

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0055.11
(605 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
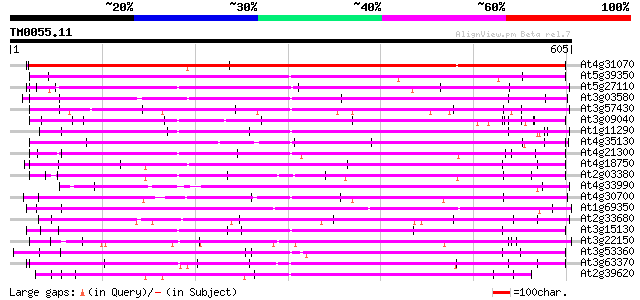
Sequences producing significant alignments: (bits) Value
At4g31070 putative protein 568 e-162
At5g39350 putative protein 371 e-103
At5g27110 putative protein 369 e-102
At3g03580 unknown protein (At3g03580) 352 3e-97
At3g57430 putative protein 351 6e-97
At3g09040 hypothetical protein 351 6e-97
At1g11290 hypothetical protein 348 4e-96
At4g35130 putative protein 347 8e-96
At4g21300 putative protein 347 1e-95
At4g18750 putative protein 347 1e-95
At2g03380 unknown protein 339 2e-93
At4g33990 putative protein 333 1e-91
At4g30700 unknown protein 332 4e-91
At1g69350 hypothetical protein 332 5e-91
At2g33680 hypothetical protein 331 8e-91
At3g15130 hypothetical protein 330 1e-90
At3g22150 hypothetical protein 330 1e-90
At3g53360 putative protein 328 4e-90
At3g63370 putative protein 326 2e-89
At2g39620 hypothetical protein 325 6e-89
>At4g31070 putative protein
Length = 613
Score = 568 bits (1465), Expect = e-162
Identities = 291/584 (49%), Positives = 404/584 (68%), Gaps = 7/584 (1%)
Query: 21 QIKSLISQGLYHQTLELFT-QLHFSAPHSISSVLPSVIKACS-SLHSHTLGTLLHGLALT 78
++K L+S Y + L L+ ++H + +++LPSVIKAC+ LG LH L L
Sbjct: 16 KLKGLVSDQFYDEALRLYKLKIHSLGTNGFTAILPSVIKACAFQQEPFLLGAQLHCLCLK 75
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G+ D VVSNSLIS+YAKFS + R+VFD M HRD +S+ S+IN+ Q+G L EA+++
Sbjct: 76 AGADCDTVVSNSLISMYAKFSRKYAVRKVFDEMLHRDTVSYCSIINSCCQDGLLYEAMKL 135
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRK-MGSRIGRQIHALVVVDGRVEHSVVLSTALVDFY 197
+K +Y G +PK EL+AS+++LC R S++ R HALV+VD R++ SV+LSTALVD Y
Sbjct: 136 IKEMYFYGFIPKSELVASLLALCTRMGSSSKVARMFHALVLVDERMQESVLLSTALVDMY 195
Query: 198 FRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLI 257
+ DD A VFD MEVKNEVSWTAMISGC+AN +Y+ FRAMQ E + PNRVTL+
Sbjct: 196 LKFDDHAAAFHVFDQMEVKNEVSWTAMISGCVANQNYEMGVDLFRAMQRENLRPNRVTLL 255
Query: 258 ALLPACAKPGFVKH-GKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSS 316
++LPAC + + KEIHG++FRHG + ++A + MYC+ G ++ L+ +FE S
Sbjct: 256 SVLPACVELNYGSSLVKEIHGFSFRHGCHADERLTAAFMTMYCRCG-NVSLSRVLFETSK 314
Query: 317 FRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCG 376
RDVV+WSS+I Y++ GD + + L N+M E E N VTLLA++SACTN + L
Sbjct: 315 VRDVVMWSSMISGYAETGDCSEVMNLLNQMRKEGIEANSVTLLAIVSACTNSTLLSFAST 374
Query: 377 LHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGH 436
+H ILK G I L NAL++MYAKCG L + ++F E+ +D +SWS+MI+AYGLHGH
Sbjct: 375 VHSQILKCGFMSHILLGNALIDMYAKCGSLSAAREVFYELTEKDLVSWSSMINAYGLHGH 434
Query: 437 GEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYA 496
G +AL++F M + G + D + FLA+LSACNHAGLV E Q +F Q Y +P+T+EHYA
Sbjct: 435 GSEALEIFKGMIKGGHEVDDMAFLAILSACNHAGLVEEAQTIFTQA-GKYHMPVTLEHYA 493
Query: 497 CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIA-EMLGPQLIRSEP 555
C ++LLGR GK++DA E+ MPMKPS RIWSSL+SAC+ HGRLD+A +++ +L++SEP
Sbjct: 494 CYINLLGRFGKIDDAFEVTINMPMKPSARIWSSLLSACETHGRLDVAGKIIANELMKSEP 553
Query: 556 DNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
DN ANY LL+ I+ E G++ E+V M+ ++L KCYGFS+IE
Sbjct: 554 DNPANYVLLSKIHTESGNYHAAEEVRRVMQRRKLNKCYGFSKIE 597
Score = 72.4 bits (176), Expect = 7e-13
Identities = 57/225 (25%), Positives = 100/225 (44%), Gaps = 12/225 (5%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
S I G + + L Q+ + S L +++ AC++ + + +H L
Sbjct: 322 SSMISGYAETGDCSEVMNLLNQMRKEGIEANSVTLLAIVSACTNSTLLSFASTVHSQILK 381
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G S ++ N+LI +YAK + +AR+VF + +D +SW+SMINAY +G EALE+
Sbjct: 382 CGFMSHILLGNALIDMYAKCGSLSAAREVFYELTEKDLVSWSSMINAYGLHGHGSEALEI 441
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLS-----TAL 193
KG+ G +++S C + + + + G+ V L L
Sbjct: 442 FKGMIKGGHEVDDMAFLAILSACNH---AGLVEEAQTIFTQAGKYHMPVTLEHYACYINL 498
Query: 194 VDFYFRCDDALMASRVFDGMEVKNEVS-WTAMISGCIANLDYDAA 237
+ + + DDA V M +K W++++S C + D A
Sbjct: 499 LGRFGKIDDAF---EVTINMPMKPSARIWSSLLSACETHGRLDVA 540
>At5g39350 putative protein
Length = 677
Score = 371 bits (953), Expect = e-103
Identities = 205/584 (35%), Positives = 323/584 (55%), Gaps = 8/584 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISS--VLPSVIKACSSLHSHTLGTLLHGLALTT 79
I+ + +GLYH + +F ++ + P V KA L S LG ++HG L +
Sbjct: 87 IRMYVREGLYHDAISVFIRMVSEGVKCVPDGYTYPFVAKAAGELKSMKLGLVVHGRILRS 146
Query: 80 GSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEML 139
D V N+L+++Y F ++ AR VFD M +RD ISWN+MI+ Y +NG + +AL M
Sbjct: 147 WFGRDKYVQNALLAMYMNFGKVEMARDVFDVMKNRDVISWNTMISGYYRNGYMNDALMMF 206
Query: 140 KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
+ + + SM+ +CG +GR +H LV + R+ + + ALV+ Y +
Sbjct: 207 DWMVNESVDLDHATIVSMLPVCGHLKDLEMGRNVHKLVE-EKRLGDKIEVKNALVNMYLK 265
Query: 200 CDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIAL 259
C A VFD ME ++ ++WT MI+G + D + A R MQ EGV PN VT+ +L
Sbjct: 266 CGRMDEARFVFDRMERRDVITWTCMINGYTEDGDVENALELCRLMQFEGVRPNAVTIASL 325
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
+ C V GK +HG+A R + S I ++LI+MY + + + L ++F +S
Sbjct: 326 VSVCGDALKVNDGKCLHGWAVRQQVYSDIIIETSLISMYAKC-KRVDLCFRVFSGASKYH 384
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
WS+II Q AL LF +M E+ EPN TL +++ A L+ L+ +H
Sbjct: 385 TGPWSAIIAGCVQNELVSDALGLFKRMRREDVEPNIATLNSLLPAYAALADLRQAMNIHC 444
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQ----NRDSISWSTMISAYGLHG 435
++ K G S+ + L+++Y+KCG LE + +IF +Q ++D + W +IS YG+HG
Sbjct: 445 YLTKTGFMSSLDAATGLVHVYSKCGTLESAHKIFNGIQEKHKSKDVVLWGALISGYGMHG 504
Query: 436 HGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHY 495
G AL++F EM GV P+ +TF + L+AC+H+GLV EG +F+ + Y+ HY
Sbjct: 505 DGHNALQVFMEMVRSGVTPNEITFTSALNACSHSGLVEEGLTLFRFMLEHYKTLARSNHY 564
Query: 496 ACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEP 555
C+VDLLGR+G++++A ++ T+P +P++ +W +L++AC H + + EM +L EP
Sbjct: 565 TCIVDLLGRAGRLDEAYNLITTIPFEPTSTVWGALLAACVTHENVQLGEMAANKLFELEP 624
Query: 556 DNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
+N NY LL IYA G W E+V M+ L+K G S IE
Sbjct: 625 ENTGNYVLLANIYAALGRWKDMEKVRSMMENVGLRKKPGHSTIE 668
Score = 168 bits (426), Expect = 7e-42
Identities = 130/519 (25%), Positives = 234/519 (45%), Gaps = 30/519 (5%)
Query: 42 HFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDI 101
HF+A SIS +LH H + TG + ++L YA I
Sbjct: 24 HFAATQSISKT--------KALHCHVI----------TGGRVSGHILSTLSVTYALCGHI 65
Query: 102 QSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGL--VPKPELLASMVS 159
AR++F+ MP +S+N +I Y++ G +A+ + + G+ VP +
Sbjct: 66 TYARKLFEEMPQSSLLSYNIVIRMYVREGLYHDAISVFIRMVSEGVKCVPDGYTYPFVAK 125
Query: 160 LCGRKMGSRIGRQIHALVVVD--GRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKN 217
G ++G +H ++ GR ++ + AL+ Y MA VFD M+ ++
Sbjct: 126 AAGELKSMKLGLVVHGRILRSWFGRDKY---VQNALLAMYMNFGKVEMARDVFDVMKNRD 182
Query: 218 EVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHG 277
+SW MISG N + A F M E VD + T++++LP C ++ G+ +H
Sbjct: 183 VISWNTMISGYYRNGYMNDALMMFDWMVNESVDLDHATIVSMLPVCGHLKDLEMGRNVHK 242
Query: 278 YAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSY 337
+ +AL+NMY + G + A +F+R RDV+ W+ +I Y++ GD
Sbjct: 243 LVEEKRLGDKIEVKNALVNMYLKCGR-MDEARFVFDRMERRDVITWTCMINGYTEDGDVE 301
Query: 338 KALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALM 397
AL L M E PN VT+ +++S C + + G LHG+ ++ + I + +L+
Sbjct: 302 NALELCRLMQFEGVRPNAVTIASLVSVCGDALKVNDGKCLHGWAVRQQVYSDIIIETSLI 361
Query: 398 NMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDAL 457
+MYAKC ++ ++F + WS +I+ + AL LF M+ V+P+
Sbjct: 362 SMYAKCKRVDLCFRVFSGASKYHTGPWSAIIAGCVQNELVSDALGLFKRMRREDVEPNIA 421
Query: 458 TFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRT 517
T ++L A + + + + +++ LV + + G +E A +I
Sbjct: 422 TLNSLLPAYAALADLRQAMNIHCYLTKT-GFMSSLDAATGLVHVYSKCGTLESAHKIFNG 480
Query: 518 MPMKPSTR---IWSSLVSACKLHGRLDIAEMLGPQLIRS 553
+ K ++ +W +L+S +HG A + +++RS
Sbjct: 481 IQEKHKSKDVVLWGALISGYGMHGDGHNALQVFMEMVRS 519
>At5g27110 putative protein
Length = 691
Score = 369 bits (947), Expect = e-102
Identities = 210/578 (36%), Positives = 338/578 (58%), Gaps = 8/578 (1%)
Query: 30 LYHQTLELFTQL---HFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPV 86
++H TLE+F +L P S + P+VIKA +L LG ++H L + +G D V
Sbjct: 86 MFHDTLEVFKRLLNCSICVPDSFT--FPNVIKAYGALGREFLGRMIHTLVVKSGYVCDVV 143
Query: 87 VSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLG 146
V++SL+ +YAKF+ +++ QVFD MP RD SWN++I+ + Q+G E+ALE+ + G
Sbjct: 144 VASSLVGMYAKFNLFENSLQVFDEMPERDVASWNTVISCFYQSGEAEKALELFGRMESSG 203
Query: 147 LVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMA 206
P L +S C R + G++IH V G E +++ALVD Y +CD +A
Sbjct: 204 FEPNSVSLTVAISACSRLLWLERGKEIHRKCVKKG-FELDEYVNSALVDMYGKCDCLEVA 262
Query: 207 SRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKP 266
VF M K+ V+W +MI G +A D + M +EG P++ TL ++L AC++
Sbjct: 263 REVFQKMPRKSLVAWNSMIKGYVAKGDSKSCVEILNRMIIEGTRPSQTTLTSILMACSRS 322
Query: 267 GFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSI 326
+ HGK IHGY R + + + +LI++Y + GE+ +LAE +F ++ W+ +
Sbjct: 323 RNLLHGKFIHGYVIRSVVNADIYVNCSLIDLYFKCGEA-NLAETVFSKTQKDVAESWNVM 381
Query: 327 IGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGL 386
I SY G+ +KA+ ++++M++ +P+ VT +V+ AC+ L++L+ G +H I + L
Sbjct: 382 ISSYISVGNWFKAVEVYDQMVSVGVKPDVVTFTSVLPACSQLAALEKGKQIHLSISESRL 441
Query: 387 SFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSE 446
L +AL++MY+KCG + + +IF + +D +SW+ MISAYG HG +AL F E
Sbjct: 442 ETDELLLSALLDMYSKCGNEKEAFRIFNSIPKKDVVSWTVMISAYGSHGQPREALYQFDE 501
Query: 447 MKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSG 506
M++ G+KPD +T LAVLSAC HAGL+ EG + F Q+ + Y I IEHY+C++D+LGR+G
Sbjct: 502 MQKFGLKPDGVTLLAVLSACGHAGLIDEGLKFFSQMRSKYGIEPIIEHYSCMIDILGRAG 561
Query: 507 KVEDALEIMRTMP-MKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLN 565
++ +A EI++ P + + S+L SAC LH + + + L+ + PD+A+ Y +L
Sbjct: 562 RLLEAYEIIQQTPETSDNAELLSTLFSACCLHLEHSLGDRIARLLVENYPDDASTYMVLF 621
Query: 566 MIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQ 603
+YA W +V MK L+K G S IE ++
Sbjct: 622 NLYASGESWDAARRVRLKMKEMGLRKKPGCSWIEMSDK 659
Score = 237 bits (605), Expect = 1e-62
Identities = 148/493 (30%), Positives = 267/493 (54%), Gaps = 7/493 (1%)
Query: 50 SSVLPSVIKACS-SLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVF 108
SS L S+++ C+ S S L+H LT G D V+ SLI++Y D SAR VF
Sbjct: 3 SSKLLSLLRECTNSTKSLRRIKLVHQRILTLGLRRDVVLCKSLINVYFTCKDHCSARHVF 62
Query: 109 DTMPHR-DPISWNSMINAYLQNGCLEEALEMLKGVYLLGL-VPKPELLASMVSLCGRKMG 166
+ R D WNS+++ Y +N + LE+ K + + VP +++ G
Sbjct: 63 ENFDIRSDVYIWNSLMSGYSKNSMFHDTLEVFKRLLNCSICVPDSFTFPNVIKAYGALGR 122
Query: 167 SRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMIS 226
+GR IH LVV G V VV++++LV Y + + + +VFD M ++ SW +IS
Sbjct: 123 EFLGRMIHTLVVKSGYV-CDVVVASSLVGMYAKFNLFENSLQVFDEMPERDVASWNTVIS 181
Query: 227 GCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIES 286
+ + + A F M+ G +PN V+L + AC++ +++ GKEIH + G E
Sbjct: 182 CFYQSGEAEKALELFGRMESSGFEPNSVSLTVAISACSRLLWLERGKEIHRKCVKKGFEL 241
Query: 287 SHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKM 346
+SAL++MY + + L +A ++F++ + +V W+S+I Y +GDS + + N+M
Sbjct: 242 DEYVNSALVDMYGKC-DCLEVAREVFQKMPRKSLVAWNSMIKGYVAKGDSKSCVEILNRM 300
Query: 347 LTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCL 406
+ E T P+ TL +++ AC+ +L HG +HG++++ ++ I ++ +L+++Y KCG
Sbjct: 301 IIEGTRPSQTTLTSILMACSRSRNLLHGKFIHGYVIRSVVNADIYVNCSLIDLYFKCGEA 360
Query: 407 EGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSAC 466
+ +F + Q + SW+ MIS+Y G+ +A++++ +M GVKPD +TF +VL AC
Sbjct: 361 NLAETVFSKTQKDVAESWNVMISSYISVGNWFKAVEVYDQMVSVGVKPDVVTFTSVLPAC 420
Query: 467 NHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRI 526
+ + +G+Q+ + ++ + + L+D+ + G ++A I ++P K
Sbjct: 421 SQLAALEKGKQIHLSI-SESRLETDELLLSALLDMYSKCGNEKEAFRIFNSIPKKDVVS- 478
Query: 527 WSSLVSACKLHGR 539
W+ ++SA HG+
Sbjct: 479 WTVMISAYGSHGQ 491
Score = 178 bits (452), Expect = 7e-45
Identities = 130/451 (28%), Positives = 219/451 (47%), Gaps = 16/451 (3%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSA--PHSISSVLPSVIKACSSLHSHTLGTLLHGLALTT 79
I G + LELF ++ S P+S+S L I ACS L G +H +
Sbjct: 180 ISCFYQSGEAEKALELFGRMESSGFEPNSVS--LTVAISACSRLLWLERGKEIHRKCVKK 237
Query: 80 GSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEML 139
G D V+++L+ +Y K ++ AR+VF MP + ++WNSMI Y+ G + +E+L
Sbjct: 238 GFELDEYVNSALVDMYGKCDCLEVAREVFQKMPRKSLVAWNSMIKGYVAKGDSKSCVEIL 297
Query: 140 KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
+ + G P L S++ C R G+ IH V+ V + ++ +L+D YF+
Sbjct: 298 NRMIIEGTRPSQTTLTSILMACSRSRNLLHGKFIHG-YVIRSVVNADIYVNCSLIDLYFK 356
Query: 200 CDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIAL 259
C +A +A VF + SW MIS I+ ++ A + M GV P+ VT ++
Sbjct: 357 CGEANLAETVFSKTQKDVAESWNVMISSYISVGNWFKAVEVYDQMVSVGVKPDVVTFTSV 416
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
LPAC++ ++ GK+IH +E+ + SAL++MY + G A +IF +D
Sbjct: 417 LPACSQLAALEKGKQIHLSISESRLETDELLLSALLDMYSKCGNEKE-AFRIFNSIPKKD 475
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
VV W+ +I +Y G +AL F++M +P+ VTLLAV+SAC + + G
Sbjct: 476 VVSWTVMISAYGSHGQPREALYQFDEMQKFGLKPDGVTLLAVLSACGHAGLIDEGLKFFS 535
Query: 380 FI-LKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSIS--WSTMISAYGL--- 433
+ K+G+ I + ++++ + G L + +I + + ST+ SA L
Sbjct: 536 QMRSKYGIEPIIEHYSCMIDILGRAGRLLEAYEIIQQTPETSDNAELLSTLFSACCLHLE 595
Query: 434 HGHGEQALKLFSEMKERGVKPDALTFLAVLS 464
H G++ +L E DA T++ + +
Sbjct: 596 HSLGDRIARLLVE----NYPDDASTYMVLFN 622
Score = 125 bits (314), Expect = 7e-29
Identities = 85/294 (28%), Positives = 144/294 (48%), Gaps = 2/294 (0%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
+ IK +++G +E+ ++ + L S++ ACS + G +HG +
Sbjct: 278 NSMIKGYVAKGDSKSCVEILNRMIIEGTRPSQTTLTSILMACSRSRNLLHGKFIHGYVIR 337
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
+ ++D V+ SLI LY K + A VF SWN MI++Y+ G +A+E+
Sbjct: 338 SVVNADIYVNCSLIDLYFKCGEANLAETVFSKTQKDVAESWNVMISSYISVGNWFKAVEV 397
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+ +G+ P S++ C + G+QIH L + + R+E +L +AL+D Y
Sbjct: 398 YDQMVSVGVKPDVVTFTSVLPACSQLAALEKGKQIH-LSISESRLETDELLLSALLDMYS 456
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
+C + A R+F+ + K+ VSWT MIS ++ A +F MQ G+ P+ VTL+A
Sbjct: 457 KCGNEKEAFRIFNSIPKKDVVSWTVMISAYGSHGQPREALYQFDEMQKFGLKPDGVTLLA 516
Query: 259 LLPACAKPGFVKHG-KEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQI 311
+L AC G + G K ++GIE S +I++ ++G L E I
Sbjct: 517 VLSACGHAGLIDEGLKFFSQMRSKYGIEPIIEHYSCMIDILGRAGRLLEAYEII 570
>At3g03580 unknown protein (At3g03580)
Length = 882
Score = 352 bits (904), Expect = 3e-97
Identities = 204/583 (34%), Positives = 328/583 (55%), Gaps = 9/583 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I S G Y + LE++ +L S S + SV+ A +L G LHG AL +G
Sbjct: 179 ISGYSSHGYYEEALEIYHELKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSGV 238
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
+S VV+N L+++Y KF AR+VFD M RD +S+N+MI YL+ +EE++ M
Sbjct: 239 NSVVVVNNGLVAMYLKFRRPTDARRVFDEMDVRDSVSYNTMICGYLKLEMVEESVRM--- 295
Query: 142 VYLLGLVP-KPELL--ASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+L L KP+LL +S++ CG + + I+ ++ G V S V + L+D Y
Sbjct: 296 -FLENLDQFKPDLLTVSSVLRACGHLRDLSLAKYIYNYMLKAGFVLESTVRNI-LIDVYA 353
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
+C D + A VF+ ME K+ VSW ++ISG I + D A F+ M + + +T +
Sbjct: 354 KCGDMITARDVFNSMECKDTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLM 413
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR 318
L+ + +K GK +H + GI S+ALI+MY + GE + + +IF
Sbjct: 414 LISVSTRLADLKFGKGLHSNGIKSGICIDLSVSNALIDMYAKCGE-VGDSLKIFSSMGTG 472
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLH 378
D V W+++I + + GD L + +M E P+ T L + C +L++ + G +H
Sbjct: 473 DTVTWNTVISACVRFGDFATGLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKEIH 532
Query: 379 GFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGE 438
+L+FG + + NAL+ MY+KCGCLE S ++F M RD ++W+ MI AYG++G GE
Sbjct: 533 CCLLRFGYESELQIGNALIEMYSKCGCLENSSRVFERMSRRDVVTWTGMIYAYGMYGEGE 592
Query: 439 QALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACL 498
+AL+ F++M++ G+ PD++ F+A++ AC+H+GLV EG F+++ Y+I IEHYAC+
Sbjct: 593 KALETFADMEKSGIVPDSVVFIAIIYACSHSGLVDEGLACFEKMKTHYKIDPMIEHYACV 652
Query: 499 VDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNA 558
VDLL RS K+ A E ++ MP+KP IW+S++ AC+ G ++ AE + ++I PD+
Sbjct: 653 VDLLSRSQKISKAEEFIQAMPIKPDASIWASVLRACRTSGDMETAERVSRRIIELNPDDP 712
Query: 559 ANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETG 601
L + YA W + +++K + + K G+S IE G
Sbjct: 713 GYSILASNAYAALRKWDKVSLIRKSLKDKHITKNPGYSWIEVG 755
Score = 258 bits (660), Expect = 5e-69
Identities = 163/518 (31%), Positives = 270/518 (51%), Gaps = 7/518 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I++ GL+ + LE + +L S PSVIKAC+ L +G L++ L G
Sbjct: 78 IRAFSKNGLFPEALEFYGKLRESKVSPDKYTFPSVIKACAGLFDAEMGDLVYEQILDMGF 137
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
SD V N+L+ +Y++ + ARQVFD MP RD +SWNS+I+ Y +G EEALE+
Sbjct: 138 ESDLFVGNALVDMYSRMGLLTRARQVFDEMPVRDLVSWNSLISGYSSHGYYEEALEIYHE 197
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ +VP ++S++ G + + G+ +H + G V VV++ LV Y +
Sbjct: 198 LKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSG-VNSVVVVNNGLVAMYLKFR 256
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
A RVFD M+V++ VS+ MI G + L+ R ++ P+ +T+ ++L
Sbjct: 257 RPTDARRVFDEMDVRDSVSYNTMICGYL-KLEMVEESVRMFLENLDQFKPDLLTVSSVLR 315
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVV 321
AC + K I+ Y + G + LI++Y + G+ + A +F +D V
Sbjct: 316 ACGHLRDLSLAKYIYNYMLKAGFVLESTVRNILIDVYAKCGDMI-TARDVFNSMECKDTV 374
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFI 381
W+SII Y Q GD +A+ LF M+ E + +++T L +IS T L+ LK G GLH
Sbjct: 375 SWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISVSTRLADLKFGKGLHSNG 434
Query: 382 LKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQAL 441
+K G+ +S+SNAL++MYAKCG + SL+IF M D+++W+T+ISA G L
Sbjct: 435 IKSGICIDLSVSNALIDMYAKCGEVGDSLKIFSSMGTGDTVTWNTVISACVRFGDFATGL 494
Query: 442 KLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVF-KQVNADYEIPLTIEHYACLVD 500
++ ++M++ V PD TFL L C G+++ + YE L I + L++
Sbjct: 495 QVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKEIHCCLLRFGYESELQIGN--ALIE 552
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
+ + G +E++ + M + W+ ++ A ++G
Sbjct: 553 MYSKCGCLENSSRVFERMSRR-DVVTWTGMIYAYGMYG 589
Score = 198 bits (504), Expect = 6e-51
Identities = 142/529 (26%), Positives = 266/529 (49%), Gaps = 10/529 (1%)
Query: 54 PSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTM-P 112
P + +A SS + +H L ++ G S S LI Y+ F + S+ VF + P
Sbjct: 8 PFISRALSSSSNLNELRRIHALVISLGLDSSDFFSGKLIDKYSHFREPASSLSVFRRVSP 67
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQ 172
++ WNS+I A+ +NG EALE + + P S++ C + +G
Sbjct: 68 AKNVYLWNSIIRAFSKNGLFPEALEFYGKLRESKVSPDKYTFPSVIKACAGLFDAEMGDL 127
Query: 173 IHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANL 232
++ ++D E + + ALVD Y R A +VFD M V++ VSW ++ISG ++
Sbjct: 128 VYE-QILDMGFESDLFVGNALVDMYSRMGLLTRARQVFDEMPVRDLVSWNSLISGYSSHG 186
Query: 233 DYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSS 292
Y+ A + ++ + P+ T+ ++LPA VK G+ +HG+A + G+ S + ++
Sbjct: 187 YYEEALEIYHELKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSGVNSVVVVNN 246
Query: 293 ALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE 352
L+ MY + A ++F+ RD V ++++I Y + +++ +F + L ++ +
Sbjct: 247 GLVAMYLKFRRPTD-ARRVFDEMDVRDSVSYNTMICGYLKLEMVEESVRMFLENL-DQFK 304
Query: 353 PNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQI 412
P+ +T+ +V+ AC +L L ++ ++LK G ++ N L+++YAKCG + + +
Sbjct: 305 PDLLTVSSVLRACGHLRDLSLAKYIYNYMLKAGFVLESTVRNILIDVYAKCGDMITARDV 364
Query: 413 FLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLV 472
F M+ +D++SW+++IS Y G +A+KLF M + D +T+L ++S +
Sbjct: 365 FNSMECKDTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISVSTRLADL 424
Query: 473 TEGQQVFKQ-VNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV 531
G+ + + + I L++ + L+D+ + G+V D+L+I +M T W++++
Sbjct: 425 KFGKGLHSNGIKSGICIDLSVSN--ALIDMYAKCGEVGDSLKIFSSMG-TGDTVTWNTVI 481
Query: 532 SACKLHGRLDIAEMLGPQLIRSE--PDNAANYTLLNMIYAEHGHWLHKE 578
SAC G + Q+ +SE PD A L M + L KE
Sbjct: 482 SACVRFGDFATGLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKE 530
Score = 108 bits (269), Expect = 1e-23
Identities = 81/346 (23%), Positives = 151/346 (43%), Gaps = 3/346 (0%)
Query: 15 TPSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHG 74
T S + I I G + ++LF + + +I + L G LH
Sbjct: 373 TVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISVSTRLADLKFGKGLHS 432
Query: 75 LALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEE 134
+ +G D VSN+LI +YAK ++ + ++F +M D ++WN++I+A ++ G
Sbjct: 433 NGIKSGICIDLSVSNALIDMYAKCGEVGDSLKIFSSMGTGDTVTWNTVISACVRFGDFAT 492
Query: 135 ALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALV 194
L++ + +VP + +C R+G++IH ++ G E + + AL+
Sbjct: 493 GLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKEIHCCLLRFG-YESELQIGNALI 551
Query: 195 DFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRV 254
+ Y +C +SRVF+ M ++ V+WT MI + + A F M+ G+ P+ V
Sbjct: 552 EMYSKCGCLENSSRVFERMSRRDVVTWTGMIYAYGMYGEGEKALETFADMEKSGIVPDSV 611
Query: 255 TLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFER 314
IA++ AC+ G V G H I A + + + AE+ +
Sbjct: 612 VFIAIIYACSHSGLVDEGLACFEKMKTHYKIDPMIEHYACVVDLLSRSQKISKAEEFIQA 671
Query: 315 SSFR-DVVLWSSIIGSYSQRGDSYKALTLFNKML-TEETEPNYVTL 358
+ D +W+S++ + GD A + +++ +P Y L
Sbjct: 672 MPIKPDASIWASVLRACRTSGDMETAERVSRRIIELNPDDPGYSIL 717
>At3g57430 putative protein
Length = 803
Score = 351 bits (901), Expect = 6e-97
Identities = 197/598 (32%), Positives = 336/598 (55%), Gaps = 18/598 (3%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSL---HSHTLGTLLHGLALT 78
I SL S + LE F + S L SV+ ACS+L +G +H L
Sbjct: 84 ISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMMGKQVHAYGLR 143
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G + ++ N+L+++Y K + S++ + + RD ++WN+++++ QN L EALE
Sbjct: 144 KGELNSFII-NTLVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVLSSLCQNEQLLEALEY 202
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
L+ + L G+ P ++S++ C R G+++HA + +G ++ + + +ALVD Y
Sbjct: 203 LREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENSFVGSALVDMYC 262
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVE-GVDPNRVTLI 257
C L RVFDGM + W AMI+G N A F M+ G+ N T+
Sbjct: 263 NCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIGMEESAGLLANSTTMA 322
Query: 258 ALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSF 317
++PAC + G + IHG+ + G++ + L++MY + G+ + +A +IF +
Sbjct: 323 GVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLGK-IDIAMRIFGKMED 381
Query: 318 RDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE-----------PNYVTLLAVISACT 366
RD+V W+++I Y AL L +KM E + PN +TL+ ++ +C
Sbjct: 382 RDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLKPNSITLMTILPSCA 441
Query: 367 NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
LS+L G +H + +K L+ +++ +AL++MYAKCGCL+ S ++F ++ ++ I+W+
Sbjct: 442 ALSALAKGKEIHAYAIKNNLATDVAVGSALVDMYAKCGCLQMSRKVFDQIPQKNVITWNV 501
Query: 427 MISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADY 486
+I AYG+HG+G++A+ L M +GVKP+ +TF++V +AC+H+G+V EG ++F + DY
Sbjct: 502 IIMAYGMHGNGQEAIDLLRMMMVQGVKPNEVTFISVFAACSHSGMVDEGLRIFYVMKPDY 561
Query: 487 EIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMK-PSTRIWSSLVSACKLHGRLDIAEM 545
+ + +HYAC+VDLLGR+G++++A ++M MP WSSL+ A ++H L+I E+
Sbjct: 562 GVEPSSDHYACVVDLLGRAGRIKEAYQLMNMMPRDFNKAGAWSSLLGASRIHNNLEIGEI 621
Query: 546 LGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQ 603
LI+ EP+ A++Y LL IY+ G W +V MK Q ++K G S IE G++
Sbjct: 622 AAQNLIQLEPNVASHYVLLANIYSSAGLWDKATEVRRNMKEQGVRKEPGCSWIEHGDE 679
Score = 216 bits (550), Expect = 3e-56
Identities = 146/522 (27%), Positives = 269/522 (50%), Gaps = 30/522 (5%)
Query: 54 PSVIKACSSLHSHTLGTLLHGLALTTGSHSDPV-VSNSLISLYAKFSDIQSARQVFDTMP 112
P+++KA + L LG +H G D V V+N+L++LY K D + +VFD +
Sbjct: 14 PALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDFGAVYKVFDRIS 73
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGR---KMGSRI 169
R+ +SWNS+I++ E ALE + + + P L S+V+ C G +
Sbjct: 74 ERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMM 133
Query: 170 GRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCI 229
G+Q+HA + G + +S +++T LV Y + + + ++ V+W ++S
Sbjct: 134 GKQVHAYGLRKGEL-NSFIINT-LVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVLSSLC 191
Query: 230 ANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHG-IESSH 288
N A R M +EGV+P+ T+ ++LPAC+ ++ GKE+H YA ++G ++ +
Sbjct: 192 QNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENS 251
Query: 289 IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLT 348
SAL++MYC + L ++F+ R + LW+++I YSQ +AL LF M
Sbjct: 252 FVGSALVDMYCNCKQVLS-GRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIGM-- 308
Query: 349 EETE---PNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGC 405
EE+ N T+ V+ AC + +HGF++K GL + N LM+MY++ G
Sbjct: 309 EESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLGK 368
Query: 406 LEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK--ERGV---------KP 454
++ +++IF +M++RD ++W+TMI+ Y H E AL L +M+ ER V KP
Sbjct: 369 IDIAMRIFGKMEDRDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLKP 428
Query: 455 DALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEI 514
+++T + +L +C + +G+++ + + + + LVD+ + G ++ + ++
Sbjct: 429 NSITLMTILPSCAALSALAKGKEIHAYAIKN-NLATDVAVGSALVDMYAKCGCLQMSRKV 487
Query: 515 MRTMPMKPSTRIWSSLVSACKLHGR----LDIAEMLGPQLIR 552
+P K + W+ ++ A +HG +D+ M+ Q ++
Sbjct: 488 FDQIPQK-NVITWNVIIMAYGMHGNGQEAIDLLRMMMVQGVK 528
Score = 164 bits (415), Expect = 1e-40
Identities = 114/399 (28%), Positives = 205/399 (50%), Gaps = 19/399 (4%)
Query: 144 LLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDA 203
+LG+ P +++ +G+QIHA V G SV ++ LV+ Y +C D
Sbjct: 3 VLGIKPDNYAFPALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDF 62
Query: 204 LMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPAC 263
+VFD + +N+VSW ++IS + ++ A FR M E V+P+ TL++++ AC
Sbjct: 63 GAVYKVFDRISERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTAC 122
Query: 264 AK---PGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
+ P + GK++H Y R G +S I ++ L+ MY + G+ L ++ + RD+
Sbjct: 123 SNLPMPEGLMMGKQVHAYGLRKGELNSFIINT-LVAMYGKLGK-LASSKVLLGSFGGRDL 180
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
V W++++ S Q +AL +M+ E EP+ T+ +V+ AC++L L+ G LH +
Sbjct: 181 VTWNTVLSSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAY 240
Query: 381 ILKFG-LSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
LK G L + + +AL++MY C + ++F M +R W+ MI+ Y + H ++
Sbjct: 241 ALKNGSLDENSFVGSALVDMYCNCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKE 300
Query: 440 ALKLFSEMKE-RGVKPDALTFLAVLSACNHAGLVT-----EGQQVFKQVNADYEIPLTIE 493
AL LF M+E G+ ++ T V+ AC +G + G V + ++ D + T
Sbjct: 301 ALLLFIGMEESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNT-- 358
Query: 494 HYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVS 532
L+D+ R GK++ A+ I M + W+++++
Sbjct: 359 ----LMDMYSRLGKIDIAMRIFGKMEDRDLV-TWNTMIT 392
Score = 115 bits (289), Expect = 5e-26
Identities = 78/240 (32%), Positives = 129/240 (53%), Gaps = 8/240 (3%)
Query: 244 MQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHG--YAFRHGIESSHIFSSALINMYCQS 301
M V G+ P+ ALL A A ++ GK+IH Y F +G++S + ++ L+N+Y +
Sbjct: 1 MIVLGIKPDNYAFPALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTV-ANTLVNLYRKC 59
Query: 302 GESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAV 361
G+ ++F+R S R+ V W+S+I S AL F ML E EP+ TL++V
Sbjct: 60 GD-FGAVYKVFDRISERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSV 118
Query: 362 ISACTNL---SSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQN 418
++AC+NL L G +H + L+ G + + N L+ MY K G L S +
Sbjct: 119 VTACSNLPMPEGLMMGKQVHAYGLRKG-ELNSFIINTLVAMYGKLGKLASSKVLLGSFGG 177
Query: 419 RDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQV 478
RD ++W+T++S+ + +AL+ EM GV+PD T +VL AC+H ++ G+++
Sbjct: 178 RDLVTWNTVLSSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKEL 237
>At3g09040 hypothetical protein
Length = 1028
Score = 351 bits (901), Expect = 6e-97
Identities = 189/580 (32%), Positives = 318/580 (54%), Gaps = 4/580 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I+ G H+ +ELF + S + S++ C++ H +G+ H + +
Sbjct: 400 IRGYAHNGESHKVMELFMDMKSSGYNIDDFTFTSLLSTCAASHDLEMGSQFHSIIIKKKL 459
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
+ V N+L+ +YAK ++ ARQ+F+ M RD ++WN++I +Y+Q+ EA ++ K
Sbjct: 460 AKNLFVGNALVDMYAKCGALEDARQIFERMCDRDNVTWNTIIGSYVQDENESEAFDLFKR 519
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ L G+V LAS + C G G+Q+H L V G ++ + ++L+D Y +C
Sbjct: 520 MNLCGIVSDGACLASTLKACTHVHGLYQGKQVHCLSVKCG-LDRDLHTGSSLIDMYSKCG 578
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
A +VF + + VS A+I+G N + + A F+ M GV+P+ +T ++
Sbjct: 579 IIKDARKVFSSLPEWSVVSMNALIAGYSQN-NLEEAVVLFQEMLTRGVNPSEITFATIVE 637
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESS-HIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
AC KP + G + HG + G S +L+ MY S E SS + +
Sbjct: 638 ACHKPESLTLGTQFHGQITKRGFSSEGEYLGISLLGMYMNSRGMTEACALFSELSSPKSI 697
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
VLW+ ++ +SQ G +AL + +M + P+ T + V+ C+ LSSL+ G +H
Sbjct: 698 VLWTGMMSGHSQNGFYEEALKFYKEMRHDGVLPDQATFVTVLRVCSVLSSLREGRAIHSL 757
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSI-SWSTMISAYGLHGHGEQ 439
I SN L++MYAKCG ++GS Q+F EM+ R ++ SW+++I+ Y +G+ E
Sbjct: 758 IFHLAHDLDELTSNTLIDMYAKCGDMKGSSQVFDEMRRRSNVVSWNSLINGYAKNGYAED 817
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
ALK+F M++ + PD +TFL VL+AC+HAG V++G+++F+ + Y I ++H AC+V
Sbjct: 818 ALKIFDSMRQSHIMPDEITFLGVLTACSHAGKVSDGRKIFEMMIGQYGIEARVDHVACMV 877
Query: 500 DLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAA 559
DLLGR G +++A + + +KP R+WSSL+ AC++HG E+ +LI EP N++
Sbjct: 878 DLLGRWGYLQEADDFIEAQNLKPDARLWSSLLGACRIHGDDIRGEISAEKLIELEPQNSS 937
Query: 560 NYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
Y LL+ IYA G W + + M+ + +KK G+S I+
Sbjct: 938 AYVLLSNIYASQGCWEKANALRKVMRDRGVKKVPGYSWID 977
Score = 207 bits (526), Expect = 2e-53
Identities = 134/508 (26%), Positives = 252/508 (49%), Gaps = 10/508 (1%)
Query: 35 LELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISL 94
+E F + S+ S S L SV+ A + + LG ++H A+ G S+ V +SL+S+
Sbjct: 312 IEYFFNMRKSSVKSTRSTLGSVLSAIGIVANLDLGLVVHAEAIKLGLASNIYVGSSLVSM 371
Query: 95 YAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELL 154
Y+K +++A +VF+ + ++ + WN+MI Y NG + +E+ + G
Sbjct: 372 YSKCEKMEAAAKVFEALEEKNDVFWNAMIRGYAHNGESHKVMELFMDMKSSGYNIDDFTF 431
Query: 155 ASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGME 214
S++S C +G Q H+ +++ ++ ++ + ALVD Y +C A ++F+ M
Sbjct: 432 TSLLSTCAASHDLEMGSQFHS-IIIKKKLAKNLFVGNALVDMYAKCGALEDARQIFERMC 490
Query: 215 VKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKE 274
++ V+W +I + + + AF F+ M + G+ + L + L AC + GK+
Sbjct: 491 DRDNVTWNTIIGSYVQDENESEAFDLFKRMNLCGIVSDGACLASTLKACTHVHGLYQGKQ 550
Query: 275 IHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRG 334
+H + + G++ S+LI+MY + G + A ++F VV +++I YSQ
Sbjct: 551 VHCLSVKCGLDRDLHTGSSLIDMYSKCG-IIKDARKVFSSLPEWSVVSMNALIAGYSQ-N 608
Query: 335 DSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSIS-LS 393
+ +A+ LF +MLT P+ +T ++ AC SL G HG I K G S L
Sbjct: 609 NLEEAVVLFQEMLTRGVNPSEITFATIVEACHKPESLTLGTQFHGQITKRGFSSEGEYLG 668
Query: 394 NALMNMYAKCGCLEGSLQIFLEMQNRDSI-SWSTMISAYGLHGHGEQALKLFSEMKERGV 452
+L+ MY + + +F E+ + SI W+ M+S + +G E+ALK + EM+ GV
Sbjct: 669 ISLLGMYMNSRGMTEACALFSELSSPKSIVLWTGMMSGHSQNGFYEEALKFYKEMRHDGV 728
Query: 453 KPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYA--CLVDLLGRSGKVED 510
PD TF+ VL C+ + EG+ + + + + ++ L+D+ + G ++
Sbjct: 729 LPDQATFVTVLRVCSVLSSLREGRAIHSLI---FHLAHDLDELTSNTLIDMYAKCGDMKG 785
Query: 511 ALEIMRTMPMKPSTRIWSSLVSACKLHG 538
+ ++ M + + W+SL++ +G
Sbjct: 786 SSQVFDEMRRRSNVVSWNSLINGYAKNG 813
Score = 192 bits (489), Expect = 3e-49
Identities = 132/492 (26%), Positives = 236/492 (47%), Gaps = 11/492 (2%)
Query: 80 GSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEML 139
G D + ++I+ Y + ++ AR +F M D ++WN MI+ + + GC A+E
Sbjct: 256 GHRPDHLAFVTVINTYIRLGKLKDARLLFGEMSSPDVVAWNVMISGHGKRGCETVAIEYF 315
Query: 140 KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
+ + L S++S G +G +HA + G + ++ + ++LV Y +
Sbjct: 316 FNMRKSSVKSTRSTLGSVLSAIGIVANLDLGLVVHAEAIKLG-LASNIYVGSSLVSMYSK 374
Query: 200 CDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIAL 259
C+ A++VF+ +E KN+V W AMI G N + F M+ G + + T +L
Sbjct: 375 CEKMEAAAKVFEALEEKNDVFWNAMIRGYAHNGESHKVMELFMDMKSSGYNIDDFTFTSL 434
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
L CA ++ G + H + + + +AL++MY + G +L A QIFER RD
Sbjct: 435 LSTCAASHDLEMGSQFHSIIIKKKLAKNLFVGNALVDMYAKCG-ALEDARQIFERMCDRD 493
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
V W++IIGSY Q + +A LF +M + L + + ACT++ L G +H
Sbjct: 494 NVTWNTIIGSYVQDENESEAFDLFKRMNLCGIVSDGACLASTLKACTHVHGLYQGKQVHC 553
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
+K GL + ++L++MY+KCG ++ + ++F + +S + +I+ Y + E+
Sbjct: 554 LSVKCGLDRDLHTGSSLIDMYSKCGIIKDARKVFSSLPEWSVVSMNALIAGYS-QNNLEE 612
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
A+ LF EM RGV P +TF ++ AC+ +T G Q Q+ + + E +
Sbjct: 613 AVVLFQEMLTRGVNPSEITFATIVEACHKPESLTLGTQFHGQIT---KRGFSSEGEYLGI 669
Query: 500 DLLG---RSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSE-- 554
LLG S + +A + + S +W+ ++S +G + A ++
Sbjct: 670 SLLGMYMNSRGMTEACALFSELSSPKSIVLWTGMMSGHSQNGFYEEALKFYKEMRHDGVL 729
Query: 555 PDNAANYTLLNM 566
PD A T+L +
Sbjct: 730 PDQATFVTVLRV 741
Score = 175 bits (444), Expect = 6e-44
Identities = 124/467 (26%), Positives = 220/467 (46%), Gaps = 42/467 (8%)
Query: 68 LGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYL 127
+G +H +L G S+ + N+++ LYAK + + A + FD + +D +WNSM++ Y
Sbjct: 78 IGKAVHSKSLILGIDSEGRLGNAIVDLYAKCAQVSYAEKQFDFL-EKDVTAWNSMLSMYS 136
Query: 128 QNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSV 187
G + L ++ + P + ++S C R+ GRQIH ++ G +E +
Sbjct: 137 SIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSMIKMG-LERNS 195
Query: 188 VLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCI-ANLDYDAAFARFRAMQV 246
ALVD Y +CD A RVF+ + N V WT + SG + A L +A F M+
Sbjct: 196 YCGGALVDMYAKCDRISDARRVFEWIVDPNTVCWTCLFSGYVKAGLPEEAVLV-FERMRD 254
Query: 247 EGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLH 306
EG P+ H+ +IN Y + G+ L
Sbjct: 255 EGHRPD-----------------------------------HLAFVTVINTYIRLGK-LK 278
Query: 307 LAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACT 366
A +F S DVV W+ +I + +RG A+ F M + TL +V+SA
Sbjct: 279 DARLLFGEMSSPDVVAWNVMISGHGKRGCETVAIEYFFNMRKSSVKSTRSTLGSVLSAIG 338
Query: 367 NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
+++L G +H +K GL+ +I + ++L++MY+KC +E + ++F ++ ++ + W+
Sbjct: 339 IVANLDLGLVVHAEAIKLGLASNIYVGSSLVSMYSKCEKMEAAAKVFEALEEKNDVFWNA 398
Query: 427 MISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADY 486
MI Y +G + ++LF +MK G D TF ++LS C + + G Q F +
Sbjct: 399 MIRGYAHNGESHKVMELFMDMKSSGYNIDDFTFTSLLSTCAASHDLEMGSQ-FHSIIIKK 457
Query: 487 EIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSA 533
++ + LVD+ + G +EDA +I M + + W++++ +
Sbjct: 458 KLAKNLFVGNALVDMYAKCGALEDARQIFERMCDRDNV-TWNTIIGS 503
Score = 107 bits (267), Expect = 2e-23
Identities = 69/284 (24%), Positives = 143/284 (50%), Gaps = 10/284 (3%)
Query: 272 GKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYS 331
GK +H + GI+S +A++++Y + + + AE+ F+ +DV W+S++ YS
Sbjct: 79 GKAVHSKSLILGIDSEGRLGNAIVDLYAKCAQ-VSYAEKQFDFLE-KDVTAWNSMLSMYS 136
Query: 332 QRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSIS 391
G K L F + + PN T V+S C ++++ G +H ++K GL +
Sbjct: 137 SIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSMIKMGLERNSY 196
Query: 392 LSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERG 451
AL++MYAKC + + ++F + + +++ W+ + S Y G E+A+ +F M++ G
Sbjct: 197 CGGALVDMYAKCDRISDARRVFEWIVDPNTVCWTCLFSGYVKAGLPEEAVLVFERMRDEG 256
Query: 452 VKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDA 511
+PD L F+ V++ G + + + +F ++++ + + ++ G+ G A
Sbjct: 257 HRPDHLAFVTVINTYIRLGKLKDARLLFGEMSSPDVVAWNV-----MISGHGKRGCETVA 311
Query: 512 LEI---MRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIR 552
+E MR +K + S++SA + LD+ ++ + I+
Sbjct: 312 IEYFFNMRKSSVKSTRSTLGSVLSAIGIVANLDLGLVVHAEAIK 355
Score = 104 bits (260), Expect = 1e-22
Identities = 85/366 (23%), Positives = 161/366 (43%), Gaps = 44/366 (12%)
Query: 168 RIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISG 227
RIG+ +H+ ++ G ++ L A+VD Y +C A + FD +E K+ +W +M+S
Sbjct: 77 RIGKAVHSKSLILG-IDSEGRLGNAIVDLYAKCAQVSYAEKQFDFLE-KDVTAWNSMLSM 134
Query: 228 CIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESS 287
+ F ++ + PN+ T +L CA+ V+ G++IH + G+E +
Sbjct: 135 YSSIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSMIKMGLERN 194
Query: 288 HIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKML 347
AL++MY + + + A ++FE + V W+ + Y + G +A+ +F +M
Sbjct: 195 SYCGGALVDMYAKC-DRISDARRVFEWIVDPNTVCWTCLFSGYVKAGLPEEAVLVFERMR 253
Query: 348 TEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLE 407
E P+++ + VI N Y + G L+
Sbjct: 254 DEGHRPDHLAFVTVI-----------------------------------NTYIRLGKLK 278
Query: 408 GSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACN 467
+ +F EM + D ++W+ MIS +G G A++ F M++ VK T +VLSA
Sbjct: 279 DARLLFGEMSSPDVVAWNVMISGHGKRGCETVAIEYFFNMRKSSVKSTRSTLGSVLSAIG 338
Query: 468 HAGLVTEGQQVFKQVNADYEIPLTIEHY--ACLVDLLGRSGKVEDALEIMRTMPMKPSTR 525
+ G V + ++ L Y + LV + + K+E A ++ + K
Sbjct: 339 IVANLDLGLVVHAEA---IKLGLASNIYVGSSLVSMYSKCEKMEAAAKVFEALEEKNDV- 394
Query: 526 IWSSLV 531
W++++
Sbjct: 395 FWNAMI 400
Score = 64.7 bits (156), Expect = 1e-10
Identities = 53/201 (26%), Positives = 97/201 (47%), Gaps = 11/201 (5%)
Query: 370 SLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMIS 429
+L+ G +H L G+ L NA++++YAKC + + + F + +D +W++M+S
Sbjct: 75 ALRIGKAVHSKSLILGIDSEGRLGNAIVDLYAKCAQVSYAEKQF-DFLEKDVTAWNSMLS 133
Query: 430 AYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIP 489
Y G + L+ F + E + P+ TF VLS C V G+Q+ + ++
Sbjct: 134 MYSSIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSM---IKMG 190
Query: 490 LTIEHY--ACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLG 547
L Y LVD+ + ++ DA + + + P+T W+ L S G + A ++
Sbjct: 191 LERNSYCGGALVDMYAKCDRISDARRVFEWI-VDPNTVCWTCLFSGYVKAGLPEEAVLVF 249
Query: 548 PQLIRSE---PDNAANYTLLN 565
++ R E PD+ A T++N
Sbjct: 250 ERM-RDEGHRPDHLAFVTVIN 269
>At1g11290 hypothetical protein
Length = 809
Score = 348 bits (894), Expect = 4e-96
Identities = 190/571 (33%), Positives = 316/571 (55%), Gaps = 2/571 (0%)
Query: 33 QTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLI 92
+ L+ F ++ + + ++K C +G +HGL + +G D L
Sbjct: 118 KALQFFVRMRYDDVEPVVYNFTYLLKVCGDEAELRVGKEIHGLLVKSGFSLDLFAMTGLE 177
Query: 93 SLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE 152
++YAK + AR+VFD MP RD +SWN+++ Y QNG ALEM+K + L P
Sbjct: 178 NMYAKCRQVNEARKVFDRMPERDLVSWNTIVAGYSQNGMARMALEMVKSMCEENLKPSFI 237
Query: 153 LLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDG 212
+ S++ +G++IH + G + V +STALVD Y +C A ++FDG
Sbjct: 238 TIVSVLPAVSALRLISVGKEIHGYAMRSG-FDSLVNISTALVDMYAKCGSLETARQLFDG 296
Query: 213 MEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHG 272
M +N VSW +MI + N + A F+ M EGV P V+++ L ACA G ++ G
Sbjct: 297 MLERNVVSWNSMIDAYVQNENPKEAMLIFQKMLDEGVKPTDVSVMGALHACADLGDLERG 356
Query: 273 KEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQ 332
+ IH + G++ + ++LI+MYC+ E + A +F + R +V W+++I ++Q
Sbjct: 357 RFIHKLSVELGLDRNVSVVNSLISMYCKCKE-VDTAASMFGKLQSRTLVSWNAMILGFAQ 415
Query: 333 RGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISL 392
G AL F++M + +P+ T ++VI+A LS H +HG +++ L ++ +
Sbjct: 416 NGRPIDALNYFSQMRSRTVKPDTFTYVSVITAIAELSITHHAKWIHGVVMRSCLDKNVFV 475
Query: 393 SNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGV 452
+ AL++MYAKCG + + IF M R +W+ MI YG HG G+ AL+LF EM++ +
Sbjct: 476 TTALVDMYAKCGAIMIARLIFDMMSERHVTTWNAMIDGYGTHGFGKAALELFEEMQKGTI 535
Query: 453 KPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDAL 512
KP+ +TFL+V+SAC+H+GLV G + F + +Y I L+++HY +VDLLGR+G++ +A
Sbjct: 536 KPNGVTFLSVISACSHSGLVEAGLKCFYMMKENYSIELSMDHYGAMVDLLGRAGRLNEAW 595
Query: 513 EIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHG 572
+ + MP+KP+ ++ +++ AC++H ++ AE +L PD+ + LL IY
Sbjct: 596 DFIMQMPVKPAVNVYGAMLGACQIHKNVNFAEKAAERLFELNPDDGGYHVLLANIYRAAS 655
Query: 573 HWLHKEQVMEAMKLQRLKKCYGFSRIETGNQ 603
W QV +M Q L+K G S +E N+
Sbjct: 656 MWEKVGQVRVSMLRQGLRKTPGCSMVEIKNE 686
Score = 224 bits (572), Expect = 8e-59
Identities = 136/483 (28%), Positives = 253/483 (52%), Gaps = 7/483 (1%)
Query: 56 VIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRD 115
+++ CSSL L +L L G + + L+SL+ ++ + A +VF+ + +
Sbjct: 43 LLERCSSLKE--LRQILP-LVFKNGLYQEHFFQTKLVSLFCRYGSVDEAARVFEPIDSKL 99
Query: 116 PISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHA 175
+ +++M+ + + L++AL+ + + P ++ +CG + R+G++IH
Sbjct: 100 NVLYHTMLKGFAKVSDLDKALQFFVRMRYDDVEPVVYNFTYLLKVCGDEAELRVGKEIHG 159
Query: 176 LVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYD 235
L+V G + T L + Y +C A +VFD M ++ VSW +++G N
Sbjct: 160 LLVKSG-FSLDLFAMTGLENMYAKCRQVNEARKVFDRMPERDLVSWNTIVAGYSQNGMAR 218
Query: 236 AAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALI 295
A ++M E + P+ +T++++LPA + + GKEIHGYA R G +S S+AL+
Sbjct: 219 MALEMVKSMCEENLKPSFITIVSVLPAVSALRLISVGKEIHGYAMRSGFDSLVNISTALV 278
Query: 296 NMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNY 355
+MY + G SL A Q+F+ R+VV W+S+I +Y Q + +A+ +F KML E +P
Sbjct: 279 DMYAKCG-SLETARQLFDGMLERNVVSWNSMIDAYVQNENPKEAMLIFQKMLDEGVKPTD 337
Query: 356 VTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLE 415
V+++ + AC +L L+ G +H ++ GL ++S+ N+L++MY KC ++ + +F +
Sbjct: 338 VSVMGALHACADLGDLERGRFIHKLSVELGLDRNVSVVNSLISMYCKCKEVDTAASMFGK 397
Query: 416 MQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEG 475
+Q+R +SW+ MI + +G AL FS+M+ R VKPD T+++V++A +
Sbjct: 398 LQSRTLVSWNAMILGFAQNGRPIDALNYFSQMRSRTVKPDTFTYVSVITAIAELSITHHA 457
Query: 476 QQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACK 535
+ + V + + LVD+ + G + A I M + T W++++
Sbjct: 458 KWIHGVVMRSC-LDKNVFVTTALVDMYAKCGAIMIARLIFDMMSERHVT-TWNAMIDGYG 515
Query: 536 LHG 538
HG
Sbjct: 516 THG 518
Score = 174 bits (440), Expect = 2e-43
Identities = 112/412 (27%), Positives = 204/412 (49%), Gaps = 10/412 (2%)
Query: 171 RQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIA 230
RQI LV +G + T LV + R A+RVF+ ++ K V + M+ G
Sbjct: 54 RQILPLVFKNGLYQEHF-FQTKLVSLFCRYGSVDEAARVFEPIDSKLNVLYHTMLKGFAK 112
Query: 231 NLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIF 290
D D A F M+ + V+P LL C ++ GKEIHG + G
Sbjct: 113 VSDLDKALQFFVRMRYDDVEPVVYNFTYLLKVCGDEAELRVGKEIHGLLVKSGFSLDLFA 172
Query: 291 SSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEE 350
+ L NMY + ++ A ++F+R RD+V W++I+ YSQ G + AL + M E
Sbjct: 173 MTGLENMYAKC-RQVNEARKVFDRMPERDLVSWNTIVAGYSQNGMARMALEMVKSMCEEN 231
Query: 351 TEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSL 410
+P+++T+++V+ A + L + G +HG+ ++ G +++S AL++MYAKCG LE +
Sbjct: 232 LKPSFITIVSVLPAVSALRLISVGKEIHGYAMRSGFDSLVNISTALVDMYAKCGSLETAR 291
Query: 411 QIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAG 470
Q+F M R+ +SW++MI AY + + ++A+ +F +M + GVKP ++ + L AC G
Sbjct: 292 QLFDGMLERNVVSWNSMIDAYVQNENPKEAMLIFQKMLDEGVKPTDVSVMGALHACADLG 351
Query: 471 LVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSL 530
+ G+ + K ++ + + + L+ + + +V+ A + + + W+++
Sbjct: 352 DLERGRFIHK-LSVELGLDRNVSVVNSLISMYCKCKEVDTAASMFGKLQSRTLVS-WNAM 409
Query: 531 VSACKLHGR-LDIAEMLGPQLIRS-EPDNAANYTLLNMI----YAEHGHWLH 576
+ +GR +D R+ +PD +++ I H W+H
Sbjct: 410 ILGFAQNGRPIDALNYFSQMRSRTVKPDTFTYVSVITAIAELSITHHAKWIH 461
Score = 125 bits (313), Expect = 9e-29
Identities = 83/330 (25%), Positives = 167/330 (50%), Gaps = 16/330 (4%)
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR 318
LL C+ +K ++I F++G+ H F + L++++C+ G S+ A ++FE +
Sbjct: 43 LLERCSS---LKELRQILPLVFKNGLYQEHFFQTKLVSLFCRYG-SVDEAARVFEPIDSK 98
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLH 378
VL+ +++ +++ D KAL F +M ++ EP ++ C + + L+ G +H
Sbjct: 99 LNVLYHTMLKGFAKVSDLDKALQFFVRMRYDDVEPVVYNFTYLLKVCGDEAELRVGKEIH 158
Query: 379 GFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGE 438
G ++K G S + L NMYAKC + + ++F M RD +SW+T+++ Y +G
Sbjct: 159 GLLVKSGFSLDLFAMTGLENMYAKCRQVNEARKVFDRMPERDLVSWNTIVAGYSQNGMAR 218
Query: 439 QALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVF-KQVNADYEIPLTIEHYAC 497
AL++ M E +KP +T ++VL A + L++ G+++ + + ++ + I
Sbjct: 219 MALEMVKSMCEENLKPSFITIVSVLPAVSALRLISVGKEIHGYAMRSGFDSLVNIS--TA 276
Query: 498 LVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDN 557
LVD+ + G +E A ++ M ++ + W+S++ A + ML Q + E
Sbjct: 277 LVDMYAKCGSLETARQLFDGM-LERNVVSWNSMIDA-YVQNENPKEAMLIFQKMLDEGVK 334
Query: 558 AANYTLLNMIYA-------EHGHWLHKEQV 580
+ +++ ++A E G ++HK V
Sbjct: 335 PTDVSVMGALHACADLGDLERGRFIHKLSV 364
>At4g35130 putative protein
Length = 804
Score = 347 bits (891), Expect = 8e-96
Identities = 202/580 (34%), Positives = 318/580 (54%), Gaps = 8/580 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
IK S GLY + ++ ++++ F+ + + P VIK+ + + S G +H + + G
Sbjct: 102 IKGFTSCGLYIEAVQFYSRMVFAGVKADTFTYPFVIKSVAGISSLEEGKKIHAMVIKLGF 161
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
SD V NSLISLY K A +VF+ MP RD +SWNSMI+ YL G +L + K
Sbjct: 162 VSDVYVCNSLISLYMKLGCAWDAEKVFEEMPERDIVSWNSMISGYLALGDGFSSLMLFKE 221
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ G P S + C ++G++IH V V++ T+++D Y +
Sbjct: 222 MLKCGFKPDRFSTMSALGACSHVYSPKMGKEIHCHAVRSRIETGDVMVMTSILDMYSKYG 281
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIA-NLDYDAAFARFRAMQVE-GVDPNRVTLIAL 259
+ A R+F+GM +N V+W MI GC A N AF F+ M + G+ P+ +T I L
Sbjct: 282 EVSYAERIFNGMIQRNIVAWNVMI-GCYARNGRVTDAFLCFQKMSEQNGLQPDVITSINL 340
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
LPA A + G+ IHGYA R G + +ALI+MY + G+ L AE IF+R + ++
Sbjct: 341 LPASA----ILEGRTIHGYAMRRGFLPHMVLETALIDMYGECGQ-LKSAEVIFDRMAEKN 395
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
V+ W+SII +Y Q G +Y AL LF ++ P+ T+ +++ A SL G +H
Sbjct: 396 VISWNSIIAAYVQNGKNYSALELFQELWDSSLVPDSTTIASILPAYAESLSLSEGREIHA 455
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
+I+K + + N+L++MYA CG LE + + F + +D +SW+++I AY +HG G
Sbjct: 456 YIVKSRYWSNTIILNSLVHMYAMCGDLEDARKCFNHILLKDVVSWNSIIMAYAVHGFGRI 515
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
++ LFSEM V P+ TF ++L+AC+ +G+V EG + F+ + +Y I IEHY C++
Sbjct: 516 SVWLFSEMIASRVNPNKSTFASLLAACSISGMVDEGWEYFESMKREYGIDPGIEHYGCML 575
Query: 500 DLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAA 559
DL+GR+G A + MP P+ RIW SL++A + H + IAE Q+ + E DN
Sbjct: 576 DLIGRTGNFSAAKRFLEEMPFVPTARIWGSLLNASRNHKDITIAEFAAEQIFKMEHDNTG 635
Query: 560 NYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
Y LL +YAE G W ++ M+ + + + S +E
Sbjct: 636 CYVLLLNMYAEAGRWEDVNRIKLLMESKGISRTSSRSTVE 675
Score = 187 bits (474), Expect = 2e-47
Identities = 135/473 (28%), Positives = 241/473 (50%), Gaps = 11/473 (2%)
Query: 83 SDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGV 142
+DP ++ +L +A ++ A Q+FD M D WN MI + G EA++ +
Sbjct: 63 NDPALTRALRG-FADSRLMEDALQLFDEMNKADAFLWNVMIKGFTSCGLYIEAVQFYSRM 121
Query: 143 YLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDD 202
G+ ++ G++IHA+V+ G V V + +L+ Y +
Sbjct: 122 VFAGVKADTFTYPFVIKSVAGISSLEEGKKIHAMVIKLGFVS-DVYVCNSLISLYMKLGC 180
Query: 203 ALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPA 262
A A +VF+ M ++ VSW +MISG +A D ++ F+ M G P+R + ++ L A
Sbjct: 181 AWDAEKVFEEMPERDIVSWNSMISGYLALGDGFSSLMLFKEMLKCGFKPDRFSTMSALGA 240
Query: 263 CAKPGFVKHGKEIHGYAFRHGIESSHIF-SSALINMYCQSGESLHLAEQIFERSSFRDVV 321
C+ K GKEIH +A R IE+ + +++++MY + GE + AE+IF R++V
Sbjct: 241 CSHVYSPKMGKEIHCHAVRSRIETGDVMVMTSILDMYSKYGE-VSYAERIFNGMIQRNIV 299
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEE-TEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
W+ +IG Y++ G A F KM + +P+ +T + ++ A S++ G +HG+
Sbjct: 300 AWNVMIGCYARNGRVTDAFLCFQKMSEQNGLQPDVITSINLLPA----SAILEGRTIHGY 355
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
++ G + L AL++MY +CG L+ + IF M ++ ISW+++I+AY +G A
Sbjct: 356 AMRRGFLPHMVLETALIDMYGECGQLKSAEVIFDRMAEKNVISWNSIIAAYVQNGKNYSA 415
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
L+LF E+ + + PD+ T ++L A + ++EG+++ + TI LV
Sbjct: 416 LELFQELWDSSLVPDSTTIASILPAYAESLSLSEGREIHAYIVKSRYWSNTI-ILNSLVH 474
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRS 553
+ G +EDA + + +K W+S++ A +HG I+ L ++I S
Sbjct: 475 MYAMCGDLEDARKCFNHILLKDVVS-WNSIIMAYAVHGFGRISVWLFSEMIAS 526
Score = 108 bits (270), Expect = 9e-24
Identities = 69/272 (25%), Positives = 131/272 (47%), Gaps = 4/272 (1%)
Query: 308 AEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTN 367
A Q+F+ + D LW+ +I ++ G +A+ +++M+ + + T VI +
Sbjct: 83 ALQLFDEMNKADAFLWNVMIKGFTSCGLYIEAVQFYSRMVFAGVKADTFTYPFVIKSVAG 142
Query: 368 LSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTM 427
+SSL+ G +H ++K G + + N+L+++Y K GC + ++F EM RD +SW++M
Sbjct: 143 ISSLEEGKKIHAMVIKLGFVSDVYVCNSLISLYMKLGCAWDAEKVFEEMPERDIVSWNSM 202
Query: 428 ISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYE 487
IS Y G G +L LF EM + G KPD + ++ L AC+H G+++
Sbjct: 203 ISGYLALGDGFSSLMLFKEMLKCGFKPDRFSTMSALGACSHVYSPKMGKEIHCHAVRSRI 262
Query: 488 IPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLG 547
+ ++D+ + G+V A I M ++ + W+ ++ +GR+ A +
Sbjct: 263 ETGDVMVMTSILDMYSKYGEVSYAERIFNGM-IQRNIVAWNVMIGCYARNGRVTDAFLCF 321
Query: 548 PQLIRS---EPDNAANYTLLNMIYAEHGHWLH 576
++ +PD + LL G +H
Sbjct: 322 QKMSEQNGLQPDVITSINLLPASAILEGRTIH 353
Score = 57.4 bits (137), Expect = 2e-08
Identities = 54/216 (25%), Positives = 98/216 (45%), Gaps = 19/216 (8%)
Query: 391 SLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKER 450
+L+ AL +A +E +LQ+F EM D+ W+ MI + G +A++ +S M
Sbjct: 66 ALTRALRG-FADSRLMEDALQLFDEMNKADAFLWNVMIKGFTSCGLYIEAVQFYSRMVFA 124
Query: 451 GVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYAC--LVDLLGRSGKV 508
GVK D T+ V+ + + EG+++ V ++ + Y C L+ L + G
Sbjct: 125 GVKADTFTYPFVIKSVAGISSLEEGKKIHAMV---IKLGFVSDVYVCNSLISLYMKLGCA 181
Query: 509 EDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRS--EPDNAANYTLLNM 566
DA ++ MP + W+S++S G + ML ++++ +PD + + L
Sbjct: 182 WDAEKVFEEMPERDIVS-WNSMISGYLALGDGFSSLMLFKEMLKCGFKPDRFSTMSALGA 240
Query: 567 IYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGN 602
+ + KE A++ SRIETG+
Sbjct: 241 CSHVYSPKMGKEIHCHAVR----------SRIETGD 266
>At4g21300 putative protein
Length = 857
Score = 347 bits (890), Expect = 1e-95
Identities = 184/545 (33%), Positives = 314/545 (56%), Gaps = 3/545 (0%)
Query: 56 VIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRD 115
V+ C+S LG LHGL + +G + + NSL+S+Y+K A ++F M D
Sbjct: 245 VLSVCASKLLIDLGVQLHGLVVVSGVDFEGSIKNSLLSMYSKCGRFDDASKLFRMMSRAD 304
Query: 116 PISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHA 175
++WN MI+ Y+Q+G +EE+L + G++P +S++ + +QIH
Sbjct: 305 TVTWNCMISGYVQSGLMEESLTFFYEMISSGVLPDAITFSSLLPSVSKFENLEYCKQIHC 364
Query: 176 LVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYD 235
++ + + L++AL+D YF+C MA +F + V +TAMISG + N Y
Sbjct: 365 YIMRHS-ISLDIFLTSALIDAYFKCRGVSMAQNIFSQCNSVDVVVFTAMISGYLHNGLYI 423
Query: 236 AAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALI 295
+ FR + + PN +TL+++LP +K G+E+HG+ + G ++ A+I
Sbjct: 424 DSLEMFRWLVKVKISPNEITLVSILPVIGILLALKLGRELHGFIIKKGFDNRCNIGCAVI 483
Query: 296 NMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNY 355
+MY + G ++LA +IFER S RD+V W+S+I +Q + A+ +F +M +
Sbjct: 484 DMYAKCGR-MNLAYEIFERLSKRDIVSWNSMITRCAQSDNPSAAIDIFRQMGVSGICYDC 542
Query: 356 VTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLE 415
V++ A +SAC NL S G +HGF++K L+ + + L++MYAKCG L+ ++ +F
Sbjct: 543 VSISAALSACANLPSESFGKAIHGFMIKHSLASDVYSESTLIDMYAKCGNLKAAMNVFKT 602
Query: 416 MQNRDSISWSTMISAYGLHGHGEQALKLFSEMKER-GVKPDALTFLAVLSACNHAGLVTE 474
M+ ++ +SW+++I+A G HG + +L LF EM E+ G++PD +TFL ++S+C H G V E
Sbjct: 603 MKEKNIVSWNSIIAACGNHGKLKDSLCLFHEMVEKSGIRPDQITFLEIISSCCHVGDVDE 662
Query: 475 GQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSAC 534
G + F+ + DY I EHYAC+VDL GR+G++ +A E +++MP P +W +L+ AC
Sbjct: 663 GVRFFRSMTEDYGIQPQQEHYACVVDLFGRAGRLTEAYETVKSMPFPPDAGVWGTLLGAC 722
Query: 535 KLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYG 594
+LH +++AE+ +L+ +P N+ Y L++ +A W +V MK + ++K G
Sbjct: 723 RLHKNVELAEVASSKLMDLDPSNSGYYVLISNAHANAREWESVTKVRSLMKEREVQKIPG 782
Query: 595 FSRIE 599
+S IE
Sbjct: 783 YSWIE 787
Score = 234 bits (597), Expect = 1e-61
Identities = 151/547 (27%), Positives = 279/547 (50%), Gaps = 6/547 (1%)
Query: 22 IKSLISQGLYHQTLEL-FTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTG 80
I S + GL +Q L F L F +S+ P ++KAC +L + L + G
Sbjct: 110 ISSFVRNGLLNQALAFYFKMLCFGVSPDVST-FPCLVKACVALKNFKGIDFLSDTVSSLG 168
Query: 81 SHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLK 140
+ V++SLI Y ++ I ++FD + +D + WN M+N Y + G L+ ++
Sbjct: 169 MDCNEFVASSLIKAYLEYGKIDVPSKLFDRVLQKDCVIWNVMLNGYAKCGALDSVIKGFS 228
Query: 141 GVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRC 200
+ + + P ++S+C K+ +G Q+H LVVV G V+ + +L+ Y +C
Sbjct: 229 VMRMDQISPNAVTFDCVLSVCASKLLIDLGVQLHGLVVVSG-VDFEGSIKNSLLSMYSKC 287
Query: 201 DDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALL 260
AS++F M + V+W MISG + + + + F M GV P+ +T +LL
Sbjct: 288 GRFDDASKLFRMMSRADTVTWNCMISGYVQSGLMEESLTFFYEMISSGVLPDAITFSSLL 347
Query: 261 PACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
P+ +K +++ K+IH Y RH I +SALI+ Y + + +A+ IF + + DV
Sbjct: 348 PSVSKFENLEYCKQIHCYIMRHSISLDIFLTSALIDAYFKC-RGVSMAQNIFSQCNSVDV 406
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
V+++++I Y G +L +F ++ + PN +TL++++ L +LK G LHGF
Sbjct: 407 VVFTAMISGYLHNGLYIDSLEMFRWLVKVKISPNEITLVSILPVIGILLALKLGRELHGF 466
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
I+K G ++ A+++MYAKCG + + +IF + RD +SW++MI+ + A
Sbjct: 467 IIKKGFDNRCNIGCAVIDMYAKCGRMNLAYEIFERLSKRDIVSWNSMITRCAQSDNPSAA 526
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
+ +F +M G+ D ++ A LSAC + + G+ + + + + + + L+D
Sbjct: 527 IDIFRQMGVSGICYDCVSISAALSACANLPSESFGKAIHGFM-IKHSLASDVYSESTLID 585
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAAN 560
+ + G ++ A+ + +TM K + W+S+++AC HG+L + L +++
Sbjct: 586 MYAKCGNLKAAMNVFKTMKEK-NIVSWNSIIAACGNHGKLKDSLCLFHEMVEKSGIRPDQ 644
Query: 561 YTLLNMI 567
T L +I
Sbjct: 645 ITFLEII 651
Score = 150 bits (380), Expect = 2e-36
Identities = 110/507 (21%), Positives = 234/507 (45%), Gaps = 9/507 (1%)
Query: 31 YHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNS 90
Y ++L L F +I L +++ACS+ + G +H + D
Sbjct: 17 YKKSLPLRNSSRF-LEETIPRRLSLLLQACSNPNLLRQGKQVHAFLIVNSISGDSYTDER 75
Query: 91 LISLYAKFSDIQSARQVFDTMPHRDPI--SWNSMINAYLQNGCLEEALEMLKGVYLLGLV 148
++ +YA ++F + R WNS+I+++++NG L +AL + G+
Sbjct: 76 ILGMYAMCGSFSDCGKMFYRLDLRRSSIRPWNSIISSFVRNGLLNQALAFYFKMLCFGVS 135
Query: 149 PKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASR 208
P +V C + G + V ++ + ++++L+ Y + S+
Sbjct: 136 PDVSTFPCLVKACVALKNFK-GIDFLSDTVSSLGMDCNEFVASSLIKAYLEYGKIDVPSK 194
Query: 209 VFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGF 268
+FD + K+ V W M++G D+ F M+++ + PN VT +L CA
Sbjct: 195 LFDRVLQKDCVIWNVMLNGYAKCGALDSVIKGFSVMRMDQISPNAVTFDCVLSVCASKLL 254
Query: 269 VKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIG 328
+ G ++HG G++ ++L++MY + G A ++F S D V W+ +I
Sbjct: 255 IDLGVQLHGLVVVSGVDFEGSIKNSLLSMYSKCGR-FDDASKLFRMMSRADTVTWNCMIS 313
Query: 329 SYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSF 388
Y Q G ++LT F +M++ P+ +T +++ + + +L++ +H +I++ +S
Sbjct: 314 GYVQSGLMEESLTFFYEMISSGVLPDAITFSSLLPSVSKFENLEYCKQIHCYIMRHSISL 373
Query: 389 SISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK 448
I L++AL++ Y KC + + IF + + D + ++ MIS Y +G +L++F +
Sbjct: 374 DIFLTSALIDAYFKCRGVSMAQNIFSQCNSVDVVVFTAMISGYLHNGLYIDSLEMFRWLV 433
Query: 449 ERGVKPDALTFLAVLSACNHAGLVTEGQQVFK-QVNADYEIPLTIEHYACLVDLLGRSGK 507
+ + P+ +T +++L + G+++ + ++ I ++D+ + G+
Sbjct: 434 KVKISPNEITLVSILPVIGILLALKLGRELHGFIIKKGFDNRCNIG--CAVIDMYAKCGR 491
Query: 508 VEDALEIMRTMPMKPSTRIWSSLVSAC 534
+ A EI + K W+S+++ C
Sbjct: 492 MNLAYEIFERL-SKRDIVSWNSMITRC 517
Score = 108 bits (269), Expect = 1e-23
Identities = 76/303 (25%), Positives = 141/303 (46%), Gaps = 17/303 (5%)
Query: 246 VEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESL 305
+E P R++L LL AC+ P ++ GK++H + + I ++ MY G
Sbjct: 30 LEETIPRRLSL--LLQACSNPNLLRQGKQVHAFLIVNSISGDSYTDERILGMYAMCGSFS 87
Query: 306 HLAEQIFE----RSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAV 361
+ + RSS R W+SII S+ + G +AL + KML P+ T +
Sbjct: 88 DCGKMFYRLDLRRSSIRP---WNSIISSFVRNGLLNQALAFYFKMLCFGVSPDVSTFPCL 144
Query: 362 ISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDS 421
+ AC L + K L + G+ + ++++L+ Y + G ++ ++F + +D
Sbjct: 145 VKACVALKNFKGIDFLSDTVSSLGMDCNEFVASSLIKAYLEYGKIDVPSKLFDRVLQKDC 204
Query: 422 ISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQ 481
+ W+ M++ Y G + +K FS M+ + P+A+TF VLS C L+ G Q+
Sbjct: 205 VIWNVMLNGYAKCGALDSVIKGFSVMRMDQISPNAVTFDCVLSVCASKLLIDLGVQLHGL 264
Query: 482 V---NADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
V D+E + L+ + + G+ +DA ++ R M + T W+ ++S G
Sbjct: 265 VVVSGVDFEGSIK----NSLLSMYSKCGRFDDASKLFRMM-SRADTVTWNCMISGYVQSG 319
Query: 539 RLD 541
++
Sbjct: 320 LME 322
>At4g18750 putative protein
Length = 871
Score = 347 bits (889), Expect = 1e-95
Identities = 194/579 (33%), Positives = 325/579 (55%), Gaps = 3/579 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
+ L G + ++ LF ++ S S V K+ SSL S G LHG L +G
Sbjct: 167 MNELAKSGDFSGSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGF 226
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
V NSL++ Y K + SAR+VFD M RD ISWNS+IN Y+ NG E+ L +
Sbjct: 227 GERNSVGNSLVAFYLKNQRVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQ 286
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ + G+ + S+ + C +GR +H++ V +T L+D Y +C
Sbjct: 287 MLVSGIEIDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNT-LLDMYSKCG 345
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
D A VF M ++ VS+T+MI+G A F M+ EG+ P+ T+ A+L
Sbjct: 346 DLDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLN 405
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVV 321
CA+ + GK +H + + + S+AL++MY + G S+ AE +F +D++
Sbjct: 406 CCARYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCG-SMQEAELVFSEMRVKDII 464
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEET-EPNYVTLLAVISACTNLSSLKHGCGLHGF 380
W++IIG YS+ + +AL+LFN +L E+ P+ T+ V+ AC +LS+ G +HG+
Sbjct: 465 SWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGY 524
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
I++ G ++N+L++MYAKCG L + +F ++ ++D +SW+ MI+ YG+HG G++A
Sbjct: 525 IMRNGYFSDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEA 584
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
+ LF++M++ G++ D ++F+++L AC+H+GLV EG + F + + +I T+EHYAC+VD
Sbjct: 585 IALFNQMRQAGIEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVD 644
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAAN 560
+L R+G + A + MP+ P IW +L+ C++H + +AE + ++ EP+N
Sbjct: 645 MLARTGDLIKAYRFIENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFELEPENTGY 704
Query: 561 YTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
Y L+ IYAE W +++ + + + L+K G S IE
Sbjct: 705 YVLMANIYAEAEKWEQVKRLRKRIGQRGLRKNPGCSWIE 743
Score = 191 bits (484), Expect = 1e-48
Identities = 135/519 (26%), Positives = 249/519 (47%), Gaps = 8/519 (1%)
Query: 53 LPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMP 112
L SV++ C+ S G + G D + + L +Y D++ A +VFD +
Sbjct: 97 LCSVLQLCADSKSLKDGKEVDNFIRGNGFVIDSNLGSKLSLMYTNCGDLKEASRVFDEVK 156
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQ 172
+ WN ++N ++G ++ + K + G+ + + G Q
Sbjct: 157 IEKALFWNILMNELAKSGDFSGSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQ 216
Query: 173 IHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANL 232
+H ++ G E + V +LV FY + A +VFD M ++ +SW ++I+G ++N
Sbjct: 217 LHGFILKSGFGERNSV-GNSLVAFYLKNQRVDSARKVFDEMTERDVISWNSIINGYVSNG 275
Query: 233 DYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSS 292
+ + F M V G++ + T++++ CA + G+ +H + F +
Sbjct: 276 LAEKGLSVFVQMLVSGIEIDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCN 335
Query: 293 ALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE 352
L++MY + G+ L A+ +F S R VV ++S+I Y++ G + +A+ LF +M E
Sbjct: 336 TLLDMYSKCGD-LDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGIS 394
Query: 353 PNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQI 412
P+ T+ AV++ C L G +H +I + L F I +SNALM+MYAKCG ++ + +
Sbjct: 395 PDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQEAELV 454
Query: 413 FLEMQNRDSISWSTMISAYGLHGHGEQALKLFS-EMKERGVKPDALTFLAVLSACNHAGL 471
F EM+ +D ISW+T+I Y + + +AL LF+ ++E+ PD T VL AC
Sbjct: 455 FSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSA 514
Query: 472 VTEGQQVFKQVNADYEIPLTIEHYA-CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSL 530
+G+++ + + + H A LVD+ + G + A + + K W+ +
Sbjct: 515 FDKGREIHGYIMRNGY--FSDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVS-WTVM 571
Query: 531 VSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYA 569
++ +HG A L Q+ R A + ++++YA
Sbjct: 572 IAGYGMHGFGKEAIALFNQM-RQAGIEADEISFVSLLYA 609
Score = 180 bits (457), Expect = 2e-45
Identities = 109/414 (26%), Positives = 214/414 (51%), Gaps = 10/414 (2%)
Query: 120 NSMINAYLQNGCLEEALEML--KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALV 177
N+ + + ++G LE A+++L G + + P L S++ LC + G+++ +
Sbjct: 65 NTQLRRFCESGNLENAVKLLCVSGKWDID----PRTLCSVLQLCADSKSLKDGKEVDNFI 120
Query: 178 VVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAA 237
+G V S L + L Y C D ASRVFD ++++ + W +++ + D+ +
Sbjct: 121 RGNGFVIDSN-LGSKLSLMYTNCGDLKEASRVFDEVKIEKALFWNILMNELAKSGDFSGS 179
Query: 238 FARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINM 297
F+ M GV+ + T + + + V G+++HG+ + G + ++L+
Sbjct: 180 IGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGFGERNSVGNSLVAF 239
Query: 298 YCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVT 357
Y ++ + + A ++F+ + RDV+ W+SII Y G + K L++F +ML E + T
Sbjct: 240 YLKN-QRVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLAT 298
Query: 358 LLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQ 417
+++V + C + + G +H +K S N L++MY+KCG L+ + +F EM
Sbjct: 299 IVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAVFREMS 358
Query: 418 NRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQ 477
+R +S+++MI+ Y G +A+KLF EM+E G+ PD T AVL+ C L+ EG++
Sbjct: 359 DRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKR 418
Query: 478 VFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV 531
V + + + ++ I L+D+ + G +++A + M +K W++++
Sbjct: 419 VHEWIK-ENDLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIIS-WNTII 470
Score = 110 bits (276), Expect = 2e-24
Identities = 93/354 (26%), Positives = 155/354 (43%), Gaps = 10/354 (2%)
Query: 17 SPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLA 76
S + I +GL + ++LF ++ + +V+ C+ G +H
Sbjct: 364 SYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWI 423
Query: 77 LTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEAL 136
D VSN+L+ +YAK +Q A VF M +D ISWN++I Y +N EAL
Sbjct: 424 KENDLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIISWNTIIGGYSKNCYANEAL 483
Query: 137 EMLKGVYLL---GLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTAL 193
+ LL P +A ++ C GR+IH ++ +G V + +L
Sbjct: 484 SLFN--LLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGYIMRNGYFSDRHV-ANSL 540
Query: 194 VDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNR 253
VD Y +C L+A +FD + K+ VSWT MI+G + A A F M+ G++ +
Sbjct: 541 VDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEAIALFNQMRQAGIEADE 600
Query: 254 VTLIALLPACAKPGFVKHGKEIHGYAFRH--GIESSHIFSSALINMYCQSGESLHLAEQI 311
++ ++LL AC+ G V G RH IE + + +++M ++G+ + I
Sbjct: 601 ISFVSLLYACSHSGLVDEGWRFFN-IMRHECKIEPTVEHYACIVDMLARTGDLIKAYRFI 659
Query: 312 FERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE-PNYVTLLAVISA 364
D +W +++ D A + K+ E E Y L+A I A
Sbjct: 660 ENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFELEPENTGYYVLMANIYA 713
>At2g03380 unknown protein
Length = 689
Score = 339 bits (870), Expect = 2e-93
Identities = 186/550 (33%), Positives = 311/550 (55%), Gaps = 6/550 (1%)
Query: 52 VLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTM 111
V +KAC+ L G +H + S D VV L+ +YAK +I+SA +VF+ +
Sbjct: 144 VFSKALKACTELQDLDNGKKIHCQLVKVPSF-DNVVLTGLLDMYAKCGEIKSAHKVFNDI 202
Query: 112 PHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGR 171
R+ + W SMI Y++N EE L + + ++ +++ C + G+
Sbjct: 203 TLRNVVCWTSMIAGYVKNDLCEEGLVLFNRMRENNVLGNEYTYGTLIMACTKLSALHQGK 262
Query: 172 QIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIAN 231
H +V G +E S L T+L+D Y +C D A RVF+ + V WTAMI G N
Sbjct: 263 WFHGCLVKSG-IELSSCLVTSLLDMYVKCGDISNARRVFNEHSHVDLVMWTAMIVGYTHN 321
Query: 232 LDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFS 291
+ A + F+ M+ + PN VT+ ++L C ++ G+ +HG + + GI +++ +
Sbjct: 322 GSVNEALSLFQKMKGVEIKPNCVTIASVLSGCGLIENLELGRSVHGLSIKVGIWDTNV-A 380
Query: 292 SALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEET 351
+AL++MY + ++ A+ +FE S +D+V W+SII +SQ G ++AL LF++M +E
Sbjct: 381 NALVHMYAKCYQNRD-AKYVFEMESEKDIVAWNSIISGFSQNGSIHEALFLFHRMNSESV 439
Query: 352 EPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGL--SFSISLSNALMNMYAKCGCLEGS 409
PN VT+ ++ SAC +L SL G LH + +K G S S+ + AL++ YAKCG + +
Sbjct: 440 TPNGVTVASLFSACASLGSLAVGSSLHAYSVKLGFLASSSVHVGTALLDFYAKCGDPQSA 499
Query: 410 LQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHA 469
IF ++ +++I+WS MI YG G +L+LF EM ++ KP+ TF ++LSAC H
Sbjct: 500 RLIFDTIEEKNTITWSAMIGGYGKQGDTIGSLELFEEMLKKQQKPNESTFTSILSACGHT 559
Query: 470 GLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSS 529
G+V EG++ F + DY + +HY C+VD+L R+G++E AL+I+ MP++P R + +
Sbjct: 560 GMVNEGKKYFSSMYKDYNFTPSTKHYTCMVDMLARAGELEQALDIIEKMPIQPDVRCFGA 619
Query: 530 LVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRL 589
+ C +H R D+ E++ +++ PD+A+ Y L++ +YA G W ++V MK + L
Sbjct: 620 FLHGCGMHSRFDLGEIVIKKMLDLHPDDASYYVLVSNLYASDGRWNQAKEVRNLMKQRGL 679
Query: 590 KKCYGFSRIE 599
K G S +E
Sbjct: 680 SKIAGHSTME 689
Score = 214 bits (544), Expect = 1e-55
Identities = 148/529 (27%), Positives = 260/529 (48%), Gaps = 20/529 (3%)
Query: 39 TQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKF 98
+ LH++A SS ++ C+++ S HG+ G D ++ L+SLY F
Sbjct: 37 SSLHYAA----SSPCFLLLSKCTNIDSLRQS---HGVLTGNGLMGDISIATKLVSLYGFF 89
Query: 99 SDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLL---GLVPKPELLA 155
+ AR VFD +P D W M+ Y N +E++E++K LL G + +
Sbjct: 90 GYTKDARLVFDQIPEPDFYLWKVMLRCYCLN---KESVEVVKLYDLLMKHGFRYDDIVFS 146
Query: 156 SMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEV 215
+ C G++IH +V ++ V+ T L+D Y +C + A +VF+ + +
Sbjct: 147 KALKACTELQDLDNGKKIHCQLVKVPSFDN--VVLTGLLDMYAKCGEIKSAHKVFNDITL 204
Query: 216 KNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEI 275
+N V WT+MI+G + N + F M+ V N T L+ AC K + GK
Sbjct: 205 RNVVCWTSMIAGYVKNDLCEEGLVLFNRMRENNVLGNEYTYGTLIMACTKLSALHQGKWF 264
Query: 276 HGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGD 335
HG + GIE S ++L++MY + G+ + A ++F S D+V+W+++I Y+ G
Sbjct: 265 HGCLVKSGIELSSCLVTSLLDMYVKCGD-ISNARRVFNEHSHVDLVMWTAMIVGYTHNGS 323
Query: 336 SYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNA 395
+AL+LF KM E +PN VT+ +V+S C + +L+ G +HG +K G+ + +++NA
Sbjct: 324 VNEALSLFQKMKGVEIKPNCVTIASVLSGCGLIENLELGRSVHGLSIKVGI-WDTNVANA 382
Query: 396 LMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPD 455
L++MYAKC + +F +D ++W+++IS + +G +AL LF M V P+
Sbjct: 383 LVHMYAKCYQNRDAKYVFEMESEKDIVAWNSIISGFSQNGSIHEALFLFHRMNSESVTPN 442
Query: 456 ALTFLAVLSACNHAGLVTEGQQVFK-QVNADYEIPLTIEHYACLVDLLGRSGKVEDALEI 514
+T ++ SAC G + G + V + ++ L+D + G + A I
Sbjct: 443 GVTVASLFSACASLGSLAVGSSLHAYSVKLGFLASSSVHVGTALLDFYAKCGDPQSARLI 502
Query: 515 MRTMPMKPSTRIWSSLVSACKLHG-RLDIAEMLGPQLIRSEPDNAANYT 562
T+ K +T WS+++ G + E+ L + + N + +T
Sbjct: 503 FDTIEEK-NTITWSAMIGGYGKQGDTIGSLELFEEMLKKQQKPNESTFT 550
Score = 82.4 bits (202), Expect = 7e-16
Identities = 57/217 (26%), Positives = 101/217 (46%), Gaps = 3/217 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTG- 80
I G H+ L LF +++ + + S+ AC+SL S +G+ LH ++ G
Sbjct: 415 ISGFSQNGSIHEALFLFHRMNSESVTPNGVTVASLFSACASLGSLAVGSSLHAYSVKLGF 474
Query: 81 -SHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEML 139
+ S V +L+ YAK D QSAR +FDT+ ++ I+W++MI Y + G +LE+
Sbjct: 475 LASSSVHVGTALLDFYAKCGDPQSARLIFDTIEEKNTITWSAMIGGYGKQGDTIGSLELF 534
Query: 140 KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
+ + P S++S CG G++ + + D S T +VD R
Sbjct: 535 EEMLKKQQKPNESTFTSILSACGHTGMVNEGKKYFSSMYKDYNFTPSTKHYTCMVDMLAR 594
Query: 200 CDDALMASRVFDGMEVKNEV-SWTAMISGCIANLDYD 235
+ A + + M ++ +V + A + GC + +D
Sbjct: 595 AGELEQALDIIEKMPIQPDVRCFGAFLHGCGMHSRFD 631
Score = 63.9 bits (154), Expect = 2e-10
Identities = 46/201 (22%), Positives = 97/201 (47%), Gaps = 18/201 (8%)
Query: 341 TLFNKMLTEETEPNYVTLLA------VISACTNLSSLKHGCGLHGFILKFGLSFSISLSN 394
T+ +LTEE + + + A ++S CTN+ SL+ HG + GL IS++
Sbjct: 24 TIKELILTEENDGSSLHYAASSPCFLLLSKCTNIDSLRQS---HGVLTGNGLMGDISIAT 80
Query: 395 ALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKP 454
L+++Y G + + +F ++ D W M+ Y L+ + +KL+ + + G +
Sbjct: 81 KLVSLYGFFGYTKDARLVFDQIPEPDFYLWKVMLRCYCLNKESVEVVKLYDLLMKHGFRY 140
Query: 455 DALTFLAVLSACNHAGLVTEGQQVFKQ---VNADYEIPLTIEHYACLVDLLGRSGKVEDA 511
D + F L AC + G+++ Q V + + LT L+D+ + G+++ A
Sbjct: 141 DDIVFSKALKACTELQDLDNGKKIHCQLVKVPSFDNVVLT-----GLLDMYAKCGEIKSA 195
Query: 512 LEIMRTMPMKPSTRIWSSLVS 532
++ + ++ + W+S+++
Sbjct: 196 HKVFNDITLR-NVVCWTSMIA 215
>At4g33990 putative protein
Length = 844
Score = 333 bits (855), Expect = 1e-91
Identities = 189/552 (34%), Positives = 315/552 (56%), Gaps = 26/552 (4%)
Query: 54 PSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPH 113
PSV+KAC ++ G +H LAL G D V+ SLI LY+++ + +AR +FD MP
Sbjct: 90 PSVLKACRTVID---GNKIHCLALKFGFMWDVYVAASLIHLYSRYKAVGNARILFDEMPV 146
Query: 114 RDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQI 173
RD SWN+MI+ Y Q+G +EAL + G+ + V + S++S C G I
Sbjct: 147 RDMGSWNAMISGYCQSGNAKEALTLSNGLRAMDSVT----VVSLLSACTEAGDFNRGVTI 202
Query: 174 HALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLD 233
H+ + G + S L D +VFD M V++ +SW ++I N
Sbjct: 203 HSYSIKHG------LESELLRD----------CQKVFDRMYVRDLISWNSIIKAYELNEQ 246
Query: 234 YDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHI-FSS 292
A + F+ M++ + P+ +TLI+L ++ G ++ + + G+ R G I +
Sbjct: 247 PLRAISLFQEMRLSRIQPDCLTLISLASILSQLGDIRACRSVQGFTLRKGWFLEDITIGN 306
Query: 293 ALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTE-ET 351
A++ MY + G + A +F DV+ W++II Y+Q G + +A+ ++N M E E
Sbjct: 307 AVVVMYAKLG-LVDSARAVFNWLPNTDVISWNTIISGYAQNGFASEAIEMYNIMEEEGEI 365
Query: 352 EPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQ 411
N T ++V+ AC+ +L+ G LHG +LK GL + + +L +MY KCG LE +L
Sbjct: 366 AANQGTWVSVLPACSQAGALRQGMKLHGRLLKNGLYLDVFVVTSLADMYGKCGRLEDALS 425
Query: 412 IFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGL 471
+F ++ +S+ W+T+I+ +G HGHGE+A+ LF EM + GVKPD +TF+ +LSAC+H+GL
Sbjct: 426 LFYQIPRVNSVPWNTLIACHGFHGHGEKAVMLFKEMLDEGVKPDHITFVTLLSACSHSGL 485
Query: 472 VTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV 531
V EGQ F+ + DY I +++HY C+VD+ GR+G++E AL+ +++M ++P IW +L+
Sbjct: 486 VDEGQWCFEMMQTDYGITPSLKHYGCMVDMYGRAGQLETALKFIKSMSLQPDASIWGALL 545
Query: 532 SACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKK 591
SAC++HG +D+ ++ L EP++ + LL+ +YA G W +++ + L+K
Sbjct: 546 SACRVHGNVDLGKIASEHLFEVEPEHVGYHVLLSNMYASAGKWEGVDEIRSIAHGKGLRK 605
Query: 592 CYGFSRIETGNQ 603
G+S +E N+
Sbjct: 606 TPGWSSMEVDNK 617
Score = 196 bits (498), Expect = 3e-50
Identities = 137/493 (27%), Positives = 249/493 (49%), Gaps = 37/493 (7%)
Query: 92 ISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLL--GLVP 149
+S + S +S FD + +RD +WN MI+ Y + G E + +++L GL P
Sbjct: 26 VSTHVSLSPNKSKMHTFDHIQNRDVYAWNLMISGYGRAGNSSEVIRCF-SLFMLSSGLTP 84
Query: 150 KPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRV 209
S++ C + G +IH L + G + V ++ +L+ Y R A +
Sbjct: 85 DYRTFPSVLKACRTVID---GNKIHCLALKFGFM-WDVYVAASLIHLYSRYKAVGNARIL 140
Query: 210 FDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFV 269
FD M V++ SW AMISG + + A ++ + VT+++LL AC + G
Sbjct: 141 FDEMPVRDMGSWNAMISGYCQSGNAKEALTLSNGLRA----MDSVTVVSLLSACTEAGDF 196
Query: 270 KHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGS 329
G IH Y+ +HG+ES E L +++F+R RD++ W+SII +
Sbjct: 197 NRGVTIHSYSIKHGLES----------------ELLRDCQKVFDRMYVRDLISWNSIIKA 240
Query: 330 YSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSF- 388
Y +A++LF +M +P+ +TL+++ S + L ++ + GF L+ G
Sbjct: 241 YELNEQPLRAISLFQEMRLSRIQPDCLTLISLASILSQLGDIRACRSVQGFTLRKGWFLE 300
Query: 389 SISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK 448
I++ NA++ MYAK G ++ + +F + N D ISW+T+IS Y +G +A+++++ M+
Sbjct: 301 DITIGNAVVVMYAKLGLVDSARAVFNWLPNTDVISWNTIISGYAQNGFASEAIEMYNIME 360
Query: 449 ERG-VKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGK 507
E G + + T+++VL AC+ AG + +G ++ ++ + + L + L D+ G+ G+
Sbjct: 361 EEGEIAANQGTWVSVLPACSQAGALRQGMKLHGRLLKN-GLYLDVFVVTSLADMYGKCGR 419
Query: 508 VEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRS--EPDNAANYTLLN 565
+EDAL + +P S W++L++ HG + A ML +++ +PD+ TLL+
Sbjct: 420 LEDALSLFYQIPRVNSVP-WNTLIACHGFHGHGEKAVMLFKEMLDEGVKPDHITFVTLLS 478
Query: 566 MI----YAEHGHW 574
+ G W
Sbjct: 479 ACSHSGLVDEGQW 491
>At4g30700 unknown protein
Length = 792
Score = 332 bits (851), Expect = 4e-91
Identities = 184/582 (31%), Positives = 311/582 (52%), Gaps = 27/582 (4%)
Query: 32 HQTLELFTQLHFSAP-HSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNS 90
H +L +F L S SS I A S G ++HG A+ G S+ ++ ++
Sbjct: 100 HSSLSVFAHLRKSTDLKPNSSTYAFAISAASGFRDDRAGRVIHGQAVVDGCDSELLLGSN 159
Query: 91 LISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVY------- 143
++ +Y KF ++ AR+VFD MP +D I WN+MI+ Y +N E++++ + +
Sbjct: 160 IVKMYFKFWRVEDARKVFDRMPEKDTILWNTMISGYRKNEMYVESIQVFRDLINESCTRL 219
Query: 144 ----LLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
LL ++P L + R+G QIH+L G H VL T + Y +
Sbjct: 220 DTTTLLDILPAVAELQEL----------RLGMQIHSLATKTGCYSHDYVL-TGFISLYSK 268
Query: 200 CDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIAL 259
C M S +F + V++ AMI G +N + + + + F+ + + G TL++L
Sbjct: 269 CGKIKMGSALFREFRKPDIVAYNAMIHGYTSNGETELSLSLFKELMLSGARLRSSTLVSL 328
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
+P G + IHGY + S S+AL +Y + E + A ++F+ S +
Sbjct: 329 VPVS---GHLMLIYAIHGYCLKSNFLSHASVSTALTTVYSKLNE-IESARKLFDESPEKS 384
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
+ W+++I Y+Q G + A++LF +M E PN VT+ ++SAC L +L G +H
Sbjct: 385 LPSWNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTITCILSACAQLGALSLGKWVHD 444
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
+ SI +S AL+ MYAKCG + + ++F M ++ ++W+TMIS YGLHG G++
Sbjct: 445 LVRSTDFESSIYVSTALIGMYAKCGSIAEARRLFDLMTKKNEVTWNTMISGYGLHGQGQE 504
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
AL +F EM G+ P +TFL VL AC+HAGLV EG ++F + Y +++HYAC+V
Sbjct: 505 ALNIFYEMLNSGITPTPVTFLCVLYACSHAGLVKEGDEIFNSMIHRYGFEPSVKHYACMV 564
Query: 500 DLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAA 559
D+LGR+G ++ AL+ + M ++P + +W +L+ AC++H ++A + +L +PDN
Sbjct: 565 DILGRAGHLQRALQFIEAMSIEPGSSVWETLLGACRIHKDTNLARTVSEKLFELDPDNVG 624
Query: 560 NYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETG 601
+ LL+ I++ ++ V + K ++L K G++ IE G
Sbjct: 625 YHVLLSNIHSADRNYPQAATVRQTAKKRKLAKAPGYTLIEIG 666
Score = 88.6 bits (218), Expect = 9e-18
Identities = 64/246 (26%), Positives = 110/246 (44%), Gaps = 3/246 (1%)
Query: 16 PSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGL 75
PS + I GL + LF ++ S + ++ AC+ L + +LG +H L
Sbjct: 386 PSWNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTITCILSACAQLGALSLGKWVHDL 445
Query: 76 ALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEA 135
+T S VS +LI +YAK I AR++FD M ++ ++WN+MI+ Y +G +EA
Sbjct: 446 VRSTDFESSIYVSTALIGMYAKCGSIAEARRLFDLMTKKNEVTWNTMISGYGLHGQGQEA 505
Query: 136 LEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVD 195
L + + G+ P P ++ C + G +I ++ E SV +VD
Sbjct: 506 LNIFYEMLNSGITPTPVTFLCVLYACSHAGLVKEGDEIFNSMIHRYGFEPSVKHYACMVD 565
Query: 196 FYFRCDDALMASRVFDGMEVKNEVS-WTAMISGCIANLDYDAAFARFRAMQVEGVDPNRV 254
R A + + M ++ S W ++ C + D AR + ++ +DP+ V
Sbjct: 566 ILGRAGHLQRALQFIEAMSIEPGSSVWETLLGAC--RIHKDTNLARTVSEKLFELDPDNV 623
Query: 255 TLIALL 260
LL
Sbjct: 624 GYHVLL 629
Score = 53.5 bits (127), Expect = 3e-07
Identities = 49/188 (26%), Positives = 87/188 (46%), Gaps = 19/188 (10%)
Query: 357 TLLAVISACTNL------SSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSL 410
T A+IS T L +S+ H H I+ G ISL L + G + +
Sbjct: 13 TTAALISKNTYLDFFKRSTSISHLAQTHAQIILHGFRNDISLLTKLTQRLSDLGAIYYAR 72
Query: 411 QIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEM-KERGVKPDALTFLAVLSAC--- 466
IFL +Q D ++ ++ + ++ +L +F+ + K +KP++ T+ +SA
Sbjct: 73 DIFLSVQRPDVFLFNVLMRGFSVNESPHSSLSVFAHLRKSTDLKPNSSTYAFAISAASGF 132
Query: 467 --NHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPST 524
+ AG V GQ V D E+ L + +V + + +VEDA ++ MP K T
Sbjct: 133 RDDRAGRVIHGQAVVD--GCDSELLLG----SNIVKMYFKFWRVEDARKVFDRMPEK-DT 185
Query: 525 RIWSSLVS 532
+W++++S
Sbjct: 186 ILWNTMIS 193
>At1g69350 hypothetical protein
Length = 787
Score = 332 bits (850), Expect = 5e-91
Identities = 193/587 (32%), Positives = 311/587 (52%), Gaps = 4/587 (0%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
S + S + G + L +F + + + SV++ C+ L + +HG
Sbjct: 171 STLVSSCLENGEVVKALRMFKCMVDDGVEPDAVTMISVVEGCAELGCLRIARSVHGQITR 230
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
D + NSL+++Y+K D+ S+ ++F+ + ++ +SW +MI++Y + E+AL
Sbjct: 231 KMFDLDETLCNSLLTMYSKCGDLLSSERIFEKIAKKNAVSWTAMISSYNRGEFSEKALRS 290
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+ G+ P L S++S CG R G+ +H V + LS ALV+ Y
Sbjct: 291 FSEMIKSGIEPNLVTLYSVLSSCGLIGLIREGKSVHGFAVRRELDPNYESLSLALVELYA 350
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
C V + +N V+W ++IS A FR M + + P+ TL +
Sbjct: 351 ECGKLSDCETVLRVVSDRNIVAWNSLISLYAHRGMVIQALGLFRQMVTQRIKPDAFTLAS 410
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR 318
+ AC G V GK+IHG+ R + S ++LI+MY +SG S+ A +F + R
Sbjct: 411 SISACENAGLVPLGKQIHGHVIRTDV-SDEFVQNSLIDMYSKSG-SVDSASTVFNQIKHR 468
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLH 378
VV W+S++ +SQ G+S +A++LF+ M E N VT LAVI AC+++ SL+ G +H
Sbjct: 469 SVVTWNSMLCGFSQNGNSVEAISLFDYMYHSYLEMNEVTFLAVIQACSSIGSLEKGKWVH 528
Query: 379 GFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGE 438
++ GL + AL++MYAKCG L + +F M +R +SWS+MI+AYG+HG
Sbjct: 529 HKLIISGLK-DLFTDTALIDMYAKCGDLNAAETVFRAMSSRSIVSWSSMINAYGMHGRIG 587
Query: 439 QALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACL 498
A+ F++M E G KP+ + F+ VLSAC H+G V EG+ F + + + + EH+AC
Sbjct: 588 SAISTFNQMVESGTKPNEVVFMNVLSACGHSGSVEEGKYYFNLMKS-FGVSPNSEHFACF 646
Query: 499 VDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNA 558
+DLL RSG +++A ++ MP +W SLV+ C++H ++DI + + L D+
Sbjct: 647 IDLLSRSGDLKEAYRTIKEMPFLADASVWGSLVNGCRIHQKMDIIKAIKNDLSDIVTDDT 706
Query: 559 ANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQCF 605
YTLL+ IYAE G W ++ AMK LKK G+S IE + F
Sbjct: 707 GYYTLLSNIYAEEGEWEEFRRLRSAMKSSNLKKVPGYSAIEIDQKVF 753
Score = 271 bits (692), Expect = 1e-72
Identities = 170/578 (29%), Positives = 301/578 (51%), Gaps = 20/578 (3%)
Query: 30 LYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSH-TLGTLLHGLALTTGSHSDPVVS 88
LYH+ + TQ+ V PSV++AC+ H ++G +HG + G D V+
Sbjct: 87 LYHRLVSETTQIS-------KFVFPSVLRACAGSREHLSVGGKVHGRIIKGGVDDDAVIE 139
Query: 89 NSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLV 148
SL+ +Y + ++ A +VFD MP RD ++W++++++ L+NG + +AL M K + G+
Sbjct: 140 TSLLCMYGQTGNLSDAEKVFDGMPVRDLVAWSTLVSSCLENGEVVKALRMFKCMVDDGVE 199
Query: 149 PKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASR 208
P + S+V C RI R +H + + L +L+ Y +C D L + R
Sbjct: 200 PDAVTMISVVEGCAELGCLRIARSVHG-QITRKMFDLDETLCNSLLTMYSKCGDLLSSER 258
Query: 209 VFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGF 268
+F+ + KN VSWTAMIS + A F M G++PN VTL ++L +C G
Sbjct: 259 IFEKIAKKNAVSWTAMISSYNRGEFSEKALRSFSEMIKSGIEPNLVTLYSVLSSCGLIGL 318
Query: 269 VKHGKEIHGYAFRHGIESSH-IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSII 327
++ GK +HG+A R ++ ++ S AL+ +Y + G+ L E + S R++V W+S+I
Sbjct: 319 IREGKSVHGFAVRRELDPNYESLSLALVELYAECGK-LSDCETVLRVVSDRNIVAWNSLI 377
Query: 328 GSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLS 387
Y+ RG +AL LF +M+T+ +P+ TL + ISAC N + G +HG +++ +S
Sbjct: 378 SLYAHRGMVIQALGLFRQMVTQRIKPDAFTLASSISACENAGLVPLGKQIHGHVIRTDVS 437
Query: 388 FSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEM 447
+ N+L++MY+K G ++ + +F ++++R ++W++M+ + +G+ +A+ LF M
Sbjct: 438 DEF-VQNSLIDMYSKSGSVDSASTVFNQIKHRSVVTWNSMLCGFSQNGNSVEAISLFDYM 496
Query: 448 KERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGK 507
++ + +TFLAV+ AC+ G + +G+ V ++ L + L+D+ + G
Sbjct: 497 YHSYLEMNEVTFLAVIQACSSIGSLEKGKWVHHKLIISGLKDLFTD--TALIDMYAKCGD 554
Query: 508 VEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMI 567
+ A + R M + S WSS+++A +HGR+ A Q++ E N + +
Sbjct: 555 LNAAETVFRAMSSR-SIVSWSSMINAYGMHGRIGSAISTFNQMV--ESGTKPNEVVFMNV 611
Query: 568 YAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQCF 605
+ G H V E L K +G S CF
Sbjct: 612 LSACG---HSGSVEEGKYYFNLMKSFGVSPNSEHFACF 646
Score = 233 bits (595), Expect = 2e-61
Identities = 156/540 (28%), Positives = 281/540 (51%), Gaps = 19/540 (3%)
Query: 56 VIKACSSLHSHTLGTLLHGLALTTGS-HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHR 114
+ ++CSSL L + LH L TG DP+ LI YA S+R VF+ P+
Sbjct: 7 LFRSCSSLR---LVSQLHAHLLVTGRLRRDPLPVTKLIESYAFMGSPDSSRLVFEAFPYP 63
Query: 115 DPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLC-GRKMGSRIGRQI 173
D + +I + L+ A+++ + + S++ C G + +G ++
Sbjct: 64 DSFMYGVLIKCNVWCHLLDAAIDLYHRLVSETTQISKFVFPSVLRACAGSREHLSVGGKV 123
Query: 174 HALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLD 233
H ++ G V+ V+ T+L+ Y + + A +VFDGM V++ V+W+ ++S C+ N +
Sbjct: 124 HGR-IIKGGVDDDAVIETSLLCMYGQTGNLSDAEKVFDGMPVRDLVAWSTLVSSCLENGE 182
Query: 234 YDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSA 293
A F+ M +GV+P+ VT+I+++ CA+ G ++ + +HG R + ++
Sbjct: 183 VVKALRMFKCMVDDGVEPDAVTMISVVEGCAELGCLRIARSVHGQITRKMFDLDETLCNS 242
Query: 294 LINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEP 353
L+ MY + G+ L +E+IFE+ + ++ V W+++I SY++ S KAL F++M+ EP
Sbjct: 243 LLTMYSKCGDLLS-SERIFEKIAKKNAVSWTAMISSYNRGEFSEKALRSFSEMIKSGIEP 301
Query: 354 NYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSI-SLSNALMNMYAKCGCLEGSLQI 412
N VTL +V+S+C + ++ G +HGF ++ L + SLS AL+ +YA+CG L +
Sbjct: 302 NLVTLYSVLSSCGLIGLIREGKSVHGFAVRRELDPNYESLSLALVELYAECGKLSDCETV 361
Query: 413 FLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLV 472
+ +R+ ++W+++IS Y G QAL LF +M + +KPDA T + +SAC +AGLV
Sbjct: 362 LRVVSDRNIVAWNSLISLYAHRGMVIQALGLFRQMVTQRIKPDAFTLASSISACENAGLV 421
Query: 473 TEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVS 532
G+Q+ V +++ L+D+ +SG V+ A + + + S W+S++
Sbjct: 422 PLGKQIHGHVIRTDVSDEFVQN--SLIDMYSKSGSVDSASTVFNQIKHR-SVVTWNSMLC 478
Query: 533 ACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYA-------EHGHWLHKEQVMEAMK 585
+G A L + S + T L +I A E G W+H + ++ +K
Sbjct: 479 GFSQNGNSVEAISLFDYMYHSYLE-MNEVTFLAVIQACSSIGSLEKGKWVHHKLIISGLK 537
>At2g33680 hypothetical protein
Length = 727
Score = 331 bits (848), Expect = 8e-91
Identities = 186/578 (32%), Positives = 315/578 (54%), Gaps = 12/578 (2%)
Query: 32 HQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSL 91
+ ++LF ++ + L + KA SSL S T+G H L + S D V SL
Sbjct: 100 YTVMQLFREMRAQDILPNAYTLAGIFKAESSLQSSTVGRQAHALVVKMSSFGDIYVDTSL 159
Query: 92 ISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKP 151
+ +Y K ++ +VF MP R+ +W++M++ Y G +EEA++ V+ L L K
Sbjct: 160 VGMYCKAGLVEDGLKVFAYMPERNTYTWSTMVSGYATRGRVEEAIK----VFNLFLREKE 215
Query: 152 E------LLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALM 205
E + +++S + +GRQIH + + +G + V LS ALV Y +C+
Sbjct: 216 EGSDSDYVFTAVLSSLAATIYVGLGRQIHCITIKNGLLGF-VALSNALVTMYSKCESLNE 274
Query: 206 ASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAK 265
A ++FD +N ++W+AM++G N + A F M G+ P+ T++ +L AC+
Sbjct: 275 ACKMFDSSGDRNSITWSAMVTGYSQNGESLEAVKLFSRMFSAGIKPSEYTIVGVLNACSD 334
Query: 266 PGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSS 325
+++ GK++H + + G E ++AL++MY ++G L A + F+ RDV LW+S
Sbjct: 335 ICYLEEGKQLHSFLLKLGFERHLFATTALVDMYAKAG-CLADARKGFDCLQERDVALWTS 393
Query: 326 IIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFG 385
+I Y Q D+ +AL L+ +M T PN T+ +V+ AC++L++L+ G +HG +K G
Sbjct: 394 LISGYVQNSDNEEALILYRRMKTAGIIPNDPTMASVLKACSSLATLELGKQVHGHTIKHG 453
Query: 386 LSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFS 445
+ + +AL MY+KCG LE +F N+D +SW+ MIS +G G++AL+LF
Sbjct: 454 FGLEVPIGSALSTMYSKCGSLEDGNLVFRRTPNKDVVSWNAMISGLSHNGQGDEALELFE 513
Query: 446 EMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRS 505
EM G++PD +TF+ ++SAC+H G V G F ++ + ++HYAC+VDLL R+
Sbjct: 514 EMLAEGMEPDDVTFVNIISACSHKGFVERGWFYFNMMSDQIGLDPKVDHYACMVDLLSRA 573
Query: 506 GKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLN 565
G++++A E + + + +W L+SACK HG+ ++ G +L+ ++ Y L+
Sbjct: 574 GQLKEAKEFIESANIDHGLCLWRILLSACKNHGKCELGVYAGEKLMALGSRESSTYVQLS 633
Query: 566 MIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQ 603
IY G E+V + M+ + K G S IE NQ
Sbjct: 634 GIYTALGRMRDVERVWKHMRANGVSKEVGCSWIELKNQ 671
Score = 224 bits (572), Expect = 8e-59
Identities = 156/532 (29%), Positives = 271/532 (50%), Gaps = 18/532 (3%)
Query: 46 PHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSAR 105
PH+ S L + S + G +HG + TG+ + +N L++ YAK + A
Sbjct: 12 PHT--STLLKKLTHHSQQRNLVAGRAVHGQIIRTGASTCIQHANVLVNFYAKCGKLAKAH 69
Query: 106 QVFDTMPHRDPISWNSMINAYLQNGCLEEA---LEMLKGVYLLGLVPKPELLASMVSLCG 162
+F+ + +D +SWNS+I Y QNG + + +++ + + ++P LA +
Sbjct: 70 SIFNAIICKDVVSWNSLITGYSQNGGISSSYTVMQLFREMRAQDILPNAYTLAGIFKAES 129
Query: 163 RKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWT 222
S +GRQ HALVV + + T+LV Y + +VF M +N +W+
Sbjct: 130 SLQSSTVGRQAHALVVKMSSF-GDIYVDTSLVGMYCKAGLVEDGLKVFAYMPERNTYTWS 188
Query: 223 AMISGCIANLDYDAA---FARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYA 279
M+SG + A F F + EG D + V A+L + A +V G++IH
Sbjct: 189 TMVSGYATRGRVEEAIKVFNLFLREKEEGSDSDYV-FTAVLSSLAATIYVGLGRQIHCIT 247
Query: 280 FRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKA 339
++G+ S+AL+ MY + ESL+ A ++F+ S R+ + WS+++ YSQ G+S +A
Sbjct: 248 IKNGLLGFVALSNALVTMYSKC-ESLNEACKMFDSSGDRNSITWSAMVTGYSQNGESLEA 306
Query: 340 LTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNM 399
+ LF++M + +P+ T++ V++AC+++ L+ G LH F+LK G + + AL++M
Sbjct: 307 VKLFSRMFSAGIKPSEYTIVGVLNACSDICYLEEGKQLHSFLLKLGFERHLFATTALVDM 366
Query: 400 YAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTF 459
YAK GCL + + F +Q RD W+++IS Y + E+AL L+ MK G+ P+ T
Sbjct: 367 YAKAGCLADARKGFDCLQERDVALWTSLISGYVQNSDNEEALILYRRMKTAGIIPNDPTM 426
Query: 460 LAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMP 519
+VL AC+ + G+QV + L + + L + + G +ED + R P
Sbjct: 427 ASVLKACSSLATLELGKQVHGH-TIKHGFGLEVPIGSALSTMYSKCGSLEDGNLVFRRTP 485
Query: 520 MKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRS--EPDNAANYTLLNMIYA 569
K W++++S +G+ D A L +++ EPD+ T +N+I A
Sbjct: 486 NKDVVS-WNAMISGLSHNGQGDEALELFEEMLAEGMEPDDV---TFVNIISA 533
Score = 118 bits (295), Expect = 1e-26
Identities = 77/236 (32%), Positives = 119/236 (49%), Gaps = 8/236 (3%)
Query: 249 VDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLA 308
++P+ TL+ L ++ + G+ +HG R G + ++ L+N Y + G+ L A
Sbjct: 10 LNPHTSTLLKKLTHHSQQRNLVAGRAVHGQIIRTGASTCIQHANVLVNFYAKCGK-LAKA 68
Query: 309 EQIFERSSFRDVVLWSSIIGSYSQRG---DSYKALTLFNKMLTEETEPNYVTLLAVISAC 365
IF +DVV W+S+I YSQ G SY + LF +M ++ PN TL + A
Sbjct: 69 HSIFNAIICKDVVSWNSLITGYSQNGGISSSYTVMQLFREMRAQDILPNAYTLAGIFKAE 128
Query: 366 TNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWS 425
++L S G H ++K I + +L+ MY K G +E L++F M R++ +WS
Sbjct: 129 SSLQSSTVGRQAHALVVKMSSFGDIYVDTSLVGMYCKAGLVEDGLKVFAYMPERNTYTWS 188
Query: 426 TMISAYGLHGHGEQALK---LFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQV 478
TM+S Y G E+A+K LF KE G D F AVLS+ V G+Q+
Sbjct: 189 TMVSGYATRGRVEEAIKVFNLFLREKEEGSDSD-YVFTAVLSSLAATIYVGLGRQI 243
Score = 85.1 bits (209), Expect = 1e-16
Identities = 60/197 (30%), Positives = 99/197 (49%), Gaps = 5/197 (2%)
Query: 350 ETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGS 409
E P+ TLL ++ + +L G +HG I++ G S I +N L+N YAKCG L +
Sbjct: 9 ELNPHTSTLLKKLTHHSQQRNLVAGRAVHGQIIRTGASTCIQHANVLVNFYAKCGKLAKA 68
Query: 410 LQIFLEMQNRDSISWSTMISAYGLHG---HGEQALKLFSEMKERGVKPDALTFLAVLSAC 466
IF + +D +SW+++I+ Y +G ++LF EM+ + + P+A T + A
Sbjct: 69 HSIFNAIICKDVVSWNSLITGYSQNGGISSSYTVMQLFREMRAQDILPNAYTLAGIFKAE 128
Query: 467 NHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRI 526
+ T G+Q V I LV + ++G VED L++ MP + +T
Sbjct: 129 SSLQSSTVGRQAHALVVKMSSFG-DIYVDTSLVGMYCKAGLVEDGLKVFAYMPER-NTYT 186
Query: 527 WSSLVSACKLHGRLDIA 543
WS++VS GR++ A
Sbjct: 187 WSTMVSGYATRGRVEEA 203
>At3g15130 hypothetical protein
Length = 689
Score = 330 bits (847), Expect = 1e-90
Identities = 175/551 (31%), Positives = 316/551 (56%), Gaps = 6/551 (1%)
Query: 53 LPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMP 112
L S+++ C+ G +H L +GS + + SN LI +Y K + A +VFD+MP
Sbjct: 9 LVSILRVCTRKGLSDQGGQVHCYLLKSGSGLNLITSNYLIDMYCKCREPLMAYKVFDSMP 68
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQ 172
R+ +SW+++++ ++ NG L+ +L + + G+ P ++ + CG G Q
Sbjct: 69 ERNVVSWSALMSGHVLNGDLKGSLSLFSEMGRQGIYPNEFTFSTNLKACGLLNALEKGLQ 128
Query: 173 IHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANL 232
IH + G E V + +LVD Y +C A +VF + ++ +SW AMI+G +
Sbjct: 129 IHGFCLKIG-FEMMVEVGNSLVDMYSKCGRINEAEKVFRRIVDRSLISWNAMIAGFVHAG 187
Query: 233 DYDAAFARFRAMQVEGVD--PNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIE--SSH 288
A F MQ + P+ TL +LL AC+ G + GK+IHG+ R G SS
Sbjct: 188 YGSKALDTFGMMQEANIKERPDEFTLTSLLKACSSTGMIYAGKQIHGFLVRSGFHCPSSA 247
Query: 289 IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLT 348
+ +L+++Y + G L A + F++ + ++ WSS+I Y+Q G+ +A+ LF ++
Sbjct: 248 TITGSLVDLYVKCGY-LFSARKAFDQIKEKTMISWSSLILGYAQEGEFVEAMGLFKRLQE 306
Query: 349 EETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEG 408
++ + L ++I + + L+ G + +K S+ N++++MY KCG ++
Sbjct: 307 LNSQIDSFALSSIIGVFADFALLRQGKQMQALAVKLPSGLETSVLNSVVDMYLKCGLVDE 366
Query: 409 SLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNH 468
+ + F EMQ +D ISW+ +I+ YG HG G++++++F EM ++PD + +LAVLSAC+H
Sbjct: 367 AEKCFAEMQLKDVISWTVVITGYGKHGLGKKSVRIFYEMLRHNIEPDEVCYLAVLSACSH 426
Query: 469 AGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWS 528
+G++ EG+++F ++ + I +EHYAC+VDLLGR+G++++A ++ TMP+KP+ IW
Sbjct: 427 SGMIKEGEELFSKLLETHGIKPRVEHYACVVDLLGRAGRLKEAKHLIDTMPIKPNVGIWQ 486
Query: 529 SLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQR 588
+L+S C++HG +++ + +G L+R + N ANY +++ +Y + G+W + E ++
Sbjct: 487 TLLSLCRVHGDIELGKEVGKILLRIDAKNPANYVMMSNLYGQAGYWNEQGNARELGNIKG 546
Query: 589 LKKCYGFSRIE 599
LKK G S +E
Sbjct: 547 LKKEAGMSWVE 557
Score = 155 bits (391), Expect = 8e-38
Identities = 93/286 (32%), Positives = 157/286 (54%), Gaps = 8/286 (2%)
Query: 251 PN-RVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAE 309
PN R L+++L C + G G ++H Y + G + I S+ LI+MYC+ E L +A
Sbjct: 3 PNQRQNLVSILRVCTRKGLSDQGGQVHCYLLKSGSGLNLITSNYLIDMYCKCREPL-MAY 61
Query: 310 QIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLS 369
++F+ R+VV WS+++ + GD +L+LF++M + PN T + AC L+
Sbjct: 62 KVFDSMPERNVVSWSALMSGHVLNGDLKGSLSLFSEMGRQGIYPNEFTFSTNLKACGLLN 121
Query: 370 SLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMIS 429
+L+ G +HGF LK G + + N+L++MY+KCG + + ++F + +R ISW+ MI+
Sbjct: 122 ALEKGLQIHGFCLKIGFEMMVEVGNSLVDMYSKCGRINEAEKVFRRIVDRSLISWNAMIA 181
Query: 430 AYGLHGHGEQALKLFSEMKERGVK--PDALTFLAVLSACNHAGLVTEGQQVFK-QVNADY 486
+ G+G +AL F M+E +K PD T ++L AC+ G++ G+Q+ V + +
Sbjct: 182 GFVHAGYGSKALDTFGMMQEANIKERPDEFTLTSLLKACSSTGMIYAGKQIHGFLVRSGF 241
Query: 487 EIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRI-WSSLV 531
P + LVDL + G + A + +K T I WSSL+
Sbjct: 242 HCPSSATITGSLVDLYVKCGYLFSARKAFD--QIKEKTMISWSSLI 285
Score = 154 bits (389), Expect = 1e-37
Identities = 105/409 (25%), Positives = 200/409 (48%), Gaps = 10/409 (2%)
Query: 34 TLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLIS 93
+L LF+++ + + +KAC L++ G +HG L G V NSL+
Sbjct: 91 SLSLFSEMGRQGIYPNEFTFSTNLKACGLLNALEKGLQIHGFCLKIGFEMMVEVGNSLVD 150
Query: 94 LYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE- 152
+Y+K I A +VF + R ISWN+MI ++ G +AL+ + + +P+
Sbjct: 151 MYSKCGRINEAEKVFRRIVDRSLISWNAMIAGFVHAGYGSKALDTFGMMQEANIKERPDE 210
Query: 153 -LLASMVSLCGRKMGSRIGRQIHALVVVDG-RVEHSVVLSTALVDFYFRCDDALMASRVF 210
L S++ C G+QIH +V G S ++ +LVD Y +C A + F
Sbjct: 211 FTLTSLLKACSSTGMIYAGKQIHGFLVRSGFHCPSSATITGSLVDLYVKCGYLFSARKAF 270
Query: 211 DGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVK 270
D ++ K +SW+++I G ++ A F+ +Q + L +++ A ++
Sbjct: 271 DQIKEKTMISWSSLILGYAQEGEFVEAMGLFKRLQELNSQIDSFALSSIIGVFADFALLR 330
Query: 271 HGKEIHGYAFR--HGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIG 328
GK++ A + G+E+S + +++++MY + G + AE+ F +DV+ W+ +I
Sbjct: 331 QGKQMQALAVKLPSGLETSVL--NSVVDMYLKCG-LVDEAEKCFAEMQLKDVISWTVVIT 387
Query: 329 SYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILK-FGLS 387
Y + G K++ +F +ML EP+ V LAV+SAC++ +K G L +L+ G+
Sbjct: 388 GYGKHGLGKKSVRIFYEMLRHNIEPDEVCYLAVLSACSHSGMIKEGEELFSKLLETHGIK 447
Query: 388 FSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSIS-WSTMISAYGLHG 435
+ ++++ + G L+ + + M + ++ W T++S +HG
Sbjct: 448 PRVEHYACVVDLLGRAGRLKEAKHLIDTMPIKPNVGIWQTLLSLCRVHG 496
Score = 55.5 bits (132), Expect = 9e-08
Identities = 46/218 (21%), Positives = 90/218 (41%), Gaps = 1/218 (0%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
S I +G + + + LF +L S L S+I + G + LA+
Sbjct: 282 SSLILGYAQEGEFVEAMGLFKRLQELNSQIDSFALSSIIGVFADFALLRQGKQMQALAVK 341
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
S + V NS++ +Y K + A + F M +D ISW +I Y ++G ++++ +
Sbjct: 342 LPSGLETSVLNSVVDMYLKCGLVDEAEKCFAEMQLKDVISWTVVITGYGKHGLGKKSVRI 401
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+ + P +++S C + G ++ + ++ ++ V +VD
Sbjct: 402 FYEMLRHNIEPDEVCYLAVLSACSHSGMIKEGEELFSKLLETHGIKPRVEHYACVVDLLG 461
Query: 199 RCDDALMASRVFDGMEVKNEVS-WTAMISGCIANLDYD 235
R A + D M +K V W ++S C + D +
Sbjct: 462 RAGRLKEAKHLIDTMPIKPNVGIWQTLLSLCRVHGDIE 499
>At3g22150 hypothetical protein
Length = 820
Score = 330 bits (846), Expect = 1e-90
Identities = 199/571 (34%), Positives = 325/571 (56%), Gaps = 17/571 (2%)
Query: 45 APHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSH--SDPVVSNSLISLYAKFSDIQ 102
+P S +V P+V S S + +GL L G D V +S IS+YA+ DI+
Sbjct: 213 SPVSFVNVFPAV----SISRSIKKANVFYGLMLKLGDEYVKDLFVVSSAISMYAELGDIE 268
Query: 103 SARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM-LKGVYLLGLVPKPELLASMVSLC 161
S+R+VFD+ R+ WN+MI Y+QN CL E++E+ L+ + +V S
Sbjct: 269 SSRRVFDSCVERNIEVWNTMIGVYVQNDCLVESIELFLEAIGSKEIVSDEVTYLLAASAV 328
Query: 162 GRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSW 221
+GRQ H V + R E +V+ +L+ Y RC + VF M ++ VSW
Sbjct: 329 SALQQVELGRQFHGFVSKNFR-ELPIVIVNSLMVMYSRCGSVHKSFGVFLSMRERDVVSW 387
Query: 222 TAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFR 281
MIS + N D MQ +G + +T+ ALL A + + GK+ H + R
Sbjct: 388 NTMISAFVQNGLDDEGLMLVYEMQKQGFKIDYITVTALLSAASNLRNKEIGKQTHAFLIR 447
Query: 282 HGIESSHIFSSALINMYCQSGESLHLAEQIFERSSF--RDVVLWSSIIGSYSQRGDSYKA 339
GI+ + +S LI+MY +SG + +++++FE S + RD W+S+I Y+Q G + K
Sbjct: 448 QGIQFEGM-NSYLIDMYSKSG-LIRISQKLFEGSGYAERDQATWNSMISGYTQNGHTEKT 505
Query: 340 LTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNM 399
+F KML + PN VT+ +++ AC+ + S+ G LHGF ++ L ++ +++AL++M
Sbjct: 506 FLVFRKMLEQNIRPNAVTVASILPACSQIGSVDLGKQLHGFSIRQYLDQNVFVASALVDM 565
Query: 400 YAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTF 459
Y+K G ++ + +F + + R+S++++TMI YG HG GE+A+ LF M+E G+KPDA+TF
Sbjct: 566 YSKAGAIKYAEDMFSQTKERNSVTYTTMILGYGQHGMGERAISLFLSMQESGIKPDAITF 625
Query: 460 LAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMP 519
+AVLSAC+++GL+ EG ++F+++ Y I + EHY C+ D+LGR G+V +A E ++ +
Sbjct: 626 VAVLSACSYSGLIDEGLKIFEEMREVYNIQPSSEHYCCITDMLGRVGRVNEAYEFVKGLG 685
Query: 520 MKPS-TRIWSSLVSACKLHGRLDIAEMLGPQLIR-SEPDNAANY-TLLNMIYAEHGHWLH 576
+ + +W SL+ +CKLHG L++AE + +L + + N + Y LL+ +YAE W
Sbjct: 686 EEGNIAELWGSLLGSCKLHGELELAETVSERLAKFDKGKNFSGYEVLLSNMYAEEQKWKS 745
Query: 577 KEQVMEAMKLQRLKKCYGFSRIETGN--QCF 605
++V M+ + LKK G S IE CF
Sbjct: 746 VDKVRRGMREKGLKKEVGRSGIEIAGYVNCF 776
Score = 171 bits (432), Expect = 1e-42
Identities = 137/536 (25%), Positives = 261/536 (48%), Gaps = 30/536 (5%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLP--SVIKACSSLHSHTLGTLLHGLALTT 79
I I L H+ L ++++ +AP + S +KAC+ + G +H +
Sbjct: 77 IIGFICNNLPHEALLFYSRMKKTAPFTNCDAYTYSSTLKACAETKNLKAGKAVHCHLIRC 136
Query: 80 GSHSDPVVSNSLISLYAK------FSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLE 133
+S VV NSL+++Y + R+VFD M ++ ++WN++I+ Y++ G
Sbjct: 137 LQNSSRVVHNSLMNMYVSCLNAPDCFEYDVVRKVFDNMRRKNVVAWNTLISWYVKTGRNA 196
Query: 134 EALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIH--------ALVVVDGRVEH 185
EA G++ + E+ S VS I R I L + D V+
Sbjct: 197 EACRQ------FGIMMRMEVKPSPVSFVNVFPAVSISRSIKKANVFYGLMLKLGDEYVKD 250
Query: 186 SVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARF-RAM 244
V+S+A + Y D + RVFD +N W MI + N + F A+
Sbjct: 251 LFVVSSA-ISMYAELGDIESSRRVFDSCVERNIEVWNTMIGVYVQNDCLVESIELFLEAI 309
Query: 245 QVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGES 304
+ + + VT + A + V+ G++ HG+ ++ E + ++L+ MY + G S
Sbjct: 310 GSKEIVSDEVTYLLAASAVSALQQVELGRQFHGFVSKNFRELPIVIVNSLMVMYSRCG-S 368
Query: 305 LHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISA 364
+H + +F RDVV W+++I ++ Q G + L L +M + + +Y+T+ A++SA
Sbjct: 369 VHKSFGVFLSMRERDVVSWNTMISAFVQNGLDDEGLMLVYEMQKQGFKIDYITVTALLSA 428
Query: 365 CTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIF--LEMQNRDSI 422
+NL + + G H F+++ G+ F +++ L++MY+K G + S ++F RD
Sbjct: 429 ASNLRNKEIGKQTHAFLIRQGIQFE-GMNSYLIDMYSKSGLIRISQKLFEGSGYAERDQA 487
Query: 423 SWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQV 482
+W++MIS Y +GH E+ +F +M E+ ++P+A+T ++L AC+ G V G+Q+
Sbjct: 488 TWNSMISGYTQNGHTEKTFLVFRKMLEQNIRPNAVTVASILPACSQIGSVDLGKQLHGFS 547
Query: 483 NADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
Y + + + LVD+ ++G ++ A E M + + ++ +++++ HG
Sbjct: 548 IRQY-LDQNVFVASALVDMYSKAGAIKYA-EDMFSQTKERNSVTYTTMILGYGQHG 601
Score = 158 bits (399), Expect = 9e-39
Identities = 127/493 (25%), Positives = 237/493 (47%), Gaps = 46/493 (9%)
Query: 79 TGSHSDPVVSNSLISLYAKFSDI------QSARQVFDTMPHRDPISWNSMINAYLQNGCL 132
+ + S P ++ S+ ++ S I Q ARQ+FD +P + WN++I ++ N
Sbjct: 27 SSTFSPPTLTPQTPSIRSRLSKICQDGNPQLARQLFDAIPKPTTVLWNTIIIGFICNNLP 86
Query: 133 EEALEMLKGVYLLGLVPKPELL--ASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLS 190
EAL + + +S + C + G+ +H ++ + S V+
Sbjct: 87 HEALLFYSRMKKTAPFTNCDAYTYSSTLKACAETKNLKAGKAVHCHLIRCLQ-NSSRVVH 145
Query: 191 TALVDFYFRCDDAL------MASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAM 244
+L++ Y C +A + +VFD M KN V+W +IS + A +F M
Sbjct: 146 NSLMNMYVSCLNAPDCFEYDVVRKVFDNMRRKNVVAWNTLISWYVKTGRNAEACRQFGIM 205
Query: 245 QVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHG---IESSHIFSSALINMYCQS 301
V P+ V+ + + PA + +K +G + G ++ + SSA I+MY +
Sbjct: 206 MRMEVKPSPVSFVNVFPAVSISRSIKKANVFYGLMLKLGDEYVKDLFVVSSA-ISMYAEL 264
Query: 302 GESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKML-TEETEPNYVTLLA 360
G+ + + ++F+ R++ +W+++IG Y Q +++ LF + + ++E + VT L
Sbjct: 265 GD-IESSRRVFDSCVERNIEVWNTMIGVYVQNDCLVESIELFLEAIGSKEIVSDEVTYLL 323
Query: 361 VISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRD 420
SA + L ++ G HGF+ K I + N+LM MY++CG + S +FL M+ RD
Sbjct: 324 AASAVSALQQVELGRQFHGFVSKNFRELPIVIVNSLMVMYSRCGSVHKSFGVFLSMRERD 383
Query: 421 SISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACN-----------HA 469
+SW+TMISA+ +G ++ L L EM+++G K D +T A+LSA + HA
Sbjct: 384 VVSWNTMISAFVQNGLDDEGLMLVYEMQKQGFKIDYITVTALLSAASNLRNKEIGKQTHA 443
Query: 470 GLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMR-TMPMKPSTRIWS 528
L+ +G Q F+ +N + L+D+ +SG + + ++ + + W+
Sbjct: 444 FLIRQGIQ-FEGMN------------SYLIDMYSKSGLIRISQKLFEGSGYAERDQATWN 490
Query: 529 SLVSACKLHGRLD 541
S++S +G +
Sbjct: 491 SMISGYTQNGHTE 503
Score = 106 bits (264), Expect = 4e-23
Identities = 90/353 (25%), Positives = 173/353 (48%), Gaps = 14/353 (3%)
Query: 205 MASRVFDGMEVKNEVSWTAMISGCIAN-LDYDAAFARFRAMQVEG-VDPNRVTLIALLPA 262
+A ++FD + V W +I G I N L ++A R + + + T + L A
Sbjct: 57 LARQLFDAIPKPTTVLWNTIIIGFICNNLPHEALLFYSRMKKTAPFTNCDAYTYSSTLKA 116
Query: 263 CAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMY--CQSGESLH---LAEQIFERSSF 317
CA+ +K GK +H + R SS + ++L+NMY C + + ++F+
Sbjct: 117 CAETKNLKAGKAVHCHLIRCLQNSSRVVHNSLMNMYVSCLNAPDCFEYDVVRKVFDNMRR 176
Query: 318 RDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGL 377
++VV W+++I Y + G + +A F M+ E +P+ V+ + V A + S+K
Sbjct: 177 KNVVAWNTLISWYVKTGRNAEACRQFGIMMRMEVKPSPVSFVNVFPAVSISRSIKKANVF 236
Query: 378 HGFILKFGLSF--SISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHG 435
+G +LK G + + + ++ ++MYA+ G +E S ++F R+ W+TMI Y +
Sbjct: 237 YGLMLKLGDEYVKDLFVVSSAISMYAELGDIESSRRVFDSCVERNIEVWNTMIGVYVQND 296
Query: 436 HGEQALKLFSE-MKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADY-EIPLTIE 493
++++LF E + + + D +T+L SA + V G+Q V+ ++ E+P+ I
Sbjct: 297 CLVESIELFLEAIGSKEIVSDEVTYLLAASAVSALQQVELGRQFHGFVSKNFRELPIVIV 356
Query: 494 HYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEML 546
+ L+ + R G V + + +M + W++++SA +G D ML
Sbjct: 357 N--SLMVMYSRCGSVHKSFGVFLSMRERDVVS-WNTMISAFVQNGLDDEGLML 406
>At3g53360 putative protein
Length = 768
Score = 328 bits (842), Expect = 4e-90
Identities = 191/549 (34%), Positives = 301/549 (54%), Gaps = 8/549 (1%)
Query: 55 SVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHR 114
S+IKAC+S LG LH + S S + N+LI++Y +F+ + A +VF +P +
Sbjct: 173 SIIKACASSSDVGLGKQLHAQVIKLESSSHLIAQNALIAMYVRFNQMSDASRVFYGIPMK 232
Query: 115 DPISWNSMINAYLQNGCLEEALEMLKGVYLLGLV-PKPELLASMVSLCGRKMGSRIGRQI 173
D ISW+S+I + Q G EAL LK + G+ P + S + C + G QI
Sbjct: 233 DLISWSSIIAGFSQLGFEFEALSHLKEMLSFGVFHPNEYIFGSSLKACSSLLRPDYGSQI 292
Query: 174 HALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLD 233
H L + + + + +L D Y RC A RVFD +E + SW +I+G N
Sbjct: 293 HGLCI-KSELAGNAIAGCSLCDMYARCGFLNSARRVFDQIERPDTASWNVIIAGLANNGY 351
Query: 234 YDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSA 293
D A + F M+ G P+ ++L +LL A KP + G +IH Y + G + ++
Sbjct: 352 ADEAVSVFSQMRSSGFIPDAISLRSLLCAQTKPMALSQGMQIHSYIIKWGFLADLTVCNS 411
Query: 294 LINMYCQSGESLHLAEQIFERSSFR---DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEE 350
L+ MY + L+ +FE FR D V W++I+ + Q + L LF ML E
Sbjct: 412 LLTMYTFCSD-LYCCFNLFE--DFRNNADSVSWNTILTACLQHEQPVEMLRLFKLMLVSE 468
Query: 351 TEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSL 410
EP+++T+ ++ C +SSLK G +H + LK GL+ + N L++MYAKCG L +
Sbjct: 469 CEPDHITMGNLLRGCVEISSLKLGSQVHCYSLKTGLAPEQFIKNGLIDMYAKCGSLGQAR 528
Query: 411 QIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAG 470
+IF M NRD +SWST+I Y G GE+AL LF EMK G++P+ +TF+ VL+AC+H G
Sbjct: 529 RIFDSMDNRDVVSWSTLIVGYAQSGFGEEALILFKEMKSAGIEPNHVTFVGVLTACSHVG 588
Query: 471 LVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSL 530
LV EG +++ + ++ I T EH +C+VDLL R+G++ +A + M ++P +W +L
Sbjct: 589 LVEEGLKLYATMQTEHGISPTKEHCSCVVDLLARAGRLNEAERFIDEMKLEPDVVVWKTL 648
Query: 531 VSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLK 590
+SACK G + +A+ +++ +P N+ + LL ++A G+W + + +MK +K
Sbjct: 649 LSACKTQGNVHLAQKAAENILKIDPFNSTAHVLLCSMHASSGNWENAALLRSSMKKHDVK 708
Query: 591 KCYGFSRIE 599
K G S IE
Sbjct: 709 KIPGQSWIE 717
Score = 207 bits (526), Expect = 2e-53
Identities = 146/533 (27%), Positives = 260/533 (48%), Gaps = 13/533 (2%)
Query: 5 TRRFLATVSPTPSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSIS-SVLPSVIKACSSL 63
T ++T+ + I SL Y + LE F ++ I S+I ACSS
Sbjct: 21 TSSVVSTIKTEELMNDHINSLCKSNFYREALEAFDFAQKNSSFKIRLRTYISLICACSSS 80
Query: 64 HSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMI 123
S G +H L + D +++N ++S+Y K ++ AR+VFD MP R+ +S+ S+I
Sbjct: 81 RSLAQGRKIHDHILNSNCKYDTILNNHILSMYGKCGSLRDAREVFDFMPERNLVSYTSVI 140
Query: 124 NAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRV 183
Y QNG EA+ + + LVP S++ C +G+Q+HA V+
Sbjct: 141 TGYSQNGQGAEAIRLYLKMLQEDLVPDQFAFGSIIKACASSSDVGLGKQLHAQVIKLESS 200
Query: 184 EHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYD-AAFARFR 242
H ++ AL+ Y R + ASRVF G+ +K+ +SW+++I+G + L ++ A + +
Sbjct: 201 SH-LIAQNALIAMYVRFNQMSDASRVFYGIPMKDLISWSSIIAG-FSQLGFEFEALSHLK 258
Query: 243 AMQVEGV-DPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQS 301
M GV PN + L AC+ +G +IHG + + + I +L +MY +
Sbjct: 259 EMLSFGVFHPNEYIFGSSLKACSSLLRPDYGSQIHGLCIKSELAGNAIAGCSLCDMYARC 318
Query: 302 GESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAV 361
G L+ A ++F++ D W+ II + G + +A+++F++M + P+ ++L ++
Sbjct: 319 G-FLNSARRVFDQIERPDTASWNVIIAGLANNGYADEAVSVFSQMRSSGFIPDAISLRSL 377
Query: 362 ISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNR-D 420
+ A T +L G +H +I+K+G +++ N+L+ MY C L +F + +N D
Sbjct: 378 LCAQTKPMALSQGMQIHSYIIKWGFLADLTVCNSLLTMYTFCSDLYCCFNLFEDFRNNAD 437
Query: 421 SISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFK 480
S+SW+T+++A H + L+LF M +PD +T +L C + G QV
Sbjct: 438 SVSWNTILTACLQHEQPVEMLRLFKLMLVSECEPDHITMGNLLRGCVEISSLKLGSQVHC 497
Query: 481 QVNADYEIPLTIEHYA--CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV 531
+ L E + L+D+ + G + A I +M + WS+L+
Sbjct: 498 Y---SLKTGLAPEQFIKNGLIDMYAKCGSLGQARRIFDSMDNRDVVS-WSTLI 546
Score = 110 bits (276), Expect = 2e-24
Identities = 79/297 (26%), Positives = 144/297 (47%), Gaps = 4/297 (1%)
Query: 255 TLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFER 314
T I+L+ AC+ + G++IH + + I ++ +++MY + G SL A ++F+
Sbjct: 69 TYISLICACSSSRSLAQGRKIHDHILNSNCKYDTILNNHILSMYGKCG-SLRDAREVFDF 127
Query: 315 SSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHG 374
R++V ++S+I YSQ G +A+ L+ KML E+ P+ ++I AC + S + G
Sbjct: 128 MPERNLVSYTSVITGYSQNGQGAEAIRLYLKMLQEDLVPDQFAFGSIIKACASSSDVGLG 187
Query: 375 CGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLH 434
LH ++K S + NAL+ MY + + + ++F + +D ISWS++I+ +
Sbjct: 188 KQLHAQVIKLESSSHLIAQNALIAMYVRFNQMSDASRVFYGIPMKDLISWSSIIAGFSQL 247
Query: 435 GHGEQALKLFSEMKERGV-KPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIE 493
G +AL EM GV P+ F + L AC+ G Q+ + E+
Sbjct: 248 GFEFEALSHLKEMLSFGVFHPNEYIFGSSLKACSSLLRPDYGSQI-HGLCIKSELAGNAI 306
Query: 494 HYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQL 550
L D+ R G + A + + +P T W+ +++ +G D A + Q+
Sbjct: 307 AGCSLCDMYARCGFLNSARRVFDQIE-RPDTASWNVIIAGLANNGYADEAVSVFSQM 362
Score = 88.6 bits (218), Expect = 9e-18
Identities = 59/229 (25%), Positives = 108/229 (46%), Gaps = 3/229 (1%)
Query: 33 QTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLI 92
+ L LF + S + ++++ C + S LG+ +H +L TG + + N LI
Sbjct: 456 EMLRLFKLMLVSECEPDHITMGNLLRGCVEISSLKLGSQVHCYSLKTGLAPEQFIKNGLI 515
Query: 93 SLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE 152
+YAK + AR++FD+M +RD +SW+++I Y Q+G EEAL + K + G+ P
Sbjct: 516 DMYAKCGSLGQARRIFDSMDNRDVVSWSTLIVGYAQSGFGEEALILFKEMKSAGIEPNHV 575
Query: 153 LLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDG 212
+++ C G +++A + + + + + +VD R A R D
Sbjct: 576 TFVGVLTACSHVGLVEEGLKLYATMQTEHGISPTKEHCSCVVDLLARAGRLNEAERFIDE 635
Query: 213 MEVKNE-VSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALL 260
M+++ + V W ++S C + A+ A + +DP T LL
Sbjct: 636 MKLEPDVVVWKTLLSAC--KTQGNVHLAQKAAENILKIDPFNSTAHVLL 682
>At3g63370 putative protein
Length = 1017
Score = 326 bits (836), Expect = 2e-89
Identities = 182/579 (31%), Positives = 322/579 (55%), Gaps = 4/579 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
+ S + G +TLELF ++H + P S + S + AC LG +H L + +
Sbjct: 219 LSSYSTSGKSLETLELFREMHMTGPAPNSYTIVSALTACDGFSYAKLGKEIHASVLKSST 278
Query: 82 HSDPV-VSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLK 140
HS + V N+LI++Y + + A ++ M + D ++WNS+I Y+QN +EALE
Sbjct: 279 HSSELYVCNALIAMYTRCGKMPQAERILRQMNNADVVTWNSLIKGYVQNLMYKEALEFFS 338
Query: 141 GVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRC 200
+ G + S+++ GR G ++HA V+ G + ++ + L+D Y +C
Sbjct: 339 DMIAAGHKSDEVSMTSIIAASGRLSNLLAGMELHAYVIKHGW-DSNLQVGNTLIDMYSKC 397
Query: 201 DDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALL 260
+ R F M K+ +SWT +I+G N + A FR + + ++ + + L ++L
Sbjct: 398 NLTCYMGRAFLRMHDKDLISWTTVIAGYAQNDCHVEALELFRDVAKKRMEIDEMILGSIL 457
Query: 261 PACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
A + + KEIH + R G+ + + + L+++Y + ++ A ++FE +DV
Sbjct: 458 RASSVLKSMLIVKEIHCHILRKGLLDT-VIQNELVDVYGKC-RNMGYATRVFESIKGKDV 515
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
V W+S+I S + G+ +A+ LF +M+ + V LL ++SA +LS+L G +H +
Sbjct: 516 VSWTSMISSSALNGNESEAVELFRRMVETGLSADSVALLCILSAAASLSALNKGREIHCY 575
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
+L+ G S++ A+++MYA CG L+ + +F ++ + + +++MI+AYG+HG G+ A
Sbjct: 576 LLRKGFCLEGSIAVAVVDMYACCGDLQSAKAVFDRIERKGLLQYTSMINAYGMHGCGKAA 635
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
++LF +M+ V PD ++FLA+L AC+HAGL+ EG+ K + +YE+ EHY CLVD
Sbjct: 636 VELFDKMRHENVSPDHISFLALLYACSHAGLLDEGRGFLKIMEHEYELEPWPEHYVCLVD 695
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAAN 560
+LGR+ V +A E ++ M +P+ +W +L++AC+ H +I E+ +L+ EP N N
Sbjct: 696 MLGRANCVVEAFEFVKMMKTEPTAEVWCALLAACRSHSEKEIGEIAAQRLLELEPKNPGN 755
Query: 561 YTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
L++ ++AE G W E+V MK ++K G S IE
Sbjct: 756 LVLVSNVFAEQGRWNDVEKVRAKMKASGMEKHPGCSWIE 794
Score = 224 bits (570), Expect = 1e-58
Identities = 147/555 (26%), Positives = 283/555 (50%), Gaps = 18/555 (3%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I + +S G L L+ + S P+++KAC+ L G+ LH L + G
Sbjct: 117 IGAYVSNGEPASALALYWNMRVEGVPLGLSSFPALLKACAKLRDIRSGSELHSLLVKLGY 176
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHR-DPISWNSMINAYLQNGCLEEALEMLK 140
HS + N+L+S+YAK D+ +AR++FD + D + WNS++++Y +G E LE+ +
Sbjct: 177 HSTGFIVNALVSMYAKNDDLSAARRLFDGFQEKGDAVLWNSILSSYSTSGKSLETLELFR 236
Query: 141 GVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRC 200
+++ G P + S ++ C +++G++IHA V+ + + AL+ Y RC
Sbjct: 237 EMHMTGPAPNSYTIVSALTACDGFSYAKLGKEIHASVLKSSTHSSELYVCNALIAMYTRC 296
Query: 201 DDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALL 260
A R+ M + V+W ++I G + NL Y A F M G + V++ +++
Sbjct: 297 GKMPQAERILRQMNNADVVTWNSLIKGYVQNLMYKEALEFFSDMIAAGHKSDEVSMTSII 356
Query: 261 PACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
A + + G E+H Y +HG +S+ + LI+MY + + ++ + F R +D+
Sbjct: 357 AASGRLSNLLAGMELHAYVIKHGWDSNLQVGNTLIDMYSKCNLTCYMG-RAFLRMHDKDL 415
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
+ W+++I Y+Q +AL LF + + E + + L +++ A + L S+ +H
Sbjct: 416 ISWTTVIAGYAQNDCHVEALELFRDVAKKRMEIDEMILGSILRASSVLKSMLIVKEIHCH 475
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
IL+ GL ++ + N L+++Y KC + + ++F ++ +D +SW++MIS+ L+G+ +A
Sbjct: 476 ILRKGLLDTV-IQNELVDVYGKCRNMGYATRVFESIKGKDVVSWTSMISSSALNGNESEA 534
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVF-----KQVNADYEIPL-TIEH 494
++LF M E G+ D++ L +LSA + +G+++ K + I + ++
Sbjct: 535 VELFRRMVETGLSADSVALLCILSAAASLSALNKGREIHCYLLRKGFCLEGSIAVAVVDM 594
Query: 495 YACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSE 554
YAC DL V D +E + ++S+++A +HG A L ++ R E
Sbjct: 595 YACCGDLQSAKA-VFDRIE-------RKGLLQYTSMINAYGMHGCGKAAVELFDKM-RHE 645
Query: 555 PDNAANYTLLNMIYA 569
+ + + L ++YA
Sbjct: 646 NVSPDHISFLALLYA 660
Score = 189 bits (479), Expect = 5e-48
Identities = 125/438 (28%), Positives = 227/438 (51%), Gaps = 7/438 (1%)
Query: 103 SARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCG 162
S +VFD MP R +WN+MI AY+ NG AL + + + G+ +++ C
Sbjct: 97 SQEKVFDEMPDRTAFAWNTMIGAYVSNGEPASALALYWNMRVEGVPLGLSSFPALLKACA 156
Query: 163 RKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNE-VSW 221
+ R G ++H+L+V G +++ ALV Y + DD A R+FDG + K + V W
Sbjct: 157 KLRDIRSGSELHSLLVKLGYHSTGFIVN-ALVSMYAKNDDLSAARRLFDGFQEKGDAVLW 215
Query: 222 TAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFR 281
+++S + FR M + G PN T+++ L AC + K GKEIH +
Sbjct: 216 NSILSSYSTSGKSLETLELFREMHMTGPAPNSYTIVSALTACDGFSYAKLGKEIHASVLK 275
Query: 282 HGIESSHIF-SSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKAL 340
SS ++ +ALI MY + G+ + AE+I + + DVV W+S+I Y Q +AL
Sbjct: 276 SSTHSSELYVCNALIAMYTRCGK-MPQAERILRQMNNADVVTWNSLIKGYVQNLMYKEAL 334
Query: 341 TLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMY 400
F+ M+ + + V++ ++I+A LS+L G LH +++K G ++ + N L++MY
Sbjct: 335 EFFSDMIAAGHKSDEVSMTSIIAASGRLSNLLAGMELHAYVIKHGWDSNLQVGNTLIDMY 394
Query: 401 AKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFL 460
+KC + FL M ++D ISW+T+I+ Y + +AL+LF ++ ++ ++ D +
Sbjct: 395 SKCNLTCYMGRAFLRMHDKDLISWTTVIAGYAQNDCHVEALELFRDVAKKRMEIDEMILG 454
Query: 461 AVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPM 520
++L A + + +++ + + I++ LVD+ G+ + A + ++
Sbjct: 455 SILRASSVLKSMLIVKEIHCHILRKGLLDTVIQNE--LVDVYGKCRNMGYATRVFESIKG 512
Query: 521 KPSTRIWSSLVSACKLHG 538
K W+S++S+ L+G
Sbjct: 513 KDVVS-WTSMISSSALNG 529
Score = 146 bits (368), Expect = 4e-35
Identities = 87/282 (30%), Positives = 151/282 (52%), Gaps = 3/282 (1%)
Query: 203 ALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPA 262
A+ +VFD M + +W MI ++N + +A A + M+VEGV + ALL A
Sbjct: 95 AVSQEKVFDEMPDRTAFAWNTMIGAYVSNGEPASALALYWNMRVEGVPLGLSSFPALLKA 154
Query: 263 CAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR-DVV 321
CAK ++ G E+H + G S+ +AL++MY ++ + L A ++F+ + D V
Sbjct: 155 CAKLRDIRSGSELHSLLVKLGYHSTGFIVNALVSMYAKN-DDLSAARRLFDGFQEKGDAV 213
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFI 381
LW+SI+ SYS G S + L LF +M PN T+++ ++AC S K G +H +
Sbjct: 214 LWNSILSSYSTSGKSLETLELFREMHMTGPAPNSYTIVSALTACDGFSYAKLGKEIHASV 273
Query: 382 LKFGL-SFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
LK S + + NAL+ MY +CG + + +I +M N D ++W+++I Y + ++A
Sbjct: 274 LKSSTHSSELYVCNALIAMYTRCGKMPQAERILRQMNNADVVTWNSLIKGYVQNLMYKEA 333
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQV 482
L+ FS+M G K D ++ ++++A + G ++ V
Sbjct: 334 LEFFSDMIAAGHKSDEVSMTSIIAASGRLSNLLAGMELHAYV 375
Score = 64.3 bits (155), Expect = 2e-10
Identities = 57/253 (22%), Positives = 113/253 (44%), Gaps = 24/253 (9%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
+ I S G + +ELF ++ + + S L ++ A +SL + G +H L
Sbjct: 519 TSMISSSALNGNESEAVELFRRMVETGLSADSVALLCILSAAASLSALNKGREIHCYLLR 578
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G + ++ +++ +YA D+QSA+ VFD + + + + SMINAY +GC + A+E+
Sbjct: 579 KGFCLEGSIAVAVVDMYACCGDLQSAKAVFDRIERKGLLQYTSMINAYGMHGCGKAAVEL 638
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGR-----VEHSVVLS--- 190
+ + P +++ C HA ++ +GR +EH L
Sbjct: 639 FDKMRHENVSPDHISFLALLYACS-----------HAGLLDEGRGFLKIMEHEYELEPWP 687
Query: 191 ---TALVDFYFRCDDALMASRVFDGMEVKNEVS-WTAMISGCIANLDYD-AAFARFRAMQ 245
LVD R + + A M+ + W A+++ C ++ + + A R ++
Sbjct: 688 EHYVCLVDMLGRANCVVEAFEFVKMMKTEPTAEVWCALLAACRSHSEKEIGEIAAQRLLE 747
Query: 246 VEGVDPNRVTLIA 258
+E +P + L++
Sbjct: 748 LEPKNPGNLVLVS 760
>At2g39620 hypothetical protein
Length = 836
Score = 325 bits (832), Expect = 6e-89
Identities = 170/535 (31%), Positives = 307/535 (56%), Gaps = 3/535 (0%)
Query: 29 GLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVS 88
G + + LELF + S ++A + + G +H A+ G D V+
Sbjct: 279 GFFEEVLELFDLMRNYDVRMNKVAAASALQAAAYVGDLVKGIAIHDYAVQQGLIGDVSVA 338
Query: 89 NSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLV 148
SL+S+Y+K +++ A Q+F + RD +SW++MI +Y Q G +EA+ + + + + +
Sbjct: 339 TSLMSMYSKCGELEIAEQLFINIEDRDVVSWSAMIASYEQAGQHDEAISLFRDMMRIHIK 398
Query: 149 PKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASR 208
P L S++ C SR+G+ IH + +E + +TA++ Y +C A +
Sbjct: 399 PNAVTLTSVLQGCAGVAASRLGKSIHCYAI-KADIESELETATAVISMYAKCGRFSPALK 457
Query: 209 VFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGF 268
F+ + +K+ V++ A+ G D + AF ++ M++ GV P+ T++ +L CA
Sbjct: 458 AFERLPIKDAVAFNALAQGYTQIGDANKAFDVYKNMKLHGVCPDSRTMVGMLQTCAFCSD 517
Query: 269 VKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSF-RDVVLWSSII 327
G ++G +HG +S + ALINM+ + ++L A +F++ F + V W+ ++
Sbjct: 518 YARGSCVYGQIIKHGFDSECHVAHALINMFTKC-DALAAAIVLFDKCGFEKSTVSWNIMM 576
Query: 328 GSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLS 387
Y G + +A+ F +M E+ +PN VT + ++ A LS+L+ G +H +++ G
Sbjct: 577 NGYLLHGQAEEAVATFRQMKVEKFQPNAVTFVNIVRAAAELSALRVGMSVHSSLIQCGFC 636
Query: 388 FSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEM 447
+ N+L++MYAKCG +E S + F+E+ N+ +SW+TM+SAY HG A+ LF M
Sbjct: 637 SQTPVGNSLVDMYAKCGMIESSEKCFIEISNKYIVSWNTMLSAYAAHGLASCAVSLFLSM 696
Query: 448 KERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGK 507
+E +KPD+++FL+VLSAC HAGLV EG+++F+++ ++I +EHYAC+VDLLG++G
Sbjct: 697 QENELKPDSVSFLSVLSACRHAGLVEEGKRIFEEMGERHKIEAEVEHYACMVDLLGKAGL 756
Query: 508 VEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYT 562
+A+E+MR M +K S +W +L+++ ++H L ++ QL++ EP N ++Y+
Sbjct: 757 FGEAVEMMRRMRVKTSVGVWGALLNSSRMHCNLWLSNAALCQLVKLEPLNPSHYS 811
Score = 229 bits (583), Expect = 4e-60
Identities = 138/488 (28%), Positives = 243/488 (49%), Gaps = 7/488 (1%)
Query: 57 IKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDP 116
+KAC+ G +H L G SD + +L+ +Y K D+ SARQVFD M +D
Sbjct: 107 LKACAGSMDFKKGLRIHDLIAEMGLESDVYIGTALVEMYCKARDLVSARQVFDKMHVKDV 166
Query: 117 ISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHAL 176
++WN+M++ QNGC AL + + + L +++ + S + R +H L
Sbjct: 167 VTWNTMVSGLAQNGCSSAALLLFHDMRSCCVDIDHVSLYNLIPAVSKLEKSDVCRCLHGL 226
Query: 177 VVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDA 236
V+ G + S+ L+D Y C D A VF+ + K+E SW M++ N ++
Sbjct: 227 VIKKGFI---FAFSSGLIDMYCNCADLYAAESVFEEVWRKDESSWGTMMAAYAHNGFFEE 283
Query: 237 AFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALIN 296
F M+ V N+V + L A A G + G IH YA + G+ +++L++
Sbjct: 284 VLELFDLMRNYDVRMNKVAAASALQAAAYVGDLVKGIAIHDYAVQQGLIGDVSVATSLMS 343
Query: 297 MYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYV 356
MY + GE L +AEQ+F RDVV WS++I SY Q G +A++LF M+ +PN V
Sbjct: 344 MYSKCGE-LEIAEQLFINIEDRDVVSWSAMIASYEQAGQHDEAISLFRDMMRIHIKPNAV 402
Query: 357 TLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEM 416
TL +V+ C +++ + G +H + +K + + + A+++MYAKCG +L+ F +
Sbjct: 403 TLTSVLQGCAGVAASRLGKSIHCYAIKADIESELETATAVISMYAKCGRFSPALKAFERL 462
Query: 417 QNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQ 476
+D+++++ + Y G +A ++ MK GV PD+ T + +L C G
Sbjct: 463 PIKDAVAFNALAQGYTQIGDANKAFDVYKNMKLHGVCPDSRTMVGMLQTCAFCSDYARGS 522
Query: 477 QVFKQ-VNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACK 535
V+ Q + ++ + H L+++ + + A+ + + ST W+ +++
Sbjct: 523 CVYGQIIKHGFDSECHVAH--ALINMFTKCDALAAAIVLFDKCGFEKSTVSWNIMMNGYL 580
Query: 536 LHGRLDIA 543
LHG+ + A
Sbjct: 581 LHGQAEEA 588
Score = 211 bits (536), Expect = 1e-54
Identities = 139/454 (30%), Positives = 228/454 (49%), Gaps = 16/454 (3%)
Query: 72 LHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGC 131
+HG + +G N LI+ Y+ F +R +FD++ + WNSMI Y + G
Sbjct: 24 VHGSLIVSGLKPH----NQLINAYSLFQRQDLSRVIFDSVRDPGVVLWNSMIRGYTRAGL 79
Query: 132 LEEALEMLKGVYLL---GLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVV 188
EAL Y+ G+ P + C M + G +IH L+ G +E V
Sbjct: 80 HREALGFFG--YMSEEKGIDPDKYSFTFALKACAGSMDFKKGLRIHDLIAEMG-LESDVY 136
Query: 189 LSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEG 248
+ TALV+ Y + D + A +VFD M VK+ V+W M+SG N AA F M+
Sbjct: 137 IGTALVEMYCKARDLVSARQVFDKMHVKDVVTWNTMVSGLAQNGCSSAALLLFHDMRSCC 196
Query: 249 VDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLA 308
VD + V+L L+PA +K + +HG + G + FSS LI+MYC + L+ A
Sbjct: 197 VDIDHVSLYNLIPAVSKLEKSDVCRCLHGLVIKKGFIFA--FSSGLIDMYCNCAD-LYAA 253
Query: 309 EQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNL 368
E +FE +D W +++ +Y+ G + L LF+ M + N V + + A +
Sbjct: 254 ESVFEEVWRKDESSWGTMMAAYAHNGFFEEVLELFDLMRNYDVRMNKVAAASALQAAAYV 313
Query: 369 SSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMI 428
L G +H + ++ GL +S++ +LM+MY+KCG LE + Q+F+ +++RD +SWS MI
Sbjct: 314 GDLVKGIAIHDYAVQQGLIGDVSVATSLMSMYSKCGELEIAEQLFINIEDRDVVSWSAMI 373
Query: 429 SAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVF-KQVNADYE 487
++Y G ++A+ LF +M +KP+A+T +VL C G+ + + AD E
Sbjct: 374 ASYEQAGQHDEAISLFRDMMRIHIKPNAVTLTSVLQGCAGVAASRLGKSIHCYAIKADIE 433
Query: 488 IPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMK 521
L E ++ + + G+ AL+ +P+K
Sbjct: 434 SEL--ETATAVISMYAKCGRFSPALKAFERLPIK 465
Score = 61.2 bits (147), Expect = 2e-09
Identities = 55/237 (23%), Positives = 110/237 (46%), Gaps = 24/237 (10%)
Query: 46 PHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSAR 105
P++++ V ++++A + L + +G +H + G S V NSL+ +YAK I+S+
Sbjct: 602 PNAVTFV--NIVRAAAELSALRVGMSVHSSLIQCGFCSQTPVGNSLVDMYAKCGMIESSE 659
Query: 106 QVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGR-- 163
+ F + ++ +SWN+M++AY +G A+ + + L P S++S C
Sbjct: 660 KCFIEISNKYIVSWNTMLSAYAAHGLASCAVSLFLSMQENELKPDSVSFLSVLSACRHAG 719
Query: 164 --KMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVS- 220
+ G RI ++ ++ VEH +VD + A + M VK V
Sbjct: 720 LVEEGKRIFEEMGERHKIEAEVEH----YACMVDLLGKAGLFGEAVEMMRRMRVKTSVGV 775
Query: 221 WTAMISGCIANLD-YDAAFARFRAMQVEGVDP------------NRVTLIALLPACA 264
W A+++ + + + + A + +++E ++P N V+ I +PAC+
Sbjct: 776 WGALLNSSRMHCNLWLSNAALCQLVKLEPLNPSHYSQDRRLGEVNNVSRIKKVPACS 832
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,034,702
Number of Sequences: 26719
Number of extensions: 532237
Number of successful extensions: 9325
Number of sequences better than 10.0: 455
Number of HSP's better than 10.0 without gapping: 413
Number of HSP's successfully gapped in prelim test: 42
Number of HSP's that attempted gapping in prelim test: 1328
Number of HSP's gapped (non-prelim): 2124
length of query: 605
length of database: 11,318,596
effective HSP length: 105
effective length of query: 500
effective length of database: 8,513,101
effective search space: 4256550500
effective search space used: 4256550500
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0055.11


