

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0042.13
(250 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
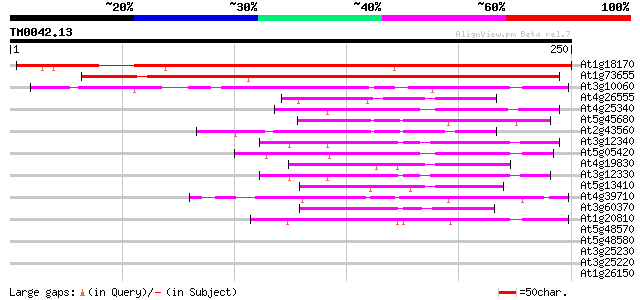
Sequences producing significant alignments: (bits) Value
At1g18170 unknown protein 287 3e-78
At1g73655 peptidylprolyl isomerase like protein 238 2e-63
At3g10060 unknown protein 73 1e-13
At4g26555 unknown protein 60 8e-10
At4g25340 unknown protein 57 9e-09
At5g45680 unknown protein 57 1e-08
At2g43560 putative FKBP type peptidyl-prolyl cis-trans isomerase 56 2e-08
At3g12340 hypothetical protein 55 4e-08
At5g05420 putative protein 55 5e-08
At4g19830 unknown protein 54 1e-07
At3g12330 hypothetical protein 53 1e-07
At5g13410 unknown protein 50 1e-06
At4g39710 unknown protein 47 7e-06
At3g60370 hypothetical protein 47 1e-05
At1g20810 immunophilin / FKBP-type peptidyl-prolyl cis-trans iso... 46 2e-05
At5g48570 peptidylprolyl isomerase 40 0.001
At5g48580 peptidyl-prolyl cis-trans isomerase-like protein 40 0.001
At3g25230 rotamase FKBP (ROF1) 40 0.002
At3g25220 immunophilin (FKBP15-1) 38 0.004
At1g26150 Pto kinase interactor, putative 38 0.004
>At1g18170 unknown protein
Length = 247
Score = 287 bits (735), Expect = 3e-78
Identities = 152/258 (58%), Positives = 192/258 (73%), Gaps = 26/258 (10%)
Query: 4 FFGSPPVFSL--PLTRN-SHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPI 60
F + P SL P T+ S+H +S + PP P S P P P
Sbjct: 5 FTATAPFLSLSKPFTKTASYHQCYASSSNPPEPESSSPPP---------------PPPPP 49
Query: 61 KPVTSSTK----VDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEAN 116
+P+ S K V++TDW+A+SLTRRFG+GAGLAW GFLAFGV+SEQIKTR+EVSQ+ AN
Sbjct: 50 QPLASQQKRKKNVETTDWVASSLTRRFGIGAGLAWAGFLAFGVISEQIKTRIEVSQEVAN 109
Query: 117 TRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTF----EG 172
TRDVE+E+E+ LPNGIRY + +VGGGA+PR GDLVVID+ G+V+ TG+VFV+TF +
Sbjct: 110 TRDVEEEKEIVLPNGIRYYDQRVGGGATPRAGDLVVIDLKGQVQGTGQVFVDTFGTKDKK 169
Query: 173 DKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPP 232
KPLALV+GS+PYSKG+CEGI+YVL++MKAGGKR+VIVPP LGF +GA+L SG+QIPP
Sbjct: 170 KMKPLALVVGSKPYSKGLCEGIDYVLRSMKAGGKRRVIVPPSLGFGVDGAELESGLQIPP 229
Query: 233 LATLEYIVEVDKVSIAPA 250
A+LEYIVE+D+VSIAPA
Sbjct: 230 NASLEYIVEIDRVSIAPA 247
>At1g73655 peptidylprolyl isomerase like protein
Length = 234
Score = 238 bits (608), Expect = 2e-63
Identities = 121/214 (56%), Positives = 154/214 (71%), Gaps = 5/214 (2%)
Query: 33 PPQQSQSPTASSSQSVQGTAKAQPQNPIKPVTSSTKVDSTDWIATSLTRRFGLGAGLAWV 92
P Q + S S SV T QP V + + TDWI + +TRRFG+GAG W
Sbjct: 18 PSQHPKQSLLSQSLSVTFTENPQPT----AVVTLQEQQLTDWITSPVTRRFGIGAGFTWA 73
Query: 93 GFLAFGVVSEQIK-TRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGGASPRPGDLV 151
GFLAFGVVSEQ+K +RL+V Q+E NTR +EK+EE+ LPNGIRY +L+VG GA+P G LV
Sbjct: 74 GFLAFGVVSEQMKKSRLDVFQEEDNTRGLEKQEEIILPNGIRYYDLQVGSGATPSSGYLV 133
Query: 152 VIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIV 211
V D+ G+V T +VFV+TF G K LA+VM SRPYSKG+C+GIE+VL++MKAGGKRKVI+
Sbjct: 134 VFDVKGQVHGTEQVFVDTFGGKGKSLAMVMDSRPYSKGLCQGIEHVLRSMKAGGKRKVII 193
Query: 212 PPQLGFRENGADLGSGVQIPPLATLEYIVEVDKV 245
PP LGF + + G G++IPP ATL+YI+EVD V
Sbjct: 194 PPSLGFGDRNVEFGQGLEIPPSATLDYIIEVDTV 227
>At3g10060 unknown protein
Length = 230
Score = 73.2 bits (178), Expect = 1e-13
Identities = 67/249 (26%), Positives = 110/249 (43%), Gaps = 39/249 (15%)
Query: 10 VFSLPLTRNSHHIASSSPTPPPIPPQQSQSPTASSSQSVQGTAKA-QPQNPIKPVTSSTK 68
+ ++ L HH SSS P P ++ QS S G K + + + P S +
Sbjct: 2 ILTMKLVHPLHHSLSSSI---PFPSRKRQSKPYRCSLPSPGCEKVIRTETVLPPAPVSCE 58
Query: 69 VDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDVEKEEEVTL 128
RR LG LA A G++S + S++ + + + TL
Sbjct: 59 -----------GRRVLLGCLLA----TASGILSTGSAEAVSTSRRALRASKLPESDFTTL 103
Query: 129 PNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYS- 187
PNG++Y ++KVG GA G V + + K + G F+ + +G V G PY
Sbjct: 104 PNGLKYYDIKVGNGAEAVKGSRVAVHYVAKWK--GITFMTSRQG-----LGVGGGTPYGF 156
Query: 188 -------KGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIV 240
V +G++ ++ M+ GG+R VIVPP+L + + G +IPP AT+E +
Sbjct: 157 DVGQSERGNVLKGLDLGVEGMRVGGQRLVIVPPELAYGKKGVQ-----EIPPNATIELDI 211
Query: 241 EVDKVSIAP 249
E+ + +P
Sbjct: 212 ELLSIKQSP 220
>At4g26555 unknown protein
Length = 207
Score = 60.5 bits (145), Expect = 8e-10
Identities = 39/106 (36%), Positives = 58/106 (53%), Gaps = 18/106 (16%)
Query: 122 KEEEVT----LPNGIRYCELKVGGGASPRPGDLVVIDIMGK------VESTGEVFVNTFE 171
KE EV LP+G+RY E+ G G GDLV ++ + + V ST V+ F
Sbjct: 74 KEPEVIRTLKLPSGVRYQEIIEGEGREAHEGDLVELNYVCRRANGYFVHST----VDQFS 129
Query: 172 GDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGF 217
G+ P+ L++ V EG++ VL MKAGGKR+ ++PP +G+
Sbjct: 130 GESSPVKLILDEND----VIEGLKEVLVGMKAGGKRRALIPPSVGY 171
>At4g25340 unknown protein
Length = 477
Score = 57.0 bits (136), Expect = 9e-09
Identities = 40/129 (31%), Positives = 63/129 (48%), Gaps = 12/129 (9%)
Query: 119 DVEKEEEVTLPNGIRYCELKVG--GGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKP 176
D + + T PNG+ EL +G G PG V + +GK++ G++F +
Sbjct: 358 DSKSSQVRTYPNGLIVEELSMGKPNGKRADPGKTVSVRYIGKLQKNGKIFDSNIGKSPFK 417
Query: 177 LALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATL 236
L +GS V +G + + M+ G KRK+ +PP +G+ GA G QIPP + L
Sbjct: 418 FRLGIGS------VIKGWDVGVNGMRVGDKRKLTIPPSMGYGVKGA----GGQIPPNSWL 467
Query: 237 EYIVEVDKV 245
+ VE+ V
Sbjct: 468 TFDVELINV 476
>At5g45680 unknown protein
Length = 208
Score = 56.6 bits (135), Expect = 1e-08
Identities = 38/116 (32%), Positives = 61/116 (51%), Gaps = 5/116 (4%)
Query: 129 PNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSK 188
P+G+ +C+ VG G G L+ +GK+E+ G+VF +++ K PL +G K
Sbjct: 90 PSGLAFCDKVVGYGPEAVKGQLIKAHYVGKLEN-GKVFDSSYNRGK-PLTFRIGVGEVIK 147
Query: 189 GVCEGI--EYVLKTMKAGGKRKVIVPPQLGFRENGADL-GSGVQIPPLATLEYIVE 241
G +GI + M GGKR + +PP+L + + GA G IPP + L + +E
Sbjct: 148 GWDQGILGSDGIPPMLTGGKRTLRIPPELAYGDRGAGCKGGSCLIPPASVLLFDIE 203
>At2g43560 putative FKBP type peptidyl-prolyl cis-trans isomerase
Length = 167
Score = 55.8 bits (133), Expect = 2e-08
Identities = 44/135 (32%), Positives = 71/135 (52%), Gaps = 10/135 (7%)
Query: 84 GLGAGLAWVGFLAFGV-VSEQIKTRLEVSQQEANTRDVEKEEEVTLPNGIRYCELKVGGG 142
G+ GL+ A+ + + K RL ++ E +++E VT +G++Y ++KVG G
Sbjct: 6 GVSGGLSMSSLAAYAAGLPPEDKPRLCEAECE---KELENVPMVTTESGLQYKDIKVGRG 62
Query: 143 ASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMK 202
SP G V + + V S G++F ++ E P +GS KG+ EGI +MK
Sbjct: 63 PSPPVGFQVAANYVAMVPS-GQIFDSSLEKGL-PYLFRVGSGQVIKGLDEGI----LSMK 116
Query: 203 AGGKRKVIVPPQLGF 217
AGGKR++ +P L F
Sbjct: 117 AGGKRRLYIPGPLAF 131
>At3g12340 hypothetical protein
Length = 647
Score = 55.1 bits (131), Expect = 4e-08
Identities = 44/143 (30%), Positives = 72/143 (49%), Gaps = 19/143 (13%)
Query: 112 QQEANTRDVEKE-------EEVTLPNGIRYCELKVG--GGASPRPGDLVVIDIMGKVEST 162
+++A +++EKE E TL NG+ +++ G G S G V I GK++ T
Sbjct: 514 KKQAIDKNIEKEAGTKKPLETRTLSNGVIIEDIEKGKLDGKSAVKGKKVSILYTGKLKDT 573
Query: 163 GEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGA 222
G +F + D PL +G + V EG+ ++ M+ G KR++I+PP LG+ + G
Sbjct: 574 GNLFDSNLGED--PLRFRLGG----ENVIEGLSIGVEGMRVGDKRRLIIPPALGYSKRGL 627
Query: 223 DLGSGVQIPPLATLEYIVEVDKV 245
++P A L Y VE K+
Sbjct: 628 K----EKVPKSAWLVYEVEAVKI 646
>At5g05420 putative protein
Length = 143
Score = 54.7 bits (130), Expect = 5e-08
Identities = 41/147 (27%), Positives = 70/147 (46%), Gaps = 16/147 (10%)
Query: 101 SEQIKTRLEVSQQ---EANTRDVEKEEEVTLPNGIRYCELKVGG--GASPRPGDLVVIDI 155
SE K ++S++ E+ + E++ +G+ EL +G G PG V +
Sbjct: 4 SESAKKNEKISEEATVESKAFSISVEKQTPDLDGLIVEELCMGNPNGKKAEPGKRVSVHY 63
Query: 156 MGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQL 215
GK++ G++F +T + L G V +G++ L M GGKRK+ +PP++
Sbjct: 64 TGKLQGNGKIFDSTVGKSRYKFRLDAGK------VIKGLDVGLNGMLVGGKRKLTIPPEM 117
Query: 216 GFRENGADLGSGVQIPPLATLEYIVEV 242
G+ GA IPP + L + VE+
Sbjct: 118 GYGAEGAG-----SIPPDSWLVFDVEL 139
>At4g19830 unknown protein
Length = 229
Score = 53.5 bits (127), Expect = 1e-07
Identities = 31/105 (29%), Positives = 55/105 (51%), Gaps = 10/105 (9%)
Query: 125 EVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVEST-GEVFVNTFE-----GDKKPLA 178
E+ G++ +L++G G P GD + I G++ + G F +T++ G+ P
Sbjct: 82 EIPNSGGVKALDLRIGDGDVPIEGDQIEIHYYGRLAAKQGWRFDSTYDHKDSNGEAVPFT 141
Query: 179 LVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGAD 223
V+GS V GIE +++MK GG R+V++PP G++ +
Sbjct: 142 FVLGSSK----VIPGIETAVRSMKVGGIRRVVIPPSQGYQNTSQE 182
>At3g12330 hypothetical protein
Length = 248
Score = 53.1 bits (126), Expect = 1e-07
Identities = 43/139 (30%), Positives = 70/139 (49%), Gaps = 19/139 (13%)
Query: 112 QQEANTRDVEKE-------EEVTLPNGIRYCELKVG--GGASPRPGDLVVIDIMGKVEST 162
+++A +++EKE E TL NG+ +++ G G S G V I GK++ T
Sbjct: 55 KKQAIDKNIEKEAGTKKPLETRTLSNGVIIEDIEKGKLDGKSAVKGKKVSILYTGKLKDT 114
Query: 163 GEVFVNTFEGDKKPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGA 222
G +F + D PL +G + V EG+ ++ M+ G KR++I+PP LG+ + G
Sbjct: 115 GNLFDSNLGED--PLRFRLGG----ENVIEGLSIGVEGMRVGDKRRLIIPPALGYSKRGL 168
Query: 223 DLGSGVQIPPLATLEYIVE 241
++P A L Y VE
Sbjct: 169 K----EKVPKSAWLVYEVE 183
>At5g13410 unknown protein
Length = 256
Score = 50.1 bits (118), Expect = 1e-06
Identities = 33/100 (33%), Positives = 50/100 (50%), Gaps = 13/100 (13%)
Query: 130 NGIRYCELKVGGGASPRPGDLVVIDIMGKV--------ESTGEVFVNTFEGDKKPL-ALV 180
+G++Y +L+VG G + GD VV+D G E+ + +FEGD K
Sbjct: 117 SGLQYKDLRVGTGPIAKKGDKVVVDWDGYTIGYYGRIFEARNKTKGGSFEGDDKEFFKFT 176
Query: 181 MGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFREN 220
+GS V E + M GG R++IVPP+LG+ +N
Sbjct: 177 LGSNE----VIPAFEEAVSGMALGGIRRIIVPPELGYPDN 212
>At4g39710 unknown protein
Length = 217
Score = 47.4 bits (111), Expect = 7e-06
Identities = 51/180 (28%), Positives = 87/180 (48%), Gaps = 28/180 (15%)
Query: 81 RRFGLGAGLAWVGFLAFGVVSEQIKTRLEVSQQEANTRDVEKEEEVTLP--------NGI 132
R FG+G +GFLA ++S ++ +A+ ++ V P +G+
Sbjct: 50 RVFGVG-----LGFLASSILS--------LTPLDADATRIDYYATVGDPLCEYSYAKSGL 96
Query: 133 RYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKGVCE 192
+C+L VG G G LV I + + G +F ++++ +PL + +G +G+ +
Sbjct: 97 GFCDLDVGFGDEAPRGVLVNIHYTARF-ADGTLFDSSYKR-ARPLTMRIGVGKVIRGLDQ 154
Query: 193 GI--EYVLKTMKAGGKRKVIVPPQLGFRENGADLGSG-VQIPPLATLEYIVEVDKVSIAP 249
GI + M+ GGKRK+ +PP+L + A SG IP ATL Y +++ V I P
Sbjct: 155 GILGGEGVPPMRVGGKRKLQIPPKLAYGPEPAGCFSGDCNIPGNATLLY--DINFVEIYP 212
>At3g60370 hypothetical protein
Length = 242
Score = 47.0 bits (110), Expect = 1e-05
Identities = 27/87 (31%), Positives = 44/87 (50%), Gaps = 6/87 (6%)
Query: 130 NGIRYCELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFEGDKKPLALVMGSRPYSKG 189
+G+ Y + VG G P+ G V +G ES + +G P + MG+
Sbjct: 120 SGLIYRDFNVGQGDFPKDGQQVTFHYIGYNESGRRIDSTYIQGS--PARIRMGTN----A 173
Query: 190 VCEGIEYVLKTMKAGGKRKVIVPPQLG 216
+ G E ++ MK GG+R++I+PP+LG
Sbjct: 174 LVPGFEMGIRDMKPGGRRRIIIPPELG 200
>At1g20810 immunophilin / FKBP-type peptidyl-prolyl cis-trans
isomerase
Length = 232
Score = 45.8 bits (107), Expect = 2e-05
Identities = 43/171 (25%), Positives = 72/171 (41%), Gaps = 34/171 (19%)
Query: 108 LEVSQQEANTRDVEK----EEEVTLPNGIRYCELKVGGGASPRPGDLVVIDIMGKVESTG 163
L S EA R K EE T P G+++ +++ G G G + + S
Sbjct: 64 LTPSSSEARERRSRKVIPLEEYSTGPEGLKFYDIEEGKGPVATEGSTAQVHFDCRYRSIT 123
Query: 164 EVFVNTFE---GDK---KPLALVMGSRPYSKGVCEGIE-------------------YVL 198
+ + G++ +P +GS P + E ++ ++
Sbjct: 124 AISTRESKLLAGNRSIAQPYEFKVGSTPGKERKREFVDNPNGLFSAQAAPKPPPAMYFIT 183
Query: 199 KTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKVSIAP 249
+ MK GGKR VIVPP+ G+ + G + +IPP AT E +E+ +V+ P
Sbjct: 184 EGMKVGGKRTVIVPPEAGYGQKGMN-----EIPPGATFELNIELLRVTPPP 229
>At5g48570 peptidylprolyl isomerase
Length = 578
Score = 40.4 bits (93), Expect = 0.001
Identities = 23/54 (42%), Positives = 30/54 (54%), Gaps = 2/54 (3%)
Query: 191 CEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQ--IPPLATLEYIVEV 242
C + +KTMK G K + V PQ GF E G G+Q IPP ATL+ +E+
Sbjct: 214 CPALSKAVKTMKRGEKVLLTVKPQYGFGEFGRPASDGLQAAIPPNATLQIDLEL 267
Score = 38.9 bits (89), Expect = 0.003
Identities = 32/108 (29%), Positives = 52/108 (47%), Gaps = 7/108 (6%)
Query: 137 LKVGGGAS-PRPGDLVVIDIMGKVESTGEVFVNT-FEGDKKPLALVMGSRPYSKGVCEGI 194
LK G G P G +V + ++GK++ VFV E D++P + + V EG+
Sbjct: 287 LKEGEGYERPNEGAIVKLKLIGKLQDGTTVFVKKGHEEDEEPFEFKID----EEQVIEGL 342
Query: 195 EYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
E + MK G + + P+ F + + V IPP +T+ Y VE+
Sbjct: 343 EKAVMGMKKGEVALITISPEYAFGSSESKQELAV-IPPNSTVYYEVEL 389
Score = 36.6 bits (83), Expect = 0.013
Identities = 21/53 (39%), Positives = 30/53 (55%), Gaps = 4/53 (7%)
Query: 190 VCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEV 242
V +G + +KTMK G +PP+L + E GS IPP ATL++ VE+
Sbjct: 101 VIKGWDLGIKTMKKGENAIFTIPPELAYGET----GSPPTIPPNATLQFDVEL 149
>At5g48580 peptidyl-prolyl cis-trans isomerase-like protein
Length = 163
Score = 40.0 bits (92), Expect = 0.001
Identities = 41/123 (33%), Positives = 59/123 (47%), Gaps = 19/123 (15%)
Query: 136 ELKVGGGASPRP-------GDLVVIDIMGKVESTGEVFVNTFE-GDKKPLALVMGSRPYS 187
EL++G P+ GD + + GK+ + G VF ++FE GD P +GS
Sbjct: 33 ELQIGVKFKPKTCEVQAHKGDTIKVHYRGKL-TDGTVFDSSFERGD--PFEFKLGSGQVI 89
Query: 188 KGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEYIVEVDKVSI 247
KG +G L G KRK+ +P +LG+ E GS IP ATL + E+ V+
Sbjct: 90 KGWDQG----LLGACVGEKRKLKIPAKLGYGEQ----GSPPTIPGGATLIFDTELIAVNE 141
Query: 248 APA 250
PA
Sbjct: 142 KPA 144
>At3g25230 rotamase FKBP (ROF1)
Length = 551
Score = 39.7 bits (91), Expect = 0.002
Identities = 24/54 (44%), Positives = 30/54 (55%), Gaps = 3/54 (5%)
Query: 191 CEGIEYVLKTMKAGGKRKVIVPPQLGFRENG--ADLGSGVQIPPLATLEYIVEV 242
C + +KTMK G K + V PQ GF E G A G G +PP ATLE +E+
Sbjct: 206 CPALTKAVKTMKKGEKVLLTVKPQYGFGEKGKPASAGEGA-VPPNATLEINLEL 258
Score = 36.6 bits (83), Expect = 0.013
Identities = 40/126 (31%), Positives = 56/126 (43%), Gaps = 19/126 (15%)
Query: 124 EEVTLPNGIRYCELKVGGG-ASPRPGDLVVIDIMGKVESTGEVFVNT-FEGDKK---PLA 178
EE + G++ LK G G +P GD V +V TG + T F+ + P
Sbjct: 32 EEKEIQQGLKKKLLKEGEGYETPENGDEV------EVHYTGTLLDGTKFDSSRDRATPFK 85
Query: 179 LVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLATLEY 238
+G KG GI KTMK G +P +L + E+ GS IP ATL++
Sbjct: 86 FTLGQGQVIKGWDIGI----KTMKKGENAVFTIPAELAYGES----GSPPTIPANATLQF 137
Query: 239 IVEVDK 244
VE+ K
Sbjct: 138 DVELLK 143
>At3g25220 immunophilin (FKBP15-1)
Length = 153
Score = 38.1 bits (87), Expect = 0.004
Identities = 42/136 (30%), Positives = 68/136 (49%), Gaps = 23/136 (16%)
Query: 121 EKEEEVT-LPNGIRY----CELKVGGGASPRPGDLVVIDIMGKVESTGEVFVNTFE-GDK 174
+K +VT L G++Y C+L+ GD + + GK+ + G VF ++FE GD
Sbjct: 26 KKSGDVTELQIGVKYKPQKCDLQA------HKGDKIKVHYRGKL-TDGTVFDSSFERGD- 77
Query: 175 KPLALVMGSRPYSKGVCEGIEYVLKTMKAGGKRKVIVPPQLGFRENGADLGSGVQIPPLA 234
P+ +G+ G +G L G KRK+ +P +LG+ +N GS +IP A
Sbjct: 78 -PIEFELGTGQVIPGWDQG----LLGACVGEKRKLKIPSKLGYGDN----GSPPKIPGGA 128
Query: 235 TLEYIVEVDKVSIAPA 250
TL + E+ V+ P+
Sbjct: 129 TLIFDTELVAVNGEPS 144
>At1g26150 Pto kinase interactor, putative
Length = 760
Score = 38.1 bits (87), Expect = 0.004
Identities = 33/103 (32%), Positives = 46/103 (44%), Gaps = 8/103 (7%)
Query: 7 SPPVFSLPLTRNSHHI--ASSSPTPPPIPPQQSQSPTASSSQSVQGTAKAQPQNPIKPVT 64
SPP +LP + S ++SS P +PP SP + +SV G P NP PVT
Sbjct: 259 SPPEETLPPPKPSPDPLPSNSSSPPTLLPPSSVVSPPSPPRKSVSGPDNPSPNNP-TPVT 317
Query: 65 SSTKVDSTDWIATSLTRRFGLGAGLAWVGFLAFGVVSEQIKTR 107
D++ S+ G+ G+A V GVV +K R
Sbjct: 318 -----DNSSSSGISIAAVVGVSIGVALVLLTLIGVVVCCLKKR 355
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,957,381
Number of Sequences: 26719
Number of extensions: 272232
Number of successful extensions: 2518
Number of sequences better than 10.0: 143
Number of HSP's better than 10.0 without gapping: 78
Number of HSP's successfully gapped in prelim test: 69
Number of HSP's that attempted gapping in prelim test: 1789
Number of HSP's gapped (non-prelim): 558
length of query: 250
length of database: 11,318,596
effective HSP length: 97
effective length of query: 153
effective length of database: 8,726,853
effective search space: 1335208509
effective search space used: 1335208509
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0042.13


