

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046a.5
(327 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
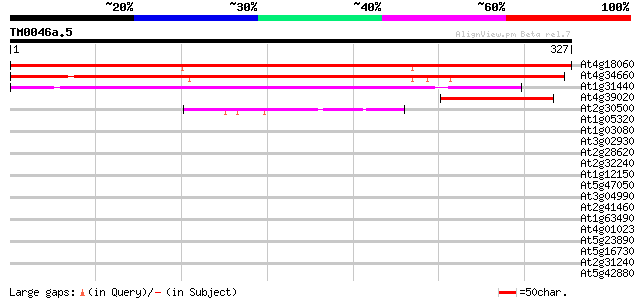
Sequences producing significant alignments: (bits) Value
At4g18060 unknown protein 464 e-131
At4g34660 SH3 domain-containing protein 2 308 2e-84
At1g31440 unknown protein 199 2e-51
At4g39020 putative protein 77 2e-14
At2g30500 unknown protein 43 3e-04
At1g05320 unknown protein 39 0.005
At1g03080 unknown protein 38 0.009
At3g02930 unknown protein 36 0.025
At2g28620 putative kinesin-like spindle protein 36 0.025
At2g32240 putative myosin heavy chain 36 0.033
At1g12150 hypothetical protein 35 0.043
At5g47050 unknown protein 35 0.056
At3g04990 hypothetical protein 35 0.056
At2g41460 DNA-(apurinic or apyrimidinic site) lyase (ARP) 35 0.073
At1g63490 RB-binding protein -like 35 0.073
At4g01023 unknown protein 34 0.095
At5g23890 unknown protein 33 0.16
At5g16730 putative protein 33 0.16
At2g31240 putative kinesin light chain protein (At2g31240) 33 0.21
At5g42880 putative protein 33 0.28
>At4g18060 unknown protein
Length = 351
Score = 464 bits (1193), Expect = e-131
Identities = 239/351 (68%), Positives = 282/351 (80%), Gaps = 24/351 (6%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA R+QASKLR+QV+KQQ AVIKQFS +GYESSDV+VIDE EMQ H QL+KLY++TR+
Sbjct: 1 MDAFRRQASKLRDQVAKQQLAVIKQFSGTGYESSDVMVIDELEMQRHHQLDKLYRSTRSA 60
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAEN--NID-NILAKAASVLGDARK 117
++FQ++IVKAAE FT IG +HIE GTKLSEDCC+YG EN NID NILAKAA++ GDARK
Sbjct: 61 KEFQRDIVKAAEAFTTIGLRHIEAGTKLSEDCCRYGNENSQNIDENILAKAAAIYGDARK 120
Query: 118 HVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVR 177
HV+KE E+ N+LLASQVLDPLR M+ G PLEDARHLAQRYSRMRQEAET E+S+RQ R
Sbjct: 121 HVDKEQEDFNKLLASQVLDPLRAMVAGSPLEDARHLAQRYSRMRQEAETHATEVSRRQAR 180
Query: 178 VRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
VRE+P E VAKL AEA+M+ELKANMAVLGKEA AALA+V++QQ RLTFQRLVAM
Sbjct: 181 VREAPIPENVAKLQLAEAKMQELKANMAVLGKEATAALAAVESQQHRLTFQRLVAMVEGE 240
Query: 234 -----------------MVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKEL 276
MV+++Q KESAPP + +ENGS KT YFLAE HPF SEKEL
Sbjct: 241 KNYHLRIAAILSDIEADMVTEKQHKESAPPAIPTENGSEKTSYFLAEVIHPFSAASEKEL 300
Query: 277 SFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSSNFSSEVY 327
KGD+IVVRKV+ TGW+EGEC GKAGWFP AY+EKRQR+P++NF++EVY
Sbjct: 301 DLDKGDYIVVRKVSQTGWAEGECKGKAGWFPMAYIEKRQRLPTTNFAAEVY 351
>At4g34660 SH3 domain-containing protein 2
Length = 368
Score = 308 bits (790), Expect = 2e-84
Identities = 174/368 (47%), Positives = 232/368 (62%), Gaps = 48/368 (13%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA+RKQAS+LREQV++QQQAV KQF GY S + DE E+ HQ+LEKLY +TR
Sbjct: 1 MDAIRKQASRLREQVARQQQAVFKQFGGGGYGSG---LADEAELNQHQKLEKLYISTRAA 57
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDN--ILAKAASVLGDARKH 118
+ +Q++IV+ E + G K +E GTKLSED KYG+EN N +L +AA G AR
Sbjct: 58 KHYQRDIVRGVEGYIVTGSKQVEIGTKLSEDSRKYGSENTCTNGNVLTRAALNYGRARAQ 117
Query: 119 VEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRV 178
+EKE + + L +QV +PLR M+ G PLEDARHLAQRY RMRQEAE E+++RQ +
Sbjct: 118 MEKERGNMLKALGTQVAEPLRAMVLGAPLEDARHLAQRYDRMRQEAEAQATEVARRQAKA 177
Query: 179 RESPTSEQVA-KLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
RES + + KL +AEA++ +LK+NM +LGKEAA+ALASV+ QQQ+LT +RL++M
Sbjct: 178 RESQGNPDILMKLESAEAKLHDLKSNMTILGKEAASALASVEDQQQKLTLERLLSMVESE 237
Query: 234 -----------------MVSDRQKKE---------SAPPVVVSENGSG------------ 255
MVS+RQ+ E S PP E +G
Sbjct: 238 RAYHQRVLQILDQLEGEMVSERQRIEAPSTPSSADSMPPPPSYEEANGVFASQMHDTSTD 297
Query: 256 KTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQ 315
YFL E P++G ++ ELS S G+++VVRKVT +GW+EGEC GKAGWFP Y+E+R+
Sbjct: 298 SMGYFLGEVLFPYHGVTDVELSLSTGEYVVVRKVTGSGWAEGECKGKAGWFPYGYIERRE 357
Query: 316 RVPSSNFS 323
RV +S S
Sbjct: 358 RVLASKVS 365
>At1g31440 unknown protein
Length = 439
Score = 199 bits (506), Expect = 2e-51
Identities = 121/300 (40%), Positives = 175/300 (58%), Gaps = 12/300 (4%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
M+A+RKQA+KLREQV++QQQAV+K G+ ++D VV+DE E+ HQ+L+ LY +T+
Sbjct: 1 MEAIRKQAAKLREQVARQQQAVLKHL---GHVNADAVVVDEEELHCHQKLQDLYSSTKAA 57
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNI-LAKAASVLGDARKHV 119
+ Q+ IV+ E F A G K +E G K +ED KYG EN N L++ A G + K V
Sbjct: 58 KRLQRNIVRGLEGFIATGTKVVEIGLKFAEDFKKYGDENPDANTPLSRVAHHFGTSYKSV 117
Query: 120 EKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVR 179
E E E L +L+ QV +P+R MI PLEDARHL Y R+RQE E ++ +R+ +++
Sbjct: 118 EGERETLLGVLSEQVCEPIRTMIYSAPLEDARHLVNHYDRLRQEVEAQATDVLRRRSKLK 177
Query: 180 ESPTSEQV-AKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDR 238
ES SE+ KL +E+R+ ELK++M LGKEA A+ VD QQQ +T QRL A++ ++R
Sbjct: 178 ESDISEEAYIKLKNSESRLAELKSSMKTLGKEATKAMLEVDDQQQNVTSQRLRALVEAER 237
Query: 239 QKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGE 298
+A ++ ++ A G S K L D + + P S GE
Sbjct: 238 SYHRNALDIL-------DKLHSEMIAEEEAIGSSPKSLPLHIEDSASLPQQEPNSNSSGE 290
>At4g39020 putative protein
Length = 169
Score = 76.6 bits (187), Expect = 2e-14
Identities = 34/67 (50%), Positives = 45/67 (66%), Gaps = 1/67 (1%)
Query: 252 NGSGKTM-YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAY 310
NG+ M YFL E P+ DS+ ELS S GD++V+R+V + W+EGEC G AGWF Y
Sbjct: 94 NGTSDAMGYFLGEVMFPYQADSDFELSLSVGDYVVIREVVSSVWAEGECKGNAGWFTYIY 153
Query: 311 VEKRQRV 317
+E+R RV
Sbjct: 154 IERRDRV 160
>At2g30500 unknown protein
Length = 517
Score = 42.7 bits (99), Expect = 3e-04
Identities = 37/144 (25%), Positives = 71/144 (48%), Gaps = 18/144 (12%)
Query: 102 DNILAKAASVLGDARKHVEKEHE----ELNRLLA--SQVLDPLRQMINGVPL-------- 147
DN + + + DA + + E E++++L SQ+ + LR++ + + L
Sbjct: 346 DNEIRALKTAVSDAEQKIFPEKAQIKGEMSKMLEERSQLGEQLRELESHIRLIKEEKAET 405
Query: 148 -EDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAV 206
E R ++ S MR E+ LREEI KR+ +++E T + + +LH + R++ + +
Sbjct: 406 EEKLRGGTEKISGMRDESNVLREEIGKREEKIKE--TEKHMEELHMEQVRLRRRSSELTE 463
Query: 207 LGKEAAAALASVDAQQQRLTFQRL 230
E AS A+Q+R ++L
Sbjct: 464 -EVERTRVSASEMAEQKREAIRQL 486
>At1g05320 unknown protein
Length = 841
Score = 38.5 bits (88), Expect = 0.005
Identities = 45/222 (20%), Positives = 99/222 (44%), Gaps = 18/222 (8%)
Query: 6 KQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEG---------EMQIHQQLEKLYKA 56
K L ++V+K+ + K+ + ++ V V EG + + +QL+ L A
Sbjct: 21 KYCDDLLQEVTKEDTVMEKEEEDTIFDGGFVKVEKEGINKKYDDDDDEKAEKQLKSLEDA 80
Query: 57 TRTMRDFQKEIVKAAETFTAIGYKHIETGTKL--SEDCCKYGA--ENNIDNILAKAASVL 112
+ KE+ + E F +G + + K+ ED + A ++ + ++AS L
Sbjct: 81 LQLHDVKHKELTEVKEAFDGLGLELENSRKKMIELEDRIRISALEAEKLEELQKQSASEL 140
Query: 113 GDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEIS 172
+ K ++ + + + LL SQ L + + L+ L+++ S ++ EE
Sbjct: 141 EEKLKISDERYSKTDALL-SQALS--QNSVLEQKLKSLEELSEKVSELKSALIVAEEEGK 197
Query: 173 KRQVRVRE--SPTSEQVAKLHAAEARMKELKANMAVLGKEAA 212
K ++++E S+ + L+ + AR EL+ ++ + ++ A
Sbjct: 198 KSSIQMQEYQEKVSKLESSLNQSSARNSELEEDLRIALQKGA 239
Score = 28.1 bits (61), Expect = 6.8
Identities = 24/98 (24%), Positives = 47/98 (47%), Gaps = 21/98 (21%)
Query: 4 LRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDF 63
L+++ K+ E SK Q+ +++SD V++E +Q+H++L+ A+ T
Sbjct: 627 LKEEVEKVAELTSKLQE--------HKHKASDRDVLEEKAIQLHKELQ----ASHTAISE 674
Query: 64 QKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNI 101
QKE A+ +KH E L + + A+ ++
Sbjct: 675 QKE---------ALSHKHSELEATLKKSQEELDAKKSV 703
>At1g03080 unknown protein
Length = 1744
Score = 37.7 bits (86), Expect = 0.009
Identities = 49/165 (29%), Positives = 72/165 (42%), Gaps = 17/165 (10%)
Query: 112 LGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPL--EDARHLAQRYSRMRQEAETLRE 169
L DA V+ E +E + Q L+ L + + V ED+R L +R +R E ETLRE
Sbjct: 218 LKDALSKVQAE-KEASLAQFDQNLEKLSNLESEVSRAQEDSRVLIERATRAEAEVETLRE 276
Query: 170 EISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQR 229
+SK +V S Q + A +L+ +++ KEA VD + R +
Sbjct: 277 SLSKVEVEKESSLLQYQQCLQNIA-----DLEDRISLAQKEA----GEVDERANRAEAET 327
Query: 230 LV--AMMVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDS 272
L +VS KE+A +V KT+ L E H DS
Sbjct: 328 LALKQSLVSSETDKEAA---LVQYQQCLKTISNLEERLHKAEEDS 369
Score = 33.1 bits (74), Expect = 0.21
Identities = 45/199 (22%), Positives = 80/199 (39%), Gaps = 24/199 (12%)
Query: 43 EMQIHQQLEKLYKATRTMRDFQKEIVKAAETFTAIGYKHIETGT---KLSEDCCKYGAEN 99
E+Q Q L+ T+ D + ++ A E + + IE G K +E+ C +
Sbjct: 401 ELQYQQCLD-------TIADLKLKLFHAQEETQRLS-REIEDGVAKLKFAEEKCVVLERS 452
Query: 100 N------IDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHL 153
N +D +L K LG+ + ++ +EL RL + LR M + L
Sbjct: 453 NQNLHSELDGLLEK----LGNQSHELTEKQKELGRLWTCVQEENLRFMEAETAFQT---L 505
Query: 154 AQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAA 213
Q +S+ ++E TL E+ R +++ + EA+ + N L A+
Sbjct: 506 QQLHSQSQEELSTLALELQNRSQILKDMEARNNGLQEEVQEAKDQSKSLNELNLSSAASI 565
Query: 214 ALASVDAQQQRLTFQRLVA 232
+ + R T Q+L A
Sbjct: 566 KSLQEEVSKLRETIQKLEA 584
>At3g02930 unknown protein
Length = 806
Score = 36.2 bits (82), Expect = 0.025
Identities = 33/150 (22%), Positives = 75/150 (50%), Gaps = 14/150 (9%)
Query: 100 NIDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSR 159
+++N AKA L +ARK E+ E+L+ L +Q ++ + +E +
Sbjct: 105 SLENEKAKALDQLKEARKEAEEASEKLDEALEAQ-----KKSLENFEIEKFEVVEAGIEA 159
Query: 160 MRQEAETLREEISK-RQVRVRESPT----SEQVAKLHAAEARMKELKANMAVLGKEAAAA 214
++++ E L++E+ + ES T ++++ ++ A K+ K+ A+ + A+
Sbjct: 160 VQRKEEELKKELENVKNQHASESATLLLVTQELENVNQELANAKDAKSK-ALCRADDASK 218
Query: 215 LASVDAQQQRLTFQ---RLVAMMVSDRQKK 241
+A++ A++ + RL A++ S R+K+
Sbjct: 219 MAAIHAEKVEILSSELIRLKALLDSTREKE 248
>At2g28620 putative kinesin-like spindle protein
Length = 1076
Score = 36.2 bits (82), Expect = 0.025
Identities = 45/206 (21%), Positives = 96/206 (45%), Gaps = 26/206 (12%)
Query: 11 LREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDFQKEIVKA 70
L +V+K A+ F G+ S +++ + +H Q EKL T+ RD + +
Sbjct: 657 LNSEVTKHSCALEDMFK--GFTSEAYTLLEGLQGSLHNQEEKLSAFTQQQRDLHSRSMDS 714
Query: 71 AETFTAI---GYKHIETG----TKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKEH 123
A++ + + +K ++T TKL+ED A+N + L+ ++ + EK+
Sbjct: 715 AKSVSTVMLDFFKTLDTHANKLTKLAED-----AQNVNEQKLSAFTKKFEESIANEEKQM 769
Query: 124 -EELNRLLASQVLDPLRQMINGVPLEDARH--------LAQRYSRMRQEAETLREEISKR 174
E++ LLAS + ++ + + ++D R L Q S M+ A +++ + +
Sbjct: 770 LEKVAELLASS--NARKKELVQIAVQDIRQGSSSQTGALQQEMSAMQDSASSIKVQWNSH 827
Query: 175 QVRVRESPTSEQVAKLHAAEARMKEL 200
V+ ES + ++ + A+ M+++
Sbjct: 828 IVQA-ESHHLDNISAVEVAKEDMQKM 852
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 35.8 bits (81), Expect = 0.033
Identities = 41/189 (21%), Positives = 82/189 (42%), Gaps = 19/189 (10%)
Query: 39 IDEGEMQIHQQLEKLYKATRTMRDFQKEIVKAAETFTAIGYKHIETGTKL--SEDCCKYG 96
I EGE + QL+ L A ++ KE+ + E F A+G + + KL E+ K
Sbjct: 151 IVEGEERHSSQLKSLEDALQSHDAKDKELTEVKEAFDALGIELESSRKKLIELEEGLKRS 210
Query: 97 AENNIDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQR 156
AE K + + H + E ++ L S++L + E A+ + ++
Sbjct: 211 AEE-----AQKFEELHKQSASHADSESQK--ALEFSELLKSTK--------ESAKEMEEK 255
Query: 157 YSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALA 216
+ ++QE + L E++S+ + E+ +L A + + K+ + ++ ++ A
Sbjct: 256 MASLQQEIKELNEKMSENE--KVEAALKSSAGELAAVQEELALSKSRLLETEQKVSSTEA 313
Query: 217 SVDAQQQRL 225
+D Q L
Sbjct: 314 LIDELTQEL 322
Score = 32.0 bits (71), Expect = 0.47
Identities = 51/258 (19%), Positives = 95/258 (36%), Gaps = 44/258 (17%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVV----------------------V 38
++ L S +E K+ + I++F+ ESSD+V V
Sbjct: 909 LEGLIGSGSVEKETALKRLEEAIERFNQKETESSDLVEKLKTHENQIEEYKKLAHEASGV 968
Query: 39 IDEGEMQIHQQLEKLYKATRTMRD-------FQKEIVKAAETFTAIGYKHIETGTKLSED 91
D ++++ L KL T+ + +KE AE + + G++ +E
Sbjct: 969 ADTRKVELEDALSKLKNLESTIEELGAKCQGLEKESGDLAEVNLKLNLELANHGSEANEL 1028
Query: 92 CCKYGA----ENNIDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPL 147
K A + N L + + + D K + E E+L ++S + +N +
Sbjct: 1029 QTKLSALEAEKEQTANELEASKTTIEDLTKQLTSEGEKLQSQISSHTEE--NNQVNAMFQ 1086
Query: 148 EDARHLAQRYSRMRQE-------AETLREEISKRQVRVRESPTSEQVAKLHAAEARMKEL 200
L +++ ++ A+TL EI K + E E + E + E+
Sbjct: 1087 STKEELQSVIAKLEEQLTVESSKADTLVSEIEKLRAVAAEKSVLE--SHFEELEKTLSEV 1144
Query: 201 KANMAVLGKEAAAALASV 218
KA + + AA A V
Sbjct: 1145 KAQLKENVENAATASVKV 1162
>At1g12150 hypothetical protein
Length = 548
Score = 35.4 bits (80), Expect = 0.043
Identities = 24/90 (26%), Positives = 44/90 (48%), Gaps = 2/90 (2%)
Query: 138 LRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARM 197
LR ++N + +E +R ++EAE L E +K+ +++ + K A EAR
Sbjct: 324 LRSLVNSLRMELEDLRREREELQQKEAERLEIEETKKLEALKQESLKLEQMKTEAIEARN 383
Query: 198 KELKANMAV--LGKEAAAALASVDAQQQRL 225
+ N + L KE AA+ + + ++RL
Sbjct: 384 EAANMNRKIESLKKETEAAMIAAEEAEKRL 413
>At5g47050 unknown protein
Length = 300
Score = 35.0 bits (79), Expect = 0.056
Identities = 35/144 (24%), Positives = 61/144 (42%), Gaps = 18/144 (12%)
Query: 82 IETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDPLRQM 141
+ TG +LS + ++N L+ + GD ++ + +ELNR L
Sbjct: 79 VSTGLRLSRE----QSQNQEQRFLS--FPITGDVAGEIKSQTDELNRFL----------Q 122
Query: 142 INGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELK 201
I G L+ R LA+ R +E EE +R++R +E+ + + EAR +++
Sbjct: 123 IQGEQLK--RMLAENSERNYRELLRTTEESVRRRLREKEAEIEKATRRHVELEARATQIE 180
Query: 202 ANMAVLGKEAAAALASVDAQQQRL 225
AAA A + Q +L
Sbjct: 181 TEARAWQMRAAAREAEATSLQAQL 204
>At3g04990 hypothetical protein
Length = 227
Score = 35.0 bits (79), Expect = 0.056
Identities = 37/134 (27%), Positives = 61/134 (44%), Gaps = 23/134 (17%)
Query: 110 SVLGDARKHVEKEHEELN--RLLASQVLDPLRQMINGVPLED----------------AR 151
S +GD +K VE+ EEL R L + LD L ++ + L+D AR
Sbjct: 65 SEVGDLKKLVEECTEELRSKRNLLTVKLDSLIRVQRELELKDNQLVQVMAELKRRYSEAR 124
Query: 152 HLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEA 211
H+ +R M E T ++E+S +++ES + K E + KE++ GKE
Sbjct: 125 HVQKRKREMEDETATKKKELSMTVDQIQES-GKQLEKKSREVELKDKEIEEK----GKEL 179
Query: 212 AAALASVDAQQQRL 225
+ V A +++L
Sbjct: 180 DLVKSQVKAWERKL 193
>At2g41460 DNA-(apurinic or apyrimidinic site) lyase (ARP)
Length = 536
Score = 34.7 bits (78), Expect = 0.073
Identities = 42/200 (21%), Positives = 78/200 (39%), Gaps = 22/200 (11%)
Query: 61 RDFQKEIVKAAETFTAIGYKHIET---GTKLSEDCCKYGAENNIDNILAKAASVLGDARK 117
R F K ++ A F+ K E G + ++C + G++ + D + L D RK
Sbjct: 41 RSFNKRLMSNATAFSINNSKRKELKIPGAAIDQNCHQMGSDTDRDEM-----GTLQDDRK 95
Query: 118 HVE----KEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREE--I 171
+E +E R L V +++I+ + L H+ ++ + + R +
Sbjct: 96 EIEAMTVQELRSTLRKLGVPVKGRKQELISTLRL----HMDSNLPDQKETSSSTRSDSVT 151
Query: 172 SKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLV 231
KR++ RE PT ++ A + E + + A A A++Q+ L+
Sbjct: 152 IKRKISNREEPTEDECTNSEAYDIEHGEKRVKQSTEKNLKAKVSAKAIAKEQK----SLM 207
Query: 232 AMMVSDRQKKESAPPVVVSE 251
Q KE + SE
Sbjct: 208 RTGKQQIQSKEETSSTISSE 227
>At1g63490 RB-binding protein -like
Length = 1458
Score = 34.7 bits (78), Expect = 0.073
Identities = 50/219 (22%), Positives = 91/219 (40%), Gaps = 27/219 (12%)
Query: 17 KQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDFQKEIVKAAETFTA 76
K Q+ I Q +SG + S + + MQ+ Q R K +++A++ A
Sbjct: 681 KTQETKISQRPSSGTKRSIALNKKQEGMQVSQA-----------RPADKWLLRASKVLDA 729
Query: 77 IGYKHIETGTKLSEDCCKYGAENNIDNI------LAKA---ASVLGDARKHVEKEHEELN 127
+ +E T L E A + +D + L KA A + D VE E + +
Sbjct: 730 -AFSSVEYATLLKESEQFLWAGSEMDRVRDVTKSLNKAKIWAEAVSDCLSKVEGEVNDDS 788
Query: 128 RLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQV 187
+ + +D L + +N VP ++ +L +++ AE R+ K + SPT Q+
Sbjct: 789 MKVHLEFIDELLR-VNPVPCFNSGYL-----KLKDYAEEARKLSEKIDSALSSSPTITQL 842
Query: 188 AKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLT 226
LH+ +R +L K+ ++A ++ LT
Sbjct: 843 ELLHSEVSRSPISLKKHEILSKKISSAKMLAKRAKRYLT 881
>At4g01023 unknown protein
Length = 255
Score = 34.3 bits (77), Expect = 0.095
Identities = 24/92 (26%), Positives = 48/92 (52%), Gaps = 4/92 (4%)
Query: 114 DARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISK 173
+ ++ +E+E E+ R + +V L + +DA+ +AQ+ +M +E ET+ E+++
Sbjct: 155 EIKERIEEEREKA-RAMVKEVTAMLEKA--STMADDAKGVAQKVVKMVEEIETMVEKVAA 211
Query: 174 RQVRVRESPTSEQVAKLHAAEARMKELKANMA 205
+ E+ T + AE M+ KANM+
Sbjct: 212 MATKAGETATM-AADMVKEAEETMETAKANMS 242
>At5g23890 unknown protein
Length = 946
Score = 33.5 bits (75), Expect = 0.16
Identities = 53/243 (21%), Positives = 100/243 (40%), Gaps = 29/243 (11%)
Query: 15 VSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM-RDFQKEIVKAAET 73
V+K Q A+ + S E+SD+V + ++ EK A + + +K++ + E
Sbjct: 618 VTKGQAAI----ALSSGEASDIVSEELARIEAESMAEKAVSAHNALVAEVEKDVNASFEK 673
Query: 74 FTAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAAS---VLGDARKHVEKEHEELNRLL 130
++ + IE K++E A+ ++ + K L R VE E E L+RL
Sbjct: 674 ELSMEREKIEAVEKMAEL-----AKVELEQLREKREEENLALVKERAAVESEMEVLSRLR 728
Query: 131 --ASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVA 188
A + L+ L + E +R +R+EAE + ISK Q + + +A
Sbjct: 729 RDAEEKLEDLMSNKAEITFEK-----ERVFNLRKEAEEESQRISKLQYELEVERKALSMA 783
Query: 189 KLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDRQKKESAPPVV 248
+ A E K +E AL + + + +V + + +E+ +V
Sbjct: 784 RSWAEEEAKK---------AREQGRALEEARKRWETNGLRVVVDKDLQETSSRETEQSIV 834
Query: 249 VSE 251
++E
Sbjct: 835 LNE 837
>At5g16730 putative protein
Length = 853
Score = 33.5 bits (75), Expect = 0.16
Identities = 49/219 (22%), Positives = 86/219 (38%), Gaps = 12/219 (5%)
Query: 7 QASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDFQKE 66
+ + L+E++ + V KQ + ++E + +++EKL T+++ +
Sbjct: 367 EITDLKERIVTLETTVAKQKEDLEVSEQRLGSVEEEVSKNEKEVEKLKSELETVKEEKNR 426
Query: 67 IVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEEL 126
+K + T+ + E +KL D E KA L A V E EL
Sbjct: 427 ALKKEQDATSRVQRLSEEKSKLLSDLESSKEEEEKSK---KAMESLASALHEVSSEGREL 483
Query: 127 NRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQ 186
L SQ I+ + L + ++Y M EA R EI V ++ +
Sbjct: 484 KEKLLSQGDHEYETQIDDLKLV-IKATNEKYENMLDEA---RHEIDVLVSAVEQTKKHFE 539
Query: 187 VAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRL 225
+K + MKE AN+ K+ +AS+ + RL
Sbjct: 540 SSK---KDWEMKE--ANLVNYVKKMEEDVASMGKEMNRL 573
Score = 32.0 bits (71), Expect = 0.47
Identities = 49/234 (20%), Positives = 88/234 (36%), Gaps = 27/234 (11%)
Query: 13 EQVSKQQQAVIKQFST-SGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDFQKEIVKAA 71
E V ++ + K+ T +SD + + + Q+LEK+ + D + + + A
Sbjct: 169 EAVQNNEEELKKELETVKNQHASDSAAL----VAVRQELEKINEELAAAFDAKSKALSQA 224
Query: 72 ETFTAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEELNRLLA 131
E + H E LS + + A +D+ K A + +E E L R L
Sbjct: 225 EDASKTAEIHAEKVDILSSELTRLKAL--LDSTREKTAISDNEMVAKLEDEIVVLKRDLE 282
Query: 132 S------------QVLDPLRQMINGVPLED--ARHLAQRYSRMRQEAETLREEISK--RQ 175
S +++ L + + + A L+ + +E E EE +K R
Sbjct: 283 SARGFEAEVKEKEMIVEKLNVDLEAAKMAESNAHSLSNEWQSKAKELEEQLEEANKLERS 342
Query: 176 VRVRESPTSEQVA----KLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRL 225
V +Q+ KLH E + +LK + L A ++ +QRL
Sbjct: 343 ASVSLESVMKQLEGSNDKLHDTETEITDLKERIVTLETTVAKQKEDLEVSEQRL 396
>At2g31240 putative kinesin light chain protein (At2g31240)
Length = 617
Score = 33.1 bits (74), Expect = 0.21
Identities = 52/233 (22%), Positives = 90/233 (38%), Gaps = 13/233 (5%)
Query: 95 YGAENNIDNILAKAASVLGDARKHVEKEHEEL--NRLLASQVLDPLRQMINGVPLEDARH 152
Y A N + L A L +K + E+ +R L + L Q + LE R
Sbjct: 262 YVAVLNFNEALPYALKALEIHKKELGNNSAEVAQDRRLLGVIYSGLEQ--HDKALEQNRL 319
Query: 153 LAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAA 212
+ + E +R EI ++V E + L + + + A++ +
Sbjct: 320 SQRVLKNWGMKLELIRAEIDAANMKVALGKYEEAIDILKSVVQQTDKDSEMRAMVFISMS 379
Query: 213 AALASVDAQQQRLTFQRLVAMMVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDS 272
AL + QQ+ +R + +KKE+A PV V+E S M + E+ + F
Sbjct: 380 KALVN---QQKFAESKRCLEFACEILEKKETALPVEVAEAYSEVAMQY--ESMNEF---- 430
Query: 273 EKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSSNFSSE 325
E +S + ++ K+ SEG + + GW Q VP ++E
Sbjct: 431 ETAISLLQKTLGILEKLPQEQHSEGSVSARIGWLLLFSGRVSQAVPYLESAAE 483
>At5g42880 putative protein
Length = 751
Score = 32.7 bits (73), Expect = 0.28
Identities = 50/250 (20%), Positives = 101/250 (40%), Gaps = 42/250 (16%)
Query: 20 QAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRDFQKEIVKAAETFTAI-- 77
+A K+FS + DV ++ + +E+L + R D + ++ ET +A+
Sbjct: 335 EAEEKRFSVAMARDQDVYNWEKELKMVENDIERLNQEVRAADDVKAKL----ETASALQH 390
Query: 78 -------GYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEELNRL- 129
+ I +G L E +N+I + A L + + ++EK E+ +L
Sbjct: 391 DLKTELAAFTDISSGNLLLE-------KNDIHAAVESARRELEEVKANIEKAASEVKKLK 443
Query: 130 -LASQVLDPL---RQMINGVPLEDARHLAQ---------------RYSRMRQEAETLRE- 169
+A + L RQ + +++ LA+ + + +EAE +
Sbjct: 444 IIAGSLQSELGRERQDLEETKQKESTGLARTNDKDAGEELVETAKKLEQATKEAEDAKAL 503
Query: 170 -EISKRQVRVRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQ 228
S+ ++R+ + + + + E+R+ E K M ALA++ A Q+ + Q
Sbjct: 504 ATASRDELRMAKELSEQAKRGMSTIESRLVEAKKEMEAARASEKLALAAIKALQETESSQ 563
Query: 229 RLVAMMVSDR 238
R + S R
Sbjct: 564 RFEEINNSPR 573
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.127 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,444,666
Number of Sequences: 26719
Number of extensions: 254533
Number of successful extensions: 964
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 64
Number of HSP's that attempted gapping in prelim test: 920
Number of HSP's gapped (non-prelim): 96
length of query: 327
length of database: 11,318,596
effective HSP length: 100
effective length of query: 227
effective length of database: 8,646,696
effective search space: 1962799992
effective search space used: 1962799992
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0046a.5


