

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0050.15
(95 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
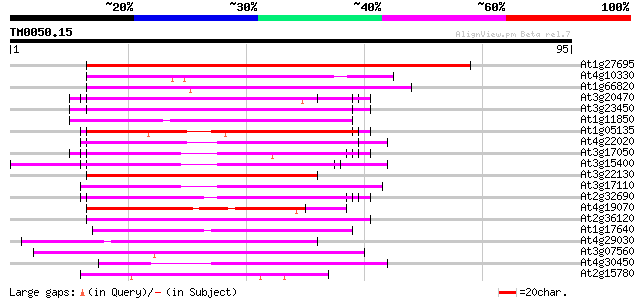
Sequences producing significant alignments: (bits) Value
At1g27695 unknown protein 121 5e-29
At4g10330 unknown protein (At4g10330) 53 2e-08
At1g66820 unknown protein 46 3e-06
At3g20470 glycine-rich protein, putative 45 6e-06
At3g23450 hypothetical protein 45 8e-06
At1g11850 unknown protein 45 8e-06
At1g05135 unknown protein 43 2e-05
At4g22020 glycine-rich protein 43 3e-05
At3g17050 unknown protein 43 3e-05
At3g15400 anther development protein, ATA20 43 3e-05
At3g22130 hypothetical protein 42 7e-05
At3g17110 hypothetical protein 41 9e-05
At2g32690 unknown protein 41 1e-04
At4g19070 cadmium-induced protein 40 2e-04
At2g36120 unknown protein 40 3e-04
At1g17640 RNA-binding protein, putative 39 3e-04
At4g29030 glycine-rich protein like 39 4e-04
At3g07560 unknown protein 39 4e-04
At4g30450 glycine-rich protein 39 6e-04
At2g15780 unknown protein 39 6e-04
>At1g27695 unknown protein
Length = 91
Score = 121 bits (304), Expect = 5e-29
Identities = 49/65 (75%), Positives = 60/65 (91%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEF 73
GFGVGCGFGVGWGFGGMP+N+LG+G GGGCGVG+GLGWGFG+AFGS YRSS++ FQG+E
Sbjct: 14 GFGVGCGFGVGWGFGGMPMNILGVGAGGGCGVGLGLGWGFGTAFGSHYRSSRLTFQGIEL 73
Query: 74 DSKEK 78
++ +K
Sbjct: 74 ETADK 78
>At4g10330 unknown protein (At4g10330)
Length = 130
Score = 53.1 bits (126), Expect = 2e-08
Identities = 30/59 (50%), Positives = 35/59 (58%), Gaps = 9/59 (15%)
Query: 14 GFGVGCGFGVGWG------FG-GMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQ 65
G GVGCGFG G G FG G+P GLG G GCG+GVG G+G G G+ Y S+
Sbjct: 34 GLGVGCGFGFGAGLIGGVGFGPGVPGLQFGLGFGAGCGIGVGFGYGVGR--GAAYDHSR 90
Score = 25.4 bits (54), Expect = 5.1
Identities = 10/17 (58%), Positives = 13/17 (75%)
Query: 39 VGGGCGVGVGLGWGFGS 55
VG G+GVG G+GFG+
Sbjct: 29 VGPAFGLGVGCGFGFGA 45
>At1g66820 unknown protein
Length = 109
Score = 46.2 bits (108), Expect = 3e-06
Identities = 24/64 (37%), Positives = 33/64 (51%), Gaps = 9/64 (14%)
Query: 14 GFGVGCGFGVGWGFGG---------MPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G G+GCG G+G G G + + + LG G GCG+G G G+GFG G +
Sbjct: 44 GGGIGCGAGIGIGLSGGLGIGASEGLDHSNVVLGFGIGCGIGFGFGYGFGVGGGYSFDDI 103
Query: 65 QIRF 68
+ RF
Sbjct: 104 KERF 107
Score = 29.3 bits (64), Expect = 0.35
Identities = 11/24 (45%), Positives = 14/24 (57%)
Query: 35 LGLGVGGGCGVGVGLGWGFGSAFG 58
+G G+GGG G G G+G G G
Sbjct: 39 VGPGIGGGIGCGAGIGIGLSGGLG 62
Score = 28.9 bits (63), Expect = 0.46
Identities = 11/24 (45%), Positives = 15/24 (61%)
Query: 36 GLGVGGGCGVGVGLGWGFGSAFGS 59
G+G G GCG G+G+G G G+
Sbjct: 42 GIGGGIGCGAGIGIGLSGGLGIGA 65
>At3g20470 glycine-rich protein, putative
Length = 174
Score = 45.1 bits (105), Expect = 6e-06
Identities = 25/48 (52%), Positives = 26/48 (54%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
GFG G G G G GFGG GLG GGG G G G G G GS G +
Sbjct: 112 GFGGGAGGGSGGGFGGGAGAGGGLGGGGGAGGGGGFGGGGGSGIGGGF 159
Score = 43.9 bits (102), Expect = 1e-05
Identities = 24/49 (48%), Positives = 27/49 (54%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
SG G+G G G G GFGG G G GGG G G GLG G G+ G +
Sbjct: 99 SGGGLGGGGGAGGGFGGGAGGGSGGGFGGGAGAGGGLGGGGGAGGGGGF 147
Score = 42.0 bits (97), Expect = 5e-05
Identities = 24/45 (53%), Positives = 25/45 (55%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G GFGG + GLG GGG G G G G G GS G
Sbjct: 80 GLGGGAGGGAGGGFGGGAGSGGGLGGGGGAGGGFGGGAGGGSGGG 124
Score = 41.6 bits (96), Expect = 7e-05
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G G G GFGG G G+GGG G G GLG G G G +
Sbjct: 52 GIGGGSGLGGGGGFGG------GGGLGGGAGGGGGLGGGAGGGAGGGF 93
Score = 40.4 bits (93), Expect = 2e-04
Identities = 22/45 (48%), Positives = 24/45 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G G G G G GG G G GGG G+G G G G G+ G
Sbjct: 124 GFGGGAGAGGGLGGGGGAGGGGGFGGGGGSGIGGGFGGGAGAGGG 168
Score = 40.0 bits (92), Expect = 2e-04
Identities = 25/49 (51%), Positives = 25/49 (51%), Gaps = 8/49 (16%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVG--LGWGFGSAFGS 59
SG G G GFG G G GG G G GGG G G G G GFG GS
Sbjct: 57 SGLGGGGGFGGGGGLGG------GAGGGGGLGGGAGGGAGGGFGGGAGS 99
Score = 39.3 bits (90), Expect = 3e-04
Identities = 23/48 (47%), Positives = 25/48 (51%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G+G G G G G GG G G GGG G G GLG G G+ G
Sbjct: 65 FGGGGGLGGGAGGGGGLGGGAGGGAGGGFGGGAGSGGGLGGGGGAGGG 112
Score = 38.9 bits (89), Expect = 4e-04
Identities = 22/45 (48%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G G G G G GG G G GGG G G G G G G G
Sbjct: 92 GFGGGAGSGGGLGGGGGAGGGFGGGAGGGSGGGFGGGAGAGGGLG 136
Score = 38.1 bits (87), Expect = 8e-04
Identities = 22/49 (44%), Positives = 26/49 (52%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
SG G G G G G G GG G G GGG G G+G G+G G+ G +
Sbjct: 121 SGGGFGGGAGAGGGLGGGGGAGGGGGFGGGGGSGIGGGFGGGAGAGGGF 169
Score = 38.1 bits (87), Expect = 8e-04
Identities = 20/39 (51%), Positives = 22/39 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
G G G G G G GFGG + +G G GGG G G G G G
Sbjct: 134 GLGGGGGAGGGGGFGGGGGSGIGGGFGGGAGAGGGFGGG 172
Score = 38.1 bits (87), Expect = 8e-04
Identities = 22/46 (47%), Positives = 23/46 (49%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
GFG G G G G G GG G G GGG G G G G G G G+
Sbjct: 64 GFGGGGGLGGGAGGGGGLGGGAGGGAGGGFGGGAGSGGGLGGGGGA 109
Score = 35.0 bits (79), Expect = 0.006
Identities = 22/49 (44%), Positives = 24/49 (48%), Gaps = 12/49 (24%)
Query: 14 GFGVGCGFGV--------GWGFGGMPLNLLGLGVGGGCGVGVGLGWGFG 54
G G G G G+ G GFGG G G+GGG G G G G GFG
Sbjct: 126 GGGAGAGGGLGGGGGAGGGGGFGGGG----GSGIGGGFGGGAGAGGGFG 170
Score = 30.4 bits (67), Expect = 0.16
Identities = 16/35 (45%), Positives = 19/35 (53%), Gaps = 6/35 (17%)
Query: 24 GWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G GG G G+GGG G+G G G+G G G
Sbjct: 44 GGGLGG------GGGIGGGSGLGGGGGFGGGGGLG 72
>At3g23450 hypothetical protein
Length = 452
Score = 44.7 bits (104), Expect = 8e-06
Identities = 25/45 (55%), Positives = 27/45 (59%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G GFG G GFGG +G GVGGG G GVG G+G G G
Sbjct: 87 GGGGGGGFGGGGGFGGGHGGGVGGGVGGGHGGGVGGGFGKGGGIG 131
Score = 43.9 bits (102), Expect = 1e-05
Identities = 22/45 (48%), Positives = 25/45 (54%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G G G GG G+G GGG G G+G G G G FG
Sbjct: 177 GGGIGGGIGKGGGIGG------GIGKGGGIGGGIGKGGGVGGGFG 215
Score = 43.9 bits (102), Expect = 1e-05
Identities = 24/48 (50%), Positives = 26/48 (54%), Gaps = 4/48 (8%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G+G G G G GFGG G G GGG G G+G G GFG G
Sbjct: 368 FGKGGGIGGGIGGGGGFGGGG----GFGKGGGIGGGIGKGGGFGGGGG 411
Score = 43.1 bits (100), Expect = 2e-05
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G GVG G G G G GG G+G GGG G G+G G G G G
Sbjct: 234 FGKGGGVGGGIGKGGGIGG------GIGKGGGIGGGIGKGGGIGGGIG 275
Score = 43.1 bits (100), Expect = 2e-05
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 4/48 (8%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G+G G G G G GG G G GGG G G+G G G G FG
Sbjct: 326 FGKGGGIGGGIGKGGGIGGGG----GFGKGGGIGGGIGKGGGIGGGFG 369
Score = 42.0 bits (97), Expect = 5e-05
Identities = 23/48 (47%), Positives = 24/48 (49%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G GVG G G G G GG G G GGG G G+G G G G G
Sbjct: 214 FGKGGGVGGGIGKGGGVGG------GFGKGGGVGGGIGKGGGIGGGIG 255
Score = 40.8 bits (94), Expect = 1e-04
Identities = 24/45 (53%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G G GVG G GG G GVGGG G G G+G G G G
Sbjct: 99 GFGGGHGGGVGGGVGGGH----GGGVGGGFGKGGGIGGGIGKGGG 139
Score = 40.8 bits (94), Expect = 1e-04
Identities = 23/48 (47%), Positives = 26/48 (53%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G+G G G G GFGG G G G G G+G G G+G G FG
Sbjct: 390 FGKGGGIGGGIGKGGGFGG------GGGFGKGGGIGGGGGFGKGGGFG 431
Score = 40.4 bits (93), Expect = 2e-04
Identities = 23/48 (47%), Positives = 25/48 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
GFG G GFG G G GG G G GGG G G G+G G G G +
Sbjct: 383 GFGGGGGFGKGGGIGGGIGKGGGFGGGGGFGKGGGIGGGGGFGKGGGF 430
Score = 40.0 bits (92), Expect = 2e-04
Identities = 24/48 (50%), Positives = 24/48 (50%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G G G G G GFGG G G GGG G G G G GFG G
Sbjct: 52 FGGGAGGGVGGGAGGGFGGGAGGGFGGGGGGGGGGGGGGGGGFGGGGG 99
Score = 39.7 bits (91), Expect = 3e-04
Identities = 23/48 (47%), Positives = 24/48 (49%), Gaps = 2/48 (4%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G G G GVG G GG G G GGG G G+G G G G G
Sbjct: 100 FGGGHGGGVGGGVGGGHGGGVGG--GFGKGGGIGGGIGKGGGVGGGIG 145
Score = 39.3 bits (90), Expect = 3e-04
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G G G G G GG G+G GGG G G+G G G G G
Sbjct: 123 GFGKGGGIGGGIGKGGGVGG--GIGKGGGIGGGIGKGGGVGGGIG 165
Score = 39.3 bits (90), Expect = 3e-04
Identities = 21/45 (46%), Positives = 24/45 (52%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G GFG G G GG G+GGG G G G G+G G G
Sbjct: 361 GGGIGGGFGKGGGIGG--------GIGGGGGFGGGGGFGKGGGIG 397
Score = 39.3 bits (90), Expect = 3e-04
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G+G GG GVGGG G G G+G GFG G
Sbjct: 203 GIGKGGGVGGGFGKGG--------GVGGGIGKGGGVGGGFGKGGG 239
Score = 38.9 bits (89), Expect = 4e-04
Identities = 23/45 (51%), Positives = 24/45 (53%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G G G G G GG G G GGG G G+G G GFG G
Sbjct: 347 GFGKGGGIGGGIGKGGGIGG--GFGKGGGIGGGIGGGGGFGGGGG 389
Score = 38.5 bits (88), Expect = 6e-04
Identities = 21/45 (46%), Positives = 24/45 (52%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G+G GG G+G GGG G G+G G G G G
Sbjct: 223 GIGKGGGVGGGFGKGGGVGG--GIGKGGGIGGGIGKGGGIGGGIG 265
Score = 38.5 bits (88), Expect = 6e-04
Identities = 22/45 (48%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G GGG G G+G G G G FG
Sbjct: 193 GIGKGGGIGGGIGKGGGVGG--GFGKGGGVGGGIGKGGGVGGGFG 235
Score = 38.1 bits (87), Expect = 8e-04
Identities = 22/45 (48%), Positives = 26/45 (56%), Gaps = 4/45 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G+G G GG G G+GGG G G G+G GFG G
Sbjct: 333 GGGIGKGGGIGGG-GGFG---KGGGIGGGIGKGGGIGGGFGKGGG 373
Score = 37.7 bits (86), Expect = 0.001
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+GGG G G G+G GFG G
Sbjct: 183 GIGKGGGIGGGIGKGG--------GIGGGIGKGGGVGGGFGKGGG 219
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/48 (45%), Positives = 23/48 (47%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
GFG G G G G G GG G G GGG G G G G G G G +
Sbjct: 47 GFGGGFGGGAGGGVGGGAGGGFGGGAGGGFGGGGGGGGGGGGGGGGGF 94
Score = 37.7 bits (86), Expect = 0.001
Identities = 23/48 (47%), Positives = 23/48 (47%), Gaps = 8/48 (16%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F GFG G G GVG G GG G GGG G G G G G G G
Sbjct: 48 FGGGFGGGAGGGVGGGAGG--------GFGGGAGGGFGGGGGGGGGGG 87
Score = 37.4 bits (85), Expect = 0.001
Identities = 21/43 (48%), Positives = 23/43 (52%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G GFGG +G G GGG G G G G+G G G
Sbjct: 41 GGGKGGGFGGGFGGGAGGGVGGGAGGGFGGGAGGGFGGGGGGG 83
Score = 37.4 bits (85), Expect = 0.001
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G GGG G G+G G G G G
Sbjct: 143 GIGKGGGIGGGIGKGGGVGG--GIGKGGGIGGGIGKGGGIGGGIG 185
Score = 37.4 bits (85), Expect = 0.001
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G GVG GFG G G GG G+G G GVG G+G G G G
Sbjct: 117 GGGVGGGFGKGGGIGG--------GIGKGGGVGGGIGKGGGIGGG 153
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G GGG G G+G G G G G
Sbjct: 163 GIGKGGGIGGGIGKGGGIGG--GIGKGGGIGGGIGKGGGIGGGIG 205
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G GGG G G+G G G G G
Sbjct: 263 GIGKGGGIGGGIGKGGGIGG--GIGKGGGIGGGIGKGGGIGGGIG 305
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G GGG G G+G G G G G
Sbjct: 253 GIGKGGGIGGGIGKGGGIGG--GIGKGGGIGGGIGKGGGIGGGIG 295
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G GGG G G+G G G G G
Sbjct: 273 GIGKGGGIGGGIGKGGGIGG--GIGKGGGIGGGIGKGGGIGGGIG 315
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G GGG G G+G G G G G
Sbjct: 243 GIGKGGGIGGGIGKGGGIGG--GIGKGGGIGGGIGKGGGIGGGIG 285
Score = 36.6 bits (83), Expect = 0.002
Identities = 22/47 (46%), Positives = 23/47 (48%), Gaps = 8/47 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGL--GWGFGSAFG 58
G G G GFG G G GG G+G GGG G G G G G G G
Sbjct: 319 GIGGGGGFGKGGGIGG------GIGKGGGIGGGGGFGKGGGIGGGIG 359
Score = 36.6 bits (83), Expect = 0.002
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G GGG G G+G G G G G
Sbjct: 153 GIGKGGGVGGGIGKGGGIGG--GIGKGGGIGGGIGKGGGIGGGIG 195
Score = 36.2 bits (82), Expect = 0.003
Identities = 22/45 (48%), Positives = 23/45 (50%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G GVG GFG G G GG G+G G GVG G G G G G
Sbjct: 207 GGGVGGGFGKGGGVGG--------GIGKGGGVGGGFGKGGGVGGG 243
Score = 35.8 bits (81), Expect = 0.004
Identities = 20/45 (44%), Positives = 22/45 (48%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+GGG G G G+G G G G
Sbjct: 283 GIGKGGGIGGGIGKGG--------GIGGGIGKGGGIGGGIGKGGG 319
Score = 35.8 bits (81), Expect = 0.004
Identities = 23/45 (51%), Positives = 23/45 (51%), Gaps = 5/45 (11%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G G GG G G GGG G G G G G G G
Sbjct: 405 GFGGGGGFGKGGGIGGGG----GFGKGGGFG-GGGFGGGGGGGGG 444
Score = 35.8 bits (81), Expect = 0.004
Identities = 19/45 (42%), Positives = 25/45 (55%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G+G G G +G G+G G G+G G G+G G G
Sbjct: 311 GGGIGKGGGIGGGGGFGKGGGIGGGIGKGGGIGGGGGFGKGGGIG 355
Score = 35.4 bits (80), Expect = 0.005
Identities = 19/45 (42%), Positives = 23/45 (50%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G G G GG G+G G G+G G G+G G G
Sbjct: 297 GGGIGGGIGKGGGIGG--------GIGKGGGIGGGGGFGKGGGIG 333
Score = 35.4 bits (80), Expect = 0.005
Identities = 22/48 (45%), Positives = 24/48 (49%), Gaps = 4/48 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G G G GFGG G G G G GVG G+G G G G +
Sbjct: 81 GGGGGGGGGGGGGFGGGG----GFGGGHGGGVGGGVGGGHGGGVGGGF 124
Score = 35.0 bits (79), Expect = 0.006
Identities = 21/41 (51%), Positives = 21/41 (51%), Gaps = 7/41 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFG 54
G G G GFG G GFGG G GGG G G G G G G
Sbjct: 417 GIGGGGGFGKGGGFGGG-------GFGGGGGGGGGGGGGIG 450
Score = 34.7 bits (78), Expect = 0.008
Identities = 20/48 (41%), Positives = 24/48 (49%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G+G G G G G GG G G G G G+G G+G G G G +
Sbjct: 307 GGGIGGGIGKGGGIGG------GGGFGKGGGIGGGIGKGGGIGGGGGF 348
Score = 34.7 bits (78), Expect = 0.008
Identities = 21/48 (43%), Positives = 24/48 (49%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G+G G G G G GG G+G G G G GVG G G G G +
Sbjct: 187 GGGIGGGIGKGGGIGGGIGKGGGVGGGFGKGGGVGGGIGKGGGVGGGF 234
Score = 34.7 bits (78), Expect = 0.008
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G+G G G G G+G G GFG G
Sbjct: 293 GIGKGGGIGGGIGKGG------GIGGGIGKGGGIGGGGGFGKGGG 331
Score = 34.7 bits (78), Expect = 0.008
Identities = 19/48 (39%), Positives = 24/48 (49%), Gaps = 8/48 (16%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G+G G G G G GG G+G G G+G G+G G G G +
Sbjct: 287 GGGIGGGIGKGGGIGG--------GIGKGGGIGGGIGKGGGIGGGGGF 326
Score = 32.7 bits (73), Expect = 0.032
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 4/45 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G GVG G G G G GG +G G G G G+G G G G G G
Sbjct: 137 GGGVGGGIGKGGGIGGG----IGKGGGVGGGIGKGGGIGGGIGKG 177
Score = 32.0 bits (71), Expect = 0.054
Identities = 22/49 (44%), Positives = 23/49 (46%), Gaps = 4/49 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVG----LGWGFGSAFG 58
G G G GFG G G GG G G GGG G G G +G G G G
Sbjct: 69 GGGAGGGFGGGGGGGGGGGGGGGGGFGGGGGFGGGHGGGVGGGVGGGHG 117
Score = 32.0 bits (71), Expect = 0.054
Identities = 22/45 (48%), Positives = 25/45 (54%), Gaps = 5/45 (11%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G G G GG G G+GGG G G G G+G G FG
Sbjct: 397 GGGIGKGGGFGGG-GGFG---KGGGIGGGGGFGKGGGFG-GGGFG 436
Score = 24.6 bits (52), Expect = 8.7
Identities = 14/31 (45%), Positives = 16/31 (51%), Gaps = 6/31 (19%)
Query: 34 LLGLGVGGGC------GVGVGLGWGFGSAFG 58
L+G G GGG G G G+G G G FG
Sbjct: 39 LVGGGKGGGFGGGFGGGAGGGVGGGAGGGFG 69
>At1g11850 unknown protein
Length = 108
Score = 44.7 bits (104), Expect = 8e-06
Identities = 25/48 (52%), Positives = 28/48 (58%), Gaps = 1/48 (2%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
FL G G G G G+G G GG L LG+G G G G G+GLG G G G
Sbjct: 39 FLGGGGSGDGLGLGLG-GGAGLGGLGIGAGIGAGAGLGLGGGGGGLGG 85
Score = 36.6 bits (83), Expect = 0.002
Identities = 20/41 (48%), Positives = 23/41 (55%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFG 54
G G+G G G+G G GG L G G+ GG G G G G G G
Sbjct: 65 GAGIGAGAGLGLGGGGGGLGGGGGGLLGGGGFGGGAGGGLG 105
Score = 35.4 bits (80), Expect = 0.005
Identities = 26/57 (45%), Positives = 29/57 (50%), Gaps = 11/57 (19%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNL-------LGLGVGGG--CGVGVGL--GWGFGSAFG 58
SG G+G G G G G GG+ + LGLG GGG G G GL G GFG G
Sbjct: 45 SGDGLGLGLGGGAGLGGLGIGAGIGAGAGLGLGGGGGGLGGGGGGLLGGGGFGGGAG 101
Score = 35.0 bits (79), Expect = 0.006
Identities = 19/40 (47%), Positives = 22/40 (54%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
+G G G G G+G G GG+ GL GGG G G G G G
Sbjct: 66 AGIGAGAGLGLGGGGGGLGGGGGGLLGGGGFGGGAGGGLG 105
>At1g05135 unknown protein
Length = 384
Score = 43.1 bits (100), Expect = 2e-05
Identities = 25/46 (54%), Positives = 29/46 (62%), Gaps = 4/46 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G+G G GVG G GG +G+G+G G GVGVG G G GS GS
Sbjct: 334 GGGLGHGGGVGGGHGGG----VGIGIGIGIGVGVGGGSGQGSGSGS 375
Score = 42.0 bits (97), Expect = 5e-05
Identities = 25/46 (54%), Positives = 27/46 (58%), Gaps = 3/46 (6%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G GVG G GG+ G G GGG G GVG G G GS FG+
Sbjct: 153 GGGFGAGGGVGGGAGGIGG---GGGAGGGGGGGVGGGSGHGSGFGA 195
Score = 40.4 bits (93), Expect = 2e-04
Identities = 24/48 (50%), Positives = 26/48 (54%), Gaps = 3/48 (6%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G GFG G G GG G+G GGG G G G G G GS GS +
Sbjct: 149 GSGHGGGFGAGGGVGG---GAGGIGGGGGAGGGGGGGVGGGSGHGSGF 193
Score = 40.0 bits (92), Expect = 2e-04
Identities = 26/49 (53%), Positives = 27/49 (55%), Gaps = 8/49 (16%)
Query: 13 SGFGVGCGFG--VGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SGFG G G G VG G GG G G GGG G GVG G G G FG+
Sbjct: 269 SGFGAGGGVGGGVGGGAGG------GGGGGGGGGGGVGGGSGHGGGFGA 311
Score = 40.0 bits (92), Expect = 2e-04
Identities = 24/48 (50%), Positives = 26/48 (54%), Gaps = 2/48 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLL--GLGVGGGCGVGVGLGWGFGSAFGS 59
G G G GFG G G GG L G G GGG G G+G G G G FG+
Sbjct: 73 GGGQGGGFGAGGGVGGGAGGGLGGGGGAGGGGGGGIGGGSGHGGGFGA 120
Score = 39.7 bits (91), Expect = 3e-04
Identities = 24/48 (50%), Positives = 25/48 (52%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G GVG G GG GLG GGG G G G G G GS G +
Sbjct: 77 GGGFGAGGGVGGGAGG------GLGGGGGAGGGGGGGIGGGSGHGGGF 118
Score = 39.7 bits (91), Expect = 3e-04
Identities = 24/48 (50%), Positives = 26/48 (54%), Gaps = 2/48 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLL--GLGVGGGCGVGVGLGWGFGSAFGS 59
G G G GFG G G GG + G G GGG G GVG G G G FG+
Sbjct: 111 GSGHGGGFGAGGGVGGGAGGGIGGGGGAGGGGGGGVGGGSGHGGGFGA 158
Score = 38.9 bits (89), Expect = 4e-04
Identities = 23/48 (47%), Positives = 25/48 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G GFG G G GG +G G GGG G G G G GS GS +
Sbjct: 186 GSGHGSGFGAGGGIGGGAGGGVGGGGGGGGGGGGGGGANGGSGHGSGF 233
Score = 38.9 bits (89), Expect = 4e-04
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G GVG G GG G+G GGG G G G G G GS G +
Sbjct: 115 GGGFGAGGGVGGGAGG------GIGGGGGAGGGGGGGVGGGSGHGGGF 156
Score = 38.5 bits (88), Expect = 6e-04
Identities = 23/45 (51%), Positives = 23/45 (51%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G GFG G G GG GLG GGG G G G G G G G
Sbjct: 302 GSGHGGGFGAGGGLGGGAGG--GLGGGGGAGGGGGGGLGHGGGVG 344
Score = 38.1 bits (87), Expect = 8e-04
Identities = 21/45 (46%), Positives = 23/45 (50%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G+G GG G G+GGG G G G G G G G
Sbjct: 107 GIGGGSGHGGGFGAGGGVGGGAGGGIGGGGGAGGGGGGGVGGGSG 151
Score = 38.1 bits (87), Expect = 8e-04
Identities = 21/46 (45%), Positives = 27/46 (58%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G+G G G G G GG + G+G G G GVG+G+G G G G
Sbjct: 319 AGGGLGGGGGAGGGGGGGLGHGGGVGGGHGGGVGIGIGIGIGVGVG 364
Score = 37.7 bits (86), Expect = 0.001
Identities = 24/47 (51%), Positives = 25/47 (53%), Gaps = 2/47 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGL--GVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG LG GVGGG G GVG+G G G G
Sbjct: 316 GGGAGGGLGGGGGAGGGGGGGLGHGGGVGGGHGGGVGIGIGIGIGVG 362
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/45 (48%), Positives = 22/45 (48%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G GFG G G GG GVGGG G G G G G G G
Sbjct: 264 GSGHGSGFGAGGGVGG--------GVGGGAGGGGGGGGGGGGGVG 300
Score = 37.7 bits (86), Expect = 0.001
Identities = 25/49 (51%), Positives = 26/49 (53%), Gaps = 8/49 (16%)
Query: 13 SGFGVGCGF--GVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SGFG G G GVG G GG G G GGG G G G G GS FG+
Sbjct: 231 SGFGAGGGVGGGVGGGAGG------GGGGGGGGGGGANGGSGHGSGFGA 273
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/46 (47%), Positives = 24/46 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G+G G GG G G GGG G G G G GS FG+
Sbjct: 190 GSGFGAGGGIGGGAGGGVGGGGGGGGGGGGGGGANGGSGHGSGFGA 235
Score = 37.4 bits (85), Expect = 0.001
Identities = 23/48 (47%), Positives = 24/48 (49%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G GVG G GG G G GGG G G G G G GS G +
Sbjct: 268 GSGFGAGGGVGGGVGG------GAGGGGGGGGGGGGGVGGGSGHGGGF 309
Score = 37.4 bits (85), Expect = 0.001
Identities = 22/45 (48%), Positives = 23/45 (50%), Gaps = 4/45 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G+G GG LG G GGG G G G G G G G
Sbjct: 298 GVGGGSGHGGGFGAGGG----LGGGAGGGLGGGGGAGGGGGGGLG 338
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/46 (47%), Positives = 24/46 (51%), Gaps = 2/46 (4%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SG G+G G G G GFG +G G GGG G G G G G G G
Sbjct: 66 SGGGIGGGGGQGGGFGAG--GGVGGGAGGGLGGGGGAGGGGGGGIG 109
Score = 36.6 bits (83), Expect = 0.002
Identities = 24/49 (48%), Positives = 26/49 (52%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
F +G GVG G G G G GG G GVGGG G G G G G G G+
Sbjct: 271 FGAGGGVGGGVGGGAGGGGGGGGGGGGGVGGGSGHGGGFGAGGGLGGGA 319
Score = 36.6 bits (83), Expect = 0.002
Identities = 23/48 (47%), Positives = 24/48 (49%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G GVG G GG G G GGG G G G G GS GS +
Sbjct: 230 GSGFGAGGGVGGGVGG------GAGGGGGGGGGGGGGANGGSGHGSGF 271
Score = 36.6 bits (83), Expect = 0.002
Identities = 24/49 (48%), Positives = 26/49 (52%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
F +G GVG G G G G GG G GVGGG G G G G G G G+
Sbjct: 118 FGAGGGVGGGAGGGIGGGGGAGGGGGGGVGGGSGHGGGFGAGGGVGGGA 166
Score = 36.6 bits (83), Expect = 0.002
Identities = 23/49 (46%), Positives = 26/49 (52%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
F +G GVG G G G G GG G G+GGG G G G G G G G+
Sbjct: 80 FGAGGGVGGGAGGGLGGGGGAGGGGGGGIGGGSGHGGGFGAGGGVGGGA 128
Score = 36.2 bits (82), Expect = 0.003
Identities = 23/50 (46%), Positives = 26/50 (52%), Gaps = 6/50 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G G GFG G G GG G+G G G G G G G G G+ GS + S
Sbjct: 226 GSGHGSGFGAGGGVGG------GVGGGAGGGGGGGGGGGGGANGGSGHGS 269
Score = 35.8 bits (81), Expect = 0.004
Identities = 22/45 (48%), Positives = 23/45 (50%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G G G G G GG G G G G G G GLG G G+ G
Sbjct: 59 GFGGGRGSGGGIGGGGGQGGGFGAGGGVGGGAGGGLGGGGGAGGG 103
Score = 35.8 bits (81), Expect = 0.004
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG + G G GGG G G G G G G G
Sbjct: 95 GGGGGAGGGGGGGIGGGSGHGGGFGAGGGVGGGAGGGIGGGGGAG 139
Score = 35.8 bits (81), Expect = 0.004
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG + G G GGG G G G G G G G
Sbjct: 170 GGGGGAGGGGGGGVGGGSGHGSGFGAGGGIGGGAGGGVGGGGGGG 214
Score = 35.0 bits (79), Expect = 0.006
Identities = 21/46 (45%), Positives = 23/46 (49%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G G G G G GG + G G GGG G G G G G G G
Sbjct: 285 AGGGGGGGGGGGGGVGGGSGHGGGFGAGGGLGGGAGGGLGGGGGAG 330
Score = 34.7 bits (78), Expect = 0.008
Identities = 20/46 (43%), Positives = 23/46 (49%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G+G G G G G GG G+G G G G G G G G G G
Sbjct: 128 AGGGIGGGGGAGGGGGG------GVGGGSGHGGGFGAGGGVGGGAG 167
Score = 33.9 bits (76), Expect = 0.014
Identities = 21/46 (45%), Positives = 23/46 (49%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G G G G G G + G G GGG G GVG G G G G
Sbjct: 247 AGGGGGGGGGGGGGANGGSGHGSGFGAGGGVGGGVGGGAGGGGGGG 292
Score = 33.9 bits (76), Expect = 0.014
Identities = 18/39 (46%), Positives = 20/39 (51%), Gaps = 6/39 (15%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
G G+G G G+G G GG G G G G G G G G G
Sbjct: 350 GVGIGIGIGIGVGVGG------GSGQGSGSGSGSGGGGG 382
Score = 33.9 bits (76), Expect = 0.014
Identities = 19/43 (44%), Positives = 21/43 (48%), Gaps = 6/43 (13%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G+G G G G G GG G+G G G G G G G G G G
Sbjct: 168 GIGGGGGAGGGGGG------GVGGGSGHGSGFGAGGGIGGGAG 204
Score = 33.5 bits (75), Expect = 0.019
Identities = 21/46 (45%), Positives = 23/46 (49%), Gaps = 8/46 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G GFG G G GG G+GGG G G G G G G G+
Sbjct: 53 GRGGGGGFGGGRGSGG--------GIGGGGGQGGGFGAGGGVGGGA 90
Score = 33.5 bits (75), Expect = 0.019
Identities = 21/46 (45%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGV-GLGWGFGSAFG 58
G G G G G G G GG + G G GGG G G G+G G G+ G
Sbjct: 133 GGGGGAGGGGGGGVGGGSGHGGGFGAGGGVGGGAGGIGGGGGAGGG 178
Score = 33.5 bits (75), Expect = 0.019
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G G + G G GGG G GVG G G G G
Sbjct: 210 GGGGGGGGGGGGGANGGSGHGSGFGAGGGVGGGVGGGAGGGGGGG 254
Score = 32.0 bits (71), Expect = 0.054
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G G G G GVG G G G+ G
Sbjct: 244 GGGAGGGGGGGGGGGGGANGGSGHGSGFGAGGGVGGGVGGGAGGG 288
Score = 31.6 bits (70), Expect = 0.071
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G G G G GVG G G G+ G
Sbjct: 206 GVGGGGGGGGGGGGGGGANGGSGHGSGFGAGGGVGGGVGGGAGGG 250
Score = 31.6 bits (70), Expect = 0.071
Identities = 20/45 (44%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G G + G G G G GVG G+G G G G
Sbjct: 246 GAGGGGGGGGGGGGGANGGSGHGSGFGAGGGVGGGVGGGAGGGGG 290
Score = 31.2 bits (69), Expect = 0.092
Identities = 20/45 (44%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G G + G G G G GVG G+G G G G
Sbjct: 208 GGGGGGGGGGGGGGGANGGSGHGSGFGAGGGVGGGVGGGAGGGGG 252
Score = 25.4 bits (54), Expect = 5.1
Identities = 14/34 (41%), Positives = 14/34 (41%)
Query: 25 WGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
WG G G GGG G G G G G G G
Sbjct: 42 WGRRCAGRGRFGRGGGGGFGGGRGSGGGIGGGGG 75
>At4g22020 glycine-rich protein
Length = 396
Score = 42.7 bits (99), Expect = 3e-05
Identities = 24/52 (46%), Positives = 30/52 (57%), Gaps = 6/52 (11%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
SG G G G G G G GG G GVG G G+G G+G G GS G+ Y+++
Sbjct: 343 SGEGYGMGGGAGTGNGG------GGGVGFGMGIGFGIGIGGGSGGGTTYQTN 388
Score = 42.4 bits (98), Expect = 4e-05
Identities = 25/47 (53%), Positives = 27/47 (57%), Gaps = 5/47 (10%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SGFG G GFG G GG G G GGG G G G G+G GS +GS
Sbjct: 273 SGFGGGVGFGNSGGGGGG-----GGGGGGGGGGGNGSGYGSGSGYGS 314
Score = 40.8 bits (94), Expect = 1e-04
Identities = 24/46 (52%), Positives = 26/46 (56%), Gaps = 5/46 (10%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G+G G GFG G GFG G G GGG G G G G G GS +GS
Sbjct: 268 GYGHGSGFGGGVGFGNS-----GGGGGGGGGGGGGGGGGNGSGYGS 308
Score = 37.0 bits (84), Expect = 0.002
Identities = 20/45 (44%), Positives = 23/45 (50%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G+G GG G G G G G+G+G G G G G
Sbjct: 336 GNGSGSGSGEGYGMGGGAGTGNGGGGGVGFGMGIGFGIGIGGGSG 380
Score = 36.6 bits (83), Expect = 0.002
Identities = 21/46 (45%), Positives = 25/46 (53%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SG G G G G G G+G +G G G G G G G+G+G G FG
Sbjct: 333 SGGGNGSGSGSGEGYG------MGGGAGTGNGGGGGVGFGMGIGFG 372
Score = 35.4 bits (80), Expect = 0.005
Identities = 21/47 (44%), Positives = 23/47 (48%), Gaps = 5/47 (10%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG G G G G G G GG G G G G G G G G+G G G+
Sbjct: 314 SGMGKGSGSGGGGGGGGG-----GSGGGNGSGSGSGEGYGMGGGAGT 355
Score = 35.4 bits (80), Expect = 0.005
Identities = 23/47 (48%), Positives = 24/47 (50%), Gaps = 3/47 (6%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG G G G G G G GG + G G GGG G G G G GS GS
Sbjct: 156 SGHGSGSGAGAGAGVGG---SSGGAGGGGGGGGGEGGGANGGSGHGS 199
Score = 35.0 bits (79), Expect = 0.006
Identities = 24/50 (48%), Positives = 26/50 (52%), Gaps = 5/50 (10%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G G G G G G GG GVGGG G G G G G GS+ GS + S
Sbjct: 116 GSGHGSGSGAGAGVGGTTG-----GVGGGGGGGGGGGEGGGSSGGSGHGS 160
Score = 35.0 bits (79), Expect = 0.006
Identities = 23/50 (46%), Positives = 24/50 (48%), Gaps = 5/50 (10%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVG-----VGLGWGFGSAFG 58
G G G G G G GG G G GGG G G VG G+G GS FG
Sbjct: 227 GANGGSGHGSGSGAGGGVSGAAGGGGGGGGGGGSGGSKVGGGYGHGSGFG 276
Score = 35.0 bits (79), Expect = 0.006
Identities = 22/47 (46%), Positives = 25/47 (52%), Gaps = 3/47 (6%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG+G G G G G G GG G G GGG G G G G G+G G+
Sbjct: 310 SGYGSGMGKGSGSGGGG---GGGGGGSGGGNGSGSGSGEGYGMGGGA 353
Score = 35.0 bits (79), Expect = 0.006
Identities = 23/49 (46%), Positives = 25/49 (50%), Gaps = 1/49 (2%)
Query: 14 GFGVGCGFGVGWGFG-GMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G+G G G+G GM G GGG G G G G G GS G Y
Sbjct: 299 GGGNGSGYGSGSGYGSGMGKGSGSGGGGGGGGGGSGGGNGSGSGSGEGY 347
Score = 34.3 bits (77), Expect = 0.011
Identities = 20/45 (44%), Positives = 21/45 (46%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G G G G G G+G G A G
Sbjct: 135 GVGGGGGGGGGGGEGGGSSGGSGHGSGSGAGAGAGVGGSSGGAGG 179
Score = 33.9 bits (76), Expect = 0.014
Identities = 21/46 (45%), Positives = 22/46 (47%), Gaps = 5/46 (10%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G GVG GG +G G GGG G G G G G GS
Sbjct: 198 GSGAGAGAGVGGAAGG-----VGGGGGGGGGEGGGANGGSGHGSGS 238
Score = 33.5 bits (75), Expect = 0.019
Identities = 20/45 (44%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G G + G G+GGG G G G G G G G
Sbjct: 324 GGGGGGGGGSGGGNGSGSGSGEGYGMGGGAGTGNGGGGGVGFGMG 368
Score = 33.1 bits (74), Expect = 0.024
Identities = 22/47 (46%), Positives = 23/47 (48%), Gaps = 3/47 (6%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG G G G GVG GG G G GGG G G G G GS G+
Sbjct: 160 SGSGAGAGAGVGGSSGGAGG---GGGGGGGEGGGANGGSGHGSGAGA 203
Score = 32.3 bits (72), Expect = 0.042
Identities = 21/48 (43%), Positives = 24/48 (49%), Gaps = 1/48 (2%)
Query: 12 LSGFG-VGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+SG+G VG G G G G GG G G G G G G G+G G G
Sbjct: 91 MSGYGGVGGGGGRGGGEGGGSSANGGSGHGSGSGAGAGVGGTTGGVGG 138
Score = 31.2 bits (69), Expect = 0.092
Identities = 21/47 (44%), Positives = 22/47 (46%), Gaps = 5/47 (10%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG G G G G G G G+G GGG G G G G GS GS
Sbjct: 195 SGHGSGAGAGAGVGGAAG-----GVGGGGGGGGGEGGGANGGSGHGS 236
Score = 30.4 bits (67), Expect = 0.16
Identities = 21/48 (43%), Positives = 24/48 (49%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGF--GSAFGS 59
G+G G G+G G G G G+G GGG G G G G GS GS
Sbjct: 78 GYGHGEGYGAGAGMSGYG----GVGGGGGRGGGEGGGSSANGGSGHGS 121
Score = 30.4 bits (67), Expect = 0.16
Identities = 23/52 (44%), Positives = 24/52 (45%), Gaps = 7/52 (13%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGV-------GLGWGFGSAFGSK 60
GVG G G G G GG G G G G G GV G G G G + GSK
Sbjct: 213 GVGGGGGGGGGEGGGANGGSGHGSGSGAGGGVSGAAGGGGGGGGGGGSGGSK 264
Score = 30.4 bits (67), Expect = 0.16
Identities = 22/49 (44%), Positives = 25/49 (50%), Gaps = 3/49 (6%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGV---GGGCGVGVGLGWGFGSAFGS 59
G G G G G G+G G + +G G GGG G G G G G GS GS
Sbjct: 295 GGGGGGGNGSGYGSGSGYGSGMGKGSGSGGGGGGGGGGSGGGNGSGSGS 343
Score = 29.6 bits (65), Expect = 0.27
Identities = 16/37 (43%), Positives = 20/37 (53%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
G G G G G GG + +G G G G G G G+G+G
Sbjct: 246 GAAGGGGGGGGGGGSGGSKVGGGYGHGSGFGGGVGFG 282
Score = 29.3 bits (64), Expect = 0.35
Identities = 21/51 (41%), Positives = 22/51 (42%), Gaps = 5/51 (9%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGV-----GLGWGFGSAFG 58
S G G G G G G GG G G G G G GV G+G G G G
Sbjct: 173 SSGGAGGGGGGGGGEGGGANGGSGHGSGAGAGAGVGGAAGGVGGGGGGGGG 223
Score = 29.3 bits (64), Expect = 0.35
Identities = 21/46 (45%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G GVG GG+ G G GGG G G G G GS G+
Sbjct: 120 GSGSGAGAGVGGTTGGVGGGGGG-GGGGGEGGGSSGGSGHGSGSGA 164
Score = 28.9 bits (63), Expect = 0.46
Identities = 19/46 (41%), Positives = 20/46 (43%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G G + G G GGG G G G G GS
Sbjct: 215 GGGGGGGGGEGGGANGGSGHGSGSGAGGGVSGAAGGGGGGGGGGGS 260
Score = 28.9 bits (63), Expect = 0.46
Identities = 23/52 (44%), Positives = 25/52 (47%), Gaps = 7/52 (13%)
Query: 14 GFGVGCGFGV-----GWGFGGMPLNLLGLGVGGGCGVGVGL--GWGFGSAFG 58
G G G G GV G G GG G VGGG G G G G GFG++ G
Sbjct: 235 GSGSGAGGGVSGAAGGGGGGGGGGGSGGSKVGGGYGHGSGFGGGVGFGNSGG 286
Score = 28.1 bits (61), Expect = 0.78
Identities = 23/58 (39%), Positives = 25/58 (42%), Gaps = 4/58 (6%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGV 71
G G G G G G G G GVGGG G G G G G + GS + S GV
Sbjct: 76 GDGYGHGEGYGAGAGMSGYG----GVGGGGGRGGGEGGGSSANGGSGHGSGSGAGAGV 129
Score = 27.7 bits (60), Expect = 1.0
Identities = 18/45 (40%), Positives = 19/45 (42%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G G + G G G G GVG G G G
Sbjct: 139 GGGGGGGGGEGGGSSGGSGHGSGSGAGAGAGVGGSSGGAGGGGGG 183
Score = 26.6 bits (57), Expect = 2.3
Identities = 20/49 (40%), Positives = 22/49 (44%), Gaps = 3/49 (6%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCG---VGVGLGWGFGSAFG 58
+G G G G G G G G + G G G G G GVG G G G G
Sbjct: 177 AGGGGGGGGGEGGGANGGSGHGSGAGAGAGVGGAAGGVGGGGGGGGGEG 225
Score = 26.2 bits (56), Expect = 3.0
Identities = 21/53 (39%), Positives = 24/53 (44%), Gaps = 4/53 (7%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGV----GLGWGFGSAFGSKYRSS 64
G G G G G G GG + G G G G G G+ G+G G G G SS
Sbjct: 60 GGGGGGGGGGGGGGEGGDGYGHGEGYGAGAGMSGYGGVGGGGGRGGGEGGGSS 112
Score = 25.8 bits (55), Expect = 3.9
Identities = 19/45 (42%), Positives = 19/45 (42%), Gaps = 7/45 (15%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G G GVGG G G G G G G G
Sbjct: 151 GSSGGSGHGSGSGAGA------GAGVGGSSG-GAGGGGGGGGGEG 188
>At3g17050 unknown protein
Length = 349
Score = 42.7 bits (99), Expect = 3e-05
Identities = 24/51 (47%), Positives = 26/51 (50%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
F G G G G GVG GFGG G G GGG G G G G+G G G +
Sbjct: 295 FGGGAGGGHGGGVGGGFGGGSGGGFGGGAGGGAGGGAGGGFGGGGGAGGGF 345
Score = 42.7 bits (99), Expect = 3e-05
Identities = 23/48 (47%), Positives = 25/48 (51%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G G G G G GFGG G G+GGG G G G+G G G G
Sbjct: 43 FGGGKGFGGGIGAGGGFGGGAGGGAGGGLGGGAGGGGGIGGGAGGGAG 90
Score = 42.4 bits (98), Expect = 4e-05
Identities = 22/45 (48%), Positives = 24/45 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G G G GG LG G GGG G+G G G G G G
Sbjct: 50 GGGIGAGGGFGGGAGGGAGGGLGGGAGGGGGIGGGAGGGAGGGLG 94
Score = 42.0 bits (97), Expect = 5e-05
Identities = 22/46 (47%), Positives = 26/46 (55%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
GFG G G G+G G GG G G GGG G G G G+G G+ G+
Sbjct: 230 GFGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGGAGGGFGGGAGGGA 275
Score = 42.0 bits (97), Expect = 5e-05
Identities = 22/45 (48%), Positives = 24/45 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G+G G GG LG G+GGG G G G G G G G
Sbjct: 120 GLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGLGGGHG 164
Score = 42.0 bits (97), Expect = 5e-05
Identities = 23/44 (52%), Positives = 23/44 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAF 57
G G G G G G GFGG G G GGG G G G G GFG F
Sbjct: 306 GVGGGFGGGSGGGFGGGAGGGAGGGAGGGFGGGGGAGGGFGGGF 349
Score = 41.2 bits (95), Expect = 9e-05
Identities = 24/49 (48%), Positives = 27/49 (54%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
F G G G G G G GFGG G GVGGG G G G G+G G+ G+
Sbjct: 279 FGGGAGGGAGGGAGGGFGGGAGGGHGGGVGGGFGGGSGGGFGGGAGGGA 327
Score = 41.2 bits (95), Expect = 9e-05
Identities = 24/47 (51%), Positives = 24/47 (51%), Gaps = 2/47 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGC--GVGVGLGWGFGSAFG 58
GFG G G G G GFGG G G GGG G G G G G G FG
Sbjct: 266 GFGGGAGGGAGGGFGGGAGGGAGGGAGGGFGGGAGGGHGGGVGGGFG 312
Score = 40.4 bits (93), Expect = 2e-04
Identities = 22/45 (48%), Positives = 24/45 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G+G G GG LG G+GGG G G G G G G G
Sbjct: 178 GLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGAGGGGG 222
Score = 40.4 bits (93), Expect = 2e-04
Identities = 24/45 (53%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G G G G G GG G G GGG G GVG G+G GS G
Sbjct: 278 GFGGGAGGGAGGGAGGG----FGGGAGGGHGGGVGGGFGGGSGGG 318
Score = 40.4 bits (93), Expect = 2e-04
Identities = 22/45 (48%), Positives = 23/45 (50%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG +G G GGG G G GLG G G G
Sbjct: 124 GHGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGLGGGHGGGIG 168
Score = 40.4 bits (93), Expect = 2e-04
Identities = 23/46 (50%), Positives = 26/46 (56%), Gaps = 6/46 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G+G GFG G GFGG G+G GGG G G G G G G G+
Sbjct: 36 GGGLGGGFGGGKGFGG------GIGAGGGFGGGAGGGAGGGLGGGA 75
Score = 40.0 bits (92), Expect = 2e-04
Identities = 23/45 (51%), Positives = 24/45 (53%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G+G G GG G G GGG G G GLG G G FG
Sbjct: 190 GAGGGSGGGLGGGIGGGAGG--GAGGGGGAGGGGGLGGGHGGGFG 232
Score = 39.7 bits (91), Expect = 3e-04
Identities = 24/50 (48%), Positives = 24/50 (48%), Gaps = 2/50 (4%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGC--GVGVGLGWGFGSAFG 58
F G G G G G G GFGG G G GGG G G G G GFG G
Sbjct: 251 FGGGAGGGAGGGAGGGFGGGAGGGAGGGFGGGAGGGAGGGAGGGFGGGAG 300
Score = 39.3 bits (90), Expect = 3e-04
Identities = 23/47 (48%), Positives = 23/47 (48%), Gaps = 2/47 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGC--GVGVGLGWGFGSAFG 58
G G G G G G GFGG G G GGG G G G G GFG G
Sbjct: 238 GLGGGAGGGTGGGFGGGAGGGAGGGAGGGFGGGAGGGAGGGFGGGAG 284
Score = 39.3 bits (90), Expect = 3e-04
Identities = 23/47 (48%), Positives = 26/47 (54%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
+G G G G G G GFGG LG G GGG G G G G G G+ G+
Sbjct: 217 AGGGGGLGGGHGGGFGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGGA 263
Score = 38.9 bits (89), Expect = 4e-04
Identities = 21/46 (45%), Positives = 25/46 (53%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G GG +G G GGG G G+G G G G+ G+
Sbjct: 166 GIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGA 211
Score = 38.9 bits (89), Expect = 4e-04
Identities = 21/46 (45%), Positives = 25/46 (53%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G GG +G G GGG G G+G G G G+ G+
Sbjct: 108 GIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGA 153
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/46 (47%), Positives = 25/46 (53%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G+G G GG LG G GGG G G G G G G+ G+
Sbjct: 72 GGGAGGGGGIGGGAGGGAGGGLGGGAGGGLGGGHGGGIGGGAGGGA 117
Score = 37.7 bits (86), Expect = 0.001
Identities = 21/46 (45%), Positives = 24/46 (51%), Gaps = 8/46 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
GFG G G G G G GG G GGG G G G G+G G+ G+
Sbjct: 250 GFGGGAGGGAGGGAGG--------GFGGGAGGGAGGGFGGGAGGGA 287
Score = 37.4 bits (85), Expect = 0.001
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G GGG G+G G G G G G
Sbjct: 128 GIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGLGGGHGGGIGGGAG 172
Score = 37.4 bits (85), Expect = 0.001
Identities = 21/46 (45%), Positives = 24/46 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G GG +G G GGG G G+G G G G G+
Sbjct: 88 GAGGGLGGGAGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGA 133
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/45 (48%), Positives = 24/45 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G+GGG G G+G G G GS G
Sbjct: 96 GAGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGG 140
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/48 (45%), Positives = 22/48 (45%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F G G G G G G G GG G G G G G G G G GFG G
Sbjct: 231 FGGGAGGGLGGGAGGGTGG------GFGGGAGGGAGGGAGGGFGGGAG 272
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/46 (47%), Positives = 24/46 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G+G G GG G G GGG G G G G G G+ GS
Sbjct: 92 GLGGGAGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGS 137
Score = 37.0 bits (84), Expect = 0.002
Identities = 21/47 (44%), Positives = 26/47 (54%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
+G G+G G G G G GG G G+GGG G G+G G G G G+
Sbjct: 67 AGGGLGGGAGGGGGIGGGAGGGAGGGLGGGAGGGLGGGHGGGIGGGA 113
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/45 (48%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG LG G GGG G G G G G G G
Sbjct: 80 GIGGGAGGGAGGGLGGGAGGGLGGGHGGGIGGGAGGGAGGGLGGG 124
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/46 (47%), Positives = 25/46 (53%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G G G G G GG G G+GGG G G+G G G GS G
Sbjct: 153 AGGGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGG 198
Score = 36.6 bits (83), Expect = 0.002
Identities = 21/46 (45%), Positives = 24/46 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G GG +G G GGG G G+G G G G G+
Sbjct: 146 GGGAGGGAGGGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGA 191
Score = 36.2 bits (82), Expect = 0.003
Identities = 21/45 (46%), Positives = 24/45 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G+G G GG G G GGG G G+G G G G+ G
Sbjct: 112 GAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGG 156
Score = 36.2 bits (82), Expect = 0.003
Identities = 22/47 (46%), Positives = 25/47 (52%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
+G G G G G+G G GG G G GGG G G G G G G+ GS
Sbjct: 149 AGGGAGGGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGS 195
Score = 36.2 bits (82), Expect = 0.003
Identities = 22/45 (48%), Positives = 23/45 (50%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G+G G GG LG G GGG G G G G G G G
Sbjct: 158 GLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGG 202
Score = 36.2 bits (82), Expect = 0.003
Identities = 22/45 (48%), Positives = 23/45 (50%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G+G G GG LG G GGG G G G G G G G
Sbjct: 100 GLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGG 144
Score = 36.2 bits (82), Expect = 0.003
Identities = 21/45 (46%), Positives = 24/45 (52%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G+G G GG G G GGG G G+G G G G+ G
Sbjct: 170 GAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGG 214
Score = 35.8 bits (81), Expect = 0.004
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G+GGG G G G G G G G
Sbjct: 116 GAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGLG 160
Score = 35.4 bits (80), Expect = 0.005
Identities = 21/46 (45%), Positives = 24/46 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G GG +G G GGG G G G G G G+ G+
Sbjct: 286 GAGGGAGGGFGGGAGGGHGGGVGGGFGGGSGGGFGGGAGGGAGGGA 331
Score = 35.0 bits (79), Expect = 0.006
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G+GGG G G G G G G G
Sbjct: 174 GAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGAG 218
Score = 34.7 bits (78), Expect = 0.008
Identities = 22/47 (46%), Positives = 22/47 (46%), Gaps = 2/47 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGC--GVGVGLGWGFGSAFG 58
G G G G G G G GG G G GGG G G G G GFG G
Sbjct: 210 GAGGGGGAGGGGGLGGGHGGGFGGGAGGGLGGGAGGGTGGGFGGGAG 256
Score = 34.7 bits (78), Expect = 0.008
Identities = 21/46 (45%), Positives = 23/46 (49%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G G LG G GGG G G G G G G+ G+
Sbjct: 202 GIGGGAGGGAGGGGGAGGGGGLGGGHGGGFGGGAGGGLGGGAGGGT 247
Score = 34.3 bits (77), Expect = 0.011
Identities = 23/53 (43%), Positives = 24/53 (44%), Gaps = 8/53 (15%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLG------VGGGC--GVGVGLGWGFGSAFG 58
GFG G G G G G GG G+G GGG G G GLG G G G
Sbjct: 58 GFGGGAGGGAGGGLGGGAGGGGGIGGGAGGGAGGGLGGGAGGGLGGGHGGGIG 110
Score = 33.5 bits (75), Expect = 0.019
Identities = 22/46 (47%), Positives = 23/46 (49%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SG G+G G G G G G LG G GGG G G G G G G G
Sbjct: 137 SGGGLGGGIGGGAGGGAGGGGGLGGGHGGGIGGGAGGGAGGGLGGG 182
Score = 33.1 bits (74), Expect = 0.024
Identities = 19/43 (44%), Positives = 21/43 (48%), Gaps = 10/43 (23%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G+G GFGG G G G G+G G GFG G
Sbjct: 32 GGGGGGGLGGGFGG----------GKGFGGGIGAGGGFGGGAG 64
Score = 33.1 bits (74), Expect = 0.024
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 2/48 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGG--GCGVGVGLGWGFGSAFGS 59
GFG G GFG G G GG G G GG G G G G G G G+ G+
Sbjct: 42 GFGGGKGFGGGIGAGGGFGGGAGGGAGGGLGGGAGGGGGIGGGAGGGA 89
Score = 29.3 bits (64), Expect = 0.35
Identities = 14/25 (56%), Positives = 15/25 (60%)
Query: 35 LGLGVGGGCGVGVGLGWGFGSAFGS 59
LG G GGG G G G G GFG G+
Sbjct: 31 LGGGGGGGLGGGFGGGKGFGGGIGA 55
>At3g15400 anther development protein, ATA20
Length = 416
Score = 42.7 bits (99), Expect = 3e-05
Identities = 23/44 (52%), Positives = 26/44 (58%), Gaps = 6/44 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSA 56
SGFG G G G GFG G+G GGG G+G+G G G GSA
Sbjct: 254 SGFGEGIGSSGGSGFGE------GIGSGGGTGIGIGEGIGSGSA 291
Score = 39.3 bits (90), Expect = 3e-04
Identities = 23/51 (45%), Positives = 26/51 (50%), Gaps = 6/51 (11%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G+GVG G G GFG G+G GG G G G+G GS FG SS
Sbjct: 219 GYGVGIGSSGGSGFGE------GIGSSGGNGFGEGIGSSGGSGFGEGIGSS 263
Score = 38.5 bits (88), Expect = 6e-04
Identities = 21/55 (38%), Positives = 27/55 (48%)
Query: 1 KSLSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGS 55
+ + SG F G G G G G G G + G G+G G G G+G+G G GS
Sbjct: 234 EGIGSSGGNGFGEGIGSSGGSGFGEGIGSSGGSGFGEGIGSGGGTGIGIGEGIGS 288
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/54 (40%), Positives = 29/54 (52%), Gaps = 9/54 (16%)
Query: 14 GFGVGCGFGVGW---GFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G+GVG G+G G G+G +G+G GG G G G+G G+ FG SS
Sbjct: 204 GYGVGRGYGSGGSGVGYG------VGIGSSGGSGFGEGIGSSGGNGFGEGIGSS 251
Score = 37.0 bits (84), Expect = 0.002
Identities = 20/46 (43%), Positives = 24/46 (51%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+GFG G G G GFG G+G GG G G G+G G G+ G
Sbjct: 242 NGFGEGIGSSGGSGFGE------GIGSSGGSGFGEGIGSGGGTGIG 281
Score = 31.6 bits (70), Expect = 0.071
Identities = 19/46 (41%), Positives = 22/46 (47%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G+G G G+G G G G G G G G G+G G G GS
Sbjct: 245 GEGIGSSGGSGFGEGIGSSGGSGFGEGIGSGGGTGIGIGEGIGSGS 290
Score = 31.2 bits (69), Expect = 0.092
Identities = 21/52 (40%), Positives = 21/52 (40%), Gaps = 6/52 (11%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNL------LGLGVGGGCGVGVGLGWGFGSAFG 58
SG G G G G G G G N G G G G G G G G G G G
Sbjct: 274 SGGGTGIGIGEGIGSGSAQPNCGPVTGAPGSGFGEGIGQGSGPGEGIGIGIG 325
Score = 31.2 bits (69), Expect = 0.092
Identities = 20/56 (35%), Positives = 23/56 (40%)
Query: 3 LSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+ SG F G G G G G G G + G G+G G G G G G G G
Sbjct: 224 IGSSGGSGFGEGIGSSGGNGFGEGIGSSGGSGFGEGIGSSGGSGFGEGIGSGGGTG 279
Score = 26.9 bits (58), Expect = 1.7
Identities = 22/66 (33%), Positives = 26/66 (39%), Gaps = 8/66 (12%)
Query: 1 KSLSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGG-------CGVGVGL-GWG 52
+ + SG F G G G G G G G +G+G G G CG G G G
Sbjct: 246 EGIGSSGGSGFGEGIGSSGGSGFGEGIGSGGGTGIGIGEGIGSGSAQPNCGPVTGAPGSG 305
Query: 53 FGSAFG 58
FG G
Sbjct: 306 FGEGIG 311
Score = 26.9 bits (58), Expect = 1.7
Identities = 12/25 (48%), Positives = 15/25 (60%)
Query: 18 GCGFGVGWGFGGMPLNLLGLGVGGG 42
G GFG G G G P +G+G+G G
Sbjct: 303 GSGFGEGIGQGSGPGEGIGIGIGQG 327
>At3g22130 hypothetical protein
Length = 166
Score = 41.6 bits (96), Expect = 7e-05
Identities = 21/39 (53%), Positives = 24/39 (60%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
G GVG G G GG+ +G+GVGGG GVG GL WG
Sbjct: 24 GVGVGGVMTGGVGVGGVTTGGVGVGVGGGHGVGGGLWWG 62
Score = 30.8 bits (68), Expect = 0.12
Identities = 22/47 (46%), Positives = 25/47 (52%), Gaps = 4/47 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGC--GVGVGLGWGFGSAFG 58
G GVG G G GG+ G+GVGG GVGVG+G G G G
Sbjct: 14 GVGVGGVTTGGVGVGGVMTG--GVGVGGVTTGGVGVGVGGGHGVGGG 58
Score = 26.6 bits (57), Expect = 2.3
Identities = 20/49 (40%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGG----GCGVGVGLGWGFGSAFG 58
G G G G G G +GG ++ GL VGG G G+ GLG G G
Sbjct: 48 GVGGGHGVGGGLWWGGFGVDGGGLVVGGLGVDGGGLVGGLGVDGGGLVG 96
Score = 26.6 bits (57), Expect = 2.3
Identities = 17/43 (39%), Positives = 22/43 (50%), Gaps = 6/43 (13%)
Query: 7 GFCLFLSGFGVGCGFG-VGWGFGGMPLNLLGLGVGGGCGVGVG 48
G+ L + G G G G +GWGF + G GVGG G+G
Sbjct: 112 GWGLTVGGLGWGLTVGGLGWGF-----TVGGFGVGGFTAGGLG 149
>At3g17110 hypothetical protein
Length = 175
Score = 41.2 bits (95), Expect = 9e-05
Identities = 26/51 (50%), Positives = 28/51 (53%), Gaps = 6/51 (11%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG+G G G G G G GG G G GGG G G G G G GS +GS Y S
Sbjct: 37 SGYGSGSGGGNGGGGGG------GGGGGGGGGGGGGDGSGSGSGYGSGYGS 81
Score = 37.7 bits (86), Expect = 0.001
Identities = 27/51 (52%), Positives = 29/51 (55%), Gaps = 3/51 (5%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG G G G+G G G+GG N G G GGG G G G G G GS GS Y S
Sbjct: 71 SGSGYGSGYGSGSGYGGG--NNGGGGGGGGRGGGGGGGNG-GSGNGSGYGS 118
Score = 37.0 bits (84), Expect = 0.002
Identities = 23/48 (47%), Positives = 27/48 (55%), Gaps = 1/48 (2%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCG-VGVGLGWGFGSAFGS 59
SG+G G G+G G GG G G GGG G G G G+G GS +GS
Sbjct: 77 SGYGSGSGYGGGNNGGGGGGGGRGGGGGGGNGGSGNGSGYGSGSGYGS 124
Score = 36.6 bits (83), Expect = 0.002
Identities = 24/48 (50%), Positives = 25/48 (52%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G G G G+G G N G G GGG G G G G G GS GS Y S
Sbjct: 30 GRGSGSGSGYGSGSGGGNGGGGGGGGGGGGGGGGGGGDGSGSGSGYGS 77
Score = 35.4 bits (80), Expect = 0.005
Identities = 29/72 (40%), Positives = 33/72 (45%), Gaps = 12/72 (16%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGS-AFGSKYRSSQIRFQGV 71
SG+G G G+G G G GG GGG G G G G G GS GS Y S G
Sbjct: 114 SGYGSGSGYGSGAGRGG----------GGGGGGGGGGGGGGGSRGNGSGYGSGYGEGYGS 163
Query: 72 EFDSKEK-GDSP 82
+ + GDSP
Sbjct: 164 GYGGGDNYGDSP 175
Score = 32.0 bits (71), Expect = 0.054
Identities = 22/47 (46%), Positives = 22/47 (46%), Gaps = 8/47 (17%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG G G G G G G GG G GGG G G G G G G GS
Sbjct: 33 SGSGSGYGSGSGGGNGG--------GGGGGGGGGGGGGGGGGDGSGS 71
Score = 31.2 bits (69), Expect = 0.092
Identities = 20/46 (43%), Positives = 24/46 (51%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G G G G G GG + G G G G G G G G+G G+ G
Sbjct: 47 NGGGGGGGGGGGGGGGGGGGDGSGSGSGYGSGYGSGSGYGGGNNGG 92
Score = 30.8 bits (68), Expect = 0.12
Identities = 21/45 (46%), Positives = 21/45 (46%), Gaps = 1/45 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG N G G G G G G G G G G G
Sbjct: 94 GGGGGRGGGGGGGNGGSG-NGSGYGSGSGYGSGAGRGGGGGGGGG 137
Score = 30.8 bits (68), Expect = 0.12
Identities = 20/51 (39%), Positives = 23/51 (44%), Gaps = 8/51 (15%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G G G G+G G G+G G G G G G G G G G GS+ S
Sbjct: 109 GSGNGSGYGSGSGYGS--------GAGRGGGGGGGGGGGGGGGGGSRGNGS 151
Score = 28.9 bits (63), Expect = 0.46
Identities = 20/51 (39%), Positives = 22/51 (42%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G G G G G G+G G + G G GG G G G G G G G S
Sbjct: 64 GGGDGSGSGSGYGSGYGSGSGYGGGNNGGGGGGGGRGGGGGGGNGGSGNGS 114
Score = 26.6 bits (57), Expect = 2.3
Identities = 18/45 (40%), Positives = 19/45 (42%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G G + G G G G G G G G G G
Sbjct: 54 GGGGGGGGGGGGGDGSGSGSGYGSGYGSGSGYGGGNNGGGGGGGG 98
Score = 24.6 bits (52), Expect = 8.7
Identities = 12/24 (50%), Positives = 12/24 (50%)
Query: 35 LGLGVGGGCGVGVGLGWGFGSAFG 58
LG G G G G G G G G G G
Sbjct: 29 LGRGSGSGSGYGSGSGGGNGGGGG 52
>At2g32690 unknown protein
Length = 201
Score = 40.8 bits (94), Expect = 1e-04
Identities = 21/44 (47%), Positives = 23/44 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAF 57
G G G G G G GFGG G G GGG G G+G G G G +
Sbjct: 152 GLGGGGGIGGGGGFGGGGGGGFGGGAGGGFGKGIGGGGGLGGGY 195
Score = 39.7 bits (91), Expect = 3e-04
Identities = 23/45 (51%), Positives = 24/45 (53%), Gaps = 2/45 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G+GG GLG GGG G G G G G G FG
Sbjct: 132 GGGGGLGGGAGGGYGGGAGG--GLGGGGGIGGGGGFGGGGGGGFG 174
Score = 39.3 bits (90), Expect = 3e-04
Identities = 23/49 (46%), Positives = 25/49 (50%), Gaps = 6/49 (12%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
SG G G G G G G GG G G+GGG G G+G G G G G Y
Sbjct: 103 SGLGGGGGLGGGSGLGG------GGGLGGGGGGGLGGGGGLGGGAGGGY 145
Score = 38.5 bits (88), Expect = 6e-04
Identities = 24/46 (52%), Positives = 24/46 (52%), Gaps = 4/46 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G GG GLG GGG G G GLG G G GS
Sbjct: 74 GLGGGGGLGGGGGLGGGG----GLGGGGGLGGGSGLGGGGGLGGGS 115
Score = 38.5 bits (88), Expect = 6e-04
Identities = 22/45 (48%), Positives = 22/45 (48%), Gaps = 4/45 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G GGG G G G G GFG G
Sbjct: 146 GGGAGGGLGGGGGIGGGG----GFGGGGGGGFGGGAGGGFGKGIG 186
Score = 38.5 bits (88), Expect = 6e-04
Identities = 22/48 (45%), Positives = 24/48 (49%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G G G G GG G G+GGG G G G G GFG G +
Sbjct: 140 GAGGGYGGGAGGGLGG------GGGIGGGGGFGGGGGGGFGGGAGGGF 181
Score = 38.1 bits (87), Expect = 8e-04
Identities = 21/45 (46%), Positives = 24/45 (52%), Gaps = 4/45 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G+GGG G+G G G G+G G
Sbjct: 110 GLGGGSGLGGGGGLGGGG----GGGLGGGGGLGGGAGGGYGGGAG 150
Score = 38.1 bits (87), Expect = 8e-04
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G+GGG G+G G G G GS G
Sbjct: 80 GLGGGGGLGGGGGLGG------GGGLGGGSGLGGGGGLGGGSGLG 118
Score = 37.4 bits (85), Expect = 0.001
Identities = 21/45 (46%), Positives = 26/45 (57%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G+G G GG G G+GGG G+G G G+G G G
Sbjct: 128 GGGLGGGGGLGGGAGGGYGGGAGGGLGGGGGIGGGGGFGGGGGGG 172
Score = 37.4 bits (85), Expect = 0.001
Identities = 21/45 (46%), Positives = 23/45 (50%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G+GGG G+G G G G G G
Sbjct: 86 GLGGGGGLGGGGGLGG------GSGLGGGGGLGGGSGLGGGGGLG 124
Score = 36.6 bits (83), Expect = 0.002
Identities = 21/49 (42%), Positives = 25/49 (50%), Gaps = 8/49 (16%)
Query: 10 LFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
++ GFG G G G G G GG G+GGG G+G G G G G G
Sbjct: 54 IYKKGFGKGLGGGGGLGGGG--------GLGGGGGLGGGGGLGGGGGLG 94
Score = 35.4 bits (80), Expect = 0.005
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G GGG G G G+G G G G
Sbjct: 124 GGGGGGGLGGGGGLGGGAGGGYGGGAGGGLGGGGGIGGGGGFGGG 168
Score = 35.0 bits (79), Expect = 0.006
Identities = 21/45 (46%), Positives = 23/45 (50%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G G G GG G G GGG G G+G G G G G
Sbjct: 120 GGGLGGGGGGGLGGGGGLGGGAGGGYGGGAGGGLGGGGGIGGGGG 164
Score = 28.5 bits (62), Expect = 0.60
Identities = 17/38 (44%), Positives = 19/38 (49%), Gaps = 8/38 (21%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVG 48
F G G G G G G GF G G+GGG G+G G
Sbjct: 165 FGGGGGGGFGGGAGGGF--------GKGIGGGGGLGGG 194
>At4g19070 cadmium-induced protein
Length = 171
Score = 40.0 bits (92), Expect = 2e-04
Identities = 21/47 (44%), Positives = 26/47 (54%), Gaps = 3/47 (6%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGV---GLGWGFGSAF 57
G G+GCG G G GFG + +G G G G VG+ GLG G + F
Sbjct: 30 GIGIGCGVGWGPGFGPEVIGYVGAGCGVGFSVGITLAGLGIGLPTNF 76
Score = 39.7 bits (91), Expect = 3e-04
Identities = 20/37 (54%), Positives = 24/37 (64%), Gaps = 2/37 (5%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLG 50
G G+G G GVGWG G P ++G VG GCGVG +G
Sbjct: 28 GMGIGIGCGVGWGPGFGP-EVIGY-VGAGCGVGFSVG 62
Score = 28.9 bits (63), Expect = 0.46
Identities = 11/17 (64%), Positives = 14/17 (81%)
Query: 36 GLGVGGGCGVGVGLGWG 52
G+G+G GCGVG G G+G
Sbjct: 28 GMGIGIGCGVGWGPGFG 44
Score = 26.2 bits (56), Expect = 3.0
Identities = 10/21 (47%), Positives = 13/21 (61%)
Query: 38 GVGGGCGVGVGLGWGFGSAFG 58
G G G+G+G G G+G FG
Sbjct: 24 GAFWGMGIGIGCGVGWGPGFG 44
>At2g36120 unknown protein
Length = 255
Score = 39.7 bits (91), Expect = 3e-04
Identities = 23/48 (47%), Positives = 25/48 (51%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G+G G G G G G+GG G G GGG G G G G G G A G Y
Sbjct: 146 GYGGGQGAGAGGGYGGGGAGGHGGGGGGGNGGGGGGGSGEGGAHGGGY 193
Score = 36.6 bits (83), Expect = 0.002
Identities = 23/52 (44%), Positives = 25/52 (47%), Gaps = 3/52 (5%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQ 65
G+G G G G G G+GG G G GGG G G G G G A G Y Q
Sbjct: 103 GYGGGSGEGGGAGYGGGEAGGHGGGGGGGAGGG---GGGGGGAHGGGYGGGQ 151
Score = 35.4 bits (80), Expect = 0.005
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 1/48 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G+G G G G G+G GG G G GGG G G G G G G A S Y
Sbjct: 192 GYGAGGGAGEGYG-GGAGAGGHGGGGGGGGGSGGGGGGGGGYAAASGY 238
Score = 34.7 bits (78), Expect = 0.008
Identities = 21/45 (46%), Positives = 22/45 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG +G G GGG G G G G G G G
Sbjct: 59 GSGYGGGSGEGGGAGGHGEGHIGGGGGGGHGGGAGGGGGGGPGGG 103
Score = 34.7 bits (78), Expect = 0.008
Identities = 22/45 (48%), Positives = 23/45 (50%), Gaps = 1/45 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G GGG G G G G+G G A G
Sbjct: 123 GHGGGGGGGAGGGGGGGG-GAHGGGYGGGQGAGAGGGYGGGGAGG 166
Score = 31.6 bits (70), Expect = 0.071
Identities = 22/49 (44%), Positives = 23/49 (46%), Gaps = 7/49 (14%)
Query: 14 GFGVG-CGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G G G G G G GG G G GGG G G G G G G G+ Y
Sbjct: 74 GHGEGHIGGGGGGGHGG------GAGGGGGGGPGGGYGGGSGEGGGAGY 116
Score = 31.6 bits (70), Expect = 0.071
Identities = 20/46 (43%), Positives = 21/46 (45%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G G G G GG G G GGG G G G G G G G
Sbjct: 175 NGGGGGGGSGEGGAHGG------GYGAGGGAGEGYGGGAGAGGHGG 214
Score = 31.6 bits (70), Expect = 0.071
Identities = 17/48 (35%), Positives = 23/48 (47%), Gaps = 10/48 (20%)
Query: 12 LSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
++ +GV G G LG+G+GGG G G G G G G G+
Sbjct: 35 VAAYGVNSGLSAG----------LGVGIGGGPGGGSGYGGGSGEGGGA 72
Score = 30.4 bits (67), Expect = 0.16
Identities = 21/46 (45%), Positives = 22/46 (47%), Gaps = 5/46 (10%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G+G GG G G GGG G G G G G G GS
Sbjct: 182 GSGEGGAHGGGYGAGGG----AGEGYGGGAGAG-GHGGGGGGGGGS 222
Score = 30.4 bits (67), Expect = 0.16
Identities = 20/46 (43%), Positives = 22/46 (47%), Gaps = 2/46 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G G G GG G GGG G G G G G+G G+
Sbjct: 166 GHGGGGGGGNGGGGGGGSGE--GGAHGGGYGAGGGAGEGYGGGAGA 209
Score = 30.0 bits (66), Expect = 0.21
Identities = 20/45 (44%), Positives = 20/45 (44%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG P G G G G G G G G G G
Sbjct: 83 GGGGGHGGGAGGGGGGGPGGGYGGGSGEGGGAGYGGGEAGGHGGG 127
Score = 30.0 bits (66), Expect = 0.21
Identities = 18/37 (48%), Positives = 18/37 (48%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLG 50
G G G G G G G GG G G GGG G G G G
Sbjct: 215 GGGGGGGSGGGGGGGGGYAAASGYGHGGGAGGGEGSG 251
Score = 30.0 bits (66), Expect = 0.21
Identities = 19/45 (42%), Positives = 21/45 (46%), Gaps = 1/45 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G+G G G G G G GG G G GGG G G+G G G
Sbjct: 202 GYGGGAGAG-GHGGGGGGGGGSGGGGGGGGGYAAASGYGHGGGAG 245
Score = 27.7 bits (60), Expect = 1.0
Identities = 19/45 (42%), Positives = 21/45 (46%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G+GG G G GGG G G G G G+ G
Sbjct: 134 GGGGGGGGAHGGGYGGGQGAGAGGGYGGGGAGGHGGGGGGGNGGG 178
Score = 27.3 bits (59), Expect = 1.3
Identities = 18/39 (46%), Positives = 18/39 (46%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
G G G G G G G GG G G G G G G G G G
Sbjct: 172 GGGNGGGGGGGSGEGGAHGGGYGAGGGAGEGYGGGAGAG 210
Score = 25.0 bits (53), Expect = 6.6
Identities = 19/45 (42%), Positives = 19/45 (42%), Gaps = 1/45 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G G G G G G GG G G G G G G G G G G
Sbjct: 127 GGGGGAGGGGGGG-GGAHGGGYGGGQGAGAGGGYGGGGAGGHGGG 170
Score = 25.0 bits (53), Expect = 6.6
Identities = 17/39 (43%), Positives = 17/39 (43%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
G G G G G G G GG G G G G G G G G
Sbjct: 213 GGGGGGGGGSGGGGGGGGGYAAASGYGHGGGAGGGEGSG 251
>At1g17640 RNA-binding protein, putative
Length = 369
Score = 39.3 bits (90), Expect = 3e-04
Identities = 18/44 (40%), Positives = 26/44 (58%), Gaps = 1/44 (2%)
Query: 15 FGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G+G G+G+GG N +G G GG +G G G +A+G
Sbjct: 294 YGYGGGYGYGYGYGGQMFN-MGYGAGGYSHMGSGYGVAAAAAYG 336
Score = 29.6 bits (65), Expect = 0.27
Identities = 18/59 (30%), Positives = 28/59 (46%), Gaps = 11/59 (18%)
Query: 6 SGFCLFLSGFGVGCGFG-----VGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
+GF + G+G G G+G +G+G GG +G G GV +G G + G+
Sbjct: 291 AGFYGYGGGYGYGYGYGGQMFNMGYGAGGYS------HMGSGYGVAAAAAYGGGKSHGN 343
>At4g29030 glycine-rich protein like
Length = 115
Score = 38.9 bits (89), Expect = 4e-04
Identities = 23/50 (46%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Query: 3 LSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
+S G L GFG G G G+G G GG + LG G+GGG G+G G G
Sbjct: 59 VSGPGGNLGYGGFG-GAGGGLGGGLGGGAGSGLGGGLGGGSGIGAGTSGG 107
Score = 36.6 bits (83), Expect = 0.002
Identities = 23/48 (47%), Positives = 28/48 (57%), Gaps = 4/48 (8%)
Query: 12 LSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
+SG G G+G GFGG L G G+GGG G G+G G G GS G+
Sbjct: 59 VSGPGGNLGYG---GFGGAGGGLGG-GLGGGAGSGLGGGLGGGSGIGA 102
Score = 29.6 bits (65), Expect = 0.27
Identities = 20/39 (51%), Positives = 21/39 (53%), Gaps = 3/39 (7%)
Query: 22 GVGWGFGGMPLNLLGLGVGGGC--GVGVGLGWGFGSAFG 58
GVG G G P LG G GG G+G GLG G GS G
Sbjct: 54 GVGGGVSG-PGGNLGYGGFGGAGGGLGGGLGGGAGSGLG 91
Score = 26.9 bits (58), Expect = 1.7
Identities = 20/40 (50%), Positives = 23/40 (57%), Gaps = 3/40 (7%)
Query: 22 GVGWGFGGMPLNLLGL-GVGGG-CGVGVGLGW-GFGSAFG 58
G G G+ G+ N L GVGGG G G LG+ GFG A G
Sbjct: 37 GYGGGYSGVGDNGLPFGGVGGGVSGPGGNLGYGGFGGAGG 76
Score = 25.4 bits (54), Expect = 5.1
Identities = 19/48 (39%), Positives = 22/48 (45%), Gaps = 6/48 (12%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGV---GGGCGVGV--GLGWGFGSAFG 58
G G G+ G G G+P +G GV GG G G G G G G G
Sbjct: 37 GYGGGYS-GVGDNGLPFGGVGGGVSGPGGNLGYGGFGGAGGGLGGGLG 83
>At3g07560 unknown protein
Length = 304
Score = 38.9 bits (89), Expect = 4e-04
Identities = 20/61 (32%), Positives = 32/61 (51%), Gaps = 5/61 (8%)
Query: 5 HSGFCLFLSGFGVGCGFGV-----GWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
+ G ++ G+G G +G G GG + G G+G G G+G+G+G G G +GS
Sbjct: 119 YGGSSMYRGGYGGGGLYGSSGMYGGGAMGGYGGTMGGYGMGMGTGMGMGMGMGMGGPYGS 178
Query: 60 K 60
+
Sbjct: 179 Q 179
>At4g30450 glycine-rich protein
Length = 107
Score = 38.5 bits (88), Expect = 6e-04
Identities = 22/49 (44%), Positives = 25/49 (50%), Gaps = 10/49 (20%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G+G G GVG LG+G+GGG G G G G G GS S SS
Sbjct: 37 GIGIGLGVG----------LGIGLGGGSGSGAGAGSGSGSGSRSSSSSS 75
Score = 37.7 bits (86), Expect = 0.001
Identities = 22/53 (41%), Positives = 25/53 (46%), Gaps = 10/53 (18%)
Query: 12 LSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
L G G+G G G+G +GLG G G G G G G G GS S SS
Sbjct: 35 LGGIGIGLGVGLG----------IGLGGGSGSGAGAGSGSGSGSRSSSSSSSS 77
Score = 33.9 bits (76), Expect = 0.014
Identities = 23/59 (38%), Positives = 28/59 (46%), Gaps = 1/59 (1%)
Query: 23 VGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEFDSKEKGDS 81
+G GG+ + L G+G+G G G G G G G GS GS RSS S G S
Sbjct: 31 LGIDLGGIGIGL-GVGLGIGLGGGSGSGAGAGSGSGSGSRSSSSSSSSSSSSSSGSGGS 88
>At2g15780 unknown protein
Length = 257
Score = 38.5 bits (88), Expect = 6e-04
Identities = 24/57 (42%), Positives = 29/57 (50%), Gaps = 15/57 (26%)
Query: 13 SGFGVGC---GFGVGWGFGGMPLNLLGLGVGG-GCGV-----------GVGLGWGFG 54
SG+G G G G GWG+GG+P N G GG G G+ G G GWG+G
Sbjct: 72 SGWGFGSSRNGSGWGWGWGGVPNNTHSSGSGGSGWGMGPNNNYSSGSGGSGSGWGYG 128
Score = 32.7 bits (73), Expect = 0.032
Identities = 17/38 (44%), Positives = 19/38 (49%), Gaps = 2/38 (5%)
Query: 22 GVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G GWG+GG N G G G G GWG GS + S
Sbjct: 29 GFGWGWGGHSNN--SSGSGSNSRSGWGWGWGQGSNYSS 64
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.147 0.481
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,580,187
Number of Sequences: 26719
Number of extensions: 120616
Number of successful extensions: 2424
Number of sequences better than 10.0: 145
Number of HSP's better than 10.0 without gapping: 108
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 406
Number of HSP's gapped (non-prelim): 879
length of query: 95
length of database: 11,318,596
effective HSP length: 71
effective length of query: 24
effective length of database: 9,421,547
effective search space: 226117128
effective search space used: 226117128
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0050.15


