

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0045.6
(214 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
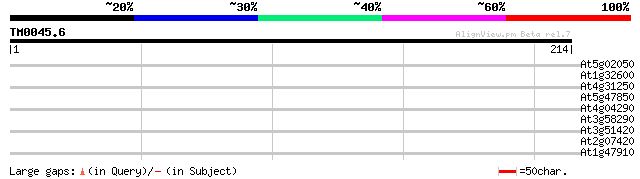
Score E
Sequences producing significant alignments: (bits) Value
At5g02050 unknown protein 30 0.73
At1g32600 hypothetical protein 29 2.1
At4g31250 receptor kinase - like protein 28 2.8
At5g47850 receptor kinase-like protein 28 4.7
At4g04290 putative reverse transcriptase 28 4.7
At3g58290 putative protein 28 4.7
At3g51420 mucin-like protein 28 4.7
At2g07420 putative retroelement pol polyprotein 28 4.7
At1g47910 reverse transcriptase, putative 27 8.1
>At5g02050 unknown protein
Length = 267
Score = 30.4 bits (67), Expect = 0.73
Identities = 14/37 (37%), Positives = 21/37 (55%)
Query: 39 IGAYEDGSVRDMITLAKYNGGDLDTYFEHPDFEPLVE 75
+ AY D V D +++ + G D D +E PDF+ L E
Sbjct: 183 VSAYPDEIVIDSLSIKQPQGSDNDLAYEGPDFDDLDE 219
>At1g32600 hypothetical protein
Length = 293
Score = 28.9 bits (63), Expect = 2.1
Identities = 12/30 (40%), Positives = 20/30 (66%)
Query: 113 RLNLWKLGATRGLISFGYNRSKDEFLVVLI 142
R+ + K G T+ L+ FGY+R D++ +V I
Sbjct: 74 RVPMIKRGQTQNLLGFGYDRVLDDYKIVTI 103
>At4g31250 receptor kinase - like protein
Length = 951
Score = 28.5 bits (62), Expect = 2.8
Identities = 23/77 (29%), Positives = 35/77 (44%), Gaps = 6/77 (7%)
Query: 85 GFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSKDEFLVV---- 140
G W SN FA R + + L++ LG+ RGL S + R+ E +
Sbjct: 61 GSDSKWKGVMCSNGSVFALRLENMSLSGELDVQALGSIRGLKSISFMRNHFEGKIPRGID 120
Query: 141 -LIKLCNLLHLAKNKIT 156
L+ L + L+LA N+ T
Sbjct: 121 GLVSLAH-LYLAHNQFT 136
>At5g47850 receptor kinase-like protein
Length = 751
Score = 27.7 bits (60), Expect = 4.7
Identities = 14/43 (32%), Positives = 22/43 (50%), Gaps = 1/43 (2%)
Query: 31 NFQNMWRNIGAYEDGSVRDMITLAKYNGGDLDTYFEHPDFEPL 73
N +N+ R +G YED R ++ G L + +P F+PL
Sbjct: 507 NHKNLVRLLGFYEDTEER-ILVYEYMKNGSLADHLHNPQFDPL 548
>At4g04290 putative reverse transcriptase
Length = 403
Score = 27.7 bits (60), Expect = 4.7
Identities = 19/84 (22%), Positives = 42/84 (49%), Gaps = 8/84 (9%)
Query: 133 SKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQFLC-- 190
S+ +V+L+ + +++ +K+ I + +++ FDI D Y ++G +C
Sbjct: 160 SQKGIVVILVYVDDIIISGNDKVGIQDTKTFLKSVFDIKDLGELKY-----FLGIEVCRS 214
Query: 191 -DSLYWLKTKKATNVLVVVRKLGL 213
+ L+ + K ++L V KLG+
Sbjct: 215 KEGLFLSQRKYTLDLLAQVGKLGV 238
>At3g58290 putative protein
Length = 282
Score = 27.7 bits (60), Expect = 4.7
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 36 WRNIGAYEDGSVRDMITLAKYNGGDLDTYFEHPDFEPLVEGPEALAQVLGFSHVWHFAAP 95
WR + AY +GS D + K NG L Y E DFE L G Q F+ V +
Sbjct: 40 WR-LFAYPEGSNGDHLLFKKNNGDHLSLYLE-VDFESLPCGWRQYTQ-FRFTVVNQISEH 96
Query: 96 SNVKAFAWRFLLVKIP 111
S+VK ++ K P
Sbjct: 97 SSVKREGRKWFDKKAP 112
>At3g51420 mucin-like protein
Length = 370
Score = 27.7 bits (60), Expect = 4.7
Identities = 26/92 (28%), Positives = 42/92 (45%), Gaps = 16/92 (17%)
Query: 121 ATRGLISFGYNRSKDEFLV-----VLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEY 175
A +GL+S K E L V KL + + +A N + YF A ++Y
Sbjct: 131 ANKGLLSISDGGKKTELLTDEADGVRFKLTDAVTVADNGVL----------YFTDASSKY 180
Query: 176 YFYKFNLDYI-GQFLCDSLYWLKTKKATNVLV 206
FY+F D++ G+ + + T +AT VL+
Sbjct: 181 DFYQFIFDFLEGKPHGRVMSFDPTTRATRVLL 212
>At2g07420 putative retroelement pol polyprotein
Length = 1664
Score = 27.7 bits (60), Expect = 4.7
Identities = 19/84 (22%), Positives = 42/84 (49%), Gaps = 8/84 (9%)
Query: 133 SKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQFLC-- 190
S+ +V+L+ + +++ +K+ I + +++ FDI D Y ++G +C
Sbjct: 1075 SQKGIVVILVYVDDIIISGNDKVGIQDTKTFLKSVFDIKDLGELKY-----FLGIEVCRS 1129
Query: 191 -DSLYWLKTKKATNVLVVVRKLGL 213
+ L+ + K ++L V KLG+
Sbjct: 1130 KEGLFLSQRKYTLDLLAQVGKLGV 1153
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 26.9 bits (58), Expect = 8.1
Identities = 13/45 (28%), Positives = 22/45 (48%), Gaps = 2/45 (4%)
Query: 87 SHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLG--ATRGLISFG 129
+++W P ++ F W+ L +P NL K G +G +S G
Sbjct: 826 AYIWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSCG 870
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.144 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,878,605
Number of Sequences: 26719
Number of extensions: 199870
Number of successful extensions: 502
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 500
Number of HSP's gapped (non-prelim): 9
length of query: 214
length of database: 11,318,596
effective HSP length: 95
effective length of query: 119
effective length of database: 8,780,291
effective search space: 1044854629
effective search space used: 1044854629
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0045.6


