

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0051b.7
(189 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
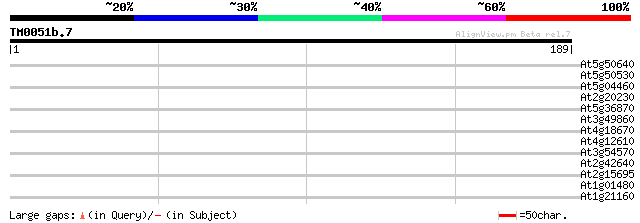
Score E
Sequences producing significant alignments: (bits) Value
At5g50640 putative protein 29 1.7
At5g50530 putative protein 29 1.7
At5g04460 putative protein 29 1.7
At2g20230 unknown protein 28 3.8
At5g36870 putative glucan synthase 27 4.9
At3g49860 putative protein 27 4.9
At4g18670 extensin-like protein 27 6.5
At4g12610 putative protein 27 6.5
At3g54570 putative protein 27 6.5
At2g42640 hypothetical protein 27 6.5
At2g15695 unknown protein 27 6.5
At1g01480 1-aminocyclopropane-1-carboxylate synthase (ACC2) 27 6.5
At1g21160 transcription factor, putative 27 8.4
>At5g50640 putative protein
Length = 548
Score = 28.9 bits (63), Expect = 1.7
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 9/70 (12%)
Query: 16 GGSGRFHRNRLRQNEDEQAISDQRSMEEEGKKLRRCSG-----GVARVVAAQRMAAQVMR 70
GGS R L + ++S +RS E K+LR C A +RMAA
Sbjct: 34 GGSDLLPRRSLTSSRSSISLSGERSGERTVKRLRLCKALTVPDSTTLFEACRRMAA---- 89
Query: 71 RRVVAVVRSD 80
RRV A++ +D
Sbjct: 90 RRVDALLLTD 99
>At5g50530 putative protein
Length = 548
Score = 28.9 bits (63), Expect = 1.7
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 9/70 (12%)
Query: 16 GGSGRFHRNRLRQNEDEQAISDQRSMEEEGKKLRRCSG-----GVARVVAAQRMAAQVMR 70
GGS R L + ++S +RS E K+LR C A +RMAA
Sbjct: 34 GGSDLLPRRSLTSSRSSISLSGERSGERTVKRLRLCKALTVPDSTTLFEACRRMAA---- 89
Query: 71 RRVVAVVRSD 80
RRV A++ +D
Sbjct: 90 RRVDALLLTD 99
>At5g04460 putative protein
Length = 831
Score = 28.9 bits (63), Expect = 1.7
Identities = 21/72 (29%), Positives = 30/72 (41%), Gaps = 12/72 (16%)
Query: 5 GLDRFRVGRVT-----GGSGRFHRNRLRQNEDEQAISDQRSMEEEGKKLRRCSGGVARVV 59
GL+ F G + G H NED+Q + +R E EG L S
Sbjct: 24 GLEEFMRGHLDECISFGSCSSVHNPEDEDNEDDQLVRRRRRSELEGDNLAESS------- 76
Query: 60 AAQRMAAQVMRR 71
AA+R +Q++ R
Sbjct: 77 AARRRQSQILSR 88
>At2g20230 unknown protein
Length = 270
Score = 27.7 bits (60), Expect = 3.8
Identities = 17/73 (23%), Positives = 34/73 (46%), Gaps = 8/73 (10%)
Query: 93 WSRFFPSLNPPSDPSEPSGPVFG-------RGPVGFVQEVMVKENVHINIFDIQGAPLFY 145
+SR P ++PP S SG + P+ FV +++ N + F+++ L
Sbjct: 39 YSRHLP-VDPPPSASSSSGTEIATSVSEPLKNPIDFVASIILGSNGGDHGFNLRSLDLPA 97
Query: 146 QWIVITSVVLGII 158
W + + + +GI+
Sbjct: 98 PWFIYSFMAVGIL 110
>At5g36870 putative glucan synthase
Length = 1662
Score = 27.3 bits (59), Expect = 4.9
Identities = 10/23 (43%), Positives = 16/23 (69%)
Query: 83 PNRNRPIQLIWSRFFPSLNPPSD 105
PNR + +Q ++S FFP +P S+
Sbjct: 4 PNRGQILQTVFSHFFPVASPDSE 26
>At3g49860 putative protein
Length = 165
Score = 27.3 bits (59), Expect = 4.9
Identities = 12/26 (46%), Positives = 16/26 (61%)
Query: 119 VGFVQEVMVKENVHINIFDIQGAPLF 144
VGF + KENV I ++D+ G P F
Sbjct: 33 VGFNMRKVTKENVAIRLWDLGGQPRF 58
>At4g18670 extensin-like protein
Length = 839
Score = 26.9 bits (58), Expect = 6.5
Identities = 15/38 (39%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Query: 82 APNRNRPIQLIWSRFFPSLNPPSDPSEPSGPVFGRGPV 119
+P +N P +I S F +PPS PS P PV P+
Sbjct: 572 SPGQNSP-PIIPSPPFTGPSPPSSPSPPLPPVIPSPPI 608
>At4g12610 putative protein
Length = 649
Score = 26.9 bits (58), Expect = 6.5
Identities = 14/46 (30%), Positives = 24/46 (51%)
Query: 12 GRVTGGSGRFHRNRLRQNEDEQAISDQRSMEEEGKKLRRCSGGVAR 57
G +GG GR + +E+E +SD+ +EE + R+ G+ R
Sbjct: 213 GGTSGGGGRGRKKSSGGDEEEGNVSDRGDEDEEEEASRKSRLGLNR 258
>At3g54570 putative protein
Length = 417
Score = 26.9 bits (58), Expect = 6.5
Identities = 18/47 (38%), Positives = 26/47 (55%), Gaps = 7/47 (14%)
Query: 11 VGRVTGGSGRFHR-----NRLRQNEDEQAISDQRSMEEEGKKLRRCS 52
V +VTGGS + + RQ++ QA D++S + GKKL CS
Sbjct: 44 VAKVTGGSPNYMKGTRSSEARRQSQSVQAGLDKKS--QTGKKLDSCS 88
>At2g42640 hypothetical protein
Length = 382
Score = 26.9 bits (58), Expect = 6.5
Identities = 12/35 (34%), Positives = 17/35 (48%)
Query: 89 IQLIWSRFFPSLNPPSDPSEPSGPVFGRGPVGFVQ 123
I + W F NP + P EP+ P+ GP +Q
Sbjct: 314 ISMAWQPDFVDNNPCASPLEPNSPMERSGPPSALQ 348
>At2g15695 unknown protein
Length = 408
Score = 26.9 bits (58), Expect = 6.5
Identities = 12/39 (30%), Positives = 19/39 (47%)
Query: 110 SGPVFGRGPVGFVQEVMVKENVHINIFDIQGAPLFYQWI 148
SG V+ GP+ F ++ VK +H I + G W+
Sbjct: 138 SGHVYDSGPLDFTSDLNVKFALHPTIRRMSGPSRLVSWV 176
>At1g01480 1-aminocyclopropane-1-carboxylate synthase (ACC2)
Length = 496
Score = 26.9 bits (58), Expect = 6.5
Identities = 15/78 (19%), Positives = 42/78 (53%), Gaps = 4/78 (5%)
Query: 104 SDPSEPSGPVFGRGPVGFVQEVMVKENVHINIFDIQGAPLFY--QWIVITSVVLGIIDYD 161
++PS P G + + + + + ++N+H+ + +I A +F ++ + VV + +
Sbjct: 206 TNPSNPLGTMLDKDTLTNLVRFVTRKNIHLVVDEIYAATVFAGGDFVSVAEVVNDVDISE 265
Query: 162 ISLEYISFYVVIMLNPDI 179
++++ I ++V L+ D+
Sbjct: 266 VNVDLI--HIVYSLSKDM 281
>At1g21160 transcription factor, putative
Length = 1088
Score = 26.6 bits (57), Expect = 8.4
Identities = 20/79 (25%), Positives = 36/79 (45%), Gaps = 9/79 (11%)
Query: 17 GSGRFHRNRLRQNEDEQAISDQRSMEEEG-------KKLRRCSGGVARVVAAQRMAA--Q 67
G + RLR+ E+E+ I ++R E E +K+ + G+ +R AA +
Sbjct: 238 GKKKEEEERLRKEEEERRIEEEREREAEEIRQKRKIRKMEKKQEGLILTAKQKRDAAKNE 297
Query: 68 VMRRRVVAVVRSDWAPNRN 86
R+RV+ S ++N
Sbjct: 298 AFRKRVLTDAGSLLVADKN 316
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.143 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,394,942
Number of Sequences: 26719
Number of extensions: 182685
Number of successful extensions: 704
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 697
Number of HSP's gapped (non-prelim): 15
length of query: 189
length of database: 11,318,596
effective HSP length: 94
effective length of query: 95
effective length of database: 8,807,010
effective search space: 836665950
effective search space used: 836665950
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0051b.7


