

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0047b.8
(171 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
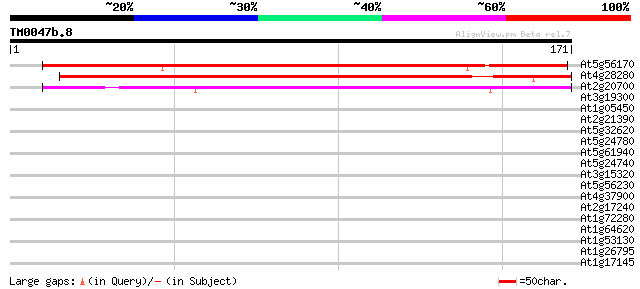
At5g56170 predicted GPI-anchored protein 207 3e-54
At4g28280 putative GPI-anchored protein 157 3e-39
At2g20700 predicted GPI-anchored protein 142 7e-35
At3g19300 unknown protein 34 0.044
At1g05450 hypothetical protein 30 0.83
At2g21390 coatomer alpha subunit 28 1.9
At5g32620 putative protein 28 3.2
At5g24780 vegetative storage protein Vsp1 27 4.1
At5g61940 putative protein 27 7.1
At5g24740 VPS13 - like protein 27 7.1
At3g15320 hypothetical protein 27 7.1
At5g56230 putative protein 26 9.2
At4g37900 putative protein 26 9.2
At2g17240 unknown protein 26 9.2
At1g72280 like disulfide bond formation protein 26 9.2
At1g64620 zinc finger protein, putative 26 9.2
At1g53130 unknown protein 26 9.2
At1g26795 unknown protein 26 9.2
At1g17145 unknown protein 26 9.2
>At5g56170 predicted GPI-anchored protein
Length = 168
Score = 207 bits (526), Expect = 3e-54
Identities = 104/164 (63%), Positives = 123/164 (74%), Gaps = 5/164 (3%)
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHT-GRNLLQAKKGCSVNFEFLNYTIIT 69
L S L L LSV SS SSS+F+SD VF SQ+ GRNLLQ KK C VNFEF+NYTIIT
Sbjct: 3 LLSRALFFFLLLSVLSSFSSSSFISDGVFESQSLVLGRNLLQTKKTCPVNFEFMNYTIIT 62
Query: 70 SKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECRE 129
SKCKGP YPPK+CCGAFK+FACPY D LNDL++DCA+TMFSYINLYGKYPPGLFA++C+E
Sbjct: 63 SKCKGPKYPPKECCGAFKDFACPYTDQLNDLSSDCATTMFSYINLYGKYPPGLFANQCKE 122
Query: 130 GKEGLACPA---LPPSALADDTSSQVVHFPSLVLVLTACFLILL 170
GKEGL CPA LPP A + ++ L L ++A L+ +
Sbjct: 123 GKEGLECPAGSQLPPETSA-EVNAATTSSSRLWLTVSAALLVFV 165
>At4g28280 putative GPI-anchored protein
Length = 160
Score = 157 bits (396), Expect = 3e-39
Identities = 83/159 (52%), Positives = 103/159 (64%), Gaps = 9/159 (5%)
Query: 16 LLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITSKCKGP 75
L+ +L++ + S + S +S F S T R LLQAK C +F NYTIITSKCKGP
Sbjct: 8 LVSLLSILLLSGFAFSHHISLDEFESHPSTSRALLQAKATCKEDFAAKNYTIITSKCKGP 67
Query: 76 SYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAHECREGKEGLA 135
+YP K CC AFK+FACP+ +VLND DCASTMFSYINLYG+YPPG+FA+ C+EGKEGL
Sbjct: 68 NYPAKVCCSAFKDFACPFAEVLNDEKTDCASTMFSYINLYGRYPPGIFANMCKEGKEGLD 127
Query: 136 CPALPPSALADDTSSQVVHFPSL---VLVLTACFLILLF 171
C + P TSS P + VL++T L LF
Sbjct: 128 CTDVTP------TSSSHASIPLVSTHVLLITVSILFHLF 160
>At2g20700 predicted GPI-anchored protein
Length = 181
Score = 142 bits (359), Expect = 7e-35
Identities = 83/180 (46%), Positives = 103/180 (57%), Gaps = 23/180 (12%)
Query: 11 LSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKG--------------- 55
+S LL +L + + S S LS F A T R LLQ +
Sbjct: 3 ISPYCLLSLLPIFLLSGFS----LSYDEFDGHAATSRALLQTRTNTLIFKYGSSVFFVGI 58
Query: 56 ---CSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYI 112
C +F NYTIITS+CKGP+YP CC AFK+FACP+ +VLND NDCASTMFSYI
Sbjct: 59 ETACKEDFANKNYTIITSRCKGPNYPANVCCSAFKDFACPFAEVLNDEKNDCASTMFSYI 118
Query: 113 NLYGKYPPGLFAHECREGKEGLACPALPPSALA-DDTSSQVVHFPSLVLVLTACFLILLF 171
NLYG+YPPG+FA+ C+EGKEGL C + SA A D+ + SL ++ T L LLF
Sbjct: 119 NLYGRYPPGIFANMCKEGKEGLDCTDVTQSASATSDSIPRASTTASLAVLSTFLVLCLLF 178
>At3g19300 unknown protein
Length = 663
Score = 33.9 bits (76), Expect = 0.044
Identities = 22/77 (28%), Positives = 33/77 (42%), Gaps = 11/77 (14%)
Query: 50 LQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPYV----------DVLND 99
L + GC ++F N+T++ S C + K CC F V V +D
Sbjct: 24 LLTEAGCPLDFTSSNFTLVASVCSNNTERAK-CCRYMNAFVAISVARYANYTADLGVTSD 82
Query: 100 LTNDCASTMFSYINLYG 116
LT C +T+ + LYG
Sbjct: 83 LTEICITTISRTMELYG 99
>At1g05450 hypothetical protein
Length = 206
Score = 29.6 bits (65), Expect = 0.83
Identities = 24/89 (26%), Positives = 41/89 (45%), Gaps = 20/89 (22%)
Query: 9 FLLSSSLLLLILALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTII 68
FL+S++L+ +L SS+S ++ L+ ++L + GC+ + +
Sbjct: 6 FLISAALIFSLL-----SSNSPTSILAQI----NTPCSPSMLSSVTGCT--------SFL 48
Query: 69 TSKCKGPSYPPKDCCGAFKEFACPYVDVL 97
T G S+P DCCGA K +D L
Sbjct: 49 TG---GGSFPTSDCCGALKSLTGTGMDCL 74
>At2g21390 coatomer alpha subunit
Length = 1218
Score = 28.5 bits (62), Expect = 1.9
Identities = 18/58 (31%), Positives = 28/58 (48%), Gaps = 11/58 (18%)
Query: 41 SQAHTGRNLLQAKK-----GCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPY 93
SQA T R ++QA + ++N++F N +I P Y + K+ ACPY
Sbjct: 1130 SQARTARQVMQAAERNMTDATTLNYDFRNPFVICGSTYVPIYKGQ------KDVACPY 1181
>At5g32620 putative protein
Length = 301
Score = 27.7 bits (60), Expect = 3.2
Identities = 12/40 (30%), Positives = 20/40 (50%)
Query: 119 PPGLFAHECREGKEGLACPALPPSALADDTSSQVVHFPSL 158
PPG+ A + + K + P+ + DD+ V HF +L
Sbjct: 215 PPGVKAAKAKGRKSAIVKEGKKPATVKDDSGQSVEHFQNL 254
>At5g24780 vegetative storage protein Vsp1
Length = 270
Score = 27.3 bits (59), Expect = 4.1
Identities = 23/83 (27%), Positives = 37/83 (43%), Gaps = 24/83 (28%)
Query: 10 LLSSSLLLLILALSVSSSSSSST------FLSDAVFGSQAHTGRNLLQAKKGCSVNF--- 60
+LS SLLLL+ A SS+S S+ +FG++A L K+G S+N+
Sbjct: 3 ILSLSLLLLLAATVSHVQSSASVPGLIELLESNTIFGNEAE-----LLEKEGLSINYPNC 57
Query: 61 ----------EFLNYTIITSKCK 73
+N+ + + CK
Sbjct: 58 RSWHLGVETSNIINFDTVPANCK 80
>At5g61940 putative protein
Length = 1094
Score = 26.6 bits (57), Expect = 7.1
Identities = 14/37 (37%), Positives = 23/37 (61%)
Query: 23 SVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVN 59
++S +SS FL + GS + ++L +AK+G SVN
Sbjct: 87 TLSPICASSLFLLGQMLGSVLYYKKSLKKAKQGLSVN 123
>At5g24740 VPS13 - like protein
Length = 3306
Score = 26.6 bits (57), Expect = 7.1
Identities = 18/68 (26%), Positives = 28/68 (40%), Gaps = 6/68 (8%)
Query: 34 LSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITSKCKGPSYPPKDCCGAFKEFACPY 93
L D+ S+AH+ +L G F Y PS+ P CC + + + P
Sbjct: 1972 LGDSYLWSEAHSISKVLSQDSGIGFRRSFACYPC------HPSHEPFRCCISVQSTSLPA 2025
Query: 94 VDVLNDLT 101
+NDL+
Sbjct: 2026 SFHINDLS 2033
>At3g15320 hypothetical protein
Length = 287
Score = 26.6 bits (57), Expect = 7.1
Identities = 12/40 (30%), Positives = 19/40 (47%)
Query: 119 PPGLFAHECREGKEGLACPALPPSALADDTSSQVVHFPSL 158
PPG+ A + + K P+ + DD+ V HF +L
Sbjct: 201 PPGVKAAKAKARKSATIKEGKKPATVKDDSGQSVEHFQNL 240
>At5g56230 putative protein
Length = 186
Score = 26.2 bits (56), Expect = 9.2
Identities = 17/53 (32%), Positives = 25/53 (47%), Gaps = 2/53 (3%)
Query: 20 LALSVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITSKC 72
L ++SS S + F LL++K + N+ F+NYTII S C
Sbjct: 28 LTTAISSHRPWSELIFSGDFSLPESFSSLLLRSKT--NFNYFFVNYTIIVSTC 78
>At4g37900 putative protein
Length = 787
Score = 26.2 bits (56), Expect = 9.2
Identities = 10/18 (55%), Positives = 12/18 (66%)
Query: 65 YTIITSKCKGPSYPPKDC 82
YT +S C+GP PP DC
Sbjct: 67 YTESSSICQGPLVPPLDC 84
>At2g17240 unknown protein
Length = 140
Score = 26.2 bits (56), Expect = 9.2
Identities = 21/57 (36%), Positives = 26/57 (44%), Gaps = 6/57 (10%)
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSS------SSSSTFLSDAVFGSQAHTGRNLLQ 51
MA + RFL S SL +L +SS SSS F +V S + RN LQ
Sbjct: 1 MASLCSARFLPSPSLDILSTKTKISSDSRSLGCSSSRVFAFSSVHRSSSRRNRNQLQ 57
>At1g72280 like disulfide bond formation protein
Length = 469
Score = 26.2 bits (56), Expect = 9.2
Identities = 26/99 (26%), Positives = 44/99 (44%), Gaps = 27/99 (27%)
Query: 13 SSLLLLILALSVSSSSSSST--FLSDAVFGSQAHTGRNLLQAKKGCSVNFEFLNYTIITS 70
++L+++ LA++VSS ++S+ F SD + CS + + T
Sbjct: 23 ATLVVVFLAVAVSSRTNSNVGFFFSD----------------RNSCSCSLQK------TG 60
Query: 71 KCKGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMF 109
K KG +DCC ++ +VLN L D +T F
Sbjct: 61 KYKGMI---EDCCCDYETVDNLNTEVLNPLLQDLVTTPF 96
>At1g64620 zinc finger protein, putative
Length = 352
Score = 26.2 bits (56), Expect = 9.2
Identities = 18/63 (28%), Positives = 25/63 (39%), Gaps = 1/63 (1%)
Query: 67 IITSKC-KGPSYPPKDCCGAFKEFACPYVDVLNDLTNDCASTMFSYINLYGKYPPGLFAH 125
I+T+ C K S PP ++ LN + +T F Y N Y P F
Sbjct: 18 IVTNTCLKQQSNPPSPATPVERKARPEKDQALNCPRCNSLNTKFCYYNNYSLTQPRYFCK 77
Query: 126 ECR 128
+CR
Sbjct: 78 DCR 80
>At1g53130 unknown protein
Length = 168
Score = 26.2 bits (56), Expect = 9.2
Identities = 13/38 (34%), Positives = 23/38 (60%)
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSSSSSSTFLSDAV 38
M + F+ ++SLL LIL + ++S S++ L+D V
Sbjct: 1 MVIKIPNTFIKATSLLSLILYFLIIATSKSNSVLADEV 38
>At1g26795 unknown protein
Length = 151
Score = 26.2 bits (56), Expect = 9.2
Identities = 14/31 (45%), Positives = 17/31 (54%)
Query: 1 MAFSLNQRFLLSSSLLLLILALSVSSSSSSS 31
MAFS NQ F+ L IL S S ++ SS
Sbjct: 1 MAFSTNQNFIFVLFLFFFILKTSASLTNHSS 31
>At1g17145 unknown protein
Length = 335
Score = 26.2 bits (56), Expect = 9.2
Identities = 12/34 (35%), Positives = 19/34 (55%)
Query: 23 SVSSSSSSSTFLSDAVFGSQAHTGRNLLQAKKGC 56
S SSSSS+S D +GS A ++ +++ C
Sbjct: 158 SYSSSSSTSAATLDTEYGSAAEDDEEIVSSQESC 191
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.137 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,773,750
Number of Sequences: 26719
Number of extensions: 151583
Number of successful extensions: 444
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 430
Number of HSP's gapped (non-prelim): 20
length of query: 171
length of database: 11,318,596
effective HSP length: 92
effective length of query: 79
effective length of database: 8,860,448
effective search space: 699975392
effective search space used: 699975392
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0047b.8


