

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0041.8
(562 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
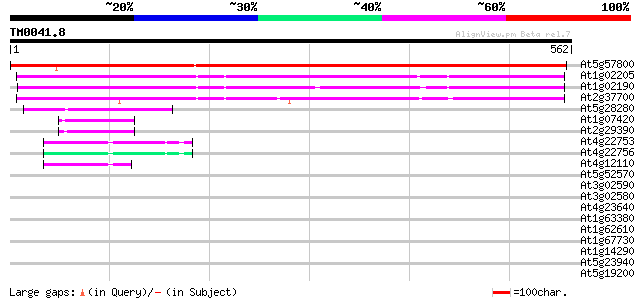
Sequences producing significant alignments: (bits) Value
At5g57800 cuticle protein (WAX2) 822 0.0
At1g02205 CER1 protein 394 e-110
At1g02190 CER1-like protein 348 5e-96
At2g37700 CER1-like protein 345 4e-95
At5g28280 CER1-like protein 97 3e-20
At1g07420 unknown protein 49 6e-06
At2g29390 sterol 4-alpha-methyl-oxidase 49 7e-06
At4g22753 C-4 sterol methyl oxidase like protein 48 1e-05
At4g22756 unknown protein 46 5e-05
At4g12110 putative C-4 sterol methyl oxidase 44 2e-04
At5g52570 beta-carotene hydroxylase 36 0.050
At3g02590 putative sterol-C5-desaturase 33 0.32
At3g02580 sterol-C5-desaturase 31 1.6
At4g23640 potassium transport like protein 31 2.1
At1g63380 reductase, putative 31 2.1
At1g62610 similar to glucose 1-dehydrogenase (AB000617); similar... 31 2.1
At1g67730 b-keto acyl reductase (glossy8) 30 2.7
At1g14290 unknown protein 30 2.7
At5g23940 acyltransferase 30 4.7
At5g19200 FVT1 - like protein 29 6.1
>At5g57800 cuticle protein (WAX2)
Length = 632
Score = 822 bits (2122), Expect = 0.0
Identities = 393/564 (69%), Positives = 459/564 (80%), Gaps = 8/564 (1%)
Query: 1 MLFLTRNRRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLP----FLQNLPSWNVK 56
MLF+TR RI +G+DFKQID EW WDN++ILQ II S+ YM P + +LP WN K
Sbjct: 67 MLFVTRTLRINPKGIDFKQIDHEWHWDNYIILQAIIVSLICYMSPPLMMMINSLPLWNTK 126
Query: 57 GVIVALVLHVGISEPLYYWVHRKFH-GDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFI 115
G+I +VLHV SEPLYY++HR FH +Y FT+YHS HHSSPVP +T G+AT++E+ +
Sbjct: 127 GLIALIVLHVTFSEPLYYFLHRSFHRNNYFFTHYHSFHHSSPVPHPMTAGNATLLENIIL 186
Query: 116 SVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYT 175
VV G+P++G + G GS S +YGY + FDF+RCLGHCNVE+ H LF+ P ++YL+YT
Sbjct: 187 CVVAGVPLIGCCLFGVGSLSAIYGYAVMFDFMRCLGHCNVEIFSHKLFEILPVLRYLIYT 246
Query: 176 PTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESF--SSGKCSRVPDFVFLAHI 233
PTYHSLHH + TNFCLFMPLFD LG T N SWEL + S+G+ RVP+FVFLAH
Sbjct: 247 PTYHSLHHQE-MGTNFCLFMPLFDVLGDTQNPNSWELQKKIRLSAGERKRVPEFVFLAHG 305
Query: 234 VDVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLH 293
VDV S+MHA F RSFAS+P+ TR FL+P WP +L MW SK FLFSFY LR+ L
Sbjct: 306 VDVMSAMHAPFVFRSFASMPYTTRIFLLPMWPFTFCVMLGMWAWSKTFLFSFYTLRNNLC 365
Query: 294 QTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVD 353
QTW VPR GFQYFLPFA +GIN +IE AIL ADKIGVKVISLAALNKNEALNGGG LFV+
Sbjct: 366 QTWGVPRFGFQYFLPFATKGINDQIEAAILRADKIGVKVISLAALNKNEALNGGGTLFVN 425
Query: 354 KHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTL 413
KHP+LRVRVVHGNTLTAAVI+ EIPKDV EVFLTGATSKLGRAIALYLC++ V+VLMLTL
Sbjct: 426 KHPDLRVRVVHGNTLTAAVILYEIPKDVNEVFLTGATSKLGRAIALYLCRRGVRVLMLTL 485
Query: 414 SADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVV 473
S +RF++IQKEAP E+Q+ LVQVTKY AAQHCKTWIVGKW+TPREQSWAP GTHFHQFVV
Sbjct: 486 SMERFQKIQKEAPVEFQNNLVQVTKYNAAQHCKTWIVGKWLTPREQSWAPAGTHFHQFVV 545
Query: 474 PPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG 533
PPIL FRR+CTYG+LAAM+LP+DVEGLG+CEYTMERGVVHACHAGGVVH LEGW HHEVG
Sbjct: 546 PPILKFRRNCTYGDLAAMKLPKDVEGLGTCEYTMERGVVHACHAGGVVHMLEGWKHHEVG 605
Query: 534 AIDVDRIDVVWKAALKHGLRPVSS 557
AIDVDRID+VW+AA+K+GL VSS
Sbjct: 606 AIDVDRIDLVWEAAMKYGLSAVSS 629
>At1g02205 CER1 protein
Length = 625
Score = 394 bits (1013), Expect = e-110
Identities = 207/549 (37%), Positives = 318/549 (57%), Gaps = 6/549 (1%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
RRI+ +G+DF Q+D+E +WD+ ++ ++ + +LP + LP W GV++A ++H G
Sbjct: 77 RRIVDKGIDFNQVDRETNWDDQILFNGVLFYIGINLLPEAKQLPWWRTDGVLMAALIHTG 136
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGAS 127
E LYYW+H+ H +L++ YHS HHSS V E +T EH ++ IP+L
Sbjct: 137 PVEFLYYWLHKALHHHFLYSRYHSHHHSSIVTEPITSVIHPFAEHIAYFILFAIPLLTTL 196
Query: 128 VMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
+ S GY++ DF+ +GHCN E++P LF FP +K+L YTP+YHSLHHT +
Sbjct: 197 LTKTASIISFAGYIIYIDFMNNMGHCNFELIPKRLFHLFPPLKFLCYTPSYHSLHHTQFR 256
Query: 188 DTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLR 247
TN+ LFMPL+D + T++ + L+E + + + D V L H+ S H + L
Sbjct: 257 -TNYSLFMPLYDYIYGTMDESTDTLYEK-TLERGDDIVDVVHLTHLTTPESIYHLRIGLA 314
Query: 248 SFASLPFRTRFFLIPFWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVVPRCGFQYFL 307
SFAS PF R+F+ WP +++ ++ F+ Q+WV+PR QY L
Sbjct: 315 SFASYPFAYRWFMRLLWPFTSLSMIFTLFYARLFVAERNSFNKLNLQSWVIPRYNLQYLL 374
Query: 308 PFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVVHGNT 367
+ E IN IE+AIL ADK GVKV+SL +N+ E LN G++++ HP+++VR+V G+
Sbjct: 375 KWRKEAINNMIEKAILEADKKGVKVLSLGLMNQGEELNRNGEVYIHNHPDMKVRLVDGSR 434
Query: 368 LTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQ 427
L AAV+I+ +PK V +TG +K+ IA LCQ+ V+V TL D +++I+ PQ
Sbjct: 435 LAAAVVINSVPKATTSVVMTGNLTKVAYTIASALCQRGVQV--STLRLDEYEKIRSCVPQ 492
Query: 428 EYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDCTYGE 487
E + +LV +T +A K W+VG+ T EQ A +GT F F P+ RRDC Y
Sbjct: 493 ECRDHLVYLTS-EALSSNKVWLVGEGTTREEQEKATKGTLFIPFSQFPLKQLRRDCIYHT 551
Query: 488 LAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG-AIDVDRIDVVWKA 546
A+ +P+ + + SCE + R + A G++H+LEGW HE G ++ + +D VW+A
Sbjct: 552 TPALIVPKSLVNVHSCENWLPRKAMSATRVAGILHALEGWEMHECGTSLLLSDLDQVWEA 611
Query: 547 ALKHGLRPV 555
L HG +P+
Sbjct: 612 CLSHGFQPL 620
>At1g02190 CER1-like protein
Length = 623
Score = 348 bits (893), Expect = 5e-96
Identities = 202/553 (36%), Positives = 304/553 (54%), Gaps = 18/553 (3%)
Query: 9 RILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGI 68
+I+ + ++F+Q+D+E WD+ +I T++ +A LP +LP W + G I+ +LH G
Sbjct: 78 KIVDKPIEFEQVDRERTWDDQVIFNTLLMYLANIKLPGASHLPPWRLDGAILMALLHAGP 137
Query: 69 SEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASV 128
E LYYW HR H +L++ YHS HHSS V E +T EH +++ IP++ AS+
Sbjct: 138 VEFLYYWFHRALHHHFLYSRYHSHHHSSIVTEPITSVVHPFAEHIAYTLLFAIPMVTASL 197
Query: 129 MGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNKD 188
G S + GY+ DF+ +GHCN E+ P LF FP +K+L YTP++HSLHHT +
Sbjct: 198 CGILSIVSIMGYITYIDFMNNMGHCNFELFPKRLFHLFPPLKFLCYTPSFHSLHHTQFR- 256
Query: 189 TNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIVDVTSSMHAQFCLRS 248
TN+ LFMP++D + T + + L+E S PD + L H+ S + S
Sbjct: 257 TNYSLFMPIYDFIYGTTDNLTDSLYER-SLEIEEESPDVIHLTHLTTHNSIYQMRLGFPS 315
Query: 249 FASLPFRTR--FFLIPF-WPIAITTLLAMW--VRSKAFLFSFYYLRDRLHQTWVVPRCGF 303
+S P +R ++L F WP + A+ + + F+F LRD + ++P+ F
Sbjct: 316 LSSCPLWSRPPWYLTCFMWPFTLLCSFALTSAIPLRTFVFERNRLRDLTVHSHLLPKFSF 375
Query: 304 QYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLFVDKHPNLRVRVV 363
E IN IEEAIL AD+ GVKV+SL +N E LNG G+++V K+P L++R+V
Sbjct: 376 HRH----HESINTIIEEAILEADEKGVKVMSLGLMNNREELNGSGEMYVQKYPKLKIRLV 431
Query: 364 HGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQK 423
G+++ A V+I+ IPK+ E+ G +K+ A+ LCQK VKV++L R + K
Sbjct: 432 DGSSMAATVVINNIPKEATEIVFRGNLTKVASAVVFALCQKGVKVVVL-----REEEHSK 486
Query: 424 EAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQFVVPPILPFRRDC 483
LV T + K W+VG I EQ A GT F F P R+DC
Sbjct: 487 LIKSGVDKNLVLSTS-NSYYSPKVWLVGDGIENEEQMKAKEGTLFVPFSHFPPNKLRKDC 545
Query: 484 TYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHEVG-AIDVDRIDV 542
Y AMR+P+ + + SCE + R V+ A GG+VH+LEGW H+ G +V R+
Sbjct: 546 FYQSTPAMRVPKSAQNIDSCENWLGRRVMSAWKIGGIVHALEGWEEHDCGNTCNVLRLHA 605
Query: 543 VWKAALKHGLRPV 555
+W+AAL+H +P+
Sbjct: 606 IWEAALRHDFQPL 618
>At2g37700 CER1-like protein
Length = 635
Score = 345 bits (885), Expect = 4e-95
Identities = 199/566 (35%), Positives = 304/566 (53%), Gaps = 28/566 (4%)
Query: 8 RRILQQGVDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVG 67
+RIL + ++F Q+D+E WD+ +I T+I + + +P W GVI+ +LH G
Sbjct: 73 KRILNKSIEFDQVDRERTWDDQIIFNTLIVYLTKVYVSGTSTIPFWRTDGVILVALLHAG 132
Query: 68 ISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHAT-------------IVEHAF 114
E +YYW HR H +L++ YHS HHSS V E +T+ EH
Sbjct: 133 PVEFIYYWFHRALHHHFLYSRYHSHHHSSIVTEPITLCATNSKPWVLIVAVVHPFAEHIG 192
Query: 115 ISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLY 174
++++G+P++ + G S + Y+ DF+ +GHCN E++P LF P +K+L Y
Sbjct: 193 YTLILGLPLITTFMCGTVSVVSIALYLTYIDFMNNMGHCNFELIPKFLFSLLPPLKFLCY 252
Query: 175 TPTYHSLHHTDNKDTNFCLFMPLFDALGHTLNTKSWELHESFSSGKCSRVPDFVFLAHIV 234
TP++HSLHHT + TN+ LFMP++D + T + S L+E+ S K PD + L H+
Sbjct: 253 TPSFHSLHHTQFR-TNYSLFMPMYDYIYGTTDECSDSLYET-SLEKEEEKPDAIHLTHLT 310
Query: 235 DVTSSMHAQFCLRSFASLPFRTRFFLIPFWPIAITTLLAMWVRS---KAFLFSFYYLRDR 291
+ S H + S +S P +R +L P A+ +L+ +RS + F+ RD
Sbjct: 311 SLDSIYHLRLGFASLSSHPLSSRCYLFLMKPFAL--ILSFILRSFSFQTFVVERNRFRDL 368
Query: 292 LHQTWVVPRCGFQYFLPFAAEGINKKIEEAILTADKIGVKVISLAALNKNEALNGGGKLF 351
+ ++P+ Y E INK IE AIL ADK GVKV+SL LN+ E LNG G+++
Sbjct: 369 TLHSHLLPKFSSHYMSHQQKECINKMIEAAILEADKKGVKVMSLGLLNQGEELNGYGEMY 428
Query: 352 VDKHPNLRVRVVHGNTLTAAVIIDEIPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLML 411
V +HP L++R+V G +L A V++ IP +EV G +K+ RAI LCQ +KV++
Sbjct: 429 VRRHPKLKIRIVDGGSLAAEVVLHSIPVGTKEVLFRGQITKVARAIVFSLCQNAIKVMV- 487
Query: 412 TLSADRFKRIQKEAPQEYQSYLVQVTKYQAAQHCKTWIVGKWITPREQSWAPRGTHFHQF 471
L + + + + + LV T Y + W+VG ++ +EQ A GT F F
Sbjct: 488 -LRKEEHSMLAEFLDDKCKENLVLTTNY----YPMIWLVGDGLSTKEQKMAKDGTLFLPF 542
Query: 472 VVPPILPFRRDCTYGELAAMRLPEDVEGLGSCEYTMERGVVHACHAGGVVHSLEGWTHHE 531
P R+DC Y AM +P + + SCE + R V+ A GG+VH+LEGW HE
Sbjct: 543 SQFPPKTLRKDCFYHTTPAMIIPHSAQNIDSCENWLGRRVMSAWRVGGIVHALEGWKEHE 602
Query: 532 VGAIDVDRIDV--VWKAALKHGLRPV 555
G D I+ VW+AAL++G +P+
Sbjct: 603 CGLDDNSIINPPRVWEAALRNGFQPL 628
>At5g28280 CER1-like protein
Length = 317
Score = 96.7 bits (239), Expect = 3e-20
Identities = 53/149 (35%), Positives = 74/149 (49%), Gaps = 2/149 (1%)
Query: 15 VDFKQIDKEWDWDNFLILQTIIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGISEPLYY 74
++F+Q+D+E WD+ +I T+ + LP +P + +LH E LYY
Sbjct: 31 IEFEQVDREQSWDDQIIFNTLFMYLVNNKLPGCSRIPPKPPS--CASFLLHAVPVEFLYY 88
Query: 75 WVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYGSA 134
W HR H +L++ YHS HHSS V E +T EH SV+ IPI+ S+ G S
Sbjct: 89 WFHRSLHHHFLYSRYHSHHHSSIVTEPITSVVHPFAEHIAYSVLFAIPIVTVSLCGILSI 148
Query: 135 SLVYGYVLGFDFLRCLGHCNVEVVPHGLF 163
YV DF+ +GH N E P LF
Sbjct: 149 VPFVAYVTYIDFMNYMGHYNFEFFPKRLF 177
>At1g07420 unknown protein
Length = 266
Score = 49.3 bits (116), Expect = 6e-06
Identities = 26/78 (33%), Positives = 41/78 (52%), Gaps = 4/78 (5%)
Query: 50 LPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATI 109
LPSW K V ++ + I + ++YW HR H +L+ N HS+HH P LT +A
Sbjct: 102 LPSW--KEVSAQILFYFIIEDFVFYWGHRILHSKWLYKNVHSVHHEYATPFGLTSEYAHP 159
Query: 110 VEHAFI--SVVIGIPILG 125
E F+ + ++G + G
Sbjct: 160 AEILFLGFATIVGPALTG 177
>At2g29390 sterol 4-alpha-methyl-oxidase
Length = 260
Score = 48.9 bits (115), Expect = 7e-06
Identities = 26/78 (33%), Positives = 41/78 (52%), Gaps = 4/78 (5%)
Query: 50 LPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHHSSPVPESLTVGHATI 109
LPSW V V ++ + I + ++YW HR H +L+ N HS+HH P LT +A
Sbjct: 102 LPSWKV--VSAQILFYFIIEDFVFYWGHRILHTKWLYKNVHSVHHEYATPFGLTSEYAHP 159
Query: 110 VEHAFI--SVVIGIPILG 125
E F+ + ++G + G
Sbjct: 160 AEILFLGFATIVGPALTG 177
>At4g22753 C-4 sterol methyl oxidase like protein
Length = 291
Score = 48.1 bits (113), Expect = 1e-05
Identities = 34/149 (22%), Positives = 61/149 (40%), Gaps = 9/149 (6%)
Query: 35 IIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHH 94
I++ + M+ LP ++ ++ LV++ I + YW+HR H + + H +HH
Sbjct: 101 IVSYPSIQMVGIRSGLPLPSLMEIVAQLVVYFLIEDYTNYWIHRWMHCKWGYEKIHRIHH 160
Query: 95 SSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCN 154
P +G+A+ H +++GIP + G + ++ F H
Sbjct: 161 EYTSP----IGYASPYAHWAEILILGIPTFLGPAIAPGHIMTFWLWISLRQFEAIETHSG 216
Query: 155 VEVVPHGLFKTFPFMKYLLYTPTYHSLHH 183
+ P + K PF P YH HH
Sbjct: 217 YD-FPWSVTKLIPFYG----GPEYHDYHH 240
>At4g22756 unknown protein
Length = 299
Score = 46.2 bits (108), Expect = 5e-05
Identities = 33/149 (22%), Positives = 59/149 (39%), Gaps = 9/149 (6%)
Query: 35 IIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHH 94
+++ + M+ LP + ++ LV++ + + YWVHR FH + + +H +HH
Sbjct: 105 LVSYPSIQMIEIRSGLPLPSCMEIVAQLVVYFLVEDYTNYWVHRFFHCKWGYEKFHHIHH 164
Query: 95 SSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFLRCLGHCN 154
P +G+A H +++GIP + G + ++ H
Sbjct: 165 EYTAP----IGYAAPYAHWAEVLLLGIPTFLGPAIAPGHMITFWLWIALRQIEAIETHSG 220
Query: 155 VEVVPHGLFKTFPFMKYLLYTPTYHSLHH 183
+ P L K PF YH HH
Sbjct: 221 YD-FPWSLTKYIPFYG----GAEYHDYHH 244
>At4g12110 putative C-4 sterol methyl oxidase
Length = 298
Score = 43.9 bits (102), Expect = 2e-04
Identities = 23/88 (26%), Positives = 42/88 (47%), Gaps = 4/88 (4%)
Query: 35 IIASMAYYMLPFLQNLPSWNVKGVIVALVLHVGISEPLYYWVHRKFHGDYLFTNYHSLHH 94
+++ + M+ LP + ++ LV++ I + YWVHR FH + + H +HH
Sbjct: 105 LVSYPSIQMIEIRSGLPLPTITEMLSQLVVYFLIEDYTNYWVHRFFHSKWGYDKIHRVHH 164
Query: 95 SSPVPESLTVGHATIVEHAFISVVIGIP 122
P +G+A H +++GIP
Sbjct: 165 EYTAP----IGYAAPYAHWAEVLLLGIP 188
>At5g52570 beta-carotene hydroxylase
Length = 303
Score = 36.2 bits (82), Expect = 0.050
Identities = 42/161 (26%), Positives = 64/161 (39%), Gaps = 26/161 (16%)
Query: 52 SWNVKGVIVALV-------LHVGISEPLYYWV---HRKFHGDYLFTNYHSLHHSSPVPES 101
SW +KG V+++ L VG + + +W HR D L+ N H HH P +
Sbjct: 120 SWQMKGGEVSVLEMFGTFALSVGAAVGMEFWARWAHRALWHDSLW-NMHESHHK-PREGA 177
Query: 102 LTVGHATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDF-LRCLGHCNVEVVPH 160
+ + +A +P +G G+ + LV G G + G + V
Sbjct: 178 FELNDVFAITNA-------VPAIGLLYYGFLNKGLVPGLCFGAGLGITMFGMAYMFVHDG 230
Query: 161 GLFKTFPF-----MKYLLYTPTYHSLHHTDN-KDTNFCLFM 195
+ K FP + YL H LHHTD K + LF+
Sbjct: 231 LVHKRFPVGPIANVPYLRKVAAAHQLHHTDKFKGVPYGLFL 271
>At3g02590 putative sterol-C5-desaturase
Length = 279
Score = 33.5 bits (75), Expect = 0.32
Identities = 35/160 (21%), Positives = 67/160 (41%), Gaps = 18/160 (11%)
Query: 34 TIIASMAYYMLPF-----LQNLPSWNVKGVIVALVLHVGISEPLYYWVHRKFHG-DYLFT 87
T++ +++ YM+ L +N + + L++ + E + YWVH++ H +L+
Sbjct: 100 TLLPAVSEYMIEHGWTKCYSTLDHFNWFLCFLYIALYLVLVEFMIYWVHKELHDIKFLYK 159
Query: 88 NYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYGSASLVYGYVLGFDFL 147
+ H+ HH +L+ A + H ++ IP V+ + L FL
Sbjct: 160 HLHATHHMYNKQNTLS-PFAGLAFHPLDGILQAIP----HVIALFIVPIHLITHLSLLFL 214
Query: 148 RCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
+ ++ HG +P M YH++HHT K
Sbjct: 215 EGIWTASIHDCIHG--NIWPIM-----GAGYHTIHHTTYK 247
>At3g02580 sterol-C5-desaturase
Length = 281
Score = 31.2 bits (69), Expect = 1.6
Identities = 27/115 (23%), Positives = 47/115 (40%), Gaps = 13/115 (11%)
Query: 74 YWVHRKFHG-DYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYG 132
YW+HR+ H L+ H+ HH +L+ A + H ++ +P V+
Sbjct: 144 YWMHRELHDIKPLYKYLHATHHIYNKQNTLS-PFAGLAFHPVDGILQAVP----HVIALF 198
Query: 133 SASLVYGYVLGFDFLRCLGHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHHTDNK 187
+ + +G F+ + N+ HG +P M YH++HHT K
Sbjct: 199 IVPIHFTTHIGLLFMEAIWTANIHDCIHG--NIWPVM-----GAGYHTIHHTTYK 246
>At4g23640 potassium transport like protein
Length = 775
Score = 30.8 bits (68), Expect = 2.1
Identities = 18/62 (29%), Positives = 34/62 (54%), Gaps = 4/62 (6%)
Query: 242 AQFCLRSFASLPFRTRFFLIP---FWPIAITTLLAMWVRSKAFLFSFYYLRDRLHQTWVV 298
A F ++F++LP + + IP FWP+ + +LA V S+A +F+ + + + +
Sbjct: 305 AAFLSKNFSALP-SSFYSSIPDPFFWPVLMMAMLAAMVASQAVIFATFSIVKQCYALGCF 363
Query: 299 PR 300
PR
Sbjct: 364 PR 365
>At1g63380 reductase, putative
Length = 287
Score = 30.8 bits (68), Expect = 2.1
Identities = 15/41 (36%), Positives = 22/41 (53%)
Query: 384 VFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKE 424
V +TGA+S +GR I L LC+ K++ DR + E
Sbjct: 27 VLVTGASSGIGREICLDLCKAGCKIVAAARRVDRLNSLCSE 67
>At1g62610 similar to glucose 1-dehydrogenase (AB000617); similar to
EST gb|T88100
Length = 307
Score = 30.8 bits (68), Expect = 2.1
Identities = 15/41 (36%), Positives = 22/41 (53%)
Query: 384 VFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKE 424
V +TGA+S +GR I L LC+ K++ DR + E
Sbjct: 49 VLVTGASSGIGREICLDLCKAGCKIVAAARRVDRLNSLCSE 89
>At1g67730 b-keto acyl reductase (glossy8)
Length = 318
Score = 30.4 bits (67), Expect = 2.7
Identities = 12/44 (27%), Positives = 24/44 (54%)
Query: 386 LTGATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQEY 429
+TG T +G+A A L QK + ++++ + D+ K + +Y
Sbjct: 56 ITGPTDGIGKAFAFQLAQKGLNLILVARNPDKLKDVSDSIRSKY 99
>At1g14290 unknown protein
Length = 259
Score = 30.4 bits (67), Expect = 2.7
Identities = 24/112 (21%), Positives = 45/112 (39%), Gaps = 12/112 (10%)
Query: 74 YWVHRKFH-GDYLFTNYHSLHHSSPVPESLTVGHATIVEHAFISVVIGIPILGASVMGYG 132
Y++HR H +L+ + HS HH VP S + + H +++ S + G
Sbjct: 110 YFIHRYMHLNKFLYKHIHSQHHRLIVPYS----YGALYNHPLEGLLLDTIGGALSFLFSG 165
Query: 133 SASLVYGYVLGFDFLRCL-GHCNVEVVPHGLFKTFPFMKYLLYTPTYHSLHH 183
+ + F ++ + HC + + + PF + YH +HH
Sbjct: 166 MSPRTAIFFFSFATIKTVDDHCGLWLPGN------PFHIFFSNNSAYHDVHH 211
>At5g23940 acyltransferase
Length = 484
Score = 29.6 bits (65), Expect = 4.7
Identities = 19/61 (31%), Positives = 29/61 (47%), Gaps = 2/61 (3%)
Query: 377 IPKDVEEVFLT--GATSKLGRAIALYLCQKKVKVLMLTLSADRFKRIQKEAPQEYQSYLV 434
IP D + F T TS + R + L K + + T+ AD +R+ P+EY L+
Sbjct: 261 IPSDSSKPFSTFQSLTSHIWRHVTLARGLKPEDITIFTVFADCRRRVDPPMPEEYFGNLI 320
Query: 435 Q 435
Q
Sbjct: 321 Q 321
>At5g19200 FVT1 - like protein
Length = 331
Score = 29.3 bits (64), Expect = 6.1
Identities = 20/69 (28%), Positives = 31/69 (43%), Gaps = 4/69 (5%)
Query: 377 IPKDVEEVFLTGATSKLGRAIALYLCQKKVKVLMLTLSADRF----KRIQKEAPQEYQSY 432
IP VF+TG +S +G A+A + KV +L S ++ + IQ E ++
Sbjct: 33 IPIKFRHVFITGGSSGIGLALAHRAVSEGAKVSILARSTEKLAEAKRSIQLATGVEVATF 92
Query: 433 LVQVTKYQA 441
V Y A
Sbjct: 93 SADVRDYDA 101
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,084,201
Number of Sequences: 26719
Number of extensions: 568788
Number of successful extensions: 1528
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1492
Number of HSP's gapped (non-prelim): 21
length of query: 562
length of database: 11,318,596
effective HSP length: 104
effective length of query: 458
effective length of database: 8,539,820
effective search space: 3911237560
effective search space used: 3911237560
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0041.8


