

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0052a.2
(68 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
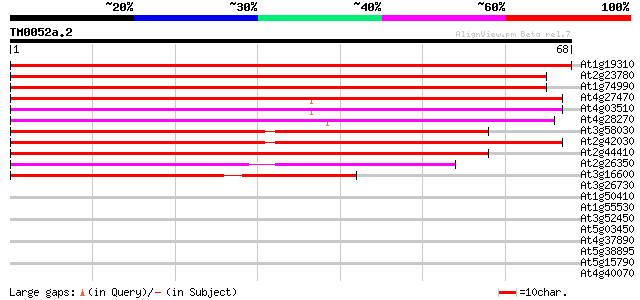
At1g19310 unknown protein 114 9e-27
At2g23780 putative RING zinc finger protein 109 2e-25
At1g74990 RING zinc finger like protein 104 9e-24
At4g27470 putative RING zinc finger protein 84 1e-17
At4g03510 RMA1 RING zinc finger protein 80 1e-16
At4g28270 unknown protein 77 2e-15
At3g58030 unknown protein 75 5e-15
At2g42030 putative RING zinc finger protein 68 7e-13
At2g44410 unknown protein 63 3e-11
At2g26350 zinc-binding peroxisomal integral membrane protein (PE... 45 9e-06
At3g16600 putative DNA-binding protein 39 5e-04
At3g26730 RING zinc finger like protein 37 0.001
At1g50410 unknown protein 37 0.001
At1g55530 unknown protein 37 0.002
At3g52450 unknown protein 37 0.002
At5g03450 unknown protein 36 0.003
At4g37890 unknown protein 36 0.003
At5g38895 unknown protein 36 0.004
At5g15790 unknown protein 35 0.005
At4g40070 unknown protein 35 0.005
>At1g19310 unknown protein
Length = 226
Score = 114 bits (285), Expect = 9e-27
Identities = 47/68 (69%), Positives = 56/68 (82%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
CNIC DLAQDP++TLCG+LF WPCLYKWLH HS S+ CPVC A+ EED+LVPL+GRG +S
Sbjct: 23 CNICLDLAQDPIVTLCGHLFCWPCLYKWLHLHSQSKDCPVCKAVIEEDRLVPLYGRGKSS 82
Query: 61 SSSSSSSI 68
+ S SI
Sbjct: 83 ADPRSKSI 90
>At2g23780 putative RING zinc finger protein
Length = 227
Score = 109 bits (273), Expect = 2e-25
Identities = 43/65 (66%), Positives = 54/65 (82%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
CNICF+LAQDP++TLCG+LF WPCLY+WLH HS SQ+CPVC A+ ++DKLVPL+GRG
Sbjct: 28 CNICFELAQDPIVTLCGHLFCWPCLYRWLHHHSHSQECPVCKAVVQDDKLVPLYGRGKNQ 87
Query: 61 SSSSS 65
+ S
Sbjct: 88 TDPRS 92
>At1g74990 RING zinc finger like protein
Length = 137
Score = 104 bits (259), Expect = 9e-24
Identities = 42/65 (64%), Positives = 51/65 (77%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
CNIC +LA++P++TLCG+LF WPCLYKWLH+HS S CPVC AL +ED LVPL+G G S
Sbjct: 19 CNICLELAREPIVTLCGHLFCWPCLYKWLHYHSKSNHCPVCKALVKEDTLVPLYGMGKPS 78
Query: 61 SSSSS 65
S S
Sbjct: 79 SDPRS 83
>At4g27470 putative RING zinc finger protein
Length = 243
Score = 84.3 bits (207), Expect = 1e-17
Identities = 39/74 (52%), Positives = 46/74 (61%), Gaps = 7/74 (9%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLS-------QQCPVCNALAEEDKLVPL 53
CNIC D A DPV+TLCG+LF WPC+YKWLH S CPVC + LVPL
Sbjct: 44 CNICLDTAHDPVVTLCGHLFCWPCIYKWLHVQLSSVSVDQHQNNCPVCKSNITITSLVPL 103
Query: 54 HGRGNTSSSSSSSS 67
+GRG +S SS+ S
Sbjct: 104 YGRGMSSPSSTFGS 117
>At4g03510 RMA1 RING zinc finger protein
Length = 249
Score = 80.5 bits (197), Expect = 1e-16
Identities = 34/75 (45%), Positives = 45/75 (59%), Gaps = 8/75 (10%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLS--------QQCPVCNALAEEDKLVP 52
CNIC D Q+PV+TLCG+LF WPC++KWL S S +QCPVC + LVP
Sbjct: 48 CNICLDSVQEPVVTLCGHLFCWPCIHKWLDVQSFSTSDEYQRHRQCPVCKSKVSHSTLVP 107
Query: 53 LHGRGNTSSSSSSSS 67
L+GRG ++ +
Sbjct: 108 LYGRGRCTTQEEGKN 122
>At4g28270 unknown protein
Length = 193
Score = 76.6 bits (187), Expect = 2e-15
Identities = 34/79 (43%), Positives = 47/79 (59%), Gaps = 13/79 (16%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQ-------------CPVCNALAEE 47
CNIC D +DPV+TLCG+LF WPC++KW + + S+Q CPVC + E
Sbjct: 21 CNICLDQVRDPVVTLCGHLFCWPCIHKWTYASNNSRQRVDQYDHKREPPKCPVCKSDVSE 80
Query: 48 DKLVPLHGRGNTSSSSSSS 66
LVP++GRG + S S+
Sbjct: 81 ATLVPIYGRGQKAPQSGSN 99
>At3g58030 unknown protein
Length = 436
Score = 75.5 bits (184), Expect = 5e-15
Identities = 29/58 (50%), Positives = 42/58 (72%), Gaps = 1/58 (1%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGN 58
CNIC DL+++PV+T CG+L+ WPCLY+WL S +++CPVC + P++GRGN
Sbjct: 139 CNICLDLSKEPVLTCCGHLYCWPCLYQWLQI-SDAKECPVCKGEVTSKTVTPIYGRGN 195
>At2g42030 putative RING zinc finger protein
Length = 425
Score = 68.2 bits (165), Expect = 7e-13
Identities = 29/67 (43%), Positives = 41/67 (60%), Gaps = 1/67 (1%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGNTS 60
C IC DL++DPV+T CG+L+ W CLY+WL S +++CPVC + P++GRG
Sbjct: 141 CYICLDLSKDPVVTNCGHLYCWSCLYQWLQV-SEAKECPVCKGEVSVKTVTPIYGRGIQK 199
Query: 61 SSSSSSS 67
S S
Sbjct: 200 RESEEVS 206
>At2g44410 unknown protein
Length = 404
Score = 62.8 bits (151), Expect = 3e-11
Identities = 22/58 (37%), Positives = 39/58 (66%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHGRGN 58
CNIC + A+DP++T CG+LF W C Y+ + ++CPVC+ + +++P++G G+
Sbjct: 116 CNICLEKAEDPILTCCGHLFCWGCFYQLPLIYLNIKECPVCDGEVTDAEVIPIYGNGD 173
>At2g26350 zinc-binding peroxisomal integral membrane protein
(PEX10)
Length = 381
Score = 44.7 bits (104), Expect = 9e-06
Identities = 18/54 (33%), Positives = 27/54 (49%), Gaps = 3/54 (5%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLH 54
C +C Q P T CG++F W C+ +W + Q+CP+C LV L+
Sbjct: 327 CTLCLSTRQHPTATPCGHVFCWSCIMEWC---NEKQECPLCRTPNTHSSLVCLY 377
>At3g16600 putative DNA-binding protein
Length = 638
Score = 38.9 bits (89), Expect = 5e-04
Identities = 15/42 (35%), Positives = 27/42 (63%), Gaps = 2/42 (4%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCN 42
C++C D +DPV+TLCG++F + C+ ++ + + CP N
Sbjct: 395 CSVCSDPPKDPVVTLCGHVFCYECVS--VNINGDNNTCPALN 434
>At3g26730 RING zinc finger like protein
Length = 772
Score = 37.4 bits (85), Expect = 0.001
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 6/60 (10%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWL------HFHSLSQQCPVCNALAEEDKLVPLH 54
C IC + P IT CG++F +PC+ ++L H ++CP+C + +L ++
Sbjct: 245 CPICLEYPLCPQITSCGHIFCFPCILQYLLTGVDNHKVDCFKRCPLCFVMISPRELYTVY 304
>At1g50410 unknown protein
Length = 981
Score = 37.4 bits (85), Expect = 0.001
Identities = 15/48 (31%), Positives = 25/48 (51%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEED 48
C +C D +DPV+TLCG++F + C+ ++ + P C D
Sbjct: 679 CCVCHDPPEDPVVTLCGHIFCYQCVSDYITGDEDTCPAPRCREQLAHD 726
>At1g55530 unknown protein
Length = 351
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 8/72 (11%)
Query: 1 CNIC---FDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVC--NALAEEDKLVPLHG 55
C++C F++ + + C + F CL WL HS CPVC A+E K +
Sbjct: 223 CSVCLDDFEIGTEAKLMPCTHKFHSDCLLPWLELHS---SCPVCRYQLPADEAKTDSVTT 279
Query: 56 RGNTSSSSSSSS 67
+ + SSS+S+
Sbjct: 280 TSDNNGSSSASA 291
>At3g52450 unknown protein
Length = 435
Score = 36.6 bits (83), Expect = 0.002
Identities = 19/54 (35%), Positives = 24/54 (44%), Gaps = 1/54 (1%)
Query: 1 CNICFDLAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLH 54
C I D+ +DPVI G + + KWL F CPV + E L P H
Sbjct: 13 CPISLDIMKDPVIVSTGITYDRESIEKWL-FSGKKNSCPVTKQVITETDLTPNH 65
>At5g03450 unknown protein
Length = 630
Score = 36.2 bits (82), Expect = 0.003
Identities = 16/49 (32%), Positives = 26/49 (52%), Gaps = 5/49 (10%)
Query: 1 CNICFDL----AQDPVITL-CGYLFRWPCLYKWLHFHSLSQQCPVCNAL 44
C+IC ++ Q V L CG+L+ + C+ KW +CP+CN +
Sbjct: 120 CSICMEVWTSGGQHQVCCLPCGHLYGYSCINKWFQQRRSGGKCPLCNKI 168
>At4g37890 unknown protein
Length = 739
Score = 36.2 bits (82), Expect = 0.003
Identities = 22/71 (30%), Positives = 30/71 (41%), Gaps = 5/71 (7%)
Query: 1 CNICFDLAQ----DPVITL-CGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDKLVPLHG 55
C IC A+ + T C + F +PC+ +L CPVC A E L+PL
Sbjct: 169 CGICLQSAKAGRGTAIFTAECSHTFHFPCVASRAGDRNLLSDCPVCGASWRETSLLPLSL 228
Query: 56 RGNTSSSSSSS 66
+ S S S
Sbjct: 229 SSSLHESGSES 239
>At5g38895 unknown protein
Length = 221
Score = 35.8 bits (81), Expect = 0.004
Identities = 17/52 (32%), Positives = 25/52 (47%), Gaps = 6/52 (11%)
Query: 1 CNICFD---LAQDPVITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAEEDK 49
C C D L +IT C + F C+Y+W+ S+ CPVC + D+
Sbjct: 169 CPTCLDDYTLENPKIITKCSHHFHLSCIYEWM---ERSETCPVCGKVMAFDE 217
>At5g15790 unknown protein
Length = 232
Score = 35.4 bits (80), Expect = 0.005
Identities = 15/49 (30%), Positives = 28/49 (56%), Gaps = 6/49 (12%)
Query: 1 CNICFD--LAQDP-VITLCGYLFRWPCLYKWLHFHSLSQQCPVCNALAE 46
C C + ++++P ++T C + F C+Y+W+ S+ CPVC + E
Sbjct: 182 CPTCLEEYISENPKIVTKCSHHFHLSCIYEWM---ERSENCPVCGKVME 227
>At4g40070 unknown protein
Length = 322
Score = 35.4 bits (80), Expect = 0.005
Identities = 21/65 (32%), Positives = 31/65 (47%), Gaps = 15/65 (23%)
Query: 1 CNICFDLAQDP----VITLCGYLFRWPCLYKWLHFHSLSQQCPVC--------NALAEED 48
C IC + +D ++ +C +LF C+ WL+ H+ CPVC N +ED
Sbjct: 123 CAICLNELEDHETVRLLPICNHLFHIDCIDTWLYSHA---TCPVCRSNLTAKSNKPGDED 179
Query: 49 KLVPL 53
VPL
Sbjct: 180 DGVPL 184
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.137 0.473
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,697,541
Number of Sequences: 26719
Number of extensions: 56223
Number of successful extensions: 519
Number of sequences better than 10.0: 221
Number of HSP's better than 10.0 without gapping: 78
Number of HSP's successfully gapped in prelim test: 143
Number of HSP's that attempted gapping in prelim test: 418
Number of HSP's gapped (non-prelim): 237
length of query: 68
length of database: 11,318,596
effective HSP length: 44
effective length of query: 24
effective length of database: 10,142,960
effective search space: 243431040
effective search space used: 243431040
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0052a.2


