

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0036b.1
(122 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
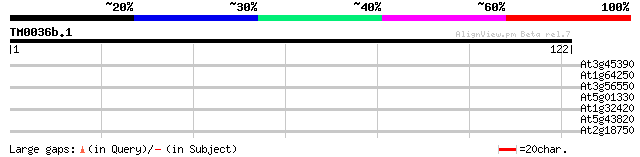
Score E
Sequences producing significant alignments: (bits) Value
At3g45390 receptor-like protein kinase 29 0.64
At1g64250 hypothetical protein 26 4.1
At3g56550 putative protein 26 5.4
At5g01330 pyruvate decarboxylase-like protein 25 7.1
At1g32420 hypothetical protein 25 7.1
At5g43820 putative protein 25 9.2
At2g18750 putative calmodulin-binding protein 25 9.2
>At3g45390 receptor-like protein kinase
Length = 604
Score = 28.9 bits (63), Expect = 0.64
Identities = 14/26 (53%), Positives = 18/26 (68%)
Query: 96 LFRSLKIGELSKMPLFKKKKFSLSPL 121
L +SL I ELS +PLF ++K SPL
Sbjct: 271 LLKSLDISELSTVPLFTEQKRKRSPL 296
>At1g64250 hypothetical protein
Length = 790
Score = 26.2 bits (56), Expect = 4.1
Identities = 12/38 (31%), Positives = 21/38 (54%)
Query: 8 YEPNDSLRSDRFIVSLKDHSSTCNFSELLEILSRHVVA 45
++ + L+ D +IV L + S TC +L + HV+A
Sbjct: 651 FQVTEPLQEDEWIVQLSEWSCTCGEFQLKKFPCLHVLA 688
>At3g56550 putative protein
Length = 581
Score = 25.8 bits (55), Expect = 5.4
Identities = 15/41 (36%), Positives = 24/41 (57%), Gaps = 3/41 (7%)
Query: 7 WYEPNDSLRSDRFIVSLKDH-SSTCNFSELLEILSRHVVAG 46
W E D + +F+V K H S +SEL E+++R ++AG
Sbjct: 450 WIEIGDQVH--KFVVDDKMHPESAVIYSELGEVINRAILAG 488
>At5g01330 pyruvate decarboxylase-like protein
Length = 592
Score = 25.4 bits (54), Expect = 7.1
Identities = 14/43 (32%), Positives = 23/43 (52%)
Query: 28 STCNFSELLEILSRHVVAGIFYKGLNIVLCILILQKDKLIRTH 70
ST SE++E ++ AG + + V L+L+K+K I H
Sbjct: 295 STLFCSEIVESADAYIFAGPIFNDYSSVGYSLLLKKEKAIIVH 337
>At1g32420 hypothetical protein
Length = 302
Score = 25.4 bits (54), Expect = 7.1
Identities = 15/52 (28%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Query: 7 WYEPNDSLRSDRFIVSLKDHSSTCNFSELLEILSRHVVAGIFYKG-LNIVLC 57
+Y+ + S F+VS S N + +IL+ H I Y+G L +++C
Sbjct: 233 YYQATNEYGSTYFLVSFDVRSEKFNHVKAPKILTDHPCTLINYQGKLGLIMC 284
>At5g43820 putative protein
Length = 680
Score = 25.0 bits (53), Expect = 9.2
Identities = 17/67 (25%), Positives = 29/67 (42%), Gaps = 8/67 (11%)
Query: 31 NFSELLEILSRH--------VVAGIFYKGLNIVLCILILQKDKLIRTHTCTLLHQLMGKS 82
++S +L L R V+ G+ +G+N L L + D +R H +L +S
Sbjct: 153 SYSVILRALGRRKLFSFMMDVLKGMVCEGVNPDLECLTIAMDSFVRVHYVRRAIELFEES 212
Query: 83 KVFDANC 89
+ F C
Sbjct: 213 ESFGVKC 219
>At2g18750 putative calmodulin-binding protein
Length = 622
Score = 25.0 bits (53), Expect = 9.2
Identities = 11/37 (29%), Positives = 19/37 (50%)
Query: 16 SDRFIVSLKDHSSTCNFSELLEILSRHVVAGIFYKGL 52
S+R +L +HS TC SE+L + G+ + +
Sbjct: 309 SNRMWETLAEHSKTCVLSEMLYVYYPEDSVGVVFNNI 345
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,800,034
Number of Sequences: 26719
Number of extensions: 100293
Number of successful extensions: 291
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 287
Number of HSP's gapped (non-prelim): 7
length of query: 122
length of database: 11,318,596
effective HSP length: 87
effective length of query: 35
effective length of database: 8,994,043
effective search space: 314791505
effective search space used: 314791505
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0036b.1


