

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0044a.6
(617 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
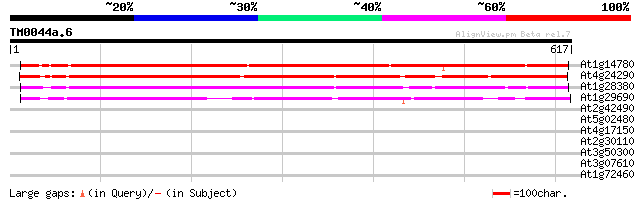
Score E
Sequences producing significant alignments: (bits) Value
At1g14780 unknown protein 518 e-147
At4g24290 unknown protein 511 e-145
At1g28380 unknown protein 446 e-125
At1g29690 unknown protein 421 e-118
At2g42490 putative copper amine oxidase 31 1.8
At5g02480 unknown protein 30 4.0
At4g17150 unknown protein 30 4.0
At2g30110 putative ubiquitin activating enzyme (UBA1) 30 4.0
At3g50300 anthranilate N-hydroxycinnamoyl/benzoyltransferase - l... 30 5.2
At3g07610 hypothetical protein 29 8.8
At1g72460 putative receptor-like protein kinase 29 8.8
>At1g14780 unknown protein
Length = 627
Score = 518 bits (1335), Expect = e-147
Identities = 277/617 (44%), Positives = 397/617 (63%), Gaps = 24/617 (3%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A++SLGKGFDL +DFRL++ K +G GS+ RLVVLD+ R++ IPG G + V
Sbjct: 12 AVKSLGKGFDLTADFRLKYCK---DGDGSAGD-DRLVVLDQTQNRELHIPGFG--VFQNV 65
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDL-SGDWFRDAAD 130
S +I CDKG+ RF+SD+L+FN+MSE NQ+S+V GKIPSG FNA F SG W DAA+
Sbjct: 66 SADINCDKGERTRFRSDILDFNKMSEYFNQRSSVTGKIPSGNFNATFGFQSGSWATDAAN 125
Query: 131 TKYLAFDGYFISLYYLHL-TASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAV 189
K L D ++L+ LH+ + + L + V+ +VP+ WDP L+RFI+ YGTH+I G++V
Sbjct: 126 VKSLGLDASVVTLFNLHIHNPNRLRLTDRVRNAVPSSWDPQLLARFIERYGTHVITGVSV 185
Query: 190 GGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLF--SDVRSPPLLQRKA--ADGKQKVPEV 245
GGQDV+ V+Q SS + LR HL DLGD LF S + S L + + + K PE
Sbjct: 186 GGQDVVVVRQDKSSDLDNDLLRHHLYDLGDQLFTGSCLLSTRRLNKAYHHSHSQPKFPEA 245
Query: 246 FNRVMQSNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVP 305
FN T++F + S +S++G+T+IC+KRGGD SHS WL TVP P+AI F F+P
Sbjct: 246 FNVFDDKQTVAFNNFS-INSQNGITVICAKRGGDGRAKSHSEWLITVPDKPDAINFNFIP 304
Query: 306 ISSLLAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKS 365
I+SLL +PGSG LSHA++LYLRYKPP DLQYFL+F PR WAP+ +LP S
Sbjct: 305 ITSLLKDVPGSGLLSHAMSLYLRYKPPLMDLQYFLDFSGPRAWAPVHNDLPFGAAPNMAS 364
Query: 366 SSLSLQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILS 425
+ +L NF+GPKL+V++T V SE+ PV G+R +LEG+KC+RLA+H+ HL + + +
Sbjct: 365 AYPALHINFMGPKLYVNTTPVTSEKNPVTGMRFFLEGKKCNRLAIHLQHLDN--TRTTVG 422
Query: 426 SETSTPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTG-------VYI 478
+ + +WRGSD ++++ EP+ K+FS+VCT VK+DPNW+ +++ +I
Sbjct: 423 EKITDEHIWRGSDQITDNDRYFEPLNGKKFSHVCTVPVKYDPNWIKTTSNHKSQNDVAFI 482
Query: 479 VTGAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEWAAAP-EASRKSSFLTNLSTTFSFT 537
VTGAQL + K+VLHLRL +T + + + + W P S+KS +++S +
Sbjct: 483 VTGAQLEVKKHGSKSVLHLRLRYTKVSDHYVVQNSWVHGPIGTSQKSGIFSSMSMPLTSG 542
Query: 538 QQSATSGPPKQAPAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAK 597
+ +L+SG++P GPPVP K++K+V+ +++ RGP +PGHWLVT +
Sbjct: 543 SVHHNMIQKDKNEVVLDSGVFPGGPPVPAN-NKIVKFVDLSQLCRGPQHSPGHWLVTGVR 601
Query: 598 LVTEGGKIGLQVKFALL 614
L + GK+ L VKFALL
Sbjct: 602 LYLDKGKLCLHVKFALL 618
>At4g24290 unknown protein
Length = 606
Score = 511 bits (1315), Expect = e-145
Identities = 276/605 (45%), Positives = 393/605 (64%), Gaps = 29/605 (4%)
Query: 11 VALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKG 70
VA+ S+G G+DLA D RL++ KG GS SR L + + + +I++PG G++I
Sbjct: 13 VAIGSIGCGYDLAIDLRLKYCKG-----GSKDSRL-LDIKEGDDNCEIVLPG--GISIPN 64
Query: 71 VSENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAAD 130
VS++I+CDKG+ +RF+SD+L F QM+E NQ+ ++ GKIPSG FNA+F+ S W +DAA
Sbjct: 65 VSKSIKCDKGERMRFRSDILPFQQMAEQFNQELSLAGKIPSGLFNAMFEFSSCWQKDAAY 124
Query: 131 TKYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVG 190
TK LAFDG FISLY + L S ++L+E VK++VP+ WDPAAL+RFI YGTH+IV + +G
Sbjct: 125 TKNLAFDGVFISLYSVALDKSQVLLREHVKQAVPSTWDPAALARFIDIYGTHIIVSVKMG 184
Query: 191 GQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVM 250
G+DVI KQ+HSSK+ P DL++ L+++ D F + + KV R+
Sbjct: 185 GKDVIYAKQQHSSKLQPEDLQKRLKEVADKRFVEASVVHNTGSERVQASSKVETKEQRLR 244
Query: 251 QSNTMSFTSISETSSKDGLTIICSKRGG-DMLKHSHSSWLQTVPSNPEAILFKFVPISSL 309
++T +S+ ++K+ +C +RGG D H+ WLQTV P+ I F+PI+SL
Sbjct: 245 FADT---SSLGSYANKEDYVFMCKRRGGNDNRNLMHNEWLQTVQMEPDVISMSFIPITSL 301
Query: 310 LAGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLS 369
L G+PGSG+LSHAINLYLRYKPP E+L FLEFQ+PRQWAP+F ELPL QR++ S S
Sbjct: 302 LNGVPGSGFLSHAINLYLRYKPPIEELHQFLEFQLPRQWAPVFSELPL-GPQRKQQSCAS 360
Query: 370 LQFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETS 429
LQF+F GPKL+V++T V ++P+ G+RLYLEGR+ +RLA+H+ HLSSLP L + +
Sbjct: 361 LQFSFFGPKLYVNTTPVDVGKRPITGMRLYLEGRRSNRLAIHLQHLSSLPKIYQLEDDLN 420
Query: 430 TPSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCS 489
R ++ E V WK +S+VCT V+ D + + +VTGAQL
Sbjct: 421 -----RSIRQESHDRRYYEKVNWKNYSHVCTEPVESDDD-------LSVVTGAQLHVESH 468
Query: 490 WPKNVLHLRLLFTNIPNCS-IRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQ 548
KNVL LRL F+ + + ++ +EW A + KS +ST S +A PP+
Sbjct: 469 GFKNVLFLRLCFSRVVGATLVKNSEWDEAVGFAPKSGL---ISTLISHHFTAAQKPPPRP 525
Query: 549 APAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQ 608
A +NS IYP GPPVP +A KLLK+V+ +E+ RGP ++PG+W+V+ A+L+ E GKI L+
Sbjct: 526 ADVNINSAIYPGGPPVPTQAPKLLKFVDTSEMTRGPQESPGYWVVSGARLLVEKGKISLK 585
Query: 609 VKFAL 613
VK++L
Sbjct: 586 VKYSL 590
>At1g28380 unknown protein
Length = 612
Score = 446 bits (1146), Expect = e-125
Identities = 255/606 (42%), Positives = 361/606 (59%), Gaps = 23/606 (3%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+ +G G+DL SD R K +G RLV +D RD++ PG G+ + V
Sbjct: 18 AVSVIGLGYDLCSDVRFSACKTTPDG-------SRLVEIDPTRNRDLIFPG--GIVVNNV 68
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
S +I+CDKG+ R +SD+L FNQMSE NQ + GKIPSG FN +F S W +DA+
Sbjct: 69 SSSIKCDKGERTRLRSDILSFNQMSEKFNQDMCLSGKIPSGMFNNMFAFSKCWPKDASSV 128
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LA+DG+FISLY + + + L++EVK+ VP+ WD AAL+ FI+ YGTH++VG+ +GG
Sbjct: 129 KTLAYDGWFISLYSVEIVRKQLTLRDEVKREVPSSWDSAALAGFIEKYGTHVVVGVTMGG 188
Query: 192 QDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQ 251
+DVI VKQ S P ++++ L+ GD F GK K + +Q
Sbjct: 189 KDVIHVKQMRKSNHEPEEIQKMLKHWGDERFCVDPVESKSPASVYSGKPKEENLLQWGLQ 248
Query: 252 SNTMSFTSISETSSK-DGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLL 310
S +S +K + + +C +RGG L SH WL TV P I FVPI+SLL
Sbjct: 249 PFGTSVSSAVVMHTKNEEIMRVCIRRGGVDLGQSHERWLSTVSQAPNVISMCFVPITSLL 308
Query: 311 AGIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSL 370
+G+PG+G+LSHA+NLYLRYKPP E+L FLEFQ+PRQWAP++ +LPL +R K SS SL
Sbjct: 309 SGLPGTGFLSHAVNLYLRYKPPIEELHQFLEFQLPRQWAPVYGDLPL-GLRRSKQSSPSL 367
Query: 371 QFNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETST 430
QF+ +GPKL+V++++V S ++PV GLR +LEG+K + LA+H+ HLS+ P + LS + +
Sbjct: 368 QFSLMGPKLYVNTSKVDSGERPVTGLRFFLEGKKGNHLAIHLQHLSACPPSLHLSHDDTY 427
Query: 431 PSMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRCSW 490
+ + + PV+W FS+VCT V++ N S IVT A L +
Sbjct: 428 EPI-----EEPVEKGYYVPVKWGIFSHVCTYPVQY--NGARSDDTASIVTKAWLEVKGMG 480
Query: 491 PKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFS--FTQQSATSGPPKQ 548
+ VL LRL F+ + RK+ W SRKS + +ST S + AT+ P Q
Sbjct: 481 MRKVLFLRLGFSLDASAVTRKSCWDNLSTNSRKSGVFSMISTRLSTGLSPNPATTKP--Q 538
Query: 549 APAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQ 608
+ +NS +YP GP PV+ KLL V+ EV+RGP + PG+W+VT AKL E GKI ++
Sbjct: 539 SKIDINSAVYPRGPSPPVKP-KLLSLVDTKEVMRGPEEQPGYWVVTGAKLCVEAGKISIK 597
Query: 609 VKFALL 614
K++LL
Sbjct: 598 AKYSLL 603
>At1g29690 unknown protein
Length = 561
Score = 421 bits (1083), Expect = e-118
Identities = 236/608 (38%), Positives = 354/608 (57%), Gaps = 73/608 (12%)
Query: 12 ALESLGKGFDLASDFRLRFAKGIREGRGSSSSRKRLVVLDEQNKRDILIPGAGGVTIKGV 71
A+++LG+GFD+ SD RL + KG + RLV ++E RD+ + + G + V
Sbjct: 24 AIQALGRGFDVTSDVRLLYCKG--------APGSRLVRIEEGQNRDLEL--SHGFLLPNV 73
Query: 72 SENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAADT 131
+I C +G+ + V F++M+E N +S V+G IP G FNA+F+ +G W DAA T
Sbjct: 74 PADIDCSRGNSGTQRISVCSFHEMAEEFNVRSGVKGNIPLGCFNAMFNYTGSWQVDAAST 133
Query: 132 KYLAFDGYFISLYYLHLTASPIVLQEEVKKSVPAQWDPAALSRFIQTYGTHLIVGMAVGG 191
K LA GYFI LY + L +VL E++++VP+ WDPA+L+ FI+ YGTH++ + +GG
Sbjct: 134 KSLALVGYFIPLYDVKLAKLTLVLHNEIRRAVPSSWDPASLASFIENYGTHIVTSVTIGG 193
Query: 192 QDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRSPPLLQRKAADGKQKVPEVFNRVMQ 251
+DV+ ++Q SS +P ++ ++ D+ + +R +
Sbjct: 194 RDVVYIRQHQSSPLPVSEIENYVNDM---------------------------IKHRFHE 226
Query: 252 SNTMSFTSISETSSKDGLTIICSKRGGDMLKHSHSSWLQTVPSNPEAILFKFVPISSLLA 311
+ + S T + KD +T+I +RGGD L+ SH+ W +TVP+ P+ I F PI SLL
Sbjct: 227 AESQSITGPLKYKDKD-ITVIFRRRGGDDLEQSHARWAETVPAAPDIINMTFTPIVSLLE 285
Query: 312 GIPGSGYLSHAINLYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQ 371
G+PG +L+ AI LYL YKPP EDLQYFL++QI R WAP L QR++ SLQ
Sbjct: 286 GVPGLRHLTRAIELYLEYKPPIEDLQYFLDYQIARAWAPEQSNL-----QRKEPVCSSLQ 340
Query: 372 FNFLGPKLHVSSTQVLSEQKPVVGLRLYLEGRKCDRLALHIHHLSSLPNKMILSSETSTP 431
F+ +GPKL +S+ QV +KPV GLRL LEG K +RL++H+ HL SLP + ++ P
Sbjct: 341 FSLMGPKLFISADQVTVGRKPVTGLRLSLEGSKQNRLSIHLQHLVSLPKILQPHWDSHVP 400
Query: 432 ---SMWRGSDDNESSNQFLEPVRWKRFSNVCTAVVKHDPNWLNSSTGVYIVTGAQLVSRC 488
W+G ++ +S ++ EP++WK FS+V T+ ++H + +GV+IVTGAQL
Sbjct: 401 IGAPKWQGPEEQDS--RWFEPIKWKNFSHVSTSPIEHTETHIGDLSGVHIVTGAQLGVWD 458
Query: 489 SWPKNVLHLRLLFTNIPNCSIRKTEWAAAPEASRKSSFLTNLSTTFSFTQQSATSGPPKQ 548
KNVLHL+LLF+ +P C+IR++ W P AS + + GP
Sbjct: 459 FGSKNVLHLKLLFSKVPGCTIRRSVWDHTPVAS---------------SGRLEPGGPSTS 503
Query: 549 APAMLNSGIYPDGPPVPVRAGKLLKYVEAAEVVRGPHDTPGHWLVTAAKLVTEGGKIGLQ 608
+ SG ++GKL K V+++E+++GP D PGHWLVT AKL E GKI L+
Sbjct: 504 SSTEEVSG----------QSGKLAKIVDSSEMLKGPQDLPGHWLVTGAKLGVEKGKIVLR 553
Query: 609 VKFALLDY 616
VK++LL+Y
Sbjct: 554 VKYSLLNY 561
>At2g42490 putative copper amine oxidase
Length = 776
Score = 31.2 bits (69), Expect = 1.8
Identities = 14/38 (36%), Positives = 21/38 (54%)
Query: 413 HHLSSLPNKMILSSETSTPSMWRGSDDNESSNQFLEPV 450
HH SLP ++S+ T ++W +DD +S LE V
Sbjct: 13 HHGGSLPPPKLVSAAPDTVAVWSDADDQRASKVSLESV 50
>At5g02480 unknown protein
Length = 508
Score = 30.0 bits (66), Expect = 4.0
Identities = 40/185 (21%), Positives = 68/185 (36%), Gaps = 27/185 (14%)
Query: 425 SSETSTPSMWRGSDDNESSNQFLEPVRWKR-----FSNVCTAVVKHDPNWLNSSTGVYIV 479
+ ++S S G +N+ +FL P + +N+ T K P+W+N TGV
Sbjct: 315 NGDSSDQSPGGGVINNKKRKEFLSPGSSEEECCLTVNNIETHHAKDPPSWVNDFTGVMKN 374
Query: 480 T-GAQLVSRCSWPKNVLHLRLLFTNIPNCSIRKTEW---------AAAPEASRKSSFLTN 529
+ G ++ + +L ++ + + K W ++ ++SF+
Sbjct: 375 SCGPVTAAKTVYEDEEAYLVVITLPFVDLNTVKVSWRNNITNGIVKVTGLSTSRASFVKR 434
Query: 530 LSTTFSFTQQSATSGP----------PKQAPAMLNSGIYPD--GPPVPVRAGKLLKYVEA 577
TF Q A P P + P N Y D GP + + KL VE
Sbjct: 435 RDRTFKLVDQMAEHCPPGEFMREIQLPNRIPEEANIEAYFDGTGPVLEIVVPKLRGGVEE 494
Query: 578 AEVVR 582
VR
Sbjct: 495 EHEVR 499
>At4g17150 unknown protein
Length = 502
Score = 30.0 bits (66), Expect = 4.0
Identities = 18/71 (25%), Positives = 37/71 (51%), Gaps = 4/71 (5%)
Query: 160 KKSVPAQWDPAALSRFIQTYGTHLIVGMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGD 219
K +PA + A+ +FIQ + + LI+ G +++I H+S P + + + +
Sbjct: 231 KTFIPALFGHASGDKFIQPHHSDLILKCYAGDKNIIKFDGDHNSSRP----QSYYDSVLV 286
Query: 220 FLFSDVRSPPL 230
F ++ +R PP+
Sbjct: 287 FFYNVLRPPPI 297
>At2g30110 putative ubiquitin activating enzyme (UBA1)
Length = 1080
Score = 30.0 bits (66), Expect = 4.0
Identities = 23/49 (46%), Positives = 24/49 (48%), Gaps = 5/49 (10%)
Query: 186 GMAVGGQDVICVKQKHSSKIPPGDLRRHLEDLGDFLFSDVRS---PPLL 231
G V G V VKQ P LR L+D GDFLFSD PPLL
Sbjct: 310 GTYVKGGIVTQVKQPKLLNFKP--LREALKDPGDFLFSDFSKFDRPPLL 356
>At3g50300 anthranilate N-hydroxycinnamoyl/benzoyltransferase - like
protein
Length = 448
Score = 29.6 bits (65), Expect = 5.2
Identities = 26/86 (30%), Positives = 36/86 (41%), Gaps = 11/86 (12%)
Query: 71 VSENIRCDKGDLLRFKSDVLEFNQMSELLNQKSAVQGKIPSGYFNALFDLSGDWFRDAAD 130
VS I CD G ++F + E +S++L G +P +F F ++G D
Sbjct: 88 VSFYIDCDDGRGVKFVHAIAESVSVSDILQP----HGSVPD-FFRLFFPMNGVRSIDGLS 142
Query: 131 TKYLAF------DGYFISLYYLHLTA 150
LA DG IS Y HL A
Sbjct: 143 EPLLALQVTEIKDGIVISFGYNHLVA 168
>At3g07610 hypothetical protein
Length = 1027
Score = 28.9 bits (63), Expect = 8.8
Identities = 31/108 (28%), Positives = 46/108 (41%), Gaps = 23/108 (21%)
Query: 325 LYLRYKPPAEDLQYFLEFQIPRQWAPMFCELPLRHHQRRKSSSLSLQFNF--------LG 376
L L+ PPA+ + +PR C LPL+H+ + L+L +G
Sbjct: 601 LKLKDWPPAK----VFKDNLPRHAEEFLCSLPLKHYTHPVNGPLNLAVKLPQNCLKPDMG 656
Query: 377 PKLHVSS--TQVLSEQKPVVGLRLYLEGRKCDRL-ALHI-HHLSSLPN 420
PK +V+S Q L V L CD A++I H+S +PN
Sbjct: 657 PKTYVASGFAQELGRGDSVTKLH-------CDMSDAVNILTHISEVPN 697
>At1g72460 putative receptor-like protein kinase
Length = 644
Score = 28.9 bits (63), Expect = 8.8
Identities = 14/43 (32%), Positives = 21/43 (48%), Gaps = 3/43 (6%)
Query: 85 FKSDVLEFNQMSELLN---QKSAVQGKIPSGYFNALFDLSGDW 124
F D+ EFN+++ L + + G IPS YF + L W
Sbjct: 102 FSGDIPEFNRLTALKSLYISGNRFSGNIPSDYFETMVSLKKAW 144
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,320,498
Number of Sequences: 26719
Number of extensions: 628405
Number of successful extensions: 1616
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1581
Number of HSP's gapped (non-prelim): 12
length of query: 617
length of database: 11,318,596
effective HSP length: 105
effective length of query: 512
effective length of database: 8,513,101
effective search space: 4358707712
effective search space used: 4358707712
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0044a.6


