

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.5
(638 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
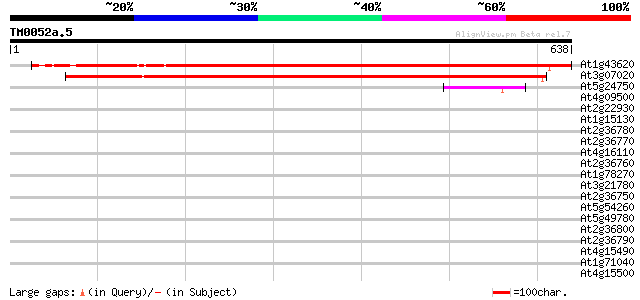
Score E
Sequences producing significant alignments: (bits) Value
At1g43620 unknown protein 877 0.0
At3g07020 UDP-glucose:sterol glucosyltransferase 622 e-178
At5g24750 sterol glucosyltransferase - like protein 70 3e-12
At4g09500 unknown protein 38 0.015
At2g22930 putative flavonol 3-O-glucosyltransferase 34 0.28
At1g15130 unknown protein 33 0.49
At2g36780 putative glucosyl transferase 32 1.1
At2g36770 putative glucosyl transferase 32 1.1
At4g16110 hypothetical protein 31 2.4
At2g36760 putative glucosyl transferase 31 2.4
At1g78270 similar to glucoronosyl transferase-like protein; simi... 31 2.4
At3g21780 putative UDP-glucose glucosyltransferase 30 3.2
At2g36750 putative glucosyl transferase 30 3.2
At5g54260 DNA repair and meiosis protein Mre11 (emb|CAB50793.1) 30 4.1
At5g49780 receptor protein kinase-like 30 4.1
At2g36800 putative glucosyl transferase 30 4.1
At2g36790 putative glucosyl transferase 30 4.1
At4g15490 indole-3-acetate beta-glucosyltransferase like protein 30 5.4
At1g71040 putative spore coat protein 30 5.4
At4g15500 indole-3-acetate beta-glucosyltransferase like protein 29 7.0
>At1g43620 unknown protein
Length = 615
Score = 877 bits (2266), Expect = 0.0
Identities = 437/620 (70%), Positives = 512/620 (82%), Gaps = 21/620 (3%)
Query: 26 LWQMDGGDDKLVRVDIKSQDGKFDLASSKEVEDWREKLGHKSSSLVTSSPRKGLGHCITA 85
L +++G D+ +KS+ L +S V+ E GH+SS +GL HC TA
Sbjct: 10 LQELEGEDN-----GVKSEKASL-LETSGSVDTTPEDSGHRSSD-----GHRGLDHCETA 58
Query: 86 PVGTERNPLIADNEIILSRSMTDKRGLPRHDLILDRLSEHEKEQLIAELVKIQNDGTVEV 145
PVG + LI D+EI SRS+T+K H+L LDRLSE EK++LI ELV+IQNDGTVEV
Sbjct: 59 PVGLYGDMLINDSEIQYSRSLTEKGSPAIHNLKLDRLSEQEKQKLIVELVRIQNDGTVEV 118
Query: 146 DMERSASVPSELLELQ-SFGEPTVS--GSLSSALKRSVPRLQIAILVVGTRGDVQPFLAI 202
++ V SEL E + + G+ T++ SL+ + RS+PRL+IAILVVGTRGDVQPFLA+
Sbjct: 119 -IDNGTPV-SELWEFEPTKGQSTITYEKSLTESF-RSIPRLKIAILVVGTRGDVQPFLAM 175
Query: 203 AKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDPRILAGYMARNKGLIPSGPGEVSI 262
AKRLQE+GHRVRLATH+NF +FV++AG++FYPLGGDPR LA YMARNKGLIPSGP E+S
Sbjct: 176 AKRLQEFGHRVRLATHANFRSFVRAAGVEFYPLGGDPRELAAYMARNKGLIPSGPSEISK 235
Query: 263 QLKQLKDIIDSLLPACTAPDLETGVPFRAQAIIANPPAYGHAHVAEALGVPLHIFFTMPW 322
Q KQLK II+SLLPAC PDLET FRAQAIIANPPAYGH HVAEALGVP+HIFFTMPW
Sbjct: 236 QRKQLKAIIESLLPACIEPDLETATSFRAQAIIANPPAYGHVHVAEALGVPIHIFFTMPW 295
Query: 323 TPTYDFPHPLARVPQSAGYWMSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRGS 382
TPT +FPHPLARVPQSA YW+SYI+VDLM WW +R INDFRK+KL L PIAYFS Y GS
Sbjct: 296 TPTNEFPHPLARVPQSAAYWLSYIVVDLMVWWSIRTYINDFRKRKLNLAPIAYFSTYHGS 355
Query: 383 ISHLPTGYMWSPHVVPKPSDWGPLVDVVGYCFLSLASKYKPREDFVQWIKKGPPPLYFGF 442
ISHLPTGYMWSPHVVPKPSDWGPLVDVVGYCFL+L SKY+PRE+F+ WI++G PP+Y GF
Sbjct: 356 ISHLPTGYMWSPHVVPKPSDWGPLVDVVGYCFLNLGSKYQPREEFLHWIERGSPPVYIGF 415
Query: 443 GSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSLA-EVPDNVFLLEECPHDWLFPQ 501
GSMPL+DP +T D+ILE LKDTEQRGI+DRGWG LG+LA EVP+NVFL+E+CPHDWLFPQ
Sbjct: 416 GSMPLDDPKQTMDIILETLKDTEQRGIVDRGWGGLGNLATEVPENVFLVEDCPHDWLFPQ 475
Query: 502 CSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQLNVENLS 561
CSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRI+EK LGPAPIPI+QL+VENLS
Sbjct: 476 CSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIYEKGLGPAPIPIAQLSVENLS 535
Query: 562 NAIKFMLQPEVKSRVMLIAKLVENEDGVAAAVDAFHRHLPDELPLPTPSPQ---EEDQIN 618
++I+FMLQPEVKS+VM +AK++ENEDGVAAAVDAFHRHLP ELPLP S + E+D+ +
Sbjct: 536 SSIRFMLQPEVKSQVMELAKVLENEDGVAAAVDAFHRHLPPELPLPESSSEKKDEDDRPD 595
Query: 619 PLQWFFLQIGKWCCAPCGGV 638
LQWFF+QIGK CC PCGGV
Sbjct: 596 LLQWFFIQIGKKCCLPCGGV 615
>At3g07020 UDP-glucose:sterol glucosyltransferase
Length = 637
Score = 622 bits (1605), Expect = e-178
Identities = 294/554 (53%), Positives = 399/554 (71%), Gaps = 8/554 (1%)
Query: 64 GHKSSSLVTSSPRKGLGHCITAPVGTERNPLIADNEIILSRSMTDKRGLPRHDLILDRLS 123
G+KS S V + P +G A + P + ++ + +T + D++S
Sbjct: 73 GNKSFSRVWTMPLEGSSSSDKAESSSTNQPRLDKSKTERQQKVTHILAEDAAKIFDDKIS 132
Query: 124 EHEKEQLIAELVKIQNDGTVEVDMERSASVPSELLELQSFGEPTVSG--SLSSALKRSVP 181
+K +L+ + +++DGTVE ++ A +P ++ + + V S+ + +P
Sbjct: 133 AGKKLKLLNRIATVKHDGTVEFEVPADA-IPQPIVVDRGESKNGVCADESIDGVDLQYIP 191
Query: 182 RLQIAILVVGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDPRI 241
+QI +L+VGTRGDVQPF+AIAKRLQ+YGHRVRLATH+NF FV +AG++FYPLGGDP++
Sbjct: 192 PMQIVMLIVGTRGDVQPFVAIAKRLQDYGHRVRLATHANFKEFVLTAGLEFYPLGGDPKV 251
Query: 242 LAGYMARNKGLIPSGPGEVSIQLKQLKDIIDSLLPACTAPDLETGVPFRAQAIIANPPAY 301
LAGYM +NKG +PSGP E+ IQ Q+KDII SLLPAC PD ++G+ F+A AIIANPPAY
Sbjct: 252 LAGYMVKNKGFLPSGPSEIPIQRNQMKDIIYSLLPACKEPDPDSGISFKADAIIANPPAY 311
Query: 302 GHAHVAEALGVPLHIFFTMPWTPTYDFPHPLARVPQSAGYWMSYIIVDLMTWWGMRGIIN 361
GH HVAEAL +P+H+FFTMPWTPT +FPHPL+RV Q AGY +SY IVD + W G+R ++N
Sbjct: 312 GHTHVAEALKIPIHVFFTMPWTPTSEFPHPLSRVKQPAGYRLSYQIVDSLIWLGIRDMVN 371
Query: 362 DFRKKKLKLPPIAYFSMYRGSISHLPTGYMWSPHVVPKPSDWGPLVDVVGYCFLSLASKY 421
D RKKKLKL P+ Y S +GS S++P GYMWSPH+VPKP DWGP +DVVG+C+L LAS Y
Sbjct: 372 DLRKKKLKLRPVTYLSGTQGSGSNIPHGYMWSPHLVPKPKDWGPQIDVVGFCYLDLASNY 431
Query: 422 KPREDFVQWIKKGPPPLYFGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGSLA 481
+P + V+W++ G P+Y GFGS+P+++P + T++I+EAL+ T+QRGII++GWG LG+L
Sbjct: 432 EPPAELVEWLEAGDKPIYIGFGSLPVQEPEKMTEIIVEALQRTKQRGIINKGWGGLGNLK 491
Query: 482 EVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIH 541
E D V+LL+ PHDWLFP+C AVVHHGGAGTTA GLKA CPTTIVPFFGDQ FWG+R+H
Sbjct: 492 EPKDFVYLLDNVPHDWLFPRCKAVVHHGGAGTTAAGLKASCPTTIVPFFGDQPFWGERVH 551
Query: 542 EKELGPAPIPISQLNVENLSNAIKFMLQPEVKSRVMLIAKLVENEDGVAAAVDAFHRHLP 601
+ +GP+PIP+ + ++ L +AI FML +VKS +AK +++EDGVA AV AF +HLP
Sbjct: 552 ARGVGPSPIPVDEFSLHKLEDAINFMLDDKVKSSAETLAKAMKDEDGVAGAVKAFFKHLP 611
Query: 602 DEL-----PLPTPS 610
P+P PS
Sbjct: 612 SAKQNISDPIPEPS 625
>At5g24750 sterol glucosyltransferase - like protein
Length = 517
Score = 70.5 bits (171), Expect = 3e-12
Identities = 34/106 (32%), Positives = 59/106 (55%), Gaps = 13/106 (12%)
Query: 494 PHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPIS 553
P++W+F C+A +HHGG+G+ A L+AG P I PF DQF+W +++ + P P+ +
Sbjct: 365 PYNWMFRTCAAAIHHGGSGSVAAALQAGIPQIICPFMLDQFYWAEKMSWLGVAPQPLKRN 424
Query: 554 QLNVEN-------------LSNAIKFMLQPEVKSRVMLIAKLVENE 586
L +E+ ++ AI L + ++R M IA+++ E
Sbjct: 425 HLLLEDSNDEKNITEAAQVVAKAIYDALSAKTRARAMEIAEILSLE 470
Score = 33.1 bits (74), Expect = 0.49
Identities = 17/59 (28%), Positives = 32/59 (53%), Gaps = 5/59 (8%)
Query: 188 LVVGTRGDVQPFLAIA---KRLQEYGHRVRLA--THSNFNTFVKSAGIDFYPLGGDPRI 241
+ GT+GDV P AIA R+Q++ V ++ H N ++ + A + ++P+ P +
Sbjct: 1 MAFGTKGDVYPLAAIAAVFARVQQHYTVVLMSHLAHENLSSHLSKAKVSYFPINSPPAL 59
>At4g09500 unknown protein
Length = 417
Score = 38.1 bits (87), Expect = 0.015
Identities = 38/166 (22%), Positives = 66/166 (38%), Gaps = 24/166 (14%)
Query: 450 PTRTTDVILEALKDTEQRGIIDRG--WGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVH 507
P R + + E L + + + DRG WG + P P V+
Sbjct: 264 PPRGSSTVQEGLPEGFEERVKDRGVVWGGW-------------VQQPLILAHPSIGCFVN 310
Query: 508 HGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQ---LNVENLSNAI 564
H G GT L + C ++PF DQ + + E+ +P + + E+LSNAI
Sbjct: 311 HCGPGTIWESLVSDCQMVLIPFLSDQVLFTRLMTEEFEVSVEVPREKTGWFSKESLSNAI 370
Query: 565 KFMLQPE------VKSRVMLIAKLVENEDGVAAAVDAFHRHLPDEL 604
K ++ + V+S + +++ + + VD F L + L
Sbjct: 371 KSVMDKDSDIGKLVRSNHTKLKEILVSPGLLTGYVDHFVEGLQENL 416
>At2g22930 putative flavonol 3-O-glucosyltransferase
Length = 442
Score = 33.9 bits (76), Expect = 0.28
Identities = 39/168 (23%), Positives = 67/168 (39%), Gaps = 28/168 (16%)
Query: 450 PTRTTDVILEALKDTEQRGIIDRG--WGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVH 507
P R + + E L + Q + RG WG + D+ P V+
Sbjct: 289 PPRGSSTVEEGLPEGFQERVKGRGVVWGGWVQQPLILDH-------------PSIGCFVN 335
Query: 508 HGGAGTTATGLKAGCPTTIVPFFGDQFFWGDRIHEKELGPAPIPISQ-----LNVENLSN 562
H G GT L C ++PF GDQ + R+ +E + +S+ + E+LS+
Sbjct: 336 HCGPGTIWECLMTDCQMVLLPFLGDQVLF-TRLMTEEF-KVSVEVSREKTGWFSKESLSD 393
Query: 563 AIKFMLQPE------VKSRVMLIAKLVENEDGVAAAVDAFHRHLPDEL 604
AIK ++ + V+S + + + + + VD F L + L
Sbjct: 394 AIKSVMDKDSDLGKLVRSNHAKLKETLGSHGLLTGYVDKFVEELQEYL 441
>At1g15130 unknown protein
Length = 846
Score = 33.1 bits (74), Expect = 0.49
Identities = 14/56 (25%), Positives = 36/56 (64%), Gaps = 1/56 (1%)
Query: 125 HEKEQLIAELVKIQNDGTVEVDMERSA-SVPSELLELQSFGEPTVSGSLSSALKRS 179
HEKE++ E+ ++++ + + ++S+ P++L+E + E +++G+L A+K +
Sbjct: 271 HEKEEIAEEIARLRSGASRLAEAKKSSRGAPAQLIEAMNTLESSINGNLDRAVKEN 326
>At2g36780 putative glucosyl transferase
Length = 496
Score = 32.0 bits (71), Expect = 1.1
Identities = 19/70 (27%), Positives = 29/70 (41%), Gaps = 16/70 (22%)
Query: 465 EQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPT 524
++RG++ +GW V +L P + H G +T G+ +G P
Sbjct: 347 KERGLLIKGWA---------PQVLILSH-------PSVGGFLTHCGWNSTLEGITSGIPL 390
Query: 525 TIVPFFGDQF 534
P FGDQF
Sbjct: 391 ITWPLFGDQF 400
>At2g36770 putative glucosyl transferase
Length = 496
Score = 32.0 bits (71), Expect = 1.1
Identities = 19/70 (27%), Positives = 29/70 (41%), Gaps = 16/70 (22%)
Query: 465 EQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPT 524
++RG++ +GW V +L P + H G +T G+ +G P
Sbjct: 347 KERGLLIKGWS---------PQVLILSH-------PSVGGFLTHCGWNSTLEGITSGIPL 390
Query: 525 TIVPFFGDQF 534
P FGDQF
Sbjct: 391 ITWPLFGDQF 400
>At4g16110 hypothetical protein
Length = 644
Score = 30.8 bits (68), Expect = 2.4
Identities = 18/52 (34%), Positives = 22/52 (41%)
Query: 190 VGTRGDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDPRI 241
V R D+ LA+ + E RV THS FN F +PL P I
Sbjct: 426 VSRRSDLTGALAVRNSIPETNSRVLPTTHSVFNNFPADLPRSSFPLASAPGI 477
>At2g36760 putative glucosyl transferase
Length = 496
Score = 30.8 bits (68), Expect = 2.4
Identities = 19/71 (26%), Positives = 29/71 (40%), Gaps = 16/71 (22%)
Query: 464 TEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCP 523
T++R ++ +GW P + L P + H G +T G+ +G P
Sbjct: 346 TKERSLLIKGWS--------PQMLILSH--------PAVGGFLTHCGWNSTLEGITSGVP 389
Query: 524 TTIVPFFGDQF 534
P FGDQF
Sbjct: 390 LITWPLFGDQF 400
>At1g78270 similar to glucoronosyl transferase-like protein; similar
to ESTs gb|AI996767.1, emb|Z46520.1, gb|T44500.1, and
gb|AI099822.1
Length = 489
Score = 30.8 bits (68), Expect = 2.4
Identities = 20/75 (26%), Positives = 29/75 (38%), Gaps = 16/75 (21%)
Query: 459 EALKDTEQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGL 518
E L +T+ RG++ +GW + + P + H G +T L
Sbjct: 347 EFLSETKNRGMLIKGWCSQEKVLS----------------HPAIGGFLTHCGWNSTLESL 390
Query: 519 KAGCPTTIVPFFGDQ 533
AG P PFF DQ
Sbjct: 391 YAGVPMICWPFFADQ 405
>At3g21780 putative UDP-glucose glucosyltransferase
Length = 431
Score = 30.4 bits (67), Expect = 3.2
Identities = 50/244 (20%), Positives = 89/244 (35%), Gaps = 60/244 (24%)
Query: 324 PTYDFPHPLARVPQ--SAGYWMSYIIVDLMTWWGMRGIINDFRKKKLKLPPIAYFSMYRG 381
P+ P+PL +P + W+++ + + +GI+ + L P A + G
Sbjct: 123 PSLTSPYPLKCLPYIFKSKEWLTFFVTQARRFRETKGILVNTVPD---LEPQALTFLSNG 179
Query: 382 SISHLPTGYMWSPHVVPKPSDWGPLVDVVGYCFLSLASKYKPREDFVQWIKKGPPP--LY 439
+I P+ GPL+ + ++ K + + ++W+ + PP ++
Sbjct: 180 NI--------------PRAYPVGPLLHLKN---VNCDYVDKKQSEILRWLDEQPPRSVVF 222
Query: 440 FGFGSMP--LEDPTRTTDVILEA--------------------------LKDTEQRGIID 471
FGSM E+ R T + L+ L++ G D
Sbjct: 223 LCFGSMGGFSEEQVRETALALDRSGHRFLWSLRRASPNILREPPGEFTNLEEILPEGFFD 282
Query: 472 RGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPTTIVPFFG 531
R N G + + V +L + P V HGG +T L G P I P +
Sbjct: 283 RT-ANRGKVIGWAEQVAILAK-------PAIGGFVSHGGWNSTLESLWFGVPMAIWPLYA 334
Query: 532 DQFF 535
+Q F
Sbjct: 335 EQKF 338
>At2g36750 putative glucosyl transferase
Length = 491
Score = 30.4 bits (67), Expect = 3.2
Identities = 19/70 (27%), Positives = 28/70 (39%), Gaps = 16/70 (22%)
Query: 465 EQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPT 524
++RG++ GW P + L P + H G +T G+ +G P
Sbjct: 342 KERGLLITGWS--------PQMLILTH--------PAVGGFLTHCGWNSTLEGITSGVPL 385
Query: 525 TIVPFFGDQF 534
P FGDQF
Sbjct: 386 LTWPLFGDQF 395
>At5g54260 DNA repair and meiosis protein Mre11 (emb|CAB50793.1)
Length = 720
Score = 30.0 bits (66), Expect = 4.1
Identities = 27/140 (19%), Positives = 59/140 (41%), Gaps = 14/140 (10%)
Query: 64 GHKSSSLVTSSPRKGLGHCITAPVGTERNPLIADNEIILSRSMTDKRGLPRHDLILDRLS 123
GH+ L+ G+G IT P + ++ S+ D P+H L+L+ +
Sbjct: 249 GHEHECLIDPQEVSGMGFHITQPGSS------------VATSLIDGESKPKHVLLLE-IK 295
Query: 124 EHEKEQLIAELVKIQNDGTVEVDMERSASV-PSELLELQSFGEPTVSGSLSSALKRSVPR 182
++ L ++ E+ ++ + + P++ + + V + A K++V R
Sbjct: 296 GNQYRPTKIPLTSVRPFEYTEIVLKDESDIDPNDQNSILEHLDKVVRNLIEKASKKAVNR 355
Query: 183 LQIAILVVGTRGDVQPFLAI 202
+I + +V + D F+ I
Sbjct: 356 SEIKLPLVRIKVDYSGFMTI 375
>At5g49780 receptor protein kinase-like
Length = 1006
Score = 30.0 bits (66), Expect = 4.1
Identities = 32/166 (19%), Positives = 69/166 (41%), Gaps = 20/166 (12%)
Query: 33 DDKLVRVDIKSQDGKFDLASSKEVEDWREKLGHKSSSLVTSSPRKGLGHCITAPVGTERN 92
D ++ D+KS + D + + +V D+ S LV + + + + +G
Sbjct: 800 DPPIIHRDVKSSNVLLDESLTAKVADFG------LSQLVEDAEKANVTAQVKGTMG---- 849
Query: 93 PLIADNEIILSRSMTDKRGLPRHDLILDRLSEHEKEQLIAELVKIQNDGTVEVDMERSAS 152
D E ++ +T+K + +++ +L+ + I+N V +M+ +
Sbjct: 850 --YLDPEYYMTNQLTEKSDVYGFGVMM--------LELLTGKIPIENGKYVVKEMKMKMN 899
Query: 153 VPSELLELQSFGEPTVSGSLSSALKRSVPRLQIAILVVGTRGDVQP 198
L +LQ F + T+S + + LK + +A+ V G +P
Sbjct: 900 KSKNLYDLQDFLDTTISATSNRNLKGFEKYVDVALRCVDPEGVKRP 945
>At2g36800 putative glucosyl transferase
Length = 495
Score = 30.0 bits (66), Expect = 4.1
Identities = 19/70 (27%), Positives = 27/70 (38%), Gaps = 16/70 (22%)
Query: 465 EQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPT 524
+ RG++ +GW P + L P + H G +T G+ AG P
Sbjct: 346 QDRGLLIKGWS--------PQMLILSH--------PSVGGFLTHCGWNSTLEGITAGLPL 389
Query: 525 TIVPFFGDQF 534
P F DQF
Sbjct: 390 LTWPLFADQF 399
>At2g36790 putative glucosyl transferase
Length = 495
Score = 30.0 bits (66), Expect = 4.1
Identities = 19/70 (27%), Positives = 27/70 (38%), Gaps = 16/70 (22%)
Query: 465 EQRGIIDRGWGNLGSLAEVPDNVFLLEECPHDWLFPQCSAVVHHGGAGTTATGLKAGCPT 524
+ RG++ +GW P + L P + H G +T G+ AG P
Sbjct: 346 QDRGLLIKGWS--------PQMLILSH--------PSVGGFLTHCGWNSTLEGITAGLPM 389
Query: 525 TIVPFFGDQF 534
P F DQF
Sbjct: 390 LTWPLFADQF 399
>At4g15490 indole-3-acetate beta-glucosyltransferase like protein
Length = 479
Score = 29.6 bits (65), Expect = 5.4
Identities = 16/54 (29%), Positives = 26/54 (47%), Gaps = 2/54 (3%)
Query: 482 EVPDNVFLLEECPHDWLF--PQCSAVVHHGGAGTTATGLKAGCPTTIVPFFGDQ 533
E+ + ++E CP + + P + + H G +T L AG P P +GDQ
Sbjct: 333 ELEEKGKIVEWCPQERVLAHPAIACFLSHCGWNSTMEALTAGVPVVCFPQWGDQ 386
>At1g71040 putative spore coat protein
Length = 581
Score = 29.6 bits (65), Expect = 5.4
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 4/67 (5%)
Query: 194 GDVQPFLAIAKRLQEYGHRVRLATHSNFNTFVKSAGIDFYPLGGDPRILAGYMARNKGLI 253
G P L + +R +Y R+ A+++ F F S G+DF +G D LA ++ L+
Sbjct: 281 GKAWPRLTVRRR--KYRFRITNASNARFFRFFFSNGLDFIVVGSDSAYLAKPVSTKSVLL 338
Query: 254 PSGPGEV 260
P E+
Sbjct: 339 --APSEI 343
>At4g15500 indole-3-acetate beta-glucosyltransferase like protein
Length = 475
Score = 29.3 bits (64), Expect = 7.0
Identities = 27/131 (20%), Positives = 49/131 (36%), Gaps = 28/131 (21%)
Query: 422 KPREDFVQWIKKGPPP--LYFGFGSMPLEDPTRTTDVILEALKDTEQRGIIDRGWGNLGS 479
KP D ++W+ P +Y FG++ + ++ GI++ G L
Sbjct: 261 KPDSDCIEWLDSREPSSVVYISFGTLAFLKQNQIDEIA---------HGILNSGLSCLWV 311
Query: 480 LA---------------EVPDNVFLLEECPHDWLF--PQCSAVVHHGGAGTTATGLKAGC 522
L E+ + ++E C + + P + + H G +T L +G
Sbjct: 312 LRPPLEGLAIEPHVLPLELEEKGKIVEWCQQEKVLAHPAVACFLSHCGWNSTMEALTSGV 371
Query: 523 PTTIVPFFGDQ 533
P P +GDQ
Sbjct: 372 PVICFPQWGDQ 382
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,633,505
Number of Sequences: 26719
Number of extensions: 726579
Number of successful extensions: 1790
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 1762
Number of HSP's gapped (non-prelim): 38
length of query: 638
length of database: 11,318,596
effective HSP length: 106
effective length of query: 532
effective length of database: 8,486,382
effective search space: 4514755224
effective search space used: 4514755224
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0052a.5


