

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0035.12
(125 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
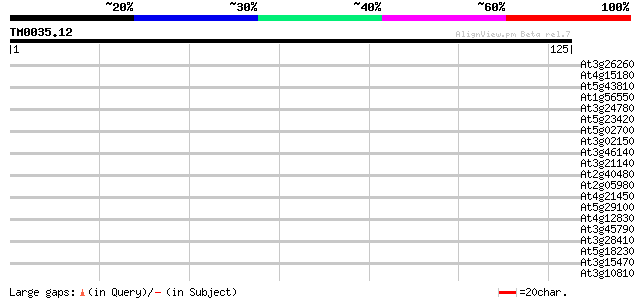
Score E
Sequences producing significant alignments: (bits) Value
At3g26260 hypothetical protein 30 0.24
At4g15180 hypothetical protein 30 0.41
At5g43810 PINHEAD (gb|AAD40098.1); translation initiation factor 29 0.53
At1g56550 unknown protein 29 0.69
At3g24780 hypothetical protein 28 0.91
At5g23420 unknown protein 28 1.2
At5g02700 putative protein 28 1.2
At3g02150 unknown protein 28 1.2
At3g46140 protein kinase - like 28 1.5
At3g21140 unknown protein 28 1.5
At2g40480 hypothetical protein 28 1.5
At2g05980 putative non-LTR retroelement reverse transcriptase 28 1.5
At4g21450 unknown protein 27 2.0
At5g29100 putative protein 27 2.6
At4g12830 hydrolase like protein 27 2.6
At3g45790 putative protein 27 2.6
At3g28410 hypothetical protein 27 2.6
At5g18230 unknown protein 26 4.5
At3g15470 unknown protein 26 4.5
At3g10810 putative RING zinc finger protein 26 4.5
>At3g26260 hypothetical protein
Length = 989
Score = 30.4 bits (67), Expect = 0.24
Identities = 19/67 (28%), Positives = 35/67 (51%), Gaps = 2/67 (2%)
Query: 28 PHEV-LAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWN-KTQIKTPLNKASTFEA 85
PH + L F+ S+ + +L + D D + +TQ+E +W+ K I +++ S +
Sbjct: 230 PHAIQLLFLESVPRIRELQFPEVDEDDDSDMEETQKEIERLWSLKLDIVWKVDRQSQTKV 289
Query: 86 TKLRSGT 92
T + SGT
Sbjct: 290 TSVMSGT 296
>At4g15180 hypothetical protein
Length = 2351
Score = 29.6 bits (65), Expect = 0.41
Identities = 16/74 (21%), Positives = 34/74 (45%), Gaps = 1/74 (1%)
Query: 45 ARRKRASDGDKSLNKTQEEKASIWNK-TQIKTPLNKASTFEATKLRSGTCIFRSGAAVRF 103
++R+ +S T ++ +W T + TP ++ T + +L G + GA
Sbjct: 851 SKRESVESHSQSTASTGQDSQGLWKTDTSVNTPRDRLCTVDDLQLHIGDWFYTDGAGQEQ 910
Query: 104 VILGFAERREMEEE 117
L F+E +++ E+
Sbjct: 911 GPLSFSELQKLVEK 924
>At5g43810 PINHEAD (gb|AAD40098.1); translation initiation factor
Length = 988
Score = 29.3 bits (64), Expect = 0.53
Identities = 17/44 (38%), Positives = 24/44 (53%), Gaps = 1/44 (2%)
Query: 31 VLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIK 74
++A + L+K +DL RR A DG KSL T E W + +K
Sbjct: 177 IIAELVRLYKESDLGRRLPAYDGRKSL-YTAGELPFTWKEFSVK 219
>At1g56550 unknown protein
Length = 383
Score = 28.9 bits (63), Expect = 0.69
Identities = 15/30 (50%), Positives = 19/30 (63%), Gaps = 2/30 (6%)
Query: 49 RASDGDKSLNKT--QEEKASIWNKTQIKTP 76
R++DG K L KT +E +A WN TQ K P
Sbjct: 236 RSTDGGKLLMKTWVEEIQAQPWNNTQAKKP 265
>At3g24780 hypothetical protein
Length = 715
Score = 28.5 bits (62), Expect = 0.91
Identities = 28/124 (22%), Positives = 48/124 (38%), Gaps = 10/124 (8%)
Query: 11 SHRRSFFTHINWAPDQHPHEVLAFMPSLFKSN------DLARRKRASDGDKSLNKTQEEK 64
S + F+T W +HP + + SL K ++ R +S+ KTQ
Sbjct: 174 SDKEGFYTAALWLHGRHPKTLACNLESLSKFGYFKDFPEILYRILQGPEIRSIQKTQRYD 233
Query: 65 ASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSGAAVRFVILGFAERREMEEEE----KR 120
+ ++ ++ G + AA R + + AER+ EE+ KR
Sbjct: 234 TIAAASLRRRSRFSRGGRGFGGGRSRGRHFLKRSAATRELRVANAERKNQEEKARASLKR 293
Query: 121 KQKQ 124
KQK+
Sbjct: 294 KQKK 297
>At5g23420 unknown protein
Length = 241
Score = 28.1 bits (61), Expect = 1.2
Identities = 22/83 (26%), Positives = 34/83 (40%), Gaps = 2/83 (2%)
Query: 40 KSNDLARRKRASDGDKS--LNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRS 97
K ++K SDG K L KT +EK S + K PL F + ++
Sbjct: 80 KKKPAEKKKTTSDGPKPKRLKKTNDEKKSSSTSNKPKRPLTAFFIFMSDFRKTFKSEHNG 139
Query: 98 GAAVRFVILGFAERREMEEEEKR 120
A +G + + + EEEK+
Sbjct: 140 SLAKDAAKIGGEKWKSLTEEEKK 162
>At5g02700 putative protein
Length = 456
Score = 28.1 bits (61), Expect = 1.2
Identities = 17/50 (34%), Positives = 27/50 (54%), Gaps = 7/50 (14%)
Query: 7 QHHHSHRR-----SFFTHINWAPDQHPHEVLAFMPS--LFKSNDLARRKR 49
+ H SHRR IN+ PD H +L+F+P+ +++ L+RR R
Sbjct: 11 RQHRSHRRIQRIIDGADFINYMPDDILHHILSFIPTDLAMRTSVLSRRWR 60
>At3g02150 unknown protein
Length = 355
Score = 28.1 bits (61), Expect = 1.2
Identities = 11/34 (32%), Positives = 18/34 (52%), Gaps = 5/34 (14%)
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPHEVLAFMP 36
HN HHH H+ H++ D H H++ + +P
Sbjct: 208 HNLEHHHHHHQ-----HLSLQADYHSHQLHSLVP 236
>At3g46140 protein kinase - like
Length = 376
Score = 27.7 bits (60), Expect = 1.5
Identities = 19/76 (25%), Positives = 34/76 (44%), Gaps = 7/76 (9%)
Query: 24 PDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEE-------KASIWNKTQIKTP 76
PDQ+P L P + K+N R S +S+ + ++ K+S W K++
Sbjct: 45 PDQNPIFTLKLSPIILKNNKRKRFTPKSSSSRSVEEITKQVFDGVVRKSSSWIKSEFLGR 104
Query: 77 LNKASTFEATKLRSGT 92
+ S + AT ++ T
Sbjct: 105 GSYGSVYLATSKKAKT 120
>At3g21140 unknown protein
Length = 387
Score = 27.7 bits (60), Expect = 1.5
Identities = 12/44 (27%), Positives = 20/44 (45%)
Query: 8 HHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRAS 51
HH F + +++APD+ H + F P + +L R S
Sbjct: 167 HHRREGYPFGSLVDFAPDRMGHPIFLFSPLAIHTRNLLNEPRCS 210
>At2g40480 hypothetical protein
Length = 541
Score = 27.7 bits (60), Expect = 1.5
Identities = 20/47 (42%), Positives = 27/47 (56%), Gaps = 10/47 (21%)
Query: 13 RRSFFTHINWAPDQHPHEVLAFMPS-LFKSN----DLARRKRASDGD 54
RRSFF+H+N +H HE L +P + KSN D+ RRK+ D
Sbjct: 426 RRSFFSHLN----KH-HEPLDILPKPVLKSNISMRDVLRRKQVPKED 467
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 27.7 bits (60), Expect = 1.5
Identities = 12/30 (40%), Positives = 21/30 (70%)
Query: 34 FMPSLFKSNDLARRKRASDGDKSLNKTQEE 63
F+ + +LAR+ R++DGD+SL K +E+
Sbjct: 1150 FLNQIKSQIELARQDRSTDGDRSLWKQKED 1179
>At4g21450 unknown protein
Length = 295
Score = 27.3 bits (59), Expect = 2.0
Identities = 23/108 (21%), Positives = 39/108 (35%), Gaps = 9/108 (8%)
Query: 2 IHNPGQHHHSHRRSFFTHINW--------APDQHPHEVLAFMPSLFKSNDLARRKRASDG 53
+HN QHHH H + + +QHP + + S+ KS +R+ D
Sbjct: 54 LHNHHQHHHQHHHQHHHQLGYNGPHGDGSGQNQHPTPSPS-VSSVAKSFLPTKRRLKLDP 112
Query: 54 DKSLNKTQEEKASIWNKTQIKTPLNKASTFEATKLRSGTCIFRSGAAV 101
+ L E + + +IK F+ +C R A+
Sbjct: 113 SEKLYFPYEPGKQVRSAIKIKNTSKSHVAFKFQTTAPKSCFMRPPGAI 160
>At5g29100 putative protein
Length = 481
Score = 26.9 bits (58), Expect = 2.6
Identities = 22/71 (30%), Positives = 38/71 (52%), Gaps = 11/71 (15%)
Query: 16 FFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEEKASIWNKTQIKT 75
FF +I+ D+ H+++ F P ++R+R S+G+ S + +E S + T T
Sbjct: 421 FFANID---DEVFHKMVPFTPG-------SKRRRRSNGE-STEEPPKEFQSKKDPTVEPT 469
Query: 76 PLNKASTFEAT 86
LNK + +EAT
Sbjct: 470 TLNKRAKWEAT 480
>At4g12830 hydrolase like protein
Length = 393
Score = 26.9 bits (58), Expect = 2.6
Identities = 15/45 (33%), Positives = 20/45 (44%), Gaps = 3/45 (6%)
Query: 2 IHNPGQHHHSHRRSFFTHINWAPDQ---HPHEVLAFMPSLFKSND 43
IHNP H H R+ T + +P H H + L KSN+
Sbjct: 5 IHNPNFFHTHHLRASLTSASASPSSYLFHRHSASKYPSFLCKSNN 49
>At3g45790 putative protein
Length = 376
Score = 26.9 bits (58), Expect = 2.6
Identities = 20/76 (26%), Positives = 33/76 (43%), Gaps = 7/76 (9%)
Query: 24 PDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLNKTQEE-------KASIWNKTQIKTP 76
PDQ+P L P L K+N R S S+ + ++ K+S W K++
Sbjct: 45 PDQNPIFTLKLSPILLKNNKRKRFTPKSSSSLSVEEITKQVFDGVVRKSSSWIKSEFLGR 104
Query: 77 LNKASTFEATKLRSGT 92
+ S + AT ++ T
Sbjct: 105 GSYGSVYLATSKKAKT 120
>At3g28410 hypothetical protein
Length = 465
Score = 26.9 bits (58), Expect = 2.6
Identities = 16/50 (32%), Positives = 27/50 (54%), Gaps = 7/50 (14%)
Query: 7 QHHHSHRR-----SFFTHINWAPDQHPHEVLAFMPS--LFKSNDLARRKR 49
+ H SHR+ IN+ PD H +L+F+P+ +++ L+RR R
Sbjct: 12 RQHRSHRKIQRIIDGADFINYMPDDILHHILSFIPTDLAMRTSVLSRRWR 61
>At5g18230 unknown protein
Length = 843
Score = 26.2 bits (56), Expect = 4.5
Identities = 17/44 (38%), Positives = 21/44 (47%), Gaps = 3/44 (6%)
Query: 45 ARRKRASDGDKSLNKTQEEKA---SIWNKTQIKTPLNKASTFEA 85
A RK + D+ L K QE SIWNK +N+ FEA
Sbjct: 3 ASRKLQGEIDRVLKKVQEGVDVFDSIWNKVYDTDNVNQKEKFEA 46
>At3g15470 unknown protein
Length = 883
Score = 26.2 bits (56), Expect = 4.5
Identities = 14/45 (31%), Positives = 22/45 (48%), Gaps = 1/45 (2%)
Query: 43 DLARRKRASDGDKSLNKTQEEKASIWNKTQIKTPLNKASTFEATK 87
+L RR+ D DK+ +K E+ + NK K S F++ K
Sbjct: 297 ELMRRQNVEDSDKNTSKENEDSGNS-NKDNASKSKKKGSWFKSIK 340
>At3g10810 putative RING zinc finger protein
Length = 684
Score = 26.2 bits (56), Expect = 4.5
Identities = 16/56 (28%), Positives = 20/56 (35%), Gaps = 5/56 (8%)
Query: 3 HNPGQHHHSHRRSFFTHINWAPDQHPHEVLAFMPSLFKSNDLARRKRASDGDKSLN 58
H+ HHH H H N +P P P+ +S RKRA N
Sbjct: 526 HHHHHHHHHHNHHHHHHHNLSPKMAPEVSPVASPAPHRS-----RKRAPSAPPPCN 576
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.129 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,870,038
Number of Sequences: 26719
Number of extensions: 103875
Number of successful extensions: 510
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 23
Number of HSP's that attempted gapping in prelim test: 496
Number of HSP's gapped (non-prelim): 33
length of query: 125
length of database: 11,318,596
effective HSP length: 87
effective length of query: 38
effective length of database: 8,994,043
effective search space: 341773634
effective search space used: 341773634
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0035.12


