMESGCGGGGGGGGGKG------KRYALLMCGEDSEYLLKK 47
LE E G GGGGGG GG G R +L +C E ++L++
Sbjct: 607 LEGEGGGGGGGGGPGGGGGGGPYCGRISLRLCLEGGYHVLEE 648
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.143 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,724,477
Number of Sequences: 26719
Number of extensions: 314591
Number of successful extensions: 5056
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 105
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 2611
Number of HSP's gapped (non-prelim): 1581
length of query: 267
length of database: 11,318,596
effective HSP length: 98
effective length of query: 169
effective length of database: 8,700,134
effective search space: 1470322646
effective search space used: 1470322646
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0055.14


