

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0047c.1
(619 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
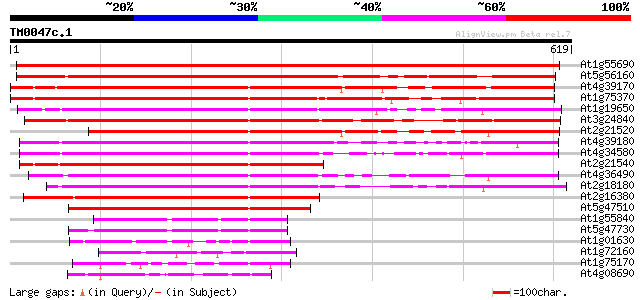
Score E
Sequences producing significant alignments: (bits) Value
At1g55690 Phosphatidylinositol Transfer Protein like 723 0.0
At5g56160 putative protein 563 e-160
At4g39170 unknown protein 504 e-143
At1g75370 sec14 cytosolic factor, putative 469 e-132
At1g19650 sec14 cytosolic factor- like protein 462 e-130
At3g24840 phosphatidylinositol transfer protein like 455 e-128
At2g21520 putative phosphatidylinositol/phosphatidylcholine tran... 417 e-117
At4g39180 SEC14 - like protein 412 e-115
At4g34580 putative protein 397 e-111
At2g21540 putative phosphatidylinositol/phophatidylcholine trans... 392 e-109
At4g36490 hypothetical protein 383 e-106
At2g18180 putative phosphatidylinositol/phophatidylcholine trans... 377 e-105
At2g16380 putative phosphatidylinositol/phosphatidylcholine tran... 367 e-101
At5g47510 putative protein 282 4e-76
At1g55840 polyphosphoinositide binding protein, putative 82 7e-16
At5g47730 putative protein 77 2e-14
At1g01630 polyphosphoinositide binding protein, putative 76 6e-14
At1g72160 cytosolic factor, putative 68 2e-11
At1g75170 unknown protein 61 2e-09
At4g08690 putative phosphoglyceride transfer protein 60 5e-09
>At1g55690 Phosphatidylinositol Transfer Protein like
Length = 625
Score = 723 bits (1865), Expect = 0.0
Identities = 376/605 (62%), Positives = 459/605 (75%), Gaps = 6/605 (0%)
Query: 8 DEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDV 67
DE RERRSD E SEDERR SKIG L+KKA+NAS+KFTHSLKKRGKRKIDYRVPAVSIEDV
Sbjct: 11 DEFRERRSDFEISEDERRRSKIGNLKKKAINASTKFTHSLKKRGKRKIDYRVPAVSIEDV 70
Query: 68 RDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEY 127
RDE EE+VV E R++L+ER LLPPRHD+Y+TLLRFLKARD NIEKT Q+WEEML WRKEY
Sbjct: 71 RDEKEESVVLEFRRKLLERDLLPPRHDEYHTLLRFLKARDLNIEKTTQLWEEMLRWRKEY 130
Query: 128 GTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKY 187
GTDTILE+F+FEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPS+LMRITTIDRYLKY
Sbjct: 131 GTDTILEDFDFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSKLMRITTIDRYLKY 190
Query: 188 HVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYY 247
HVQEFERALQEKFPACSIAAKRRI STTTILDVQGLG+KNFT TAANL+AAM+KID +YY
Sbjct: 191 HVQEFERALQEKFPACSIAAKRRICSTTTILDVQGLGIKNFTPTAANLVAAMSKIDNSYY 250
Query: 248 PETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFL 307
PETLH+MYIVNAG+GFKKMLWPAAQKF D KTIAKI +L+ KSL+KL EVIDSSQLP+FL
Sbjct: 251 PETLHRMYIVNAGTGFKKMLWPAAQKFLDAKTIAKIHVLEPKSLFKLHEVIDSSQLPEFL 310
Query: 308 GGSCTCPTE-GGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF-QVHSL 365
GGSC+C + GGCL+++KGPWNDP+I+KL++ E++L RQ +R + S+ +H
Sbjct: 311 GGSCSCFGDGGGCLRSNKGPWNDPEIMKLIYHGESSLFRQSTRKLTDPHYSSSYISIHPS 370
Query: 366 KGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEK 425
K +++S AES S + SSPT + S + EE R D+NGYYSCDD +K
Sbjct: 371 KAIQAETSAAESISCSDVPSSPTGRLCSASSHVNSAYEEARASDVNGYYSCDDKFAIPDK 430
Query: 426 VV-ENGQFHLSQKQSLQTNDME-NVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLA 483
GQ SQ Q + N + T S G + ++ +++K + + L
Sbjct: 431 ATNRKGQERQSQYQMRELNATTIGLKCETSSPGAPIIRWLHDLRVMIDKIKCENLAKRLL 490
Query: 484 SFMERLITFIRSLRFEFWRTQNNVHPSNTTEHN--INNHSAAVEAAFERDHILPCAQRLQ 541
S M +L R E R+Q V PS+ TE + + S +D ILPC +R+Q
Sbjct: 491 SLMLKLAAVFRYTPLELLRSQTTVSPSSLTEDDSRCSLISPPPREPTMKDRILPCLERIQ 550
Query: 542 RLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVE 601
+LEK +E++ NKP +P+EKE+MLMDS+DRIKSVEFDL+KTKR+LH+ V +Q+EI ++++
Sbjct: 551 KLEKSYEDIRNKPVAIPVEKERMLMDSLDRIKSVEFDLDKTKRLLHATVMKQMEITEMLQ 610
Query: 602 NLQKS 606
N++ S
Sbjct: 611 NIRDS 615
>At5g56160 putative protein
Length = 592
Score = 563 bits (1451), Expect = e-160
Identities = 294/597 (49%), Positives = 399/597 (66%), Gaps = 37/597 (6%)
Query: 8 DEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKR-GKRKIDYRVPAVSIED 66
++ E+ SD E E+E R S+IG L+KKA + S+K TH LK R GKRKID+++P IED
Sbjct: 20 EQTGEKLSDSEYIEEEPRRSRIGNLKKKAFSCSTKLTHPLKMRKGKRKIDFQIPL--IED 77
Query: 67 VRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKE 126
VRDE EE +V +LRQ+L+++ LLPP HDDY+ LLRFLK +F IEKT+ WEEML WRKE
Sbjct: 78 VRDEKEEKLVSKLRQQLLQKDLLPPVHDDYHMLLRFLKTMEFKIEKTVTAWEEMLKWRKE 137
Query: 127 YGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLK 186
+GTD I+++F F+EL+EV ++YPQGYHGVDK+GRP+YIERLGKAHP +LM +TTI+RYLK
Sbjct: 138 FGTDRIIQDFNFKELDEVTRHYPQGYHGVDKDGRPIYIERLGKAHPGKLMEVTTIERYLK 197
Query: 187 YHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNY 246
YHVQEFER LQEK PACS+AAKRR+ +TTTILDV+GLGMKNFT TAANLLA + K+D NY
Sbjct: 198 YHVQEFERTLQEKLPACSVAAKRRVTTTTTILDVEGLGMKNFTPTAANLLATIAKVDCNY 257
Query: 247 YPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDF 306
YPETLH+M+IVNAG GF+ LWPAAQK DP TIAKIQ+L+ +SL KLLE IDSSQLP+F
Sbjct: 258 YPETLHRMFIVNAGIGFRSFLWPAAQKLLDPMTIAKIQVLEPRSLSKLLEAIDSSQLPEF 317
Query: 307 LGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLK 366
LGG C CP EGGCL+++KGPWNDP+I++LVH E V Q + A +++DS
Sbjct: 318 LGGLCKCPNEGGCLRSNKGPWNDPEIVELVHHMEVNNVPQTTTAPLHVRDYDSTTC---- 373
Query: 367 GRCSDSSTAESGSDINDYSSPTRQRSSTHLRLA-PVDEEVRVPDLNGYYSCDDSAPTTEK 425
S T + + +Y S T RSS H + P+ ++ D D TT +
Sbjct: 374 -TISPKETLKEEPEPEEYYSSTGSRSSMHTCIVPPLSDKASTSD-------GDKFITTVE 425
Query: 426 VVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASF 485
+E+ +Q Q L + + ++ EG + F ++E + N ++ ++L F
Sbjct: 426 SIES-----AQSQLLDADTENTFANTSVREGGQILR-FGALREKINSENIFHLVKILLVF 479
Query: 486 MERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEK 545
+L L +W+ QN V +++ +N +L C RL+++EK
Sbjct: 480 PLKLFVLFGFLLPGYWQRQNTVVVPDSSTNN---------------KVLECFDRLKKMEK 524
Query: 546 VFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVEN 602
F E++ K +P EK+L +S++RIKS+E DL+KTK VLH + +QL+I + +E+
Sbjct: 525 EFTEISRKQVKIPEANEKLLAESLERIKSLELDLDKTKSVLHITLTKQLQITEQLES 581
>At4g39170 unknown protein
Length = 614
Score = 504 bits (1298), Expect = e-143
Identities = 279/610 (45%), Positives = 391/610 (63%), Gaps = 36/610 (5%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVP 60
+EG S+DE +ER+SD ENSEDERR+ +IG+L+KKA+NAS+KF HSLKK+ +RK D RV
Sbjct: 13 FEGILSSDEKKERKSDFENSEDERRT-RIGSLKKKAINASTKFKHSLKKK-RRKSDVRVS 70
Query: 61 AVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEM 120
+VSIEDVRD E V E RQ L+ LLP +HDDY+ +LRFLKAR F+IEK MW +M
Sbjct: 71 SVSIEDVRDVEELQAVDEFRQALVMEELLPHKHDDYHMMLRFLKARKFDIEKAKHMWADM 130
Query: 121 LLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITT 180
+ WRKE+GTDTI+++F+FEE++EVL+YYP GYH VDKEGRPVYIERLGK P++LM++TT
Sbjct: 131 IQWRKEFGTDTIIQDFQFEEIDEVLKYYPHGYHSVDKEGRPVYIERLGKVDPNKLMQVTT 190
Query: 181 IDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMT 240
+DRY++YHV+EFER+ KFPAC+IAAK+ I S+TTILDVQG+G+KNFT++A L+ +
Sbjct: 191 LDRYIRYHVKEFERSFMLKFPACTIAAKKYIDSSTTILDVQGVGLKNFTKSARELITRLQ 250
Query: 241 KIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDS 300
KID + YPETLHQM+I+NAG GF ++LW + F DPKT +KI +L K KLLE+IDS
Sbjct: 251 KIDGDNYPETLHQMFIINAGPGF-RLLWSTVKSFLDPKTTSKIHVLGCKYQSKLLEIIDS 309
Query: 301 SQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF 360
S+LP+FLGG+CTC +GGC+ + KGPW +P+I+K+V A +Q+ + N ++
Sbjct: 310 SELPEFLGGACTCADQGGCMLSDKGPWKNPEIVKMVLHGGAHRAKQVVKVLNSDGKVIAY 369
Query: 361 QVHS---LKGRCSDSSTAESGSDIND-YSSPTRQRSSTHLRLAPVDEEVRVPD-----LN 411
S +KG SD+STAESGS+ D SP +S +HLRL PV EE +V
Sbjct: 370 AKPSYPWIKG--SDTSTAESGSEAEDIVVSPKAVKSYSHLRLTPVREEAKVGSGETSFAG 427
Query: 412 GYYSCDDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVE 471
+ D+ P +K V+ +R S+G + V
Sbjct: 428 SFAGYDEYVPMVDKAVD-----------ATWKVKPTAINRAPSKGAH-------MPPNVP 469
Query: 472 KTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERD 531
K + + RVL +FM ++ + R R P ++ I +AA EA D
Sbjct: 470 KDHESFSARVLVTFMAFVMAILTFFRTVSNRVVTKQLPPPPSQPQIEGSAAAEEA----D 525
Query: 532 HILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVK 591
+ ++L LE+ L +KP MP EKE++L ++ R+ ++E +L TK+ L+ A+
Sbjct: 526 LLNSVLKKLTELEEKIGALQSKPSEMPYEKEELLNAAVCRVDALEAELIATKKALYEALM 585
Query: 592 QQLEIADLVE 601
+Q E+ ++
Sbjct: 586 RQEELLAYID 595
>At1g75370 sec14 cytosolic factor, putative
Length = 612
Score = 469 bits (1208), Expect = e-132
Identities = 273/614 (44%), Positives = 379/614 (61%), Gaps = 42/614 (6%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTL--RKKAMNASSKFTHSLKKRG--KRKID 56
+EG S DE RERRSD E SEDE+++ +IG +KKA ASSK HSLKK+G +R+
Sbjct: 13 FEGVSSNDERRERRSDFEVSEDEKKT-RIGNFNFKKKAAKASSKLRHSLKKKGSSRRRSS 71
Query: 57 YRVPAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQM 116
R +++IED+ D E V E R L+ LLPP DDY+ +LRFLKAR F+I KT M
Sbjct: 72 DRTFSLTIEDIHDVEELRAVDEFRNLLVSENLLPPTLDDYHIMLRFLKARKFDIGKTKLM 131
Query: 117 WEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLM 176
W M+ WRK++GTDTI E+FEFEE +EVL+YYP GYHGVDKEGRPVYIERLG P++LM
Sbjct: 132 WSNMIKWRKDFGTDTIFEDFEFEEFDEVLKYYPHGYHGVDKEGRPVYIERLGLVDPAKLM 191
Query: 177 RITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLL 236
++TT++R+++YHV+EFE+ + K PAC IAAKR I S+TTILDVQG+G KNF++ A +L+
Sbjct: 192 QVTTVERFIRYHVREFEKTVNIKLPACCIAAKRHIDSSTTILDVQGVGFKNFSKPARDLI 251
Query: 237 AAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLE 296
+ KID + YPETLH+M+I+N GSGF K++W ++F DPKT+ KI ++ +K KLLE
Sbjct: 252 IQLQKIDNDNYPETLHRMFIINGGSGF-KLVWATVKQFLDPKTVTKIHVIGNKYQNKLLE 310
Query: 297 VIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQN 356
+ID+SQLPDFLGG+CTC GGC+++ KGPWNDP+I+K++ S L R S A N
Sbjct: 311 IIDASQLPDFLGGTCTCADRGGCMRSDKGPWNDPEILKMLQSG-GPLCRHNS-ALNSFSR 368
Query: 357 FDSFQVHSLKG-RCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYS 415
S S G + SD+STAESGS++ + +SP R +L PV E++R ++ Y
Sbjct: 369 VSSCDKPSFSGIKASDTSTAESGSEVEEMASPKVNRELRVPKLTPVCEDIRGTAIS--YP 426
Query: 416 CDDS---APTTEKVVENG-QFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVE 471
D S +P +KVV+ H K S SE T + R +
Sbjct: 427 TDSSEYDSPMVDKVVDVAWMAHEKPKASKG------------SEDTPDSGKIRTV----- 469
Query: 472 KTNHLYVTRVLASFMERLITFIRSL----RFEFWRTQNNVHPSNTTEHNINNHSAAVEAA 527
Y+ R L F L T + SL R +++++V N E S A
Sbjct: 470 ----TYIWRWLMMFFVNLFTLLISLALPQREGHSQSESSVDGPNARES--RPPSPAFATI 523
Query: 528 FERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLH 587
ER+ RL LEK E L++K + MP EKE++L ++ R+ ++E +L TK+ LH
Sbjct: 524 AERNVFSSVVNRLGDLEKQVETLHSKRHEMPREKEELLNTAVYRVDALEAELIATKKALH 583
Query: 588 SAVKQQLEIADLVE 601
A+ +Q ++ ++
Sbjct: 584 EALMRQDDLLAYID 597
>At1g19650 sec14 cytosolic factor- like protein
Length = 608
Score = 462 bits (1188), Expect = e-130
Identities = 262/613 (42%), Positives = 371/613 (59%), Gaps = 40/613 (6%)
Query: 9 EIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVR 68
E RE++SD E SEDE+++ G L+KK+ + SKF HSLK+RG R ID R +++ ED+
Sbjct: 18 EKREKKSDFEVSEDEKKTRIGGILKKKS--SKSKFRHSLKRRGSRSID-RTLSLTFEDIH 74
Query: 69 DEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYG 128
D E V E RQ LI LLPP DDY+ +LRFL AR F++ K MW M+ WR+++G
Sbjct: 75 DAEELRYVSEFRQSLISDHLLPPNLDDYHIMLRFLFARKFDLGKAKLMWTNMIQWRRDFG 134
Query: 129 TDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYH 188
TDTILE+FEF EL+EVL+YYPQGYHGVDKEGRPVYIERLGK S+LM++TT++RYL+YH
Sbjct: 135 TDTILEDFEFPELDEVLRYYPQGYHGVDKEGRPVYIERLGKVDASKLMQVTTLERYLRYH 194
Query: 189 VQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYP 248
V+EFE+ + KFPAC IAAKR I S+TTILDVQGLG+KNFT+TA +L+ + KID++ YP
Sbjct: 195 VKEFEKTITVKFPACCIAAKRHIDSSTTILDVQGLGLKNFTKTARDLIIQLQKIDSDNYP 254
Query: 249 ETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLG 308
ETLH+M+I+NAGSGF K+LW + F DPKT++KI +L +K KLLE+ID+SQLPDF G
Sbjct: 255 ETLHRMFIINAGSGF-KLLWGTVKSFLDPKTVSKIHVLGNKYQNKLLEMIDASQLPDFFG 313
Query: 309 GSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGR 368
G+CTC +GGC+++ KGPW D +I+K+ S T R + S + +
Sbjct: 314 GTCTCADQGGCMRSDKGPWKDSEILKMGRSG-GTFCRHAGAFLSSDSQISSSDKPTYSLK 372
Query: 369 CSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDE----EVRVPDLNGYYSCDDSAPTTE 424
SD+STA+SGS++ + +SP ++ +L PV E + L+ Y C P +
Sbjct: 373 VSDTSTAKSGSELEEMASPKTNTNNHVPKLTPVSEYANGNISPTVLSEYEEC---VPMVD 429
Query: 425 KVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLAS 484
KVV+ + Q +M N SEG + I + ++ L +
Sbjct: 430 KVVD---------VAWQLQEMPNA-----SEGPQYTSSLGKIGSV------RHIWSWLTA 469
Query: 485 FMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNH--------SAAVEAAFERDHILPC 536
F T + SL + + +H S+ + S ER I
Sbjct: 470 FFISFFTLLASLALPQTKEHSQLHSSSVRAELCDERIARESRPPSPPRSTITERVIISSV 529
Query: 537 AQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEI 596
RL LEK E L+++ MP EKE++L ++ R+ ++E +L TK+ LH A+ +Q E+
Sbjct: 530 LSRLGDLEKQIENLHSRKSEMPHEKEELLNAAVYRVDALEAELITTKKALHEALIRQEEL 589
Query: 597 ADLVENLQKSNCR 609
++ +++ CR
Sbjct: 590 LGYIDRQKEAKCR 602
>At3g24840 phosphatidylinositol transfer protein like
Length = 579
Score = 455 bits (1171), Expect = e-128
Identities = 259/594 (43%), Positives = 364/594 (60%), Gaps = 58/594 (9%)
Query: 17 VENSEDERRS-SKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETV 75
+E SEDE+ + ++ +L+KKA+ AS+K THSL+KRGKR D P V IEDVRDE EE
Sbjct: 22 IEVSEDEKITRTRSRSLKKKAIKASNKLTHSLRKRGKRVADQYAPIV-IEDVRDEEEEKA 80
Query: 76 VHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE 135
V+ R+ L+ LLPPRHDDY+T+LRFLKAR F++EKT+QMWEEML WRKE G DTI+++
Sbjct: 81 VNVFRKALVSLDLLPPRHDDYHTMLRFLKARRFDLEKTVQMWEEMLKWRKENGVDTIIQD 140
Query: 136 FEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERA 195
F ++E EEV QYYP GYHGVD+EGRPVYIERLGK P +LM++TT++R+L+YHVQ FE+
Sbjct: 141 FVYDEYEEVQQYYPHGYHGVDREGRPVYIERLGKIDPGKLMKVTTLERFLRYHVQGFEKT 200
Query: 196 LQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMY 255
EKFPACSIAAKR I S+TTI+DV G+ +F + A +L+ M KID + YPETL+QMY
Sbjct: 201 FSEKFPACSIAAKRHINSSTTIIDVHGVSWMSFRKLAQDLVMRMQKIDGDNYPETLNQMY 260
Query: 256 IVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPT 315
I+NAG+GF K++W + F DPKT +KI +L +K LLE+ID S+LP+FLGG+C C
Sbjct: 261 IINAGNGF-KLVWNTVKGFLDPKTTSKIHVLGNKYRSHLLEIIDPSELPEFLGGNCKCAH 319
Query: 316 EGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTA 375
EGGC++ +KGPWNDP+I+KLV S +A + +N + ++ SL+ +D S+
Sbjct: 320 EGGCMRFNKGPWNDPEIMKLVRSRDA---MYKPKEMGLLENGEVAKLFSLRHVNTDMSSP 376
Query: 376 ESGSDINDYSSPTRQRSS--THLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQFH 433
+ G R+R S H + A + + + D ++P +
Sbjct: 377 DGGH--------VRERESHPEHDKRAQLSNQAEAVGVGRMEQSDSTSPLPN--------N 420
Query: 434 LSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFI 493
L+ ++SL T+ + V LA F+ +L+ +
Sbjct: 421 LAVERSLTTSLQK-------------------------------VASFLARFILQLLGSL 449
Query: 494 RSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEELNNK 553
L F N P N ++ + + + H PC RLQ LE + L +K
Sbjct: 450 -CLMFRILGRLVNKQPENQLRPELSVSVSQQQVPPPQVH--PCWLRLQNLETMVTVLCDK 506
Query: 554 PYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSN 607
P +P EKE +L DS+DRIKS+E DL+KTK+ L +Q+E+A+ ENL++S+
Sbjct: 507 PSSIPQEKEDILRDSLDRIKSIEQDLQKTKKALFLTASKQIELAECFENLKESS 560
>At2g21520 putative phosphatidylinositol/phosphatidylcholine
transfer protein
Length = 531
Score = 417 bits (1073), Expect = e-117
Identities = 230/541 (42%), Positives = 328/541 (60%), Gaps = 51/541 (9%)
Query: 88 LLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQY 147
LLP RHDDY+ +LRFLKAR F++EK QMW +M+ WRKE+GTDTI+++F+FEE+ EVL++
Sbjct: 4 LLPDRHDDYHMMLRFLKARKFDVEKAKQMWADMIQWRKEFGTDTIIQDFDFEEINEVLKH 63
Query: 148 YPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAA 207
YPQ YHGVDKEGRP+YIERLGK P+RLM++T++DRY++YHV+EFER+ KFP+C+I+A
Sbjct: 64 YPQCYHGVDKEGRPIYIERLGKVDPNRLMQVTSMDRYVRYHVKEFERSFMIKFPSCTISA 123
Query: 208 KRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKML 267
KR I S+TTILDVQG+G+KNF ++A +L+ + KID + YPETLHQM+I+NAG GF ++L
Sbjct: 124 KRHIDSSTTILDVQGVGLKNFNKSARDLITRLQKIDGDNYPETLHQMFIINAGPGF-RLL 182
Query: 268 WPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPW 327
W + F DPKT AKI +L K L KLLEVID ++LP+FLGG+CTC +GGC+ + KGPW
Sbjct: 183 WNTVKSFLDPKTSAKIHVLGYKYLSKLLEVIDVNELPEFLGGACTCADQGGCMLSDKGPW 242
Query: 328 NDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHS---LKGRCSDSSTAESGSDINDY 384
+P+I+K+V A RQ+ + N + ++ S +KG SD+STAESGSD D
Sbjct: 243 KNPEIVKMVLHGGAHRARQVVKVLNSEGKVIAYAKPSYTWIKG--SDTSTAESGSDAEDI 300
Query: 385 SSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQFHLSQKQSLQTND 444
SP +S +HLRL PV E + + D+ P +K V+ T
Sbjct: 301 GSPKAIKSFSHLRLTPVPGETSL--AGSFPGYDEYVPMVDKAVD------------ATWK 346
Query: 445 MENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQ 504
++ R S G ++ V K + RVL FM L+ F F+RT
Sbjct: 347 VKPAIQRVASRGA-------LMSPTVPKDHEGIKARVLVMFMAFLMAV-----FTFFRTV 394
Query: 505 NNVHPSNTTEHNINNHSAAVEAA-------------------FERDHILPCAQRLQRLEK 545
P+ TT A+E E D + ++L LE
Sbjct: 395 TKKLPATTTSSPAETQGNAIELGSNGEGVKEECRPPSPVPDLTETDLLNCVTKKLTELEG 454
Query: 546 VFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQK 605
L +KP MP EKE++L ++ R+ ++E +L TK+ L+ A+ +Q E+ ++ ++
Sbjct: 455 KIGTLQSKPNEMPYEKEELLNAAVCRVDALEAELIATKKALYEALMRQEELLAYIDRQEE 514
Query: 606 S 606
+
Sbjct: 515 A 515
>At4g39180 SEC14 - like protein
Length = 554
Score = 412 bits (1058), Expect = e-115
Identities = 242/601 (40%), Positives = 350/601 (57%), Gaps = 71/601 (11%)
Query: 11 RERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDE 70
R + DVE SED++R +K+ +L+KKA+NA++KF HS+ K+G+R RV VSI D D
Sbjct: 11 RHNKIDVEISEDDKRLTKLCSLKKKAINATNKFKHSMTKKGRRHS--RVACVSIVDEIDT 68
Query: 71 GEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTD 130
E V RQ LI LLP +HDD++ +LRFL+AR F++EK QMW +ML WRKEYG D
Sbjct: 69 EELQAVDAFRQALILDELLPSKHDDHHMMLRFLRARKFDLEKAKQMWSDMLNWRKEYGAD 128
Query: 131 TILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQ 190
TI+E+F+F+E+EEV++YYPQGYHGVDKEGRP+YIERLG+ ++LM++TTIDRY+KYHV+
Sbjct: 129 TIMEDFDFKEIEEVVKYYPQGYHGVDKEGRPIYIERLGQVDATKLMKVTTIDRYVKYHVK 188
Query: 191 EFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPET 250
EFE+ KFPACSIAAKR I +TTILDVQG+G+ NF + A +LL ++ KID + YPET
Sbjct: 189 EFEKTFNVKFPACSIAAKRHIDQSTTILDVQGVGLSNFNKAAKDLLQSIQKIDNDNYPET 248
Query: 251 LHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGS 310
L++M+I+NAG GF ++LW + F DPKT AKI +L +K KLLE+ID+++LP+FLGG
Sbjct: 249 LNRMFIINAGCGF-RLLWNTVKSFLDPKTTAKIHVLGNKYQTKLLEIIDANELPEFLGGK 307
Query: 311 CTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCS 370
CTC +GGC+++ KGPWNDP+I KLV + E +R+ E+ F+ + K +C
Sbjct: 308 CTCADKGGCMRSDKGPWNDPEIFKLVQNGEGRCLRRSLSGIEEKTIFE--YNNETKKKCE 365
Query: 371 DSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENG 430
T ++S+ + +D V D+A + +
Sbjct: 366 PEE--------------THKQSAAEMEKKFIDTNV------------DAAAAADWPTK-- 397
Query: 431 QFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLI 490
K D+++V ++VN L E+ +LY S M L+
Sbjct: 398 ----LNKAEKNPTDLKDVY-------SAVNPL--------ERKGYLY-----GSVMALLM 433
Query: 491 TFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEEL 550
+ +R T+N P TE N+ +S A ++ + Q + K +L
Sbjct: 434 GIVGVMRL----TKN--MPRRLTEANV--YSREGSAVYQDGVTVMSKQEYIAMVKKITDL 485
Query: 551 NNKPYGMP------LEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQ 604
K M +E+EK L ++ RI +E L +T + L + +Q EI +E +
Sbjct: 486 EEKCKSMEAQAAFYMEREKTLDAALRRIDQLELQLSETNKALDETMTRQHEIMAFIEKKK 545
Query: 605 K 605
K
Sbjct: 546 K 546
>At4g34580 putative protein
Length = 560
Score = 397 bits (1021), Expect = e-111
Identities = 234/600 (39%), Positives = 355/600 (59%), Gaps = 63/600 (10%)
Query: 12 ERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEG 71
E + ++E SE+ER+ KI +L+KKA+NAS++F +S KK+G+R RV +V IED D
Sbjct: 3 ETKPEIEMSEEERKIVKISSLKKKAINASNRFKNSFKKKGRRSSS-RVMSVPIEDDIDAE 61
Query: 72 EETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDT 131
+ + RQ LI LLP + DD + +LRFL+AR F+IEK QMW +M+ WRK++G DT
Sbjct: 62 DLQALDAFRQALILDELLPSKLDDLHMMLRFLRARKFDIEKAKQMWSDMIQWRKDFGADT 121
Query: 132 ILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQE 191
I+E+F+FEE++EV+++YPQGYHGVDKEGRPVYIERLG+ ++L+++TT+DRY+KYHV+E
Sbjct: 122 IIEDFDFEEIDEVMKHYPQGYHGVDKEGRPVYIERLGQIDANKLLQVTTMDRYVKYHVKE 181
Query: 192 FERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETL 251
FE+ + KFP+CS+AA + I +TTILDVQG+G+KNF+++A LL + KID YPETL
Sbjct: 182 FEKTFKVKFPSCSVAANKHIDQSTTILDVQGVGLKNFSKSARELLQRLCKIDNENYPETL 241
Query: 252 HQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSC 311
++M+I+NAGSGF ++LW + F DPKT AKI +L +K KLLEVID+S+LP+F GG+C
Sbjct: 242 NRMFIINAGSGF-RLLWSTVKSFLDPKTTAKIHVLGNKYHSKLLEVIDASELPEFFGGAC 300
Query: 312 TCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSD 371
TC +GGC+++ KGPWNDP+++K+ + EA + S S ++
Sbjct: 301 TCEDKGGCMRSDKGPWNDPEVLKIAINREA----KCSPISEDEHKH-------------- 342
Query: 372 SSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQ 431
+ G + + S R + T DE+ N Y EK +
Sbjct: 343 ---VDQGRSTSGFESLERIKKKT-------DED------NVY----------EKQIATID 376
Query: 432 FHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLIT 491
+ +T EN IS+G + +++K +K + L V V+A F+ ++
Sbjct: 377 KSMDMAWLAKTQKAENF---PISKG-----ITKLMKGAPKKGDGLLVGGVMA-FVMGIVA 427
Query: 492 FIRSLR------FEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEK 545
+R + E N+V +T+ N A A + +R+ LE
Sbjct: 428 MVRLSKDVPRKLTEAALYGNSVCYEESTKSKQNQGQFA--APVSSSEYMLMVKRMAELED 485
Query: 546 VFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQK 605
L+ KP + EKE+ L +++R++ +E +L +TK+ L A+ Q EI +E +K
Sbjct: 486 KCMFLDLKPAHVESEKEEKLQAALNRVQVLEQELTETKKALEEALVSQKEILAYIEKKKK 545
>At2g21540 putative phosphatidylinositol/phophatidylcholine transfer
protein
Length = 371
Score = 392 bits (1007), Expect = e-109
Identities = 188/336 (55%), Positives = 259/336 (76%), Gaps = 4/336 (1%)
Query: 11 RERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDE 70
R + D + SEDE+++ K+ +L+KKA+NAS+KF HS KR +R + RV +VSI D D
Sbjct: 11 RHNKLDYDGSEDEKKT-KLCSLKKKAINASNKFKHSFTKRTRR--NSRVMSVSIVDDIDL 67
Query: 71 GEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTD 130
E V RQ LI LLP +HDD++ +LRFL+AR F++EK QMW +M+ WRKE+G D
Sbjct: 68 EELQAVDAFRQALILDELLPSKHDDHHMMLRFLRARKFDLEKAKQMWTDMIHWRKEFGVD 127
Query: 131 TILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQ 190
TI+E+F+F+E++EVL+YYPQGYHGVDK+GRPVYIERLG+ ++LM++TTIDRY+KYHV+
Sbjct: 128 TIMEDFDFKEIDEVLKYYPQGYHGVDKDGRPVYIERLGQVDATKLMQVTTIDRYVKYHVR 187
Query: 191 EFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPET 250
EFE+ K PACSIAAK+ I +TTILDVQG+G+K+F++ A +LL + KID++ YPET
Sbjct: 188 EFEKTFNIKLPACSIAAKKHIDQSTTILDVQGVGLKSFSKAARDLLQRIQKIDSDNYPET 247
Query: 251 LHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGS 310
L++M+I+NAGSGF ++LW + F DPKT AKI +L +K KLLE+IDS++LP+FLGG+
Sbjct: 248 LNRMFIINAGSGF-RLLWSTVKSFLDPKTTAKIHVLGNKYQSKLLEIIDSNELPEFLGGN 306
Query: 311 CTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQ 346
CTC +GGC+++ KGPWNDPDI K+V + E R+
Sbjct: 307 CTCADKGGCMRSDKGPWNDPDIFKMVQNGEGKCPRK 342
>At4g36490 hypothetical protein
Length = 543
Score = 383 bits (983), Expect = e-106
Identities = 215/591 (36%), Positives = 341/591 (57%), Gaps = 66/591 (11%)
Query: 21 EDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETVVHELR 80
+D ++G+ +K++ + + +++ + ++R + + + IEDV D E V R
Sbjct: 5 QDAELKPRMGSFKKRSSSKNLRYSMTKRRRSSKVMSVEI----IEDVHDAEELKAVDAFR 60
Query: 81 QRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEE 140
Q LI LLP +HDDY+ +LRFLKAR F++EKT QMW EML WRKE+G DT++EEF+F+E
Sbjct: 61 QSLILDELLPEKHDDYHMMLRFLKARKFDLEKTKQMWTEMLRWRKEFGADTVMEEFDFKE 120
Query: 141 LEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKF 200
++EVL+YYPQG+HGVDKEGRPVYIERLG ++LM++TT+DRY+ YHV EFER KF
Sbjct: 121 IDEVLKYYPQGHHGVDKEGRPVYIERLGLVDSTKLMQVTTMDRYVNYHVMEFERTFNVKF 180
Query: 201 PACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAG 260
PACSIAAK+ I +TTILDVQG+G+KNF + A +L+ + K+D + YPETL++M+I+NAG
Sbjct: 181 PACSIAAKKHIDQSTTILDVQGVGLKNFNKAARDLITRLQKVDGDNYPETLNRMFIINAG 240
Query: 261 SGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCL 320
SGF +MLW + F DPKT AKI +L +K KLLE+ID S+LP+FLGGSCTC GGC+
Sbjct: 241 SGF-RMLWNTVKSFLDPKTTAKIHVLGNKYQSKLLEIIDESELPEFLGGSCTCADNGGCM 299
Query: 321 KTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTAESGSD 380
++ KGPW +P+I+K VH+ + + S+ S + + + D ST E S+
Sbjct: 300 RSDKGPWKNPEIMKRVHNGD----HKCSKGSQAENSGEKTIPE------EDDSTTEPASE 349
Query: 381 INDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQFHLSQKQSL 440
+++S + + P +A + E +F LS+K
Sbjct: 350 --------EEKASKEVEIVP------------------AAHPAWNMPEAHKFSLSKK--- 380
Query: 441 QTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEF 500
+ ++ + +EG ++ ++ + VT+ + +
Sbjct: 381 EVYAIQEACNNATTEGGRSPIFTGVMALVMGVVTMIKVTKNVPRKL-------------- 426
Query: 501 WRTQNNVHPSNTTEHNINNHSAAVEA------AFERDHILPCAQRLQRLEKVFEELNNKP 554
T++ ++ S + + + +A+++ A + + +R+ LE+ L+ +P
Sbjct: 427 --TESTLYSSPVYCDDASMNKSAMQSEKMTVPAISGEDFMAIMKRMAELEQKVTVLSAQP 484
Query: 555 YGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQK 605
MP +KE+ML ++ R +E +L TK+ L ++ +Q E+ +E +K
Sbjct: 485 TVMPPDKEEMLNAAISRSNVLEQELAATKKALDDSLGRQEELVAYIEKKKK 535
>At2g18180 putative phosphatidylinositol/phophatidylcholine transfer
protein
Length = 558
Score = 377 bits (969), Expect = e-105
Identities = 229/584 (39%), Positives = 334/584 (56%), Gaps = 72/584 (12%)
Query: 41 SKFTHSLKKRGKRKIDYRVPAVSI-EDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTL 99
SK + SL K+ + +V +V I ED D E VV RQ LI LLP +HDDY+ +
Sbjct: 26 SKLSCSLTKKRRSS---KVMSVEIFEDEHDAEELKVVDAFRQVLILDELLPDKHDDYHMM 82
Query: 100 LRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEG 159
LRFLKAR F++EKT QMW +ML WRKE+G DT++E+FEF+E++EVL+YYPQG+HGVDKEG
Sbjct: 83 LRFLKARKFDLEKTNQMWSDMLRWRKEFGADTVMEDFEFKEIDEVLKYYPQGHHGVDKEG 142
Query: 160 RPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILD 219
RPVYIERLG+ ++LM++TT+DRY+ YHV EFER KFPACSIAAK+ I +TTILD
Sbjct: 143 RPVYIERLGQVDSTKLMQVTTMDRYVNYHVMEFERTFNVKFPACSIAAKKHIDQSTTILD 202
Query: 220 VQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKT 279
VQG+G+KNF + A +L+ + K+D + YPETL++M+I+NAGSGF +MLW + F DPKT
Sbjct: 203 VQGVGLKNFNKAARDLITRLQKVDGDNYPETLNRMFIINAGSGF-RMLWNTVKSFLDPKT 261
Query: 280 IAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSA 339
AKI +L +K KLLE+ID+S+LP+FLGGSCTC GGC+++ KGPWN+PDI+K V++
Sbjct: 262 TAKIHVLGNKYQSKLLEIIDASELPEFLGGSCTCADNGGCMRSDKGPWNNPDIMKRVNNG 321
Query: 340 EATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLA 399
+ + + S+A N +N S +++A + D S P
Sbjct: 322 D-HICSKRSQADNAGENIIS----------QGNNSAVEEAPETDQSQP------------ 358
Query: 400 PVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQFHL--SQKQSLQTNDMENVTHRTISEGT 457
+P VV + +++ + K SL D+ I E
Sbjct: 359 --------------------SPCQNVVVAHPAWNIPEAHKFSLSKRDV-----YAIQEAC 393
Query: 458 SVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIR-SLRFEFWRTQNNVH--PSNTTE 514
N E T V+A F+ ++T IR + T++ ++ P E
Sbjct: 394 KATN---------ESGRSPIFTGVMA-FVMGVVTMIRVTKNVPRKLTESTIYSSPVYCDE 443
Query: 515 HNINNHS----AAVEAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMD 570
+++N S + + +R+ LE+ L+ +P MP EKE+ML ++
Sbjct: 444 NSMNKSSMHGKKMATTTISGEDFMAVMKRMAELEQKVTNLSAQPATMPPEKEEMLNAAIS 503
Query: 571 RIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSNCRVSWSL 614
R +E +L TK+ L ++ +Q ++ VE +K V + +
Sbjct: 504 RADFLEQELAATKKALDDSLTRQEDLVAYVERKKKKKKLVRFQI 547
>At2g16380 putative phosphatidylinositol/phosphatidylcholine
transfer protein
Length = 582
Score = 367 bits (941), Expect = e-101
Identities = 172/326 (52%), Positives = 244/326 (74%), Gaps = 2/326 (0%)
Query: 16 DVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETV 75
D+ENSED R+ K+ +L++KA++AS++F +S KK+ +R V + +D+ + +
Sbjct: 5 DMENSEDGRKLVKMSSLKQKAISASNRFKNSFKKKTRRTSSKIVSVANTDDINGD-DYLS 63
Query: 76 VHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE 135
V RQ L+ LLPP+HDD + +LRFL+AR F+ EK QMW +ML WR ++G DTI+E+
Sbjct: 64 VEAFRQVLVLDDLLPPKHDDLHMMLRFLRARKFDKEKAKQMWSDMLQWRMDFGVDTIIED 123
Query: 136 FEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERA 195
FEFEE+++VL++YPQGYHGVDKEGRPVYIERLG+ ++L++ TT+DRY KYHV+EFE+
Sbjct: 124 FEFEEIDQVLKHYPQGYHGVDKEGRPVYIERLGQIDANKLLQATTMDRYEKYHVKEFEKM 183
Query: 196 LQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMY 255
+ KFP+CS AAK+ I +TTI DVQG+G+KNF ++A LL + KID + YPETL++M+
Sbjct: 184 FKIKFPSCSAAAKKHIDQSTTIFDVQGVGLKNFNKSARELLQRLLKIDNDNYPETLNRMF 243
Query: 256 IVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPT 315
I+NAG GF ++LW +KF DPKT +KI +L +K KLLE ID+S+LP F GG CTC
Sbjct: 244 IINAGPGF-RLLWAPIKKFLDPKTTSKIHVLGNKYQPKLLEAIDASELPYFFGGLCTCAD 302
Query: 316 EGGCLKTSKGPWNDPDIIKLVHSAEA 341
+GGCL++ KGPWNDP+++K+ + EA
Sbjct: 303 KGGCLRSDKGPWNDPELLKIARNPEA 328
Score = 38.5 bits (88), Expect = 0.011
Identities = 19/68 (27%), Positives = 38/68 (54%)
Query: 538 QRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIA 597
+R+ LE+ ++ L++K LEK+ L +++R++ +E +L +TK+ L + Q I
Sbjct: 491 KRMAELEEKYKSLDSKSADEALEKDDKLQAALNRVQVLEHELSETKKALDETMVNQQGIL 550
Query: 598 DLVENLQK 605
+E K
Sbjct: 551 AYIEKKNK 558
>At5g47510 putative protein
Length = 403
Score = 282 bits (721), Expect = 4e-76
Identities = 131/268 (48%), Positives = 187/268 (68%), Gaps = 1/268 (0%)
Query: 65 EDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWR 124
++ + E +V R L+ G LP +H D+ TL RFLK RDF++EK+ + + + WR
Sbjct: 17 KEEQSPNNEEMVEAFRNLLLLHGHLPDKHGDHNTLRRFLKMRDFDLEKSKEAFLNYMKWR 76
Query: 125 KEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRY 184
+Y D I ++F+FEE EV ++YP G+H VDK GRP+YIERLG + ++ TTI+RY
Sbjct: 77 VDYKVDLISQKFKFEEYGEVKKHYPHGFHKVDKTGRPIYIERLGMTDLNAFLKATTIERY 136
Query: 185 LKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDT 244
+ YH++E E+ + ++PACSIA+ + + STTTILDV G+GM NF++ A +L + KID+
Sbjct: 137 VNYHIKEQEKTMSLRYPACSIASDKHVSSTTTILDVSGVGMSNFSKPARSLFMEIQKIDS 196
Query: 245 NYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLP 304
NYYPETLH++++VNA SGF +MLW A + F D +T+AK+Q+L L +LLE I+ S LP
Sbjct: 197 NYYPETLHRLFVVNASSGF-RMLWLALKTFLDARTLAKVQVLGPNYLGELLEAIEPSNLP 255
Query: 305 DFLGGSCTCPTEGGCLKTSKGPWNDPDI 332
FLGG+CTC GGCL + +GPWNDP I
Sbjct: 256 TFLGGNCTCSDHGGCLFSDEGPWNDPGI 283
>At1g55840 polyphosphoinositide binding protein, putative
Length = 325
Score = 82.4 bits (202), Expect = 7e-16
Identities = 67/218 (30%), Positives = 99/218 (44%), Gaps = 11/218 (5%)
Query: 93 HDDYYT--LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE--FEFEELEEVLQYY 148
H Y T LLRFLKARD N++K +M E L WR + D IL + + +
Sbjct: 31 HQGYPTENLLRFLKARDGNVQKAHKMLLECLEWRTQNEIDKILTKPIVPVDLYRGIRDTQ 90
Query: 149 PQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAK 208
G G KEG PV +G + + ++ Y++ H+Q E + P+ S
Sbjct: 91 LVGVSGYSKEGLPVIAIGVGLSTYDK----ASVHYYVQSHIQMNEYRDRVVLPSASKKQG 146
Query: 209 RRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLW 268
R I + ILD+ GL + ++ L+ A+T ID YPE Y+VN F W
Sbjct: 147 RPICTCLKILDMSGLKLSALSQ--IKLMTAITTIDDLNYPEKTETYYVVNVPYIF-SACW 203
Query: 269 PAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDF 306
+ +T KIQ+L +LL+++D LP F
Sbjct: 204 KTIKPLLQERTKKKIQVLKGCGKDELLKIMDYESLPHF 241
>At5g47730 putative protein
Length = 341
Score = 77.4 bits (189), Expect = 2e-14
Identities = 71/248 (28%), Positives = 113/248 (44%), Gaps = 18/248 (7%)
Query: 65 EDVRDEGEETV--VHELRQRLIERGLLPPRHDDYY--TLLRFLKARDFNIEKTIQMWEEM 120
E+ DE +E + V E ++ ER H Y L RFLKARD+N+ K M E
Sbjct: 6 EEAIDEFQELMDQVEEPLKKTYERV-----HQGYLRENLGRFLKARDWNVCKAHTMLVEC 60
Query: 121 LLWRKEYGTDTILEE--FEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRI 178
L WR + D+IL + E +V G G KEG PV+ +G + +
Sbjct: 61 LRWRVDNEIDSILSKPIVPTELYRDVRDSQLIGMSGYTKEGLPVFAIGVGLSTFDK---- 116
Query: 179 TTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAA 238
++ Y++ H+Q E + P+ S R I + +LD+ GL + ++ L+
Sbjct: 117 ASVHYYVQSHIQINEYRDRVLLPSISKKNGRPITTCVKVLDMTGLKLSALSQ--IKLVTI 174
Query: 239 MTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVI 298
++ ID YPE + Y+VNA F W + +T K+ +L +LL+++
Sbjct: 175 ISTIDDLNYPEKTNTYYVVNAPYIF-SACWKVVKPLLQERTRKKVHVLSGCGRDELLKIM 233
Query: 299 DSSQLPDF 306
D + LP F
Sbjct: 234 DFTSLPHF 241
>At1g01630 polyphosphoinositide binding protein, putative
Length = 255
Score = 75.9 bits (185), Expect = 6e-14
Identities = 77/250 (30%), Positives = 111/250 (43%), Gaps = 33/250 (13%)
Query: 67 VRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKE 126
+ DE E + V +R L +R + D + RFL+ARD +IEK M+ L W++
Sbjct: 22 IEDEIERSKVGIMRA-LCDRQDPETKEVDDLMIRRFLRARDLDIEKASTMFLNYLTWKRS 80
Query: 127 YGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLK 186
+ E E+ L + G DK GRP+ + + +PS+ D + +
Sbjct: 81 MLPKGHIPE---AEIANDLSHNKMCMQGHDKMGRPIAVAIGNRHNPSK----GNPDEFKR 133
Query: 187 YHVQEFERAL------QEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMT 240
+ V E+ QEKF A I D+QG G N LAA++
Sbjct: 134 FVVYTLEKICARMPRGQEKFVA--------------IGDLQGWGYSNC--DIRGYLAALS 177
Query: 241 KIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLY-KLLEVID 299
+ + YPE L ++YIV+A F W F D T KI +++K L LLE ID
Sbjct: 178 TLQ-DCYPERLGKLYIVHAPYIF-MTAWKVIYPFIDANTKKKIVFVENKKLTPTLLEDID 235
Query: 300 SSQLPDFLGG 309
SQLPD GG
Sbjct: 236 ESQLPDIYGG 245
>At1g72160 cytosolic factor, putative
Length = 490
Score = 67.8 bits (164), Expect = 2e-11
Identities = 59/228 (25%), Positives = 105/228 (45%), Gaps = 26/228 (11%)
Query: 99 LLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKE 158
LL+FL+AR+F ++ + M + + WRKE+ D ++EE ++L++V+ HG D+E
Sbjct: 167 LLKFLRAREFKVKDSFAMLKNTIKWRKEFKIDELVEEDLVDDLDKVV-----FMHGHDRE 221
Query: 159 GRPVYIERLGKAHPSRLMRITTID-----RYLKYHVQEFERALQE-KFPACSIAAKRRIF 212
G PV G+ L T D +L+ +Q ER++++ F + ++ IF
Sbjct: 222 GHPVCYNVYGEFQNKELYNKTFSDEEKRKHFLRTRIQFLERSIRKLDFSSGGVST---IF 278
Query: 213 STTTILDVQGLGMKNF---TRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWP 269
+ + GLG K T+ A LL + YPE + + +N + + +
Sbjct: 279 QVNDMKNSPGLGKKELRSATKQAVELL-------QDNYPEFVFKQAFINV-PWWYLVFYT 330
Query: 270 AAQKFCDPKTIAKIQIL-DSKSLYKLLEVIDSSQLPDFLGGSCTCPTE 316
F P++ +K+ S+S L + I Q+P GG P +
Sbjct: 331 VIGPFMTPRSKSKLVFAGPSRSAETLFKYISPEQVPVQYGGLSVDPCD 378
>At1g75170 unknown protein
Length = 296
Score = 60.8 bits (146), Expect = 2e-09
Identities = 65/253 (25%), Positives = 111/253 (43%), Gaps = 38/253 (15%)
Query: 70 EGEETVVHELRQRLIER--GLLPPRHDDYYT---LLRFLKARDFNIEKTIQMWEEMLLWR 124
E +E + E + + ++ G L R+ Y + L R+L+AR++N+ K +M EE L WR
Sbjct: 13 EQKEAALREAKMKELKTLIGQLSGRNSLYCSDACLKRYLEARNWNVGKAKKMLEETLKWR 72
Query: 125 KEYGTDTILEEFEFEELE---EVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTI 181
+ EE + E+ E + Y G+H D+ GR V I R G L ++
Sbjct: 73 SSFKP----EEIRWNEVSGEGETGKVYKAGFH--DRHGRTVLILRPG------LQNTKSL 120
Query: 182 DRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTK 241
+ +K+ V E A+ + + ++D G M T+ + +A
Sbjct: 121 ENQMKHLVYLIENAI--------LNLPEDQEQMSWLIDFTGWSMS----TSVPIKSARET 168
Query: 242 ID--TNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQIL---DSKSLYKLLE 296
I+ N+YPE L ++ N F + W + F D KT K++ + +S+S+ +
Sbjct: 169 INILQNHYPERLAVAFLYNPPRLF-EAFWKIVKYFIDAKTFVKVKFVYPKNSESVELMST 227
Query: 297 VIDSSQLPDFLGG 309
D LP GG
Sbjct: 228 FFDEENLPTEFGG 240
>At4g08690 putative phosphoglyceride transfer protein
Length = 301
Score = 59.7 bits (143), Expect = 5e-09
Identities = 61/228 (26%), Positives = 102/228 (43%), Gaps = 27/228 (11%)
Query: 64 IEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYT---LLRFLKARDFNIEKTIQMWEEM 120
++ V E E+ + E+R+ L G LP + + + +LR+L+AR+++++K +M +E
Sbjct: 11 VKPVPTEEEQAKIEEVRKLL---GPLPEKLSSFCSDDAVLRYLRARNWHVKKATKMLKET 67
Query: 121 LLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITT 180
L WR +Y + I E E E Y VDK GRPV I R PS + +
Sbjct: 68 LKWRVQYKPEEICWEEVAGEAETGKIYRSS---CVDKLGRPVLIMR-----PS-VENSKS 118
Query: 181 IDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMT 240
+ ++Y V E A+Q P ++D G + N + A +
Sbjct: 119 VKGQIRYLVYCMENAVQNLPPGEE--------QMVWMIDFHGYSLANVSLRTTKETAHVL 170
Query: 241 KIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDS 288
+ +YPE L + N F+ W A+ F +PKT K++ + S
Sbjct: 171 Q---EHYPERLAFAVLYNPPKFFEP-FWKVARPFLEPKTRNKVKFVYS 214
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,675,393
Number of Sequences: 26719
Number of extensions: 587437
Number of successful extensions: 2177
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 2068
Number of HSP's gapped (non-prelim): 78
length of query: 619
length of database: 11,318,596
effective HSP length: 105
effective length of query: 514
effective length of database: 8,513,101
effective search space: 4375733914
effective search space used: 4375733914
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0047c.1


