

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046b.4
(1706 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
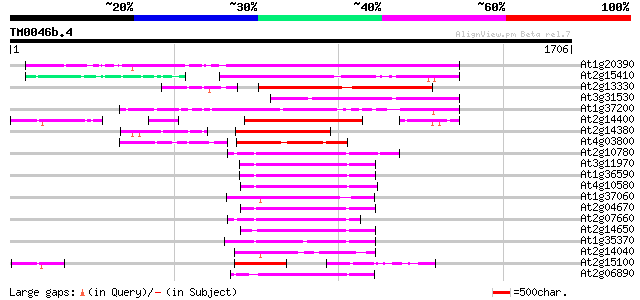
Score E
Sequences producing significant alignments: (bits) Value
At1g20390 hypothetical protein 771 0.0
At2g15410 putative retroelement pol polyprotein 544 e-154
At2g13330 F14O4.9 470 e-132
At3g31530 hypothetical protein 439 e-123
At1g37200 hypothetical protein 394 e-109
At2g14400 putative retroelement pol polyprotein 385 e-107
At2g14380 putative retroelement pol polyprotein 341 2e-93
At4g03800 321 3e-87
At2g10780 pseudogene 229 1e-59
At3g11970 hypothetical protein 219 1e-56
At1g36590 hypothetical protein 219 1e-56
At4g10580 putative reverse-transcriptase -like protein 214 3e-55
At1g37060 Athila retroelment ORF 1, putative 213 1e-54
At2g04670 putative retroelement pol polyprotein 211 2e-54
At2g07660 putative retroelement pol polyprotein 206 9e-53
At2g14650 putative retroelement pol polyprotein 203 6e-52
At1g35370 hypothetical protein 193 8e-49
At2g14040 putative retroelement pol polyprotein 184 4e-46
At2g15100 putative retroelement pol polyprotein 182 1e-45
At2g06890 putative retroelement integrase 181 2e-45
>At1g20390 hypothetical protein
Length = 1791
Score = 771 bits (1992), Expect = 0.0
Identities = 495/1371 (36%), Positives = 721/1371 (52%), Gaps = 134/1371 (9%)
Query: 49 PFVPAITRVDIPKHLQTMALDAFSGESDPMEHLRYFNTKMVIGGAT----DAVKCRLLPS 104
PF IT V I + Q + L++++G +DP E L FN + T DA +C++
Sbjct: 156 PFTRRITNVSI-RGAQKIKLESYNGRNDPKEFLTSFNVAINRAELTIDNFDAGRCQIFIE 214
Query: 105 TFKGMAMQWFIRQPPFSIDNFTDLSTKFLTQFSANKTTKATMFDLISIHQQPGEKLKTYM 164
G A WF R P SID+F L++ FL ++ + + DL SI Q E L++++
Sbjct: 215 HLTGPAHNWFSRLKPNSIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFV 274
Query: 165 ARFSKMAVQLEDENPDVCLASFKNGLRAGDLNRD-LTRRPAKDMLDLRARVQEFILIEQD 223
RF K+ V + + + +N + RD +T + D R FI +E++
Sbjct: 275 DRF-KLVVTNITVPDEAAIVALRNAVWYDSRFRDDITLHAPSTLEDALHRASRFIELEEE 333
Query: 224 DQKKQEREEGRKQSQSGGVSQDKSKAGKETRIAQTPRVPRPGPYQNSKPGFQNTWHRNPQ 283
+ K T + V + GP +++P + RNP
Sbjct: 334 KLILARKHNSTK-----------------TPACKDAVVIKVGPDDSNEP--RQHLDRNP- 373
Query: 284 GPPAATAGQTGASPAPVPLTKLNAPLSTILRAVGQTNVVQYPPPPRRPPANVDTNRWCEY 343
+AG+ S T P + +R P P +CEY
Sbjct: 374 -----SAGRKPTSFLVSTETPDAKPWNKYIRDADS---------PAAGPM------YCEY 413
Query: 344 HKALGHTTDNCWNLRREIDRLIKAGHLA---------------------NFVKDTAAP-- 380
HK+ H+T+NC L+ + K+G + ++ D P
Sbjct: 414 HKSRAHSTENCRFLQGLLMAKYKSGGITIECDRPPINNKNQRRNETTARQYLNDQTKPPT 473
Query: 381 --EVAKITQGD------KGKGKEIVEELGDPVGECSSIAGGFGGGAISSKARKRYVAAVH 432
E IT D + GK I E V + I GG + S ++ K+Y
Sbjct: 474 PAEQGIITSADDPAAKRQRNGKAIAAE-PVVVRQVHVIMGGLQNCSDSVRSIKQYRKKAE 532
Query: 433 SVQESNEGECWVNH----SPIIFTPQDFAHVIPHDNDPIVVTIRVNNYVTKKVFLDQGSS 488
V + +PI FT D + NDP+VV + +++ +V +D GSS
Sbjct: 533 MVVAWPSSTSTTRNPNQSAPISFTDVDLEGLDTPHNDPLVVELIISDSRVTRVLIDTGSS 592
Query: 489 ADIIYGDAFDRLGLKESDLKPYKGTLVGFTGDRVSVRGYVEIPTAFGEGEFVKKFQVKYL 548
D+I+ D + + + +KP L GF GD V G +++P G G VK++
Sbjct: 593 VDLIFKDVLTAMNITDRQIKPVSKPLAGFDGDFVMTIGTIKLPIFVG-GLIA---WVKFV 648
Query: 549 VLACRANYNVLLGRDTLNKVCAVISTAHLTVKYPACNGKVGILRVDQNAARECYLRSVAL 608
V+ A YNV+LG ++++ A+ ST H VK+P NG + LR + A
Sbjct: 649 VIGKPAVYNVILGTPWIHQMQAIPSTYHQCVKFPTHNG-IFTLRAPKEA----------- 696
Query: 609 YGRKAAKESHRITEIFPQEGFSLDPRDDADDFRPQPLEETKQVQVKDKFLKIGSGLTTEQ 668
K S+ +E+ E ++D D + + +G+ ++
Sbjct: 697 ---KTPSRSYEESELCRTEMVNIDESDPT------------------RCVGVGAEISPSI 735
Query: 669 EDRLITLLGDNLDLFAWTINDVPGIDPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQL 728
LI LL N FAW+I D+ GIDP + H+L + P PV Q RR++ E+ +AV
Sbjct: 736 RLELIALLKRNSKTFAWSIEDMKGIDPAITAHELNVDPTFKPVKQKRRKLGPERARAVNE 795
Query: 729 ETEKLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLV 788
E EKL+KA I EV+YP WLAN V+VKK NGKWR+C DYT LNK CPKDSYPLP++D+LV
Sbjct: 796 EVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPHIDRLV 855
Query: 789 DGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLM 848
+ SGN LLS MDA+SGYNQI+MH D+E T+F+T++ YCYK M FGLKNAGATYQR +
Sbjct: 856 EATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTDRGTYCYKVMSFGLKNAGATYQRFV 915
Query: 849 DKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGK 908
+K+ + Q+GR +EVY+DDM+VKS + DH L + FD L TY MKLNP KC+FG+ G+
Sbjct: 916 NKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHLSKCFDVLNTYGMKLNPTKCTFGVTSGE 975
Query: 909 FLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFFTC 968
FLG+++T RGIE NP + RAILE+ SP + +EVQRLTGR+AAL+RF+ + DK PF+
Sbjct: 976 FLGYVVTKRGIEANPKQIRAILELPSPRNAREVQRLTGRIAALNRFISRSTDKCLPFYNL 1035
Query: 969 LKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEEGK 1028
LK+ ++F W ++ E+AF KLK+ L+T P+L KP L LY+AV+D AVS+VL++E+
Sbjct: 1036 LKRRAQFDWDKDSEEAFEKLKDYLSTPPILVKPEVGETLYLYIAVSDHAVSSVLVREDRG 1095
Query: 1029 KQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTDVPLRQVLQKP 1088
+Q+ I++ S +L AE RY IEKAALA++ +AR+LRPYFQS + + TD PLR L P
Sbjct: 1096 EQRPIFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTDQPLRVALHSP 1155
Query: 1089 DLSGRLVSWSVELSEYDIQYEPRGQVTVQSLIDFVAELTPTEGEKTQG-----EWVLSVD 1143
SGR+ W+VELSEYDI + PR + Q L DF+ EL E+ EW L VD
Sbjct: 1156 SQSGRMTKWAVELSEYDIDFRPRPAMKSQVLADFLIELPLQSAERAVSGNRGEEWSLYVD 1215
Query: 1144 GSSNNTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGLRLAIELGVQKLFIK 1203
GSS+ GSG GI + SP ++EQS + F A+NN +EYE LIAGLRLA + + +
Sbjct: 1216 GSSSARGSGIGIRLVSPTAEVLEQSFRLRFVATNNVAEYEVLIAGLRLAAGMQITTIHAF 1275
Query: 1204 GDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRGQNERADVLAKLAST 1263
DSQL+ Q+ GEY+ K+ ++ YL++V+ + + + K+ +PRG N AD LA LA T
Sbjct: 1276 TDSQLIAGQLSGEYEAKNEKMDAYLKIVQLMTKDFENFKLSKIPRGDNAPADALAALALT 1335
Query: 1264 GRLGNYQTVIQETLPRPSI---DLVEIKLKAVKSVSEGE----ISWMESIKTFL-ESPPK 1315
+ + E++ +PSI D VEI + ++S + + W I+ +L +
Sbjct: 1336 SDSDLRRIIPVESIDKPSIDSTDAVEI-VNTIRSSNAPDPADPTDWRVEIRDYLSDGTLP 1394
Query: 1316 EDDLNTRTKRREASFYTLVGGELYRRGIMSPMLKCVDIKDALGIMAEVHEG 1366
D R R +A+ YTL+ L + ML C+ + IM E HEG
Sbjct: 1395 SDKWTARRLRIKAAKYTLMKEHLLKVSAFGAMLNCLHGTEINEIMKETHEG 1445
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 544 bits (1402), Expect = e-154
Identities = 316/792 (39%), Positives = 444/792 (55%), Gaps = 120/792 (15%)
Query: 639 DFRPQPLEETK-QVQVKD----KFLKIGSGLTTEQEDRLITLLGDNLDLFAWTINDVPGI 693
D RP P + T QV + + + + I L +E ++ L+ L N FAW++ D+PGI
Sbjct: 706 DERPNPQKGTVVQVNIDESDPSRCVGIRIDLPSELQNELVNFLRQNAATFAWSVEDMPGI 765
Query: 694 DPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVM 753
D V H+L + P P+ Q RR++ ++ K V E +KL+ A I EV+YP WL N V+
Sbjct: 766 DSAVTCHELNVDPTYKPLKQKRRKLGPDRTKDVNEEVKKLLDAGSIVEVRYPDWLRNPVV 825
Query: 754 VKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHP 813
VKK NGKWR+C D+T LNK CPKDS+PLP++D+LV+ +GNELLS MDA+SGYNQI+MH
Sbjct: 826 VKKKNGKWRVCIDFTDLNKACPKDSFPLPHIDRLVEATAGNELLSFMDAFSGYNQILMHQ 885
Query: 814 SDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSAR 873
+D E T F+T+Q YCYK MPFGLKNAGATY RL++++F+ Q+ +MEVY+DDM+VKS R
Sbjct: 886 NDREKTVFITDQGTYCYKVMPFGLKNAGATYPRLVNQMFTDQLDHSMEVYIDDMLVKSLR 945
Query: 874 ASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMK 933
A +H L++ F L Y MKLNP KC+FG+ G+FLG+++T RGIE NP + AI+++
Sbjct: 946 AEEHITHLRQCFQVLNRYNMKLNPSKCTFGVTSGEFLGYLVTRRGIEANPKQISAIIDLP 1005
Query: 934 SPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLA 993
SP + +EVQRL GR+AAL+RF+ + DK PF+ L+ N +F+W E+CE+ T
Sbjct: 1006 SPRNTREVQRLIGRIAALNRFISRSTDKCLPFYQLLRANKRFEWDEKCEEGET------- 1058
Query: 994 TLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKA 1053
L LY+AV+ AVS VL++E+ +Q I++VS TL GAELRY +EK
Sbjct: 1059 -------------LYLYIAVSTSAVSGVLVREDRGEQHPIFYVSKTLDGAELRYPTLEKL 1105
Query: 1054 ALAILKTARRLRPYFQSFQVKIKTDVPLRQVLQKPDLSGRLVSWSVELSEYDIQYEPRGQ 1113
A A++ +AR+LRPYF+S+ V++ T+ PLR +L P SGRL W+VELSEYDI Y+ R
Sbjct: 1106 AFAVVISARKLRPYFKSYTVEVLTNQPLRTILHSPSQSGRLAKWAVELSEYDIAYKNRTC 1165
Query: 1114 VTVQSLIDFVAELTPTEGEKTQGEWVLSVDGSSNNTGSGAGITIESPDKMIIEQSLKFEF 1173
+ PT + I D F
Sbjct: 1166 AK--------SHRAPTRPYR-------------------RSIPCRKVDLA--------RF 1190
Query: 1174 KASNNQSEYEALIAGLRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRR 1233
ASNN++EYEALIAGLRLA + V+K+ DSQLV Q G Y+ K+ ++ YL+VVR
Sbjct: 1191 PASNNEAEYEALIAGLRLAHGIEVKKIQAYCDSQLVASQFSGNYEAKNERMDAYLKVVRE 1250
Query: 1234 LMMEVKEIKIEHVPRGQNERADVLAKLASTGRLGNYQTV-----------IQETLPRPS- 1281
L + ++ +PR N AD LA LAST + + IQ T+ +PS
Sbjct: 1251 LSYNFEVFELTKIPRSDNAPADALAVLASTSDPDLRRVIPVECIDVPSIKIQGTITKPSE 1310
Query: 1282 ---IDLVEIKL------------------------------------------KAVKSVS 1296
IDL +L ++ S S
Sbjct: 1311 YTVIDLTAEQLSIVMVILPPATDRSEDAGPSGNLVCSSDPGQSSGCDRSDDSNRSADSDS 1370
Query: 1297 EGEISWMESIKTFLES--PPKEDDLNTRTKRREASFYTLVGGELYRRGIMSPMLKCVDIK 1354
+W+E I++++ PK+ R + R A YTL L R +L C+D +
Sbjct: 1371 PASTNWIEEIRSYIADRIVPKDKWAARRLRARSAQ-YTLPHEHLLRWSATGVLLSCLDDE 1429
Query: 1355 DALGIMAEVHEG 1366
+A +M E+HEG
Sbjct: 1430 EAQQVMREIHEG 1441
Score = 72.8 bits (177), Expect = 2e-12
Identities = 116/505 (22%), Positives = 189/505 (36%), Gaps = 79/505 (15%)
Query: 49 PFVPAITRVDIPKHLQTMALDAFSGESDPMEHLRYFNTKMVIGGA------TDAVKCRLL 102
PF I+ V I +H+ + L + G DP L + + IG A DA C+L
Sbjct: 237 PFTSCISDVRI-RHISKIKLANYEGLVDPRPFLT--SVSIAIGRAHFSDEDRDAGSCQLF 293
Query: 103 PSTFKGMAMQWFIRQPPFSIDNFTDLSTKFLTQFSANKTTKATMFDLISIHQQPGEKLKT 162
G A+ WF R SID+ L+T FL + A+ DL ++ Q E L++
Sbjct: 294 VKHLSGAALTWFSRLEANSIDSVHALTTSFLKNYGVFMEKGASNVDLWTMAQTAKESLRS 353
Query: 163 YMARFSKMAVQLEDENPDVCLASFKNGLRAGDLNRDLTR-RPAKDMLDLRARVQEFILIE 221
++ RF ++ + + D +A+ +N L GD +R T +P +A + +
Sbjct: 354 FIGRFKEIVTSVATPD-DAAIAALRNALWHGDSSRCTTLCKPFHRQGSAKAPIAK----- 407
Query: 222 QDDQKKQEREEGRKQSQSGGVSQDKSKAGKETRIAQT-PRVPRPGPYQNSKPGFQNTWHR 280
++ K+ E R+ + ++K K G + T PR +P N W+R
Sbjct: 408 --EKPKEGHHESRQHYDADYAKEEKLKKGTSYYVGDTAPRSDKP----------WNKWNR 455
Query: 281 NPQGPPAATAGQTGASPAPVP--LTKLNAPLSTILRAVGQTNVVQYPPPPRRPPANVDTN 338
+ A Q +P T L T+L + + P RR N N
Sbjct: 456 SAD----AKGDQKYYEFHKIPGHSTDECRQLQTLLLTKFKKGDLDIEPDRRRTGTNDRDN 511
Query: 339 RWCEYHKALGHT-TDNCWNLRR--EIDRLIKAGH-----LANFVKDTAAPEVAKITQGDK 390
H+ + +T DN + RR E DR + H ++ AP + K
Sbjct: 512 T----HRRVENTDRDNQDDKRRQDETDRRAERIHDDDRRAEPHKRNHDAPRQFEDEPASK 567
Query: 391 GKGKEIVEELGDPVGECSSIAGGFGGGAISSKARKRYVAAVHSVQESNEGECWVNHSPII 450
+ I+ L C + ++ ++ S E NE PI
Sbjct: 568 RRRNMIMGGL----TACRDSVRSIKSYRRQGEKQRAWIEKNTSAVECNE--------PIT 615
Query: 451 FTPQDFAHVIPHDNDPIVVTIRVNNYVTKKVFLDQGSSADIIYGDAFDRLGLKESDLKPY 510
F+ D A H NDP+VV +++ V ++ +D S D +
Sbjct: 616 FSEADIALPGSH-NDPLVVELKIGENVLLQLEIDVRSFQDKVQ----------------- 657
Query: 511 KGTLVGFTGDRVSVRGYVEIPTAFG 535
L GF GD + G + +P G
Sbjct: 658 --PLTGFDGDTIMTVGTITLPIYVG 680
>At2g13330 F14O4.9
Length = 889
Score = 470 bits (1209), Expect = e-132
Identities = 240/529 (45%), Positives = 329/529 (61%), Gaps = 32/529 (6%)
Query: 758 NGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEE 817
NGKWR+C D+ LNK CPKDS+PLP++D+LV+ +E LS MDA+ GYNQI+M +E
Sbjct: 291 NGKWRVCIDFRDLNKACPKDSFPLPHIDRLVEATVEHEKLSFMDAFYGYNQILMRRDGQE 350
Query: 818 STTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDH 877
T F+T++ Y YK MPFGLKNAG TYQRL++++F Q+G+ +EVY+DDM+VKSA DH
Sbjct: 351 KTAFITDRGTYYYKVMPFGLKNAGTTYQRLVNRMFVDQLGKTIEVYIDDMLVKSAHEKDH 410
Query: 878 GGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTS 937
L+E F L ++MKLNPEKCSF +Q +FL +++T RGIE NP + A +EM SP
Sbjct: 411 VPQLRECFKILIKFEMKLNPEKCSFEVQSREFLEYLVTERGIEANPKQIAAFIEMPSPKM 470
Query: 938 VKEVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPV 997
+EVQRLTGR+ AL+ F+ + +K PF+ L+K +F W ++CEQAF +LK L P+
Sbjct: 471 AREVQRLTGRIPALNGFISRSANKCVPFYQPLRKGKEFDWNKDCEQAFKQLKAYLTEPPI 530
Query: 998 LSKPTPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAI 1057
L+KP PL LY + R +EK AL +
Sbjct: 531 LAKPEKGEPLYLY------------------------------TNRDKRCPAMEKLALTV 560
Query: 1058 LKTARRLRPYFQSFQVKIKTDVPLRQVLQKPDLSGRLVSWSVELSEYDIQYEPRGQVTVQ 1117
+ + R+LR YFQS + + T P+R +L SGRL W++EL EYDI+Y R + Q
Sbjct: 561 VMSVRKLRLYFQSHPIVVMTSQPIRTILHSLTQSGRLAKWAIELREYDIEYRTRTSLKAQ 620
Query: 1118 SLIDFVAE--LTPTEGEKTQGEWVLSVDGSSNNTGSGAGITIESPDKMIIEQSLKFEFKA 1175
L DFV + L +G + +W+L VDGSSN GSG GI + SP +IEQSL+ F A
Sbjct: 621 VLADFVIKLPLADLDGTNSNKKWLLHVDGSSNRQGSGVGIQLTSPTGEVIEQSLQLGFNA 680
Query: 1176 SNNQSEYEALIAGLRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLM 1235
SNN+SEYEALIAG++LA E G++++ DSQLV Q GEY+ KD ++ YLE+V+ L
Sbjct: 681 SNNESEYEALIAGIKLAQEKGIREIHAYSDSQLVTSQFHGEYEAKDERMEAYLELVKTLA 740
Query: 1236 MEVKEIKIEHVPRGQNERADVLAKLASTGRLGNYQTVIQETLPRPSIDL 1284
+ + K+ +PRG+N AD LA LAST + + E + SIDL
Sbjct: 741 QQFESFKLTRIPRGENTSADTLAALASTSDPFVKRIIPVEGIEHTSIDL 789
Score = 95.1 bits (235), Expect = 3e-19
Identities = 77/246 (31%), Positives = 119/246 (48%), Gaps = 32/246 (13%)
Query: 461 PHDNDPIVVTIRVNNYVTKKVFLDQGSSADIIYGDAFDRLGLKESDLKPYKGTLVGFTGD 520
PHD D +V+ I V NY + +D GSS D+++ DAF R G +S L+ K L GF GD
Sbjct: 60 PHD-DALVIRIDVGNYELSCIMVDTGSSVDVLFYDAFKRTGHLDSKLQGRKTPLTGFAGD 118
Query: 521 RVSVRGYVEIPT-AFGEGEFVKKFQVKYLVLACRANYNVLLGRDTLNKVCAVISTAHLTV 579
G +++PT A G + +LV+ +A +N +LGR L+ + AV ST H +
Sbjct: 119 TTFSIGTIQLPTIARGVRQL-----TNFLVVDKKAPFNAILGRPWLHVMKAVPSTYHQCI 173
Query: 580 KYPACNGKVGILRVDQNAARECYLRS---------VALYGRKAAKESHRITEIFP-QEGF 629
K+P+ G + ++ Q ++R+CY+ S V L E + + P Q G
Sbjct: 174 KFPSYKG-IAVVYGSQRSSRKCYMGSYEDIKKADPVVLMIEDELAEMKTVRSLDPSQRGT 232
Query: 630 --SLDPRDDADDFRPQPLEETKQVQVKDKFLKIGSGLTTEQEDRLITLLGDNLDLFAWTI 687
SL + D+ P+ + + IG L + LIT L +N D FAW+
Sbjct: 233 RKSLITQVCIDESDPK------------RCVGIGHDLDLTVREDLITFLKENKDSFAWSS 280
Query: 688 NDVPGI 693
++ GI
Sbjct: 281 ANLQGI 286
>At3g31530 hypothetical protein
Length = 831
Score = 439 bits (1130), Expect = e-123
Identities = 243/585 (41%), Positives = 352/585 (59%), Gaps = 34/585 (5%)
Query: 794 NELL--SLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKI 851
N+LL S +++ + +MM+P D+E T F T Q +CY+ MPFGLKNAGATY+R ++KI
Sbjct: 77 NQLLETSYCHSWTLFVVMMMNPEDQEKTAFYTEQGIFCYRVMPFGLKNAGATYERFVNKI 136
Query: 852 FSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLG 911
F+ Q+G+ MEVY+DDM+VKS DH L+E F QL Y +KLNP KC FG++ G
Sbjct: 137 FALQIGKTMEVYIDDMLVKSMTEKDHISHLRECFKQLNLYNVKLNPAKCRFGVRSG---- 192
Query: 912 FMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFFTCLKK 971
IE NP + A+L M SP + +EVQ LTGR+AAL+RF+ + +K F+ L++
Sbjct: 193 -------IEANPKQIEALLGMASPQNKREVQCLTGRVAALNRFISRSTEKCLAFYDVLRR 245
Query: 972 NSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEEGKKQK 1031
N KF+WT CE+AF +LK+ LAT P+L+KP P LY+AV+D AVS L++E+ +QK
Sbjct: 246 NKKFEWTTRCEEAFQELKKYLATPPILAKPVIGEPQYLYVAVSDTAVSGELVREDRGEQK 305
Query: 1032 VIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTDVPLRQVLQKPDLS 1091
+I++VS T AE RY ++EK ALA++ +A++LRPYFQS + + +PLR +L P S
Sbjct: 306 LIFYVSQTFTSAESRYPQMEKLALAVVMSAQKLRPYFQSHSIIVMGSMPLRVILHSPSQS 365
Query: 1092 GRLVSWSVELSEYDIQYEPRGQVTVQSLIDFVAELTPTEGEKT--QGEWVLSVDGSSNNT 1149
GRL W++ELSEYDI+Y+ + Q L DF+ EL E + W+L VDGSS+
Sbjct: 366 GRLAKWTIELSEYDIEYQNKTCAKSQVLADFIVELPTKEARENPLDTTWLLHVDGSSSKQ 425
Query: 1150 GSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGLRLAIELGVQKLFIKGDSQLV 1209
GSG GI + SP +A+NN +EYEAL+AGL LA L + K DSQL+
Sbjct: 426 GSGVGIRLTSPTG-----------EATNNVAEYEALVAGLNLAWGLKIGKTRAFCDSQLI 474
Query: 1210 VKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRGQNERADVLAKLASTGRLGNY 1269
Q GEY +D ++ YL V+ L E ++ +PRG+N AD LA LAST
Sbjct: 475 ANQFNGEYTTQDKKMEAYLIHVQNLAKNFDEFELTRIPRGENTSADALAALASTSDTSLK 534
Query: 1270 QTVIQETLPRPSIDL------VEIKLKAVK-SVSEGEISWMESIKTFL-ESPPKEDDLNT 1321
+ + E + +PSI+L + I++ A + + WME I +++ E D
Sbjct: 535 RVIPVEFIEKPSIELGKEEHVLPIQISADQDDPDDCNSEWMEPIISYISEGKLPSDKWKA 594
Query: 1322 RTKRREASFYTLVGGELYRRGIMSPMLKCVDIKDALGIMAEVHEG 1366
R + +A+ + LV +LY+ + P++ CV+ + IM E+H G
Sbjct: 595 RKLKAQAARFVLVDTKLYKWRLSGPLMTCVEAEAICKIMKEIHSG 639
Score = 30.4 bits (67), Expect = 9.2
Identities = 13/27 (48%), Positives = 18/27 (66%)
Query: 556 YNVLLGRDTLNKVCAVISTAHLTVKYP 582
YN ++G LN+ AV ST HL +K+P
Sbjct: 30 YNAIIGTPWLNQFRAVASTYHLCLKFP 56
>At1g37200 hypothetical protein
Length = 1564
Score = 394 bits (1011), Expect = e-109
Identities = 318/1109 (28%), Positives = 504/1109 (44%), Gaps = 190/1109 (17%)
Query: 335 VDTNRWCEYHKALGHTTDNCWNLRREIDRLIKAGHL----ANFVKDTAAPEVAKITQGDK 390
+D ++C+YH GH+T+ C R + +LI AG +N +T P+ + Q K
Sbjct: 318 LDLGKYCKYHNKRGHSTEEC----RAVKKLIAAGGKTKKGSNPKVETPPPDEQEEEQTPK 373
Query: 391 GKGKEIVEELGD--PVGECSSIAGGFGGGAISSKARKRYVAAVHSVQESNEGECWVNHSP 448
K +E E GD P I F + K + + + N P
Sbjct: 374 QKKRERTLEGGDSPPPARRERIDLVFTELDLGGKTGRSVTTPPSAPHKKNM------RFP 427
Query: 449 II---FTPQDFAHVIPHDNDPIVVTIRVNNYVTKKVFLDQGSSADIIYGDAF-----DRL 500
+ F H+ P D I+ ++ N + Q + G + ++
Sbjct: 428 LRLSKFCRSATEHLSPKRIDFIMGGSQLCNDSINSIKTHQRKADSYTKGKSLMMGPDHKI 487
Query: 501 GLKESDL----KPYKGTLVGFTGDRVSVRGYVEIPTAFGEGEFVKKFQVKYLVLACRANY 556
L ES+ KP+ +V R+ V Y G V K A +
Sbjct: 488 TLWESETTDLDKPHDDAIV----IRIDVGNYKLSRIMIDTGSSVDK-----------APF 532
Query: 557 NVLLGRDTLNKVCAVISTAHLTVKYPACNGKVGILRVDQNAARECYLRSVALYGRKAAKE 616
N +LGR L+ + AV ST H +K+P+ G + ++ Q ++R CY+ S L K+
Sbjct: 533 NAILGRPWLHAMKAVPSTYHQCIKFPSEKG-IAVIYGSQRSSRRCYMGSYELI-----KK 586
Query: 617 SHRITEIFPQEGFSLDPRDDADDFRPQPLEET-KQVQVKDKFLK----IGSGLTTEQEDR 671
+ + + + + +D + P + + QV + + K +G L +
Sbjct: 587 AELVVLMIEDKLAEMKTVRSSDPSQCGPQKSSITQVYIDESDPKQCVGVGQDLDPAIRED 646
Query: 672 LITLLGDNLDLFAWTINDVPGIDPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLETE 731
IT L +N D FAW+ ++ GI +V +H+L + P P+ Q RR++ E+ KAVQ E +
Sbjct: 647 FITFLKENKDSFAWSSANLQGISLEVTSHELNVDPTYRPIKQKRRKLGPERAKAVQDEVD 706
Query: 732 KLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGA 791
+L+K IREV+YP WLAN V+VK NGKWR+C D+ LNK CPKDS+PLP++D+LV
Sbjct: 707 RLLKIGSIREVKYPDWLANPVVVKNKNGKWRVCIDFMDLNKACPKDSFPLPHIDRLVKAT 766
Query: 792 SGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKI 851
+G+ELLS MDA+SGYNQI+M P D+E T F+T+ CYK MPFGLKN GATYQRL++++
Sbjct: 767 AGHELLSFMDAFSGYNQILMRPDDQEKTAFITD----CYKVMPFGLKNTGATYQRLVNRM 822
Query: 852 FSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLG 911
F+ Q+G+ ME+Y+DDM+VKSA DH L+E F L ++MKLNPEKCSFG+ G+FLG
Sbjct: 823 FADQLGKTMELYIDDMLVKSAHEKDHLPQLRECFKILNKFEMKLNPEKCSFGVPSGEFLG 882
Query: 912 FMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFF----T 967
+++T RGIE NP + A ++M SP ++ R R + P + P + T
Sbjct: 883 YLVTQRGIEANPKQIAAFIDMPSPKMARD--RPYRRPQQVIYRAPPDHEGQKPIYYVSKT 940
Query: 968 CLKKNSKFQWTEECEQAFT----KLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLL 1023
+ + ++ E+ A KL++ L + P+ + +T + + T+L
Sbjct: 941 LIDAETHYRAMEKLALAVVMSARKLRQYLQSHPI-------------VVMTSQPIRTIL- 986
Query: 1024 QEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTDVPLRQ 1083
T Q + L IE + I R SF+ ++ D +
Sbjct: 987 -------------HSTTQSSRLAKWAIELSEYDIEYRTR------TSFKAQVLADFVIEL 1027
Query: 1084 VLQKPDLSGRLVSWSVELSEYDIQYEPRGQVTVQSLIDFVAELTPTEGEKTQGEWVLSVD 1143
L D + W + + G Q + + +LT GE + + L +
Sbjct: 1028 PLADLDGTNSNKKWLLHVD---------GSSNRQGSGEGI-QLTSPTGEVIEQSFRLGFN 1077
Query: 1144 GSSNNTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGLRLAIELGVQKLFIK 1203
S+N + E++A LI G++LA + ++ +
Sbjct: 1078 ASNNES----------------------EYEA---------LIDGIKLAQGMRIRDIHAH 1106
Query: 1204 GDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRGQNERADVLAKLAST 1263
DSQLV Q GEY+ KD ++ YLE+V+ L + + ++ +PRG+N D LA LAST
Sbjct: 1107 SDSQLVTSQFHGEYEAKDERMEAYLELVKTLTQQFESFELTRIPRGENTSTDALAALAST 1166
Query: 1264 GRLGNYQTVIQETLPRPSIDLV-------------------------------------- 1285
+ + E + PSIDL
Sbjct: 1167 LDPFVKRIIPVEGIEHPSIDLTVKHAGMETSATCNFTRVTRSTTAAARALAAAAEAGAAE 1226
Query: 1286 -EIKLKAVKSVSEGE----ISWMESIKTFLE---SPPKEDDLNTRTKRREASFYTLVGGE 1337
E KL+ + ++ E W I ++E +PP D TR + +++ Y ++ G
Sbjct: 1227 EEAKLRTPEQLASYEPEPYNDWRIPIIDYIERGITPP--DKWETRKLKAQSARYCIMEGR 1284
Query: 1338 LYRRGIMSPMLKCVDIKDALGIMAEVHEG 1366
L +R + P + C + +M +HEG
Sbjct: 1285 LMKRSVAGPYMVCTYGQQTKDLMKSMHEG 1313
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 385 bits (990), Expect = e-107
Identities = 189/359 (52%), Positives = 257/359 (70%), Gaps = 2/359 (0%)
Query: 715 RRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYTSLNKVC 774
RR++ E+ KAV E +KL+K IREVQYP WLAN V+VKK NGK R+C D+T LNK C
Sbjct: 469 RRKLGVERAKAVNDEVDKLLKIGSIREVQYPDWLANTVVVKKKNGKDRVCIDFTDLNKAC 528
Query: 775 PKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMP 834
PKDS+PLP++D+LV+ +GNELLS MDA+SGYNQIMM+P D+E T F+T++ YCYK MP
Sbjct: 529 PKDSFPLPHIDRLVESTAGNELLSFMDAFSGYNQIMMNPEDQEKTLFITDRGIYCYKVMP 588
Query: 835 FGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMK 894
FGL+NAGATY RL++K+FS+ VG+ MEVY+DDM++KS + DH L+E F L YQMK
Sbjct: 589 FGLRNAGATYPRLVNKMFSEHVGKTMEVYIDDMLIKSLKKEDHVKHLEECFAILNQYQMK 648
Query: 895 LNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRF 954
LNP KC+FG+ G+FLG+++T RGIE NP++ A L M SP + KEVQRLTGR+AAL+RF
Sbjct: 649 LNPAKCTFGVPSGEFLGYIVTKRGIEANPNQINAFLNMPSPKNFKEVQRLTGRIAALNRF 708
Query: 955 LPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVT 1014
+ + DK+ PF+ LK N +F W E+CE+AF +LK L + PVL KP L LY++V+
Sbjct: 709 ISRSTDKSLPFYQILKGNKEFLWDEKCEEAFGQLKAYLTSPPVLFKPELDEKLYLYVSVS 768
Query: 1015 DKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQV 1073
+ AVS VL++E+ +QK IY++ + Y+ + A L + L F+ F++
Sbjct: 769 NHAVSGVLIREDRGEQKPIYYIIVNQFNGD--YEAKDSRMEAYLHVVKDLAKNFRKFEL 825
Score = 71.2 bits (173), Expect = 5e-12
Identities = 69/292 (23%), Positives = 122/292 (41%), Gaps = 25/292 (8%)
Query: 4 QQMLDTLQKVQSQNEQLQAQVEYLTQRQDFKQEQRLEAEDVVEFQPFVPAITRVDIPKHL 63
Q+ D L+ ++ + +L+ Q + + E + PF P IT + I +
Sbjct: 100 QENFDQLETMKEVSRELKEMRSKFLQATSSEPDINRVIEKARQ-TPFTPQITSLRI-RDS 157
Query: 64 QTMALDAFSGESDPMEHLRYFNTKMVIGGATDAVK-------CRLLPSTFKGMAMQWFIR 116
+ + L++++G DP +L F ++ G D + C+L G A+ WF +
Sbjct: 158 RKLNLESYNGLEDPKGYLAAF---LIAAGRVDLNEADEDVRYCKLFSENLCGQALMWFTQ 214
Query: 117 QPPFSIDNFTDLSTKFLTQFSANKTTKATMFDLISIHQQPGEKLKTYMARFSKMAVQLED 176
P SI NF +LS FL Q+S + DL ++ Q P E L+ ++ +F + +L
Sbjct: 215 LEPGSISNFNELSVVFLKQYSILMDKSISDTDLWNLSQGPNETLRAFITKFKYVLSKLSR 274
Query: 177 ENPDVCLASFKNGL-RAGDLNRDLTRRPAKDMLDLRARVQEFILIEQDDQKKQEREEGRK 235
+ L++ + GL DL + D R ++ +E + + +R+ K
Sbjct: 275 ISQQSALSALRKGLWYDSRFKEDLILHKLDTIQDALFRANNWMEVEDEKESFAKRD---K 331
Query: 236 QSQSGGVSQDKSKAGKETRIAQTPRVPRPGPYQNS------KPGFQNTWHRN 281
Q++ K E R Q PR P+ N+ G NTW R+
Sbjct: 332 QAKPAVTFPPKK---FEPRENQGPRKFGLLPFNNNVGKQFQGKGRSNTWIRD 380
Score = 55.8 bits (133), Expect = 2e-07
Identities = 54/219 (24%), Positives = 90/219 (40%), Gaps = 45/219 (20%)
Query: 1186 IAGLRLAIELGVQKLFIKGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEH 1245
++G+ + + G QK ++V Q G+Y+ KD ++ YL VV+ L ++ ++
Sbjct: 772 VSGVLIREDRGEQKPIY----YIIVNQFNGDYEAKDSRMEAYLHVVKDLAKNFRKFELIR 827
Query: 1246 VPRGQNERADVLAKLASTGRLGNYQTVIQETLPRPSI----------------------- 1282
+PRGQN AD LA LAST + E + SI
Sbjct: 828 IPRGQNTTADALAALASTSDPEVNSIIPVECISERSIKEEKEAFVVTRSRAAARDRGDTA 887
Query: 1283 -DLVEIKLKAVKSVSEGEISWMESI------------KTFLESPP-KEDDLNTRTKRREA 1328
+L +K + K + + + E+I K E+ P ++D+ + E
Sbjct: 888 VELPPVKRRKSKEATVPKPAVQENIGIELIEEVISDAKIEAETEPWVDEDILAPNEHPEP 947
Query: 1329 SFYTLVGGELYRRGIMSPMLKCVDIKDALGIMAEVHEGV 1367
LYRRG+ P L + +M+EVHEG+
Sbjct: 948 EAL----NSLYRRGVSDPYLLSIFGPVVEIVMSEVHEGL 982
Score = 45.4 bits (106), Expect = 3e-04
Identities = 25/91 (27%), Positives = 47/91 (51%), Gaps = 1/91 (1%)
Query: 421 SKARKRYVAAVHSVQESNEGECWVNHSPIIFTPQDFAHVIPHDNDPIVVTIRVNNYVTKK 480
S K++ VH +E E + I+F ++ H+ +D +VVT+ V N+ +
Sbjct: 382 SSVIKKHSRNVHLKLSLSE-EVDFQSTSILFDEKETQHLERSHDDALVVTLDVANFEVSR 440
Query: 481 VFLDQGSSADIIYGDAFDRLGLKESDLKPYK 511
+ +D GSS D+I+ +R+G+ +D+ K
Sbjct: 441 ILIDTGSSVDLIFLSTLERMGISRADVNRRK 471
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 341 bits (874), Expect = 2e-93
Identities = 158/287 (55%), Positives = 216/287 (75%)
Query: 688 NDVPGIDPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTW 747
+D+ GIDP+V H+L + P V Q RR++ E++KAV E +KL+ A I EV+YP W
Sbjct: 475 HDMVGIDPEVACHELNVDPTFKLVKQKRRKLGPERSKAVNDEVDKLLDAGSIVEVKYPEW 534
Query: 748 LANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYN 807
LAN V+VKK N KWR+C D+T LNK CPKDS+PLP++D++V+ +GNELLS MDA+SGYN
Sbjct: 535 LANPVVVKKKNDKWRVCIDFTDLNKACPKDSFPLPHIDRMVEATTGNELLSFMDAFSGYN 594
Query: 808 QIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDM 867
QI MH D+E T+F+ ++ YCYK MPFGLKN GA YQRL++++F+ Q+G+ MEVY+DDM
Sbjct: 595 QIPMHKDDQEKTSFIIDRGTYCYKVMPFGLKNVGARYQRLVNQMFAPQLGKTMEVYIDDM 654
Query: 868 IVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGR 927
+VKS R++DH LK F+ L Y MKLNP KC FG+ G+FLG+++T RGIE NP + R
Sbjct: 655 LVKSTRSADHIDHLKACFETLNKYNMKLNPAKCLFGVTSGEFLGYIVTKRGIEANPKQIR 714
Query: 928 AILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSK 974
AIL+++SP + KEVQRLTGR+A L+RF+ + DK+ PF+ L+ +K
Sbjct: 715 AILDLQSPRNKKEVQRLTGRIAGLNRFIARSTDKSLPFYQLLRSANK 761
Score = 77.4 bits (189), Expect = 7e-14
Identities = 66/301 (21%), Positives = 127/301 (41%), Gaps = 46/301 (15%)
Query: 338 NRWCEYHKALGHTTDNCWNLRREIDRLIKAGHL---------------------ANFVKD 376
+++C++HK GH+T C +L+ + K G + N V
Sbjct: 184 DQYCDFHKRSGHSTAACRHLQSILLNKYKKGDIEVQHRQYKSHNNTYAARGGRDGNNVFH 243
Query: 377 TAAPEVAKITQGDKG-------------KGKEIVEELGD---PVGECSSIAGGFGGGAIS 420
P + + K +E ++ D P + I GG S
Sbjct: 244 RLGPHTGRQQEAPPANEEERHPDMEPPKKNRENDQQHNDAPVPRRRVNMIMGGLTACRDS 303
Query: 421 SKARKRYVAAVHSVQESNEGECWVNHSPIIFTPQDFAHV-IPHDNDPIVVTIRVNNYVTK 479
++ K Y+ + + S+ +P+ FT +D V +PH NDP+V+ + +
Sbjct: 304 FRSIKEYIKSGAATLWSSPAT--KEMTPLTFTSEDLFGVDLPH-NDPLVIELHIGESEVT 360
Query: 480 KVFLDQGSSADIIYGDAFDRLGLKESDLKPYKGTLVGFTGDRVSVRGYVEIPTAFGEGEF 539
++ +D GSS ++++ D ++ + + +KP L GF G+ + G +++P G
Sbjct: 361 RILIDTGSSVNVVFKDVLQKMKVHDRHIKPSVRPLTGFDGNTMMTNGTIKLPIYLGGAAT 420
Query: 540 VKKFQVKYLVLACRANYNVLLGRDTLNKVCAVISTAHLTVKYPACNGKVGILRVDQNAAR 599
KF +V+ YN++LG ++ + A+ S+ H +K P G + +R +QN A
Sbjct: 421 WHKF----VVVDKPTIYNIILGTPWIHDMQAIPSSYHQCIKIPTSIG-IETIRGNQNLAH 475
Query: 600 E 600
+
Sbjct: 476 D 476
>At4g03800
Length = 637
Score = 321 bits (822), Expect = 3e-87
Identities = 165/337 (48%), Positives = 229/337 (66%), Gaps = 21/337 (6%)
Query: 690 VPGIDPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLA 749
+P IDP +I H+L + P P+ Q RR++ E+ KAV + +KL+K IREVQYP W+A
Sbjct: 321 MPDIDPSIICHELNVDPRFKPLKQKRRKLGVERAKAVNGKIDKLLKIGSIREVQYPDWVA 380
Query: 750 NVVMVKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQI 809
V+VKK NGK R+C D+T LNK CPKDS+PLP++D+LV+ +GNELL+ MDA+ GYNQI
Sbjct: 381 ITVVVKKKNGKDRVCIDFTDLNKACPKDSFPLPHIDRLVESTAGNELLTFMDAFLGYNQI 440
Query: 810 MMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIV 869
MM+P D+E T+F+T++ GATYQ L++K+F++ + + MEV +DD +V
Sbjct: 441 MMNPEDQEKTSFITDR---------------GATYQWLVNKMFNEHLRKTMEVSIDDTLV 485
Query: 870 KSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAI 929
KS + DH L E F+ L YQMKLN KC+FG+ G+FLG+++T RGIE NP++ A
Sbjct: 486 KSLKKEDHVKHLGECFEILNQYQMKLNLAKCTFGVPSGEFLGYIVTKRGIEANPNQINAF 545
Query: 930 LEMKSPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLK 989
L+ S + KEVQRLTGR+A+ S DK+ PF+ LK N+ F W E+CE+AF +LK
Sbjct: 546 LKTPSLRNFKEVQRLTGRIASRST------DKSLPFYQILKGNNGFLWDEKCEEAFRQLK 599
Query: 990 ETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEE 1026
L T PVLSKP L LY+ V++ AVS VL++E+
Sbjct: 600 AYLTTPPVLSKPEADEKLYLYVFVSNHAVSGVLVRED 636
Score = 70.5 bits (171), Expect = 8e-12
Identities = 90/340 (26%), Positives = 140/340 (40%), Gaps = 49/340 (14%)
Query: 335 VDTNRWCEYHKALGHTTDNCWNLRREIDRLIKAGHLA-----NFVKDTAAPEVAKITQGD 389
VD + E + H T +C L+R + L +G L+ +FVK TQ
Sbjct: 67 VDLDEAEEDARVTEHLTKDCIVLKRYLAELWASGDLSKFNIDDFVKQYHEARDNSETQNS 126
Query: 390 KGKGKEIVEELGDPVGECSSIAGGFGGGAISSKARKRY---VAAVHSVQESNEGECWVNH 446
K EE G+ + I GG S A K++ V S+ E +C
Sbjct: 127 KRPRLTNEEEPRSSKGKINVILGGSKLCHNSVSAIKKHRPNVLLKSSLSEEVNYQC---- 182
Query: 447 SPIIFTPQDFAHVI-PHDNDPIVVTIRVNNYVTKKVFLDQGSSADIIYGDAFDRLGLKES 505
S + F ++ H+ PHD D +++T+ V N+ ++ +D GSS D+I+
Sbjct: 183 SSVSFDEEETRHIERPHD-DALIITLDVANFKISRILVDTGSSVDLIF------------ 229
Query: 506 DLKPYKGTLVGFTGDRVSVRGYVEIPTAFGEGEFVKKFQVKYLVLACRANYNVLLGRDTL 565
L P LV FT + G +++P G+ + V ++V A YN++LG +
Sbjct: 230 -LGP-PSPLVAFTSESAMSLGTIKLPVLAGKMSKI----VDFVVFDKPATYNIILGTPWI 283
Query: 566 NKVCAVISTAHLTVKYPACNGKVGILRVDQNAARECYLRSVALYGRKAAKESHRITEIFP 625
++ AV ST H +K+P NG I + + Y+ + I
Sbjct: 284 FQMKAVPSTYHQWLKFPTSNGVETIWGDLEGSRTSAYMPDID-------------PSIIC 330
Query: 626 QEGFSLDPRDD--ADDFRPQPLEETKQVQVK-DKFLKIGS 662
E ++DPR R +E K V K DK LKIGS
Sbjct: 331 HE-LNVDPRFKPLKQKRRKLGVERAKAVNGKIDKLLKIGS 369
>At2g10780 pseudogene
Length = 1611
Score = 229 bits (583), Expect = 1e-59
Identities = 158/535 (29%), Positives = 265/535 (49%), Gaps = 27/535 (5%)
Query: 663 GLTTEQEDRLITLLGDNLDLFAWTINDVPGIDP-KVITHKLAIRPGATPVIQPRRRMSEE 721
G + E +D I ++ + D+FA V G+ P + + + PG TP+ + RM+
Sbjct: 617 GASAELKD--IPIVNEFSDVFA----AVSGVPPDRSDPFTIELEPGTTPISKAPYRMAPA 670
Query: 722 KNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPL 781
+ ++ + E+L+ FIR P W A V+ VKK +G +R+C DY LNKV K+ YPL
Sbjct: 671 EMAKLKKQLEELLDKGFIRPSSSP-WGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPL 729
Query: 782 PNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAG 841
P +D+L+D G + S +D SGY+QI + P+D T F T ++ + MPFGL NA
Sbjct: 730 PRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYDHFEFVVMPFGLTNAP 789
Query: 842 ATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCS 901
A + ++M+ +F + + ++++D++V S H L+ ++LR +++ KCS
Sbjct: 790 AAFMKMMNGVFRDFLDEFVIIFINDILVYSKSWEAHQEHLRAVLERLREHELFAKLSKCS 849
Query: 902 FGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDK 961
F + FLG +++ +G+ V+P+K R+I E P + E++ G RF+
Sbjct: 850 FWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASM 909
Query: 962 AAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTV 1021
A P K++ F W++ECE++F +LK L PVL P P +Y + + V
Sbjct: 910 AQPLTRLTGKDTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCV 969
Query: 1022 LLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTD-VP 1080
L+Q K VI + S L+ E Y + A++ + R Y +V+I TD
Sbjct: 970 LMQ----KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLYGAKVQIYTDHKS 1025
Query: 1081 LRQVLQKPDLSGRLVSWSVELSEY--DIQYEPRGQVTVQSLIDFVAELTPTEGEKTQGEW 1138
L+ + +P+L+ R W +++Y DI Y P V + + E E++Q +
Sbjct: 1026 LKYIFTQPELNLRQRRWMELVADYNLDIAYHPGKANQVADALS--RRRSEVEAERSQVDL 1083
Query: 1139 V-----LSVDGSSNNT---GSGAGITIESPDKMIIEQSLKFEFK--ASNNQSEYE 1183
V L V+ S G GA + ++ + Q E K A NN++EY+
Sbjct: 1084 VNMMGTLHVNALSKEVEPLGLGAADQADLLSRIRLAQERDEEIKGWAQNNKTEYQ 1138
>At3g11970 hypothetical protein
Length = 1499
Score = 219 bits (558), Expect = 1e-56
Identities = 132/413 (31%), Positives = 215/413 (51%), Gaps = 7/413 (1%)
Query: 700 HKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANG 759
HK+ + G+ PV Q R S + + E L+ ++ P + + VV+VKK +G
Sbjct: 591 HKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSP-YASPVVLVKKKDG 649
Query: 760 KWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEEST 819
WR+C DY LN + KDS+P+P ++ L+D G + S +D +GY+Q+ M P D + T
Sbjct: 650 TWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKT 709
Query: 820 TFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGG 879
F T+ ++ Y MPFGL NA AT+Q LM+ IF + + + V+ DD++V S+ +H
Sbjct: 710 AFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQ 769
Query: 880 DLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVK 939
LK+ F+ +R ++ KC+F + ++LG ++++GIE +P K +A+ E PT++K
Sbjct: 770 HLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLK 829
Query: 940 EVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLS 999
+++ G RF+ G A P L K F+WT +QAF LK L PVLS
Sbjct: 830 QLRGFLGLAGYYRRFVRSFGVIAGPLH-ALTKTDAFEWTAVAQQAFEDLKAALCQAPVLS 888
Query: 1000 KPTPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILK 1059
P V+ + + VL+QE + ++S L+G +L EK LA++
Sbjct: 889 LPLFDKQFVVETDACGQGIGAVLMQE----GHPLAYISRQLKGKQLHLSIYEKELLAVIF 944
Query: 1060 TARRLRPYFQSFQVKIKTDV-PLRQVLQKPDLSGRLVSWSVELSEYDIQYEPR 1111
R+ R Y IKTD L+ +L++ + W +L E+D + + R
Sbjct: 945 AVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYR 997
>At1g36590 hypothetical protein
Length = 1499
Score = 219 bits (558), Expect = 1e-56
Identities = 132/413 (31%), Positives = 215/413 (51%), Gaps = 7/413 (1%)
Query: 700 HKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANG 759
HK+ + G+ PV Q R S + + E L+ ++ P + + VV+VKK +G
Sbjct: 591 HKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSP-YASPVVLVKKKDG 649
Query: 760 KWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEEST 819
WR+C DY LN + KDS+P+P ++ L+D G + S +D +GY+Q+ M P D + T
Sbjct: 650 TWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKT 709
Query: 820 TFMTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGG 879
F T+ ++ Y MPFGL NA AT+Q LM+ IF + + + V+ DD++V S+ +H
Sbjct: 710 AFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQ 769
Query: 880 DLKEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVK 939
LK+ F+ +R ++ KC+F + ++LG ++++GIE +P K +A+ E PT++K
Sbjct: 770 HLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLK 829
Query: 940 EVQRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLS 999
+++ G RF+ G A P L K F+WT +QAF LK L PVLS
Sbjct: 830 QLRGFLGLAGYYRRFVRSFGVIAGPLH-ALTKTDAFEWTAVAQQAFEDLKAALCQAPVLS 888
Query: 1000 KPTPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILK 1059
P V+ + + VL+QE + ++S L+G +L EK LA++
Sbjct: 889 LPLFDKQFVVETDACGQGIGAVLMQE----GHPLAYISRQLKGKQLHLSIYEKELLAVIF 944
Query: 1060 TARRLRPYFQSFQVKIKTDV-PLRQVLQKPDLSGRLVSWSVELSEYDIQYEPR 1111
R+ R Y IKTD L+ +L++ + W +L E+D + + R
Sbjct: 945 AVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYEIQYR 997
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 214 bits (546), Expect = 3e-55
Identities = 129/422 (30%), Positives = 217/422 (50%), Gaps = 9/422 (2%)
Query: 702 LAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKW 761
+ + PG P+ + RM+ + ++ + + L+ FIR P W A V+ VKK +G +
Sbjct: 483 IELEPGTAPLSKAPYRMAPAEMAELKKQLKDLLGKGFIRPSTSP-WGAPVLFVKKKDGSF 541
Query: 762 RMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTF 821
R+C DY LN+V K+ YPLP +D+L+D G S +D SGY+QI + +D T F
Sbjct: 542 RLCIDYRELNRVTVKNRYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAF 601
Query: 822 MTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDL 881
T ++ + MPFGL NA A + RLM+ +F + + + +++DD++V S + L
Sbjct: 602 RTRYGHFEFVVMPFGLTNAPAVFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEQEVHL 661
Query: 882 KEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEV 941
+ ++LR ++ KCSF + FLG ++++ G+ V+P+K AI + PT+ E+
Sbjct: 662 RRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEI 721
Query: 942 QRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKP 1001
+ G RF+ A P K+ F W++ECE+ F LKE L + PVL+ P
Sbjct: 722 RSFLGWAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSQECEEGFVSLKEMLTSTPVLALP 781
Query: 1002 TPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTA 1061
P ++Y + + VL+Q KVI + S L E Y + A++
Sbjct: 782 EHGQPYMVYTDASRVGLGCVLMQH----GKVIAYASRQLMKHEGNYPTHDLEMAAVIFAL 837
Query: 1062 RRLRPYFQSFQVKIKTD-VPLRQVLQKPDLSGRLVSWSVELSEYDIQ--YEP-RGQVTVQ 1117
+ R Y +V++ TD L+ + +P+L+ R W +++YD++ Y P + V V
Sbjct: 838 KIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLEIAYHPGKANVVVD 897
Query: 1118 SL 1119
+L
Sbjct: 898 AL 899
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 213 bits (541), Expect = 1e-54
Identities = 136/467 (29%), Positives = 219/467 (46%), Gaps = 47/467 (10%)
Query: 660 IGSGLTTEQEDRLITLLGDNLDLFAWTINDVPGIDPKVITHKLAIRPGATPVIQPRRRMS 719
+ LT +Q + LIT L ++++D+ GI P + TH++ + + I+P+RR++
Sbjct: 849 VNDELTADQVNLLITELMKYRKAIGYSLDDIKGISPTLCTHRIHLENESYSSIEPQRRLN 908
Query: 720 EEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKW------------------ 761
+ V+ E KL+ A I + TW++ V V K G
Sbjct: 909 PNLKEVVKKEILKLLDAGVIYPISDSTWVSPVHCVPKKGGMTVVKNSKDELIPTRTITGH 968
Query: 762 RMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTF 821
RMC +Y LN K+ +PLP +D +++ + + +D+YSG+ QI +HP+D+ TTF
Sbjct: 969 RMCIEYRKLNVASRKEHFPLPFIDHMLERLANHPYYCFLDSYSGFFQIPIHPNDQGKTTF 1028
Query: 822 MTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDL 881
+ YK MPFGL NA AT+QR M IFS + +EV++DD V + S +L
Sbjct: 1029 TCPYGTFAYKRMPFGLCNAPATFQRCMTSIFSDLIEEMVEVFMDDFSVYGSSFSSCLLNL 1088
Query: 882 KEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEV 941
+ + LN EKC F ++ G LG ++ GIEV+ K +++++ P +VK++
Sbjct: 1089 CRVLKRCEETNLVLNWEKCHFMVREGIVLGRKISEEGIEVDKAKIDVMMQLQPPKTVKDI 1148
Query: 942 QRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKP 1001
+ G F+ A P L K ++F + +EC AF +KE L T P++ P
Sbjct: 1149 RSFLGHAGFYRIFIKDFSKLARPLTRLLCKETEFAFDDECLTAFKLIKEALITAPIVQAP 1208
Query: 1002 TPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTA 1061
P F T+ A++RY EK LA++
Sbjct: 1209 NWDFP----------------------------FEIITMDDAQVRYATTEKELLAVVFAF 1240
Query: 1062 RRLRPYFQSFQVKIKTD-VPLRQVLQKPDLSGRLVSWSVELSEYDIQ 1107
+ R Y +V I TD LR + K D RL+ W + L E+D++
Sbjct: 1241 EKFRSYLVGSKVTIYTDHAALRHIYAKKDTKPRLLRWILLLQEFDME 1287
Score = 39.3 bits (90), Expect = 0.020
Identities = 34/128 (26%), Positives = 52/128 (40%), Gaps = 9/128 (7%)
Query: 76 DPMEHLRYFNTKMVIGG----ATDAVKCRLLPSTFKGMAMQWFIRQPPFSIDNFTDLSTK 131
DP++HL F+ + + D K RL P + A QW P SI ++ D
Sbjct: 61 DPLDHLDEFDRLCSLTKINRVSEDGFKLRLFPFSLGDKAHQWEKSLPQGSITSWNDCKKA 120
Query: 132 FLTQFSANKTTKATMFDLISIHQQPGEKLKTYMARFSKMAVQLEDENPDVCLASFKNGLR 191
FL +F +N T D+ Q E RF Q + + ++ L + NG
Sbjct: 121 FLAKFFSNSRTARLRNDISGFTQTNNETFYEAWERFK--GYQTQCPHHEMLLDTASNG-- 176
Query: 192 AGDLNRDL 199
LN+D+
Sbjct: 177 -NFLNKDV 183
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 211 bits (538), Expect = 2e-54
Identities = 125/412 (30%), Positives = 211/412 (50%), Gaps = 8/412 (1%)
Query: 702 LAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKW 761
+ + PG P+ + RM+ + ++ + E L+ FIR P W A V+ VKK +G +
Sbjct: 509 IELEPGTAPLSKAPYRMAPAEMTELKKQLEDLLGKGFIRPSTSP-WGAPVLFVKKKDGSF 567
Query: 762 RMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTF 821
R+C DY LN V K+ YPLP +D+L+D G S +D SGY+QI + +D T F
Sbjct: 568 RLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAF 627
Query: 822 MTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDL 881
T ++ + MPF L NA A + RLM+ +F + + + +++DD++V S +H L
Sbjct: 628 RTRYGHFEFVVMPFALTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEHEVHL 687
Query: 882 KEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEV 941
+ ++LR ++ KCSF + FLG ++++ G+ V+P+K AI + PT+ E+
Sbjct: 688 RRVMEKLREQKLFAKLSKCSFWQREIGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEI 747
Query: 942 QRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKP 1001
+ RF+ A P K+ F W+ ECE+ F LKE L + PVL+ P
Sbjct: 748 RSFLRLTGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALP 807
Query: 1002 TPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTA 1061
P ++Y + + VL+Q + KVI + S L+ E Y + ++
Sbjct: 808 EHGQPYMVYTDASRVGLGCVLMQ----RGKVIAYASRQLRKHEGNYPTHDLEMAVVIFAL 863
Query: 1062 RRLRPYFQSFQVKIKTD-VPLRQVLQKPDLSGRLVSWSVELSEYDIQ--YEP 1110
+ R Y +V++ TD L+ + +P+L+ R + W +++YD++ Y P
Sbjct: 864 KIWRSYLYGGKVQVFTDHKSLKYIFNQPELNLRQMRWMELVADYDLEIAYHP 915
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 206 bits (524), Expect = 9e-53
Identities = 126/406 (31%), Positives = 208/406 (51%), Gaps = 12/406 (2%)
Query: 663 GLTTEQEDRLITLLGDNLDLFAWTINDVPGIDP-KVITHKLAIRPGATPVIQPRRRMSEE 721
G + E +D LI + + D+FA V G+ P + + + PG TP+ + RM+
Sbjct: 112 GASAELKDILI--VNEFSDVFA----AVSGVPPDRSDPFTIELEPGTTPISKAPYRMAPA 165
Query: 722 KNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPL 781
+ ++ + E+L+ FIR P W A V+ VKK +G +R+C DY LNKV K+ YPL
Sbjct: 166 EMAELKKQLEELLAKGFIRPSSSP-WGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPL 224
Query: 782 PNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAG 841
P +D+L+D G + S +D SGY+QI + P+D T F T ++ + MPFGL NA
Sbjct: 225 PRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYGHFEFVVMPFGLTNAP 284
Query: 842 ATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCS 901
A + ++M+ +F + + +++DD++V S H L+ ++LR +++ K S
Sbjct: 285 AAFMKMMNGVFRDFLDEFVIIFIDDILVHSKSWEAHQEHLRAVLERLREHELFAKLSKFS 344
Query: 902 FGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDK 961
F + FLG +++ +G+ V+P+K R+I E P + E++ G RF+
Sbjct: 345 FWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASM 404
Query: 962 AAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTV 1021
A P K++ F W++ECE++F +LK L PVL P P +Y + + V
Sbjct: 405 AQPLTRLTGKDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVYTDASIVGLGCV 464
Query: 1022 LLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPY 1067
L+Q K VI + S L+ E Y + A++ + R Y
Sbjct: 465 LMQ----KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFALKIWRSY 506
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 203 bits (517), Expect = 6e-52
Identities = 122/412 (29%), Positives = 207/412 (49%), Gaps = 20/412 (4%)
Query: 702 LAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKW 761
+ + PG P+ + RM+ + ++ + E L+ A V+ VKK +G +
Sbjct: 470 IELEPGTAPLSKAPYRMAPAEMAELKKQLEDLLGAP-------------VLFVKKKDGSF 516
Query: 762 RMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTF 821
R+C DY LN V K+ YPLP +D+L+D G S +D SGY+ I + +D T F
Sbjct: 517 RLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEADVRKTAF 576
Query: 822 MTNQANYCYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDL 881
T ++ + MPFGL NA A + RLM+ +F + + + +++DD++V S +H L
Sbjct: 577 RTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLEEHEVHL 636
Query: 882 KEAFDQLRTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEV 941
+ ++LR ++ KCSF + FLG ++++ G+ V+P+K AI + +PT+ E+
Sbjct: 637 RRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTPTNATEI 696
Query: 942 QRLTGRMAALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKP 1001
+ G RF+ A P K+ F W+ ECE+ F LKE L + PVL+ P
Sbjct: 697 RSFLGLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALP 756
Query: 1002 TPSVPLVLYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTA 1061
P ++Y + + VL+Q + KVI + S L+ E Y + A++
Sbjct: 757 EHGEPYMVYTDASGVGLGCVLMQ----RGKVIAYASRQLRKHEGNYPTHDLEMAAVIFAL 812
Query: 1062 RRLRPYFQSFQVKIKTD-VPLRQVLQKPDLSGRLVSWSVELSEYDIQ--YEP 1110
+ R Y V++ TD L+ + +P+L+ R W +++YD++ Y P
Sbjct: 813 KIWRSYLYGGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLEIAYHP 864
>At1g35370 hypothetical protein
Length = 1447
Score = 193 bits (490), Expect = 8e-49
Identities = 139/464 (29%), Positives = 233/464 (49%), Gaps = 21/464 (4%)
Query: 653 VKDKFLKIGS--GLTTE--QEDRLITLLGDNLDLFAWTINDVPGIDPKVITHKLAIRPGA 708
V D+ +IGS LT++ +E + ++ + D+FA D+P K HK+ + GA
Sbjct: 521 VSDEEQEIGSISALTSDVVEESVVQNIVEEFPDVFAEP-TDLPPFREKH-DHKIKLLEGA 578
Query: 709 TPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYT 768
PV Q R + + + +IK+ I+ P + + VV+VKK +G WR+C DYT
Sbjct: 579 NPVNQRPYRYVVHQKDEIDKIVQDMIKSGTIQVSSSP-FASPVVLVKKKDGTWRLCVDYT 637
Query: 769 SLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANY 828
LN + KD + +P ++ L+D G+ + S +D +GY+Q+ M P D + T F T+ ++
Sbjct: 638 ELNGMTVKDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQVRMDPDDIQKTAFKTHNGHF 697
Query: 829 CYKTMPFGLKNAGATYQRLMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQL 888
Y M FGL NA AT+Q LM+ +F + + + V+ DD+++ S+ +H L+ F+ +
Sbjct: 698 EYLVMLFGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILIYSSSIEEHKEHLRLVFEVM 757
Query: 889 RTYQMKLNPEKCSFGIQGGKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRM 948
R +++ F + LG +++R IE +P K +A+ E +PT+VK+V+ G
Sbjct: 758 RLHKL--------FAKGSKEHLGHFISAREIETDPAKIQAVKEWPTPTTVKQVRGFLGFA 809
Query: 949 AALSRFLPMAGDKAAPFFTCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLV 1008
RF+ G A P L K F W+ E + AF LK L PVL+ P +
Sbjct: 810 GYYRRFVRNFGVIAGPLH-ALTKTDGFCWSLEAQSAFDTLKAVLCNAPVLALPVFDKQFM 868
Query: 1009 LYLAVTDKAVSTVLLQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYF 1068
+ + + VL+Q K + ++S L+G +L EK LA + R+ R Y
Sbjct: 869 VETDACGQGIRAVLMQ----KGHPLAYISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYL 924
Query: 1069 QSFQVKIKTDV-PLRQVLQKPDLSGRLVSWSVELSEYDIQYEPR 1111
IKTD L+ +L++ + W +L E+D + + R
Sbjct: 925 LPSHFIIKTDQRSLKYLLEQRLNTPVQQQWLPKLLEFDYEIQYR 968
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 184 bits (467), Expect = 4e-46
Identities = 122/445 (27%), Positives = 196/445 (43%), Gaps = 73/445 (16%)
Query: 685 WTINDVPGIDPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREVQY 744
++++D+ GI P + H + + + ++ +RR++ V+ E KL+ A I +
Sbjct: 349 YSLDDITGISPTLCMHMIHLEGESITSVEHQRRLNSNLRDVVKKEIMKLLDAGIIYPISD 408
Query: 745 PTWLANVVMVKKANGK------------------WRMCTDYTSLNKVCPKDSYPLPNVDK 786
TW++ V +V K G RMC DY LN KD++PL +D+
Sbjct: 409 STWVSPVHVVPKKGGVIVIKNEKNELIPTRTVTGHRMCIDYRKLNSATRKDNFPLSFIDQ 468
Query: 787 LVDGASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQR 846
+++ S +D Y G+ QI++HP D+E TTF + Y+ MPFGL NA AT+Q
Sbjct: 469 MLERLSNQPYYCFLDGYLGFFQILIHPDDQEKTTFTCPYGTFAYRRMPFGLCNAPATFQH 528
Query: 847 LMDKIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQG 906
M IFS + MEV++DD +V H + LN EK F ++
Sbjct: 529 CMKYIFSDMIEDFMEVFIDDFLVLQRCEDKH---------------LVLNWEKSHFMVRD 573
Query: 907 GKFLGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFF 966
G LG ++ +G+EV+ K E+ R G RF+ A P
Sbjct: 574 GIVLGHKISEKGVEVDRAK-------------IEIMRFLGHAGFYRRFIKDFSKIARPCT 620
Query: 967 TCLKKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEE 1026
L K K ++ + C QAF +KE+L + ++ P +P + +D V VL Q +
Sbjct: 621 QLLCKEQKSEFDDTCLQAFNIVKESLVSALIVQPPEWELPFEVICDASDYVVGAVLGQRK 680
Query: 1027 GKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTDVPLRQVLQ 1086
KK IY+ S TL GA++ + A LR +L
Sbjct: 681 DKKLHAIYYASRTLDGAQVIVHTVHAA---------------------------LRYLLS 713
Query: 1087 KPDLSGRLVSWSVELSEYDIQYEPR 1111
K D RL+ W + L ++D++ + +
Sbjct: 714 KKDAKPRLLRWILLLQDFDLEIKDK 738
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 182 bits (463), Expect = 1e-45
Identities = 82/158 (51%), Positives = 115/158 (71%)
Query: 683 FAWTINDVPGIDPKVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLETEKLIKARFIREV 742
F + + GI +VI+H+L + P PV Q RR++ ++ +AV +E +L++ IREV
Sbjct: 405 FPTPVGKMTGISTEVISHELNVDPTFKPVKQKRRKLGPDRAQAVNIEVVRLLEVGRIREV 464
Query: 743 QYPTWLANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVDGASGNELLSLMDA 802
+YP WLAN V+VKK NGKWR+C D+T LNK C KD +PLP++D+LV+ +G+E+LS MDA
Sbjct: 465 KYPEWLANPVVVKKKNGKWRVCVDFTDLNKACSKDFFPLPHIDRLVESTTGHEMLSFMDA 524
Query: 803 YSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNA 840
+SGYNQI+M+P D+E T+F+T Y YK MPFGLKNA
Sbjct: 525 FSGYNQILMNPEDQEKTSFITECGTYYYKVMPFGLKNA 562
Score = 122 bits (307), Expect = 1e-27
Identities = 95/333 (28%), Positives = 153/333 (45%), Gaps = 76/333 (22%)
Query: 964 PFFTCLKKNSK-FQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVL 1022
PF K SK F W E CE+AF +LK L+ VL+K L LY+AV++ AV+ V
Sbjct: 617 PFLHFTAKISKGFIWNESCEEAFKQLKRYLSEPSVLAKREFGEQLFLYIAVSESAVTGVQ 676
Query: 1023 LQEEGKKQKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTDVPLR 1082
++ E ++ I++V + RY +EK ALA++ AR+LRPYFQS + + T +PLR
Sbjct: 677 VRVERSDKRPIFYV-------KTRYPMMEKLALAVVTAARKLRPYFQSHPIVVLTSLPLR 729
Query: 1083 QVLQKPDLSGRLVSWSVELSEYDIQYEPRGQVTVQSLIDFVAELTPTEGEKTQGEWVLSV 1142
+L P SGRL W++ELSE+D+++ R + ++ S +D T +W
Sbjct: 730 TILHSPTQSGRLGKWAIELSEFDLEF--RARTSLNSTMD------------TSRQWGFI- 774
Query: 1143 DGSSNNTGSGAGITIESPDKMIIEQSLKFEFKASNNQSEYEALIAGLRLAIELGVQKLFI 1202
+ ++L A+ EYEA A + + L
Sbjct: 775 ------------------KTWVRSRNLSDAPTANRYSGEYEAKDACMEAYLNL------- 809
Query: 1203 KGDSQLVVKQVKGEYQVKDPQLSKYLEVVRRLMMEVKEIKIEHVPRGQNERADVLAKLAS 1262
V++V G ++ + ++ +PR +N A+ LA LAS
Sbjct: 810 -------VREVSGRFE---------------------QFELTRIPRAENSAANALAALAS 841
Query: 1263 TGRLGNYQTVIQETLPRPSIDLVEIKLKAVKSV 1295
T + + + ET+ +PSI L EI +++
Sbjct: 842 TFEVTLPRVIPVETISQPSIRLDEISFVTTRAM 874
Score = 66.6 bits (161), Expect = 1e-10
Identities = 51/170 (30%), Positives = 76/170 (44%), Gaps = 14/170 (8%)
Query: 7 LDTLQKVQSQNEQLQAQVEYLTQRQDFKQEQRLEAEDVVEFQ---PFVPAITRVDIPKHL 63
L L + S+ + LQ Q+ + + E E V+E PF P I+++ I +
Sbjct: 112 LRILSPLNSELQNLQDQIWAMNAKVHQATTSAPEVEKVIEATRRTPFTPRISKLRI-REF 170
Query: 64 QTMALDAFSGESDPMEHLRYFNTKMVIGGAT-------DAVKCRLLPSTFKGMAMQWFIR 116
+ L ++G+ D EHL F VI G DA C+L G+A+ WF +
Sbjct: 171 RDFKLPVYNGKGDLKEHLTSFQ---VIAGRVPLEPHEEDAGLCKLFSENLFGLALTWFTQ 227
Query: 117 QPPFSIDNFTDLSTKFLTQFSANKTTKATMFDLISIHQQPGEKLKTYMAR 166
SIDNF LST F+ Q+ + T L + Q E L+TY+ R
Sbjct: 228 LEEGSIDNFKQLSTAFIKQYEYFINSDITEAHLWNFSQSADEPLRTYIYR 277
Score = 32.7 bits (73), Expect = 1.9
Identities = 30/82 (36%), Positives = 39/82 (46%), Gaps = 6/82 (7%)
Query: 508 KPYKGTLVGFTGDRVSVRGYVEIP-TAFGEGEFVKKFQVKYLVLACRANYNVLLGRDTLN 566
KP GT + G V +P TA G VK V++ V A YNV+LG L
Sbjct: 336 KPAAGTCTTMLCETSMSLGTVVLPVTAQG---VVK--MVEFTVFDRPAAYNVILGTPWLY 390
Query: 567 KVCAVISTAHLTVKYPACNGKV 588
++ V ST H VK+P GK+
Sbjct: 391 EMKVVPSTYHQCVKFPTPVGKM 412
>At2g06890 putative retroelement integrase
Length = 1215
Score = 181 bits (460), Expect = 2e-45
Identities = 131/443 (29%), Positives = 211/443 (47%), Gaps = 39/443 (8%)
Query: 672 LITLLGDNLDLFAWTINDVP-GIDP-KVITHKLAIRPGATPVIQPRRRMSEEKNKAVQLE 729
+ +LL D D+F D P G+ P + I H++ PGA+ +P R + + K +Q
Sbjct: 408 MTSLLQDYKDVFP---EDNPKGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVETKELQ-- 462
Query: 730 TEKLIKARFIREVQYPTWLANVVMVKKANGKWRMCTDYTSLNKVCPKDSYPLPNVDKLVD 789
++ +G WRMC D ++N V K +P+P +D ++D
Sbjct: 463 -------------------------RQKDGSWRMCFDCRAINNVTVKYCHPIPRLDDMLD 497
Query: 790 GASGNELLSLMDAYSGYNQIMMHPSDEESTTFMTNQANYCYKTMPFGLKNAGATYQRLMD 849
G+ + S +D SGY+QI M+ DE T F T Y + MPFGL +A +T+ RLM+
Sbjct: 498 ELHGSSIFSKIDLKSGYHQIRMNEGDEWKTAFKTKHGLYEWLVMPFGLTHAPSTFMRLMN 557
Query: 850 KIFSKQVGRNMEVYVDDMIVKSARASDHGGDLKEAFDQLRTYQMKLNPEKCSFGIQGGKF 909
+ +G + VY DD++V S +H L + LR ++ N +KC+F F
Sbjct: 558 HVLRAFIGIFVIVYFDDILVYSESLREHIEHLDSVLNVLRKEELYANLKKCTFCTDNLVF 617
Query: 910 LGFMLTSRGIEVNPDKGRAILEMKSPTSVKEVQRLTGRMAALSRFLPMAGDKAAPFFTCL 969
LGF++++ G++V+ +K +AI + SP +V EV+ G RF AP +
Sbjct: 618 LGFVVSADGVKVDEEKVKAIRDWPSPKTVGEVRSFHGLAGFYRRFFKDFSTIVAPLTEVM 677
Query: 970 KKNSKFQWTEECEQAFTKLKETLATLPVLSKPTPSVPLVLYLAVTDKAVSTVLLQEEGKK 1029
KK+ F+W + E+AF LK+ L PVL + + + VL+Q+
Sbjct: 678 KKDVGFKWEKAQEEAFQSLKDKLTNAPVLILSEFLKTFEIECDASGIGIGAVLMQD---- 733
Query: 1030 QKVIYFVSHTLQGAELRYQKIEKAALAILKTARRLRPYFQSFQVKIKTD-VPLRQVLQKP 1088
QK+I F S L GA L Y +K A+++ +R + Y I TD L+ + +
Sbjct: 734 QKLIAFFSEKLGGATLNYPTYDKELYALVRALQRWQHYLWPKVFVIHTDHESLKHLKGQQ 793
Query: 1089 DLSGRLVSW--SVELSEYDIQYE 1109
L+ R W +E Y I+Y+
Sbjct: 794 KLNKRHARWVEFIETFAYVIKYK 816
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,137,525
Number of Sequences: 26719
Number of extensions: 1650984
Number of successful extensions: 6687
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 80
Number of HSP's successfully gapped in prelim test: 60
Number of HSP's that attempted gapping in prelim test: 6316
Number of HSP's gapped (non-prelim): 297
length of query: 1706
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1593
effective length of database: 8,299,349
effective search space: 13220862957
effective search space used: 13220862957
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0046b.4


