

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0034a.6
(1532 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
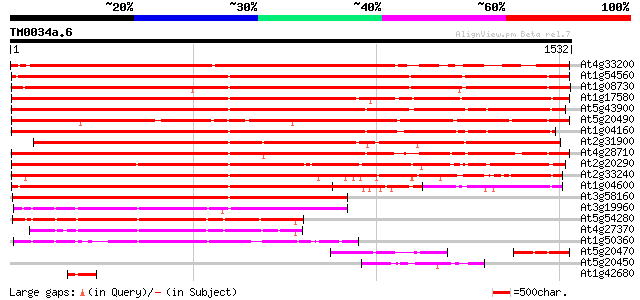
At4g33200 myosin - like protein 1897 0.0
At1g54560 1606 0.0
At1g08730 myosin MYA1, class V (Z28389) like protein 1592 0.0
At1g17580 myosin 1560 0.0
At5g43900 myosin heavy chain MYA2 (pir||S51824) 1535 0.0
At5g20490 myosin-like protein 1508 0.0
At1g04160 myosin heavy chain MYA2 1496 0.0
At2g31900 putative unconventional myosin 1446 0.0
At4g28710 myosin heavy chain - like protein (fragment) 1412 0.0
At2g20290 putative myosin heavy chain 1333 0.0
At2g33240 putative myosin heavy chain 1320 0.0
At1g04600 putative myosin heavy chain 1110 0.0
At3g58160 myosin heavy chain MYA3 992 0.0
At3g19960 myosin 629 e-180
At5g54280 myosin heavy chain 596 e-170
At4g27370 myosin heavy chain - like protein 568 e-162
At1g50360 myosin, putative 466 e-131
At5g20470 myosin-like protein 159 2e-38
At5g20450 putative protein 147 5e-35
At1g42680 myosin heavy chain ATM2, putative 101 4e-21
>At4g33200 myosin - like protein
Length = 1374
Score = 1897 bits (4915), Expect = 0.0
Identities = 988/1528 (64%), Positives = 1165/1528 (75%), Gaps = 177/1528 (11%)
Query: 1 MSFRMGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEH 60
++ R G KVWV+D+D AW+AA+VL D N++ + T +GKK+ RD D+EEH
Sbjct: 9 LNLRKGDKVWVEDKDLAWIAADVL--DSFDNKLHVETSTGKKLFR-------RDPDDEEH 59
Query: 61 GGVEDMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYK 120
GV+DMT+L YL+E GVLYNL+RRY LNDIYTYTGSILIAVNPF KLPHLY+ HMMEQY
Sbjct: 60 NGVDDMTKLTYLHEAGVLYNLQRRYALNDIYTYTGSILIAVNPFKKLPHLYNGHMMEQYM 119
Query: 121 GAPLGELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAA 180
GAP GELSPHVFAV+D +YRAM+++ +SQSILVSGESGAGKTETTKLIMQYLTFVGGRA
Sbjct: 120 GAPFGELSPHVFAVSDVAYRAMIDDSRSQSILVSGESGAGKTETTKLIMQYLTFVGGRAT 179
Query: 181 GDDRSVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLER 240
DDRSVEQQVLESNPLLEAFGNA+TVRNDNSSRFGKFVEIQFD+NGRISGAAIRTYLLER
Sbjct: 180 DDDRSVEQQVLESNPLLEAFGNAKTVRNDNSSRFGKFVEIQFDTNGRISGAAIRTYLLER 239
Query: 241 SRVVQITDPERNYHCFYQLCAFETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRT 300
SRVV+ITDPERNYHCFYQLCA DAEKY+L +P FHYLNQSK YEL+GVS+ EEY T
Sbjct: 240 SRVVRITDPERNYHCFYQLCASGNDAEKYKLSNPRQFHYLNQSKTYELEGVSSAEEYKNT 299
Query: 301 RRAMNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFM 360
RRAM+IVGIS ++QE IFRTLAAILHLGN+EFS G+E+DSSV+KD +SR H+QMAA+LF
Sbjct: 300 RRAMDIVGISQDEQEGIFRTLAAILHLGNVEFSSGREHDSSVVKDPESRHHLQMAADLFK 359
Query: 361 CDVDLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQ 420
CD +LLL++LCTRSI TREG I+KALD NAAV RDTLAKTVYA LFDWLVDKIN+SVGQ
Sbjct: 360 CDANLLLASLCTRSILTREGIIIKALDPNAAVTSRDTLAKTVYAHLFDWLVDKINKSVGQ 419
Query: 421 DINSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWS 480
D S+ QIGVLDIYGFECFK+NSFEQFCINFANEKLQQHFNEHVFKMEQ+EY +EEINWS
Sbjct: 420 DPESRFQIGVLDIYGFECFKNNSFEQFCINFANEKLQQHFNEHVFKMEQDEYRKEEINWS 479
Query: 481 YIEFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQ 540
YIEF+DNQDVL+LIEKKPIG++ALLDEACMFP+STHE+FS KLFQ+F HPRL K KFS+
Sbjct: 480 YIEFIDNQDVLDLIEKKPIGVIALLDEACMFPRSTHESFSMKLFQNFRFHPRLEKPKFSE 539
Query: 541 TDFAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYR 600
TDF +SHYAGK T FLDKNRDY +VEHCNLLSSSKCPFV+G+FP PEES+RSSY+
Sbjct: 540 TDFTLSHYAGKAT-----FLDKNRDYTIVEHCNLLSSSKCPFVAGIFPSAPEESTRSSYK 594
Query: 601 FSSVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVR 660
FSSV+SRFKQQLQALMETL+ TEPHYVRCVKPNSLNRPQ FE+ SV+HQLRCGGVLEAVR
Sbjct: 595 FSSVSSRFKQQLQALMETLSKTEPHYVRCVKPNSLNRPQKFESLSVLHQLRCGGVLEAVR 654
Query: 661 ISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRA 720
ISLAGYPTRR YS+FVDRFGL+A EFMD S D++A+ EKIL KL L N+QLGRTKVFLRA
Sbjct: 655 ISLAGYPTRRNYSDFVDRFGLLAPEFMDESNDEQALTEKILSKLGLGNYQLGRTKVFLRA 714
Query: 721 GQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASK 780
GQIGILDSRRAEVLD +AR IQR+LRTF+ + FI+ RA+A+ +QA CRG + + YA++
Sbjct: 715 GQIGILDSRRAEVLDASARLIQRRLRTFVTHQNFISARASAISIQAYCRGCLSRNAYATR 774
Query: 781 RETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQAC 840
R AAA+ VQK++R WL R ++KL S+A ++QSC+R TR +F H KEHRAA+ IQA
Sbjct: 775 RNAAAAVLVQKHVRRWLSRCAFVKLVSAAIVLQSCIRADSTRLKFSHQKEHRAASLIQAH 834
Query: 841 WRMYKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDEL 900
WR++KFRSAF Q+SI+AIQC WR++ AKRE R+LKQ ANE GALRLAK+KLEK+L++L
Sbjct: 835 WRIHKFRSAFRHRQSSIIAIQCRWRQKLAKREFRKLKQVANEAGALRLAKTKLEKRLEDL 894
Query: 901 TWRLHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELS 960
WRL LEK++R S E+AK EISKLQK +E+ +L+LDAA+LATINECNKNAVL+ Q ++S
Sbjct: 895 EWRLQLEKRLRTSGEEAKSSEISKLQKTLESFSLKLDAARLATINECNKNAVLEKQLDIS 954
Query: 961 IKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFE 1020
+KEKSA++REL + E++K+NA+LK S+++ EKK LE EL+NA+ + T++KL+E E
Sbjct: 955 MKEKSAVERELNGMVELKKDNALLKNSMNSLEKKNRVLEKELLNAKTNCNNTLQKLKEAE 1014
Query: 1021 QKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVA-SRT 1079
++CS+L+ +V+SLEEK+ LE+EN VL QK L ++ + + L E++S+AV ++
Sbjct: 1015 KRCSELQTSVQSLEEKLSHLENENQVLMQKTLI----TSPERIGQILGEKHSSAVVPAQN 1070
Query: 1080 ERKPIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAA 1139
+R+ +FE N E LSRCIKENLGF + KPLAA
Sbjct: 1071 DRRSVFE------------------------------NYELLSRCIKENLGFNDDKPLAA 1100
Query: 1140 PIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLR 1199
+IYKCLLHW AFESE TAIF+ IIEGINE LK RNLR
Sbjct: 1101 CVIYKCLLHWRAFESESTAIFNIIIEGINEALK-----------------------RNLR 1137
Query: 1200 SNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYP 1259
SN FL + QR +G A G KSP K G DDG SH+EARYP
Sbjct: 1138 SNSFLNASAQR-SGRAAY-------------------GVKSPFKLHGPDDGASHIEARYP 1177
Query: 1260 AILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQQPS 1319
A+LFKQQLTACVEKI+GL+RD+LKKELSPLLG CIQ
Sbjct: 1178 ALLFKQQLTACVEKIYGLIRDNLKKELSPLLGSCIQ------------------------ 1213
Query: 1320 GGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGE 1379
VPSFFIRKLVTQVFSFIN++LFNSLLLRRECCTFSNGE
Sbjct: 1214 ----------------------VPSFFIRKLVTQVFSFINLSLFNSLLLRRECCTFSNGE 1251
Query: 1380 YMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALT 1439
Y+KSG++ELEKWI NAKEE LT
Sbjct: 1252 YVKSGISELEKWIANAKEE--------------------------------------VLT 1273
Query: 1440 VRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAED 1499
+RQIYRISTMYWDDKYGTQSVS+EVVS+MR +V KDNQ TSNSFLLDDDMSIPFSAED
Sbjct: 1274 IRQIYRISTMYWDDKYGTQSVSSEVVSQMRVLV-DKDNQKQTSNSFLLDDDMSIPFSAED 1332
Query: 1500 IDMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
ID AIP +DP EI+ P FV EY+CAQ L
Sbjct: 1333 IDKAIPVLDPSEIEPPKFVSEYTCAQSL 1360
>At1g54560
Length = 1529
Score = 1606 bits (4158), Expect = 0.0
Identities = 832/1529 (54%), Positives = 1094/1529 (71%), Gaps = 18/1529 (1%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VW++D D AW+ + L G V++ +GKK+ A K+ P+D E GGV+
Sbjct: 12 VGSHVWIEDSDVAWI--DGLVEKINGQDVEVQATNGKKITAKLSKIYPKDM-EAPAGGVD 68
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMT+L+YL+EPGVL NLK RY LN+IYTYTG+ILIA+NPF +LPH+YD HMM+QYKGAP
Sbjct: 69 DMTKLSYLHEPGVLQNLKIRYELNEIYTYTGNILIAINPFQRLPHIYDAHMMQQYKGAPF 128
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDR 184
GELSPHVFAVAD +YRAM+NEGKS SILVSGESGAGKTETTK++M+YL ++GGRA + R
Sbjct: 129 GELSPHVFAVADVAYRAMINEGKSNSILVSGESGAGKTETTKMLMRYLAYLGGRAVTEGR 188
Query: 185 SVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVV 244
+VEQQVLESNP+LEAFGNA+TVRN+NSSRFGKFVEIQFD GRISGAA+RTYLLERSRV
Sbjct: 189 TVEQQVLESNPVLEAFGNAKTVRNNNSSRFGKFVEIQFDKQGRISGAAVRTYLLERSRVC 248
Query: 245 QITDPERNYHCFYQLCAF-ETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRA 303
QI+DPERNYHCFY LCA + + EKY+LGHP FHYLNQSK +EL G+S+ +Y+ TRRA
Sbjct: 249 QISDPERNYHCFYLLCAAPQEELEKYKLGHPKTFHYLNQSKCFELVGISDAHDYIATRRA 308
Query: 304 MNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDV 363
M+IVG+S ++QEAIFR +AAILHLGN+EF+ GKE DSSV KD+KS+FH+ A L MCDV
Sbjct: 309 MDIVGMSEKEQEAIFRVVAAILHLGNVEFTKGKEVDSSVPKDDKSKFHLNTVAELLMCDV 368
Query: 364 DLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDIN 423
L LC R + T E I ++LD +A+ RD LAKT+Y+RLFDWLV+KIN S+GQD
Sbjct: 369 KALEDALCKRVMVTPEEVIKRSLDPQSALISRDGLAKTIYSRLFDWLVEKINVSIGQDAT 428
Query: 424 SQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIE 483
S+ IGVLDIYGFE FK NSFEQFCINF NEKLQQHFN+HVFKMEQEEY +E I+WSYIE
Sbjct: 429 SRSLIGVLDIYGFESFKTNSFEQFCINFTNEKLQQHFNQHVFKMEQEEYTKEAIDWSYIE 488
Query: 484 FVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDF 543
FVDNQDVL+LIEKKP GIVALLDEACMFPKSTHETF+ KL+Q F +H R K K S+TDF
Sbjct: 489 FVDNQDVLDLIEKKPGGIVALLDEACMFPKSTHETFANKLYQTFKTHKRFIKPKLSRTDF 548
Query: 544 AISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSS 603
A++HYAG+V Y +D FLDKN+DYV+ EH +LL +SKCPFV GLFP PEE+S+SS +FSS
Sbjct: 549 AVAHYAGEVQYQSDLFLDKNKDYVIPEHQDLLGASKCPFVVGLFPPLPEETSKSS-KFSS 607
Query: 604 VASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISL 663
+ SRFK QLQ LMETLNSTEPHY+RCVKPN+L +P +FEN +++ QLRCGGVLEA+RIS
Sbjct: 608 IGSRFKLQLQQLMETLNSTEPHYIRCVKPNNLLKPAVFENVNIMQQLRCGGVLEAIRISC 667
Query: 664 AGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQI 723
AGYPTR+ + EF++RFGL+ ++G+Y++KA A+KIL + L+ +Q+G+TKVFLRAGQ+
Sbjct: 668 AGYPTRKPFFEFINRFGLLYPRALEGNYEEKAAAQKILDNIGLKGYQVGKTKVFLRAGQM 727
Query: 724 GILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRET 783
LD+RR VL AA+ IQR++RT A+R FI +R A + LQA CRG + K++ + R
Sbjct: 728 AELDARRTMVLSAAAKKIQRRIRTHQAQRRFILLRKATISLQALCRGRLSSKIFDNLRRQ 787
Query: 784 AAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRM 843
AAA+ +QK R RK+Y L +A ++Q+ +R ++F K+ +AAT+IQA +R
Sbjct: 788 AAAVKIQKNARRLHSRKSYKNLHVAALVVQTGLRAMAAHKQFRFRKQTKAATTIQAQFRC 847
Query: 844 YKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELTWR 903
++ F + + ++ Q WR + A+RELR+LK + ETGAL+ AK LEK+++ELT+R
Sbjct: 848 HRATLYFKKLKKGVILSQTRWRGKLARRELRQLKMASRETGALKEAKDMLEKKVEELTYR 907
Query: 904 LHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIKE 963
LEK+ RV E+ K EI KLQ +E + ++D + E + + E
Sbjct: 908 AQLEKRSRVDLEEEKNQEIKKLQSSLEEMRKKVDETNGLLVKEREAAKKAIEEAPPVVTE 967
Query: 964 KSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFEQKC 1023
L + ++ + +E LK +L+ +++ + AQ+ ++ +KL + E+K
Sbjct: 968 TQVLVEDTQKIEALTEEVEGLKANLEQEKQRADDATRKFDEAQESSEDRKKKLEDTEKKA 1027
Query: 1024 SQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVASRTERKP 1083
QL+++V LEEK +LE EN VLRQ+A+S G ++S+ +R S + + +P
Sbjct: 1028 QQLQESVTRLEEKCNNLESENKVLRQQAVSIAPNKFLSGRSRSILQRGSESGHLSVDARP 1087
Query: 1084 IFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAAPIIY 1143
+ + + I D + K E+ Q+N E L RCI ++LGF+ +P+ A IIY
Sbjct: 1088 SLDLHSHS--INRRDLSEVDDKPQKSLNEKQQENQELLIRCIVQHLGFQGKRPVTACIIY 1145
Query: 1144 KCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLRSNGF 1203
KCLL W +FE ERT++FD II+ I + ++ +D++++L YWLSN S LL LLQR L+++G
Sbjct: 1146 KCLLQWRSFEVERTSVFDRIIQTIGQAIETQDNNNILAYWLSNASTLLLLLQRTLKASGA 1205
Query: 1204 LTTTGQ-RYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYPAIL 1262
Q R + SA L R +N + G D + VEA+YPA+L
Sbjct: 1206 AGMAPQRRRSSSATLFGRMTQSFRGTPQGVNLA-------MINGGVDTLRQVEAKYPALL 1258
Query: 1263 FKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGK-LSRSPSG-LPQQPSG 1320
FKQQLTA VEKI+G++RD+LKKE+SPLLGLCIQAP+T R K SRS QQ
Sbjct: 1259 FKQQLTAYVEKIYGMIRDNLKKEISPLLGLCIQAPRTSRASLVKGASRSVGNTAAQQALI 1318
Query: 1321 GQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEY 1380
W IV L + ++ L NHVP F +RK+ TQ+FSFIN+ LFNSLLLRRECC+FSNGEY
Sbjct: 1319 AHWQGIVKSLTNFLNNLKSNHVPPFLVRKVFTQIFSFINVQLFNSLLLRRECCSFSNGEY 1378
Query: 1381 MKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTV 1440
+K+GLAELE W NA +EYAG+SW EL +IRQA+GFLVIHQK KK+LDEI +LCP L++
Sbjct: 1379 VKAGLAELEHWCYNATDEYAGSSWDELKHIRQAIGFLVIHQKPKKTLDEISHELCPVLSI 1438
Query: 1441 RQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDI 1500
+Q+YRISTMYWDDKYGT SVS +V++ MR ++ ++D+ N SNSFLLDDD SIPFS +D+
Sbjct: 1439 QQLYRISTMYWDDKYGTHSVSPDVIANMR-VLMTEDSNNAVSNSFLLDDDSSIPFSVDDL 1497
Query: 1501 DMAIPAIDPDEIDLPAFVPEYSCAQFLNP 1529
++ I+ +++ P + E S FL P
Sbjct: 1498 SKSMERIEIGDVEPPPLIRENSGFSFLLP 1526
>At1g08730 myosin MYA1, class V (Z28389) like protein
Length = 1572
Score = 1592 bits (4123), Expect = 0.0
Identities = 840/1569 (53%), Positives = 1100/1569 (69%), Gaps = 58/1569 (3%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VW +D + AW+ EV +G +Q T GKKV A K+ P+D E GGV+
Sbjct: 17 IGSHVWFEDPEVAWIDGEVEKINGQEVVIQATT--GKKVTAKLSKIYPKDV-EAPAGGVD 73
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMT+L+YL+EPGVL NLK RY LN+IYTYTG+ILIA+NPF +LPH+YD HMM+QYKGAPL
Sbjct: 74 DMTKLSYLHEPGVLQNLKIRYELNEIYTYTGNILIAINPFQRLPHIYDAHMMQQYKGAPL 133
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDR 184
GELSPHVFAVAD +YRAM+NEGKS SILVSGESGAGKTETTK++M+YL ++GGRA + R
Sbjct: 134 GELSPHVFAVADVAYRAMINEGKSNSILVSGESGAGKTETTKMLMRYLAYLGGRAVTEGR 193
Query: 185 SVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVV 244
+VEQQVLESNP+LEAFGNA+TVRN+NSSRFGKFVEIQFD GRISGAAIRTYLLERSRV
Sbjct: 194 TVEQQVLESNPVLEAFGNAKTVRNNNSSRFGKFVEIQFDKQGRISGAAIRTYLLERSRVC 253
Query: 245 QITDPERNYHCFYQLCAF-ETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRA 303
QI+DPERNYHCFY LCA + + EKY+LGHP FHYLNQSK +EL G+S+ +Y+ TRRA
Sbjct: 254 QISDPERNYHCFYLLCAAPQEEIEKYKLGHPKTFHYLNQSKCFELVGISDAHDYLATRRA 313
Query: 304 MNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDV 363
M+IVGIS ++QEAIFR +AAILH+GNI+F+ GKE DSSV KDEKS+FH++ AA L MCD+
Sbjct: 314 MDIVGISEKEQEAIFRVVAAILHIGNIDFTKGKEVDSSVPKDEKSKFHLKTAAELLMCDL 373
Query: 364 DLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDIN 423
L LC R + T E I ++LD +AV RD LAKTVY+RLFDWLVDKIN+S+GQD N
Sbjct: 374 KALEDALCKRVMITPEEVIKRSLDPQSAVTSRDGLAKTVYSRLFDWLVDKINKSIGQDAN 433
Query: 424 SQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIE 483
S+ IGVLDIYGFE FK NSFEQFCINF NEKLQQHFN+HVFKMEQEEY +E I+WSYIE
Sbjct: 434 SRSLIGVLDIYGFESFKTNSFEQFCINFTNEKLQQHFNQHVFKMEQEEYTKEAIDWSYIE 493
Query: 484 FVDNQDVLELIEK----------------------------------KPIGIVALLDEAC 509
FVDNQDVL+LIEK KP GIVALLDEAC
Sbjct: 494 FVDNQDVLDLIEKVISEPRKDNVNKITPHTGWILLSHFISPFIFHLQKPGGIVALLDEAC 553
Query: 510 MFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFAISHYAGKVTYHTDTFLDKNRDYVVV 569
MFPKSTHETF+ KL+Q F +H R K K S+TDFA++HYAG+V Y ++ FLDKN+DYV+
Sbjct: 554 MFPKSTHETFANKLYQTFKTHKRFIKPKLSRTDFAVAHYAGEVLYQSELFLDKNKDYVIP 613
Query: 570 EHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSSVASRFKQQLQALMETLNSTEPHYVRC 629
EH +LL +SKCPFV GLFP PEE+S+SS +FSS+ SRFK QLQ LMETLN TEPHY+RC
Sbjct: 614 EHQDLLGASKCPFVVGLFPPLPEETSKSS-KFSSIGSRFKLQLQQLMETLNCTEPHYIRC 672
Query: 630 VKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISLAGYPTRRTYSEFVDRFGLIALEFMDG 689
VKPN+L +P +FEN +++ QLRCGGVLEA+RIS AGYPTR+ + EF++RFGL++ ++G
Sbjct: 673 VKPNNLLKPAIFENVNIMQQLRCGGVLEAIRISCAGYPTRKPFFEFINRFGLLSPAALEG 732
Query: 690 SYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFI 749
++D+K +KIL + L+ +Q+G+TKVFLRAGQ+ LD+RRAEVL +AA+ IQR++RT
Sbjct: 733 NFDEKVACQKILDNMGLKGYQIGKTKVFLRAGQMAELDARRAEVLSSAAKKIQRRIRTHQ 792
Query: 750 ARRVFIAVRAAAVCLQACCRGYIGQKMYASKRETAAAISVQKYIRMWLRRKTYMKLCSSA 809
A++ FI +R A + LQA CRG + K Y + R AAA+ +QK R RK+Y KL ++
Sbjct: 793 AQKRFIVLRKATISLQAICRGRLSCKHYDNLRREAAAVKIQKNGRRHYSRKSYKKLHVAS 852
Query: 810 TIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRMYKFRSAFHRHQASIVAIQCLWRRRQA 869
++Q+ +R R++F K+ +AAT +QA WR ++ S + + + +V Q WR R A
Sbjct: 853 LVVQTGLRAMAARKQFRFRKQTKAATIVQAQWRCHRAISYYKKLKNGVVLSQTRWRGRLA 912
Query: 870 KRELRRLKQEANETGALRLAKSKLEKQLDELTWRLHLEKKIRVSNEDAKQIEISKLQKMI 929
KRELR+LK A ETGAL+ AK LEK+++ELT+R+ LEK+ R E+AK EI KL+
Sbjct: 913 KRELRKLKMAARETGALKEAKDMLEKKVEELTYRVQLEKRSRGDLEEAKTQEILKLKSSF 972
Query: 930 EALNLELDAAKLATINECNKNAVLQNQFELSIKEKSALKRELVAVDEIRKENAMLKVSLD 989
E + ++D + E + IKE L + ++ + +E +KV+L+
Sbjct: 973 EEMRKKVDETNALLLKEREAAKKAAEEAPPVIKETQILVEDTKKIELMTEELESVKVTLE 1032
Query: 990 AFEKKCTALEVELINAQKGRDETIEKLREFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQ 1049
+++ + AQ+ ++ +KL E E+K QL++++ +EEK +LE EN VLRQ
Sbjct: 1033 NEKQRADDAVRKFEEAQESLEDKKKKLEETEKKGQQLQESLTRMEEKCSNLESENKVLRQ 1092
Query: 1050 KALSAPLKSNRQGFAKSLSERYSNAVASRTERKPIFESPTPTKLIPTFTPGMSDSRRSKL 1109
+A+S G ++S+ +R S + + + + + + I P + + K
Sbjct: 1093 QAVSMAPNKFLSGRSRSILQRGSESGHLAVDARSNLDLHSHS--INHRDPSEVEDKPQKS 1150
Query: 1110 TAERHQDNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINE 1169
E+ Q+N + L R I ++LGF+ +P+ A IIYKCLL W +FE ERT++FD II+ I
Sbjct: 1151 LNEKQQENQDLLIRSIVQHLGFQGNRPITACIIYKCLLQWRSFEVERTSVFDRIIQTIGH 1210
Query: 1170 VLKARDDDDVLPYWLSNTSALLCLLQRNLRSNGFLTTTGQRYTGSAGLASRTVHVSY--- 1226
++ +D+++ L YWLSNTS LL LLQR L+++G QR S+ + S+
Sbjct: 1211 AIETQDNNNTLAYWLSNTSTLLLLLQRTLKASGAAGMAPQRRRSSSATLFGRMSQSFRGA 1270
Query: 1227 ---LVSTSINHSCGPKSPLKFIGYDDGVSHVEARYPAILFKQQLTACVEKIFGLLRDDLK 1283
+ IN + G G D VEA+YPA+LFKQQLTA VEKI+G++RD+LK
Sbjct: 1271 PPGVNLAMINGAAG--------GGADTFRQVEAKYPALLFKQQLTAYVEKIYGMIRDNLK 1322
Query: 1284 KELSPLLGLCIQAPKTGRQHGGK-LSRSPSG-LPQQPSGGQWANIVNFLDSLMSKLHGNH 1341
KE+SPLLGLCIQAP+T R K SRS QQ W IV L + ++ L N+
Sbjct: 1323 KEISPLLGLCIQAPRTSRASLVKGASRSVGNTAAQQALIAHWQGIVKSLTNFLNTLKSNN 1382
Query: 1342 VPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEYMKSGLAELEKWIVNAKEEYAG 1401
VPSF +RK+ TQ+FSFIN+ LFNSLLLRRECC+FSNGEY+K+GL+ELE W A EYAG
Sbjct: 1383 VPSFLVRKVFTQIFSFINVQLFNSLLLRRECCSFSNGEYVKAGLSELEHWCFKATNEYAG 1442
Query: 1402 TSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTVRQIYRISTMYWDDKYGTQSVS 1461
+SW EL +IRQA+GFLV+HQK KK+LDEI DLCP L+++Q+YRISTMYWDDKYGT SVS
Sbjct: 1443 SSWDELKHIRQAIGFLVVHQKPKKTLDEISHDLCPVLSIQQLYRISTMYWDDKYGTHSVS 1502
Query: 1462 NEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEY 1521
+V++ MR ++ ++D+ N SNSFLLDDD SIPFS +D+ ++ + +I+ P + E
Sbjct: 1503 PDVIANMR-VLMTEDSNNAVSNSFLLDDDSSIPFSVDDLSKSMEKFEIADIEPPPLIREN 1561
Query: 1522 SCAQFLNPI 1530
S FL P+
Sbjct: 1562 SGFSFLLPV 1570
>At1g17580 myosin
Length = 1520
Score = 1560 bits (4038), Expect = 0.0
Identities = 828/1535 (53%), Positives = 1077/1535 (69%), Gaps = 39/1535 (2%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VWV+D AW+ EV DG V + T GK V+ + P+D E GGV+
Sbjct: 8 VGSHVWVEDPHLAWIDGEVTRIDG--INVHVKTKKGKTVVTNV--YFPKDT-EAPSGGVD 62
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMT+L+YL+EPGVL NL+ RY LN+IYTYTG+ILIAVNPF +LPH+Y+ MMEQYKG L
Sbjct: 63 DMTKLSYLHEPGVLRNLETRYELNEIYTYTGNILIAVNPFQRLPHIYETDMMEQYKGIAL 122
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDR 184
GELSPHVFA+ DA+YRAM+NEGK+ SILVSGESGAGKTETTK++M+YL F+GGR+ + R
Sbjct: 123 GELSPHVFAIGDAAYRAMINEGKNNSILVSGESGAGKTETTKMLMRYLAFLGGRSGVEGR 182
Query: 185 SVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVV 244
+VEQQVLESNP+LEAFGNA+T+RN+NSSRFGKFVEIQFD NGRISGAAIRTYLLERSRV
Sbjct: 183 TVEQQVLESNPVLEAFGNAKTLRNNNSSRFGKFVEIQFDKNGRISGAAIRTYLLERSRVC 242
Query: 245 QITDPERNYHCFYQLCAFET-DAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRA 303
QI+DPERNYHCFY LCA D +KY+L +P FHYLNQS Y+LDGV + EY+ TRRA
Sbjct: 243 QISDPERNYHCFYLLCAAPPEDIKKYKLENPHKFHYLNQSSCYKLDGVDDASEYLETRRA 302
Query: 304 MNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDV 363
M++VGIS+E+QEAIFR +AAILHLGNI+F G+E DSSVIKD+ SR H+ MAA L MC+
Sbjct: 303 MDVVGISNEEQEAIFRVVAAILHLGNIDFGKGEEIDSSVIKDKDSRSHLNMAAELLMCNA 362
Query: 364 DLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDIN 423
L L R + T E I + LD + A+A RDTLAKT+Y+ LFDW+V+KIN S+GQD
Sbjct: 363 QSLEDALIRRVMVTPEEIITRTLDPDNAIASRDTLAKTIYSHLFDWIVNKINTSIGQDPR 422
Query: 424 SQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIE 483
S+ IGVLDIYGFE FK NSFEQFCINF NEKLQQHFN+HVFKMEQEEY +EEI WSYIE
Sbjct: 423 SKSIIGVLDIYGFESFKCNSFEQFCINFTNEKLQQHFNQHVFKMEQEEYTKEEIAWSYIE 482
Query: 484 FVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDF 543
F+DNQDVLELIEKKP GI++LLDEACMFPKSTHETFS KLFQ F H R K K S+TDF
Sbjct: 483 FIDNQDVLELIEKKPGGIISLLDEACMFPKSTHETFSQKLFQTFKEHERFAKPKLSRTDF 542
Query: 544 AISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSS 603
ISHYAG+VTY ++ F+DKN+DY+V EH L ++S C FV+GLF E+SSRSS +FSS
Sbjct: 543 TISHYAGEVTYQSNHFIDKNKDYIVAEHQALFTASNCKFVAGLFHALHEDSSRSS-KFSS 601
Query: 604 VASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISL 663
+ SRFKQQL +LME+LN TEPHY+RC+KPN++ +P +FEN +VIHQLRCGGVLEA+RIS
Sbjct: 602 IGSRFKQQLHSLMESLNGTEPHYIRCIKPNNVLKPGIFENFNVIHQLRCGGVLEAIRISC 661
Query: 664 AGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQI 723
AGYPTR + +F+DRFGL+A E ++G+YDDK + IL K L ++Q+G+TK+FLRAGQ+
Sbjct: 662 AGYPTRLAFYDFLDRFGLLAPEVLEGNYDDKVACQMILDKKSLTDYQIGKTKIFLRAGQM 721
Query: 724 GILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRET 783
LD+RRAEVL NAAR IQRQ RT +AR+ + ++R AA+ LQ+ RG I + ++ R
Sbjct: 722 AELDARRAEVLGNAARVIQRQFRTCMARKNYRSIRNAAIVLQSFLRGEIARAVHKKLRIE 781
Query: 784 AAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRM 843
AAA+ VQK R ++ RK+++ SS ++Q+ +R + R F ++ +AA +QA WR
Sbjct: 782 AAALRVQKNFRRYVDRKSFVTTRSSTIVLQTGLRAMIARSEFRLRRQRKAAIVLQAHWRG 841
Query: 844 YKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELTWR 903
+ S + R Q + + QC WR R A+RELR LK A +TGAL+ AK+KLE++++EL+ R
Sbjct: 842 RQAFSYYTRLQKAAIVTQCAWRCRLARRELRMLKMAARDTGALKDAKNKLEQRVEELSLR 901
Query: 904 LHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIKE 963
LHLEK++R E+AK E++KLQ+ + + L+L + E Q ++I+E
Sbjct: 902 LHLEKRLRTDLEEAKVQEVAKLQEALHTMRLQLKETTAMVVKE-------QEAARVAIEE 954
Query: 964 KSALKRELVAVDEIRKENAM------LKVSLDAFEKKCTALEVELINAQKGRDETIEKLR 1017
S++ +E V V++ K +++ LK L + K + +A +E +KL
Sbjct: 955 ASSVNKEPVVVEDTEKIDSLSNEIDRLKGLLSSETHKADEAQHAYQSALVQNEELCKKLE 1014
Query: 1018 EFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVAS 1077
E +K QL+ +V+ +EK+ SLE EN VLRQ+ L+ ++L+ R +
Sbjct: 1015 EAGRKIDQLQDSVQRFQEKVFSLESENKVLRQQTLTI------SPTTRALALRPKTTIIQ 1068
Query: 1078 RTERKPIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPL 1137
RT K F + T+L T + R K ++ Q+N E L + I E++GF GKP+
Sbjct: 1069 RTPEKDTFSNGETTQLQEPET----EDRPQKSLNQKQQENQELLLKSISEDIGFSEGKPV 1124
Query: 1138 AAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRN 1197
AA +IYKCL+HW +FE ERT+IF+ IIE I ++ +++ DVL YWLSN++ LL LQR
Sbjct: 1125 AACLIYKCLIHWRSFEVERTSIFNRIIETIASAIEMQENSDVLCYWLSNSATLLMFLQRT 1184
Query: 1198 LRSNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCG-PKSPLKFIGYD-DGVSHVE 1255
L++ + T R G V S+ S S G P + IG D + VE
Sbjct: 1185 LKAGATGSITTPRRRGMPSSLFGRVSQSFRGSP---QSAGFPFMTGRAIGGGLDELRQVE 1241
Query: 1256 ARYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQ---HGGKLSRSPS 1312
A+YPA+LFKQQLTA +EKI+G++RD +KKE+SPLL CIQ P+T R G + +
Sbjct: 1242 AKYPALLFKQQLTAFLEKIYGMIRDKMKKEISPLLASCIQVPRTPRSGLVKGRSQNTQNN 1301
Query: 1313 GLPQQPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRREC 1372
+ +P W NIV L+ + + N+VPS I K+ Q+FSFIN+ LFNSLLLRREC
Sbjct: 1302 VVAPKPMIAHWQNIVTCLNGHLRTMRANYVPSLLISKVFGQIFSFINVQLFNSLLLRREC 1361
Query: 1373 CTFSNGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQ 1432
C+FSNGEY+K+GLAELEKW +A EE+ G++W EL +IRQAVGFLVIHQK KKSL EI
Sbjct: 1362 CSFSNGEYVKTGLAELEKWCHDATEEFVGSAWDELKHIRQAVGFLVIHQKPKKSLKEITT 1421
Query: 1433 DLCPALTVRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMS 1492
+LCP L+++Q+YRISTMYWDDKYGT SVS EV++ MR VS I SNSFLLDDD S
Sbjct: 1422 ELCPVLSIQQLYRISTMYWDDKYGTHSVSTEVIATMRAEVSDVSKSAI-SNSFLLDDDSS 1480
Query: 1493 IPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
IPFS +DI ++ ++ E+D P + + S FL
Sbjct: 1481 IPFSLDDISKSMQNVEVAEVDPPPLIRQNSNFMFL 1515
>At5g43900 myosin heavy chain MYA2 (pir||S51824)
Length = 1505
Score = 1535 bits (3975), Expect = 0.0
Identities = 804/1523 (52%), Positives = 1067/1523 (69%), Gaps = 38/1523 (2%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VWV+D D+AW+ EV+ +G + ++++ SGK V+ P+D E GV+
Sbjct: 9 VGSFVWVEDPDEAWIDGEVVQVNG--DEIKVLCTSGKHVVTKISNAYPKDV-EAPASGVD 65
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMTRLAYL+EPGVL NL RY +N+IYTYTGSILIAVNPF +LPHLY +HMM QYKGA L
Sbjct: 66 DMTRLAYLHEPGVLQNLHSRYDINEIYTYTGSILIAVNPFRRLPHLYSSHMMAQYKGASL 125
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDR 184
GELSPH FAVADA+YR M+N+G SQSILVSGESGAGKTE+TKL+M+YL ++GGRAA + R
Sbjct: 126 GELSPHPFAVADAAYRQMINDGVSQSILVSGESGAGKTESTKLLMRYLAYMGGRAAAEGR 185
Query: 185 SVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVV 244
SVEQ+VLESNP+LEAFGNA+TVRN+NSSRFGKFVEIQFD GRISGAAIRTYLLERSRV
Sbjct: 186 SVEQKVLESNPVLEAFGNAKTVRNNNSSRFGKFVEIQFDEKGRISGAAIRTYLLERSRVC 245
Query: 245 QITDPERNYHCFYQLCAF-ETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRA 303
Q++DPERNYHCFY LCA + D +K++L P +HYLNQSK ELD +++ EEY TRRA
Sbjct: 246 QVSDPERNYHCFYMLCAAPQEDVKKFKLEEPKKYHYLNQSKCLELDSINDAEEYHATRRA 305
Query: 304 MNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDV 363
M++VGIS E+Q+AIF +AAILH+GNIEF+ G+E DSS+ KD+KS FH++ AA L CD
Sbjct: 306 MDVVGISTEEQDAIFSVVAAILHIGNIEFAKGEEIDSSIPKDDKSLFHLKTAAELLSCDE 365
Query: 364 DLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDIN 423
L +LC R + TR+ +I K LD AA RD LAK +Y+RLFDWLVDKIN S+GQD +
Sbjct: 366 KALEDSLCKRIMVTRDETITKTLDPEAATLSRDALAKVMYSRLFDWLVDKINSSIGQDHD 425
Query: 424 SQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIE 483
S+ IGVLDIYGFE FK NSFEQFCIN NEKLQQHFN+HVFKMEQEEY +EEINWSYIE
Sbjct: 426 SKYLIGVLDIYGFESFKTNSFEQFCINLTNEKLQQHFNQHVFKMEQEEYKKEEINWSYIE 485
Query: 484 FVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDF 543
FVDNQD+L+LIEKKP GI+ALLDEACMFP+STHETF+ KL+Q F +H R K K +++DF
Sbjct: 486 FVDNQDILDLIEKKPGGIIALLDEACMFPRSTHETFAQKLYQTFKTHKRFTKPKLARSDF 545
Query: 544 AISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSS 603
I HYAG VTY T+ FLDKN+DYV+ EH LL+SS C FV+ LFP ++S +S +FSS
Sbjct: 546 TICHYAGDVTYQTELFLDKNKDYVIAEHQALLNSSSCSFVASLFPPMSDDSKQS--KFSS 603
Query: 604 VASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISL 663
+ +RFKQQL +L+E LN+TEPHY+RC+KPN+L +P +FEN +++ QLRCGGV+EA+RIS
Sbjct: 604 IGTRFKQQLVSLLEILNTTEPHYIRCIKPNNLLKPGIFENENILQQLRCGGVMEAIRISC 663
Query: 664 AGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQI 723
AGYPTR+ + EF+ RFG++A E + + DD A +K+L K+ LE +Q+G+TKVFLRAGQ+
Sbjct: 664 AGYPTRKHFDEFLARFGILAPEVLVKNSDDPAACKKLLDKVGLEGYQIGKTKVFLRAGQM 723
Query: 724 GILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRET 783
LD+RR EVL +A IQR++R+++A++ FI +R +A +Q+ CRGY+ + +Y R
Sbjct: 724 ADLDTRRTEVLGRSASIIQRKVRSYLAKKSFIVLRNSAKQIQSVCRGYLARSVYEGMRRE 783
Query: 784 AAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRM 843
AAA+ +Q+ +R +L RK Y +L S+A +Q+ +RG + R+ ++ +AA IQ R
Sbjct: 784 AAALKIQRDLRRFLARKAYTELYSAAVSVQAGMRGMVARKELCFRRQTKAAIIIQTWCRG 843
Query: 844 YKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELTWR 903
Y R + + + + + QC WR + A+ ELR+LK A ETGAL+ AK+KLEKQ++ELTWR
Sbjct: 844 YLARLHYRKLKKAAITTQCAWRSKVARGELRKLKMAARETGALQAAKNKLEKQVEELTWR 903
Query: 904 LHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIKE 963
L LEK+IR E+AK+ E +K Q +E L L+ + I E + + IKE
Sbjct: 904 LQLEKRIRTDLEEAKKQESAKAQSSLEELQLKCKETEALLIKEREAAKKIAETAPI-IKE 962
Query: 964 KSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFEQKC 1023
+ +EL +D+I EN LK + + E K E +L K + + + E E K
Sbjct: 963 IPVVDQEL--MDKITNENEKLKSMVSSLEMKIGETEKKLQETTKISQDRLNQALEAESKL 1020
Query: 1024 SQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVASRTERK- 1082
+L+ ++ LEEK+L +E E ++ Q+ +S P+++N + + N + E++
Sbjct: 1021 VKLKTAMQRLEEKILDMEAEKKIMHQQTISTPVRTNLGHPPTAPVKNLENGHQTNLEKEF 1080
Query: 1083 PIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAAPII 1142
E TP D + K AER N + L C+K+N+GF NGKP+AA I
Sbjct: 1081 NEAEFTTPV-----------DGKAGKSAAERQIMNVDALIDCVKDNIGFSNGKPVAAFTI 1129
Query: 1143 YKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLRSNG 1202
YKCLLHW FESE+T +FD +I+ I ++ DD+ L YWL++TSALL LLQ++L++NG
Sbjct: 1130 YKCLLHWKCFESEKTNVFDRLIQMIGSAIENEDDNSHLAYWLTSTSALLFLLQKSLKTNG 1189
Query: 1203 FLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYPAIL 1262
T ++ S L R S N + ++ + V VEA+YPA+L
Sbjct: 1190 SGATQSKKPPASTSLFGRMAMSFRSSPASGNLAAAAEAAALAV-----VRPVEAKYPALL 1244
Query: 1263 FKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQQPSGGQ 1322
FKQQL A VEK+FG++RD+LK+ELS LL LCIQAP++ + G + RS +
Sbjct: 1245 FKQQLAAYVEKMFGMVRDNLKRELSTLLSLCIQAPRSSK---GGMLRSGRSFGKDSPAVH 1301
Query: 1323 WANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEYMK 1382
W +I++ L+SL+ L NHVP I+K+ +Q FS+IN+ LFNSLLLR+ECCTFSNGE++K
Sbjct: 1302 WQSIIDGLNSLLVTLKENHVPLVLIQKIYSQTFSYINVQLFNSLLLRKECCTFSNGEFVK 1361
Query: 1383 SGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTVRQ 1442
SGLAELE W AK EY+G SW EL +IRQAVGFLVIHQK + S DEI DLCP L+V+Q
Sbjct: 1362 SGLAELELWCCQAK-EYSGPSWEELKHIRQAVGFLVIHQKYRISYDEIANDLCPVLSVQQ 1420
Query: 1443 IYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDM 1502
+YRI T+YWDD Y T+SVS EV+S MR +++ + N + S+SFLLDDD SIPFS +DI
Sbjct: 1421 LYRICTLYWDDSYNTRSVSQEVISSMRTLMTEESN-DADSDSFLLDDDSSIPFSIDDISS 1479
Query: 1503 AIP-----AIDPDE--IDLPAFV 1518
++ I P E ++ PAFV
Sbjct: 1480 SMEEKDFVGIKPAEELLENPAFV 1502
>At5g20490 myosin-like protein
Length = 1544
Score = 1508 bits (3903), Expect = 0.0
Identities = 811/1562 (51%), Positives = 1064/1562 (67%), Gaps = 88/1562 (5%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VW++D AW+ EV+ +G V T +GK V+A+ + P+D E GGV+
Sbjct: 23 VGSHVWIEDPGAAWIDGEVVKING--EEVHAHTTNGKTVVANIANVFPKDT-EAPPGGVD 79
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMT+L+YL+EPGVL NL RY LN+IYTYTG+ILIAVNPF +LPHLYD HMMEQYKGA
Sbjct: 80 DMTKLSYLHEPGVLNNLAMRYELNEIYTYTGNILIAVNPFQRLPHLYDTHMMEQYKGAGF 139
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDR 184
GELSPHVFA+A+ +YRAM+NEGKS SILVSGESGAGKTETTK++M+YL ++GGR+ + R
Sbjct: 140 GELSPHVFAIAEVAYRAMINEGKSNSILVSGESGAGKTETTKMLMRYLAYLGGRSGVEGR 199
Query: 185 SVEQQVLE-----------------SNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGR 227
+VEQQVLE SNP+LEAFGNA+T+RN+NSSRFGKFVE+QFD+ GR
Sbjct: 200 TVEQQVLEACNASLIHYFGFIDGIQSNPVLEAFGNAKTLRNNNSSRFGKFVELQFDNCGR 259
Query: 228 ISGAAIRTYLLERSRVVQITDPERNYHCFYQLCAFETDA-EKYELGHPSHFHYLNQSKIY 286
ISGAA+RTYLLERSRV QI+DPERNYHCFY LCA + EK++LG P FHYLNQSK Y
Sbjct: 260 ISGAAVRTYLLERSRVCQISDPERNYHCFYLLCAAPPEEREKFKLGDPKLFHYLNQSKCY 319
Query: 287 ELDGVSNVEEYVRTRRAMNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDE 346
+LDGV + EEY+ TRRAM+IVGIS E+Q+AIFR +AAILHLGN+ F+ GKE DSSV+KDE
Sbjct: 320 KLDGVDDTEEYLATRRAMDIVGISEEEQDAIFRVVAAILHLGNVNFAKGKEIDSSVLKDE 379
Query: 347 KSRFHMQMAANLFMCDVDLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARL 406
KSR+H+ + A L CD + L R + T E I + LD ++A RD L
Sbjct: 380 KSRYHLDVCAELLRCDAKKMEDALIKRVMVTPEEVITRTLDPDSATGSRDAL-------- 431
Query: 407 FDWLVDKINRSVGQDINSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFK 466
VDKIN S+GQD NS+ IGVLDIYGFE FK NSFEQFCINF NEKLQQHFN+HVFK
Sbjct: 432 ----VDKINNSIGQDPNSKTIIGVLDIYGFESFKINSFEQFCINFTNEKLQQHFNQHVFK 487
Query: 467 MEQEEYGREEINWSYIEFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQH 526
MEQE+Y +EEINWSYIEFVDN+DVLELIEKKP G++ALLDEACMFPKSTHETF+ KL+Q
Sbjct: 488 MEQEDYTKEEINWSYIEFVDNKDVLELIEKKPGGVIALLDEACMFPKSTHETFAQKLYQT 547
Query: 527 FPSHPRLGKEKFSQTDFAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGL 586
F ++ R K K S+T FAISHYAG+ D FLDKN+DYVV EH +LL +S FV+GL
Sbjct: 548 FKNYKRFTKPKLSRTSFAISHYAGE----ADLFLDKNKDYVVAEHQDLLIASSDTFVAGL 603
Query: 587 FPLPPEESSRSSYRFSSVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSV 646
FP PEE+S S +FSS+ SRFK QLQ+LMETL+STEPHY+RCVKPN++ +P +FEN
Sbjct: 604 FPRLPEETS-SKTKFSSIGSRFKLQLQSLMETLSSTEPHYIRCVKPNNVLKPAIFEN--- 659
Query: 647 IHQLRCGGVLEAVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKL 706
GVLEA+RIS AGYPT+RT+ EF++RFG++A E ++G+YDDK + +L K+ L
Sbjct: 660 -----VNGVLEAIRISCAGYPTKRTFYEFLNRFGVLAPEVLEGNYDDKVACKMLLDKIGL 714
Query: 707 ENFQLGRTKVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQA 766
+ ++LG+TKVFLRAGQ+ LD+RRAEVL NAAR IQRQ RTFIA + F A+R AA+ LQ+
Sbjct: 715 KGYELGKTKVFLRAGQMAELDARRAEVLGNAARRIQRQSRTFIACKEFRALRGAAIVLQS 774
Query: 767 CCR-----------GYIGQKMYASKRETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSC 815
CR G + +Y R AAA+ +QK R + R++Y+++ S +Q+
Sbjct: 775 NCRVELLKRFCCMQGKLACNLYEEMRRQAAAVKIQKIFRRHIARESYLRIRHSTITVQTA 834
Query: 816 VRGFMTRQRFLHIKEHRAATSIQACWRMYKFRSAFHRHQASIVAIQCLWRRRQAKRELRR 875
+RG + R F K+ +AAT IQA R + S + + Q + ++ QC WR R A++ELR
Sbjct: 835 LRGMVARNEFRFRKQMKAATIIQARLRSHLTHSYYKQLQKAALSTQCGWRSRVARKELRT 894
Query: 876 LKQEANETGALRLAKSKLEKQLDELTWRLHLEKKIRVSNEDAKQIEISKLQKMIEALNLE 935
LK A +TGALR AK KLEK+++ELTWRL LEK+ R E+AK E +K Q+ +E + L+
Sbjct: 895 LKMAARDTGALREAKDKLEKRVEELTWRLQLEKRQRTELEEAKTQEYAKQQEALETMRLQ 954
Query: 936 LDAAKLATINECNKNAVLQNQFELSIKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKC 995
++ A A I E + IKE L + ++ + E LK SL A +
Sbjct: 955 VEEANAAVIREREAARKAIEEAPPVIKETPVLVEDTEKINSLTSEVEALKASLQAERQAA 1014
Query: 996 TALEVELINAQKGRDETIEKLREFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAP 1055
L A+ E +L +K QL ++V+ LEEK+ + E E VLRQ+AL+
Sbjct: 1015 ENLRKAFSEAEARNSELATELENATRKADQLHESVQRLEEKLSNSESEIQVLRQQALAIS 1074
Query: 1056 LKSNRQGFAKSLSERYSNAVASRTERKPIFESPTPTKLIPTFTPGM----SDSRRSKLTA 1111
S ++++ R + RT + + TK P T + S+ + K
Sbjct: 1075 PTS------RTMATRSKTMLLPRTPENGNYLN-GGTKTTPDMTLAVREPESEEKPQKHLN 1127
Query: 1112 ERHQDNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVL 1171
E+ Q+N + L +CI +NLG+ KP+AA +IYKCLLHW +FE ERT++FD II+ I +
Sbjct: 1128 EKQQENQDLLVKCISQNLGYNGDKPVAACVIYKCLLHWRSFEVERTSVFDRIIQTIATAI 1187
Query: 1172 KARDDDDVLPYWLSNTSALLCLLQRNLRSNGFLTTTGQRYTGSAGLASRTVHVSYLVSTS 1231
+ D+++VL YWLSN++ LL LLQR L++ G + T QR RT S S
Sbjct: 1188 EVPDNNEVLAYWLSNSATLLLLLQRTLKATGAASLTPQR--------RRTTSASLFGRMS 1239
Query: 1232 INHSCGPKSP-LKFIGYD-----DGVSHVEARYPAILFKQQLTACVEKIFGLLRDDLKKE 1285
P+S L F+ D + VEA+YPA+LFKQQLTA +EKI+G++RD+LKKE
Sbjct: 1240 QGLRGSPQSAGLSFLNRQGLTKLDDLRQVEAKYPALLFKQQLTAFLEKIYGMIRDNLKKE 1299
Query: 1286 LSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQQPSGGQWANIVNFLDSLMSKLHGNHVPSF 1345
+SPLLGLCIQAP+T R K + + QQ W +I L+S ++ + N+ P F
Sbjct: 1300 ISPLLGLCIQAPRTSRASLVKGRAQANAVAQQALIAHWQSIRKSLNSYLNLMKANNAPPF 1359
Query: 1346 FIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEYMKSGLAELEKWIVNAKEEYAGTSWH 1405
+RK+ TQ+FSFIN+ LFN R CC+FSNGEY+K+GLAELE+W + A +EYAG++W
Sbjct: 1360 LVRKVFTQIFSFINVQLFN-----RHCCSFSNGEYVKAGLAELEQWCIEATDEYAGSAWD 1414
Query: 1406 ELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTVRQIYRISTMYWDDKYGTQSVSNEVV 1465
EL +IRQAVGFLVIHQK KK+LDEI ++LCP L+++Q+YRISTMYWDDKYGT SVS++V+
Sbjct: 1415 ELRHIRQAVGFLVIHQKPKKTLDEITRELCPVLSIQQLYRISTMYWDDKYGTHSVSSDVI 1474
Query: 1466 SEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMAIPAIDPDEIDLPAFVPEYSCAQ 1525
+ MR ++ ++D+ N S+SFLLDDD SIPF+ EDI ++ +D ++I+ P + E S
Sbjct: 1475 ANMR-VMMTEDSNNAVSSSFLLDDDSSIPFTVEDISKSMQQVDVNDIEPPQLIRENSGFG 1533
Query: 1526 FL 1527
FL
Sbjct: 1534 FL 1535
>At1g04160 myosin heavy chain MYA2
Length = 1519
Score = 1496 bits (3873), Expect = 0.0
Identities = 781/1491 (52%), Positives = 1056/1491 (70%), Gaps = 42/1491 (2%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VWV+D D+AW+ EV+ +G ++++++ SGK+V+ + P+D E GVE
Sbjct: 9 VGSHVWVEDPDEAWLDGEVVEING--DQIKVLCASGKQVVVKDSNIYPKDV-EAPASGVE 65
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMTRLAYL+EPGVL NL+ RY +N+IYTYTGSILIAVNPF +LPHLY +HMM QYKGA L
Sbjct: 66 DMTRLAYLHEPGVLQNLQSRYDINEIYTYTGSILIAVNPFRRLPHLYSSHMMTQYKGASL 125
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGR-AAGDD 183
GELSPH FAVADA+YR M+NEG SQSILVSGESGAGKTE+TKL+M+YL F+GGR AA +
Sbjct: 126 GELSPHPFAVADAAYRQMVNEGVSQSILVSGESGAGKTESTKLLMRYLAFMGGRGAATEG 185
Query: 184 RSVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRV 243
R+VEQ+VLESNP+LEAFGNA+TV+N+NSSRFGKFVEIQFD +GRISGAAIRTYLLERSRV
Sbjct: 186 RTVEQKVLESNPVLEAFGNAKTVKNNNSSRFGKFVEIQFDQSGRISGAAIRTYLLERSRV 245
Query: 244 VQITDPERNYHCFYQLCAF-ETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRR 302
Q++DPERNYHCFY LCA E DA+K++LG P +HYLNQSK +LD +++ EEY T++
Sbjct: 246 CQVSDPERNYHCFYMLCAAPEEDAKKFKLGDPKIYHYLNQSKCIQLDAMNDAEEYHATKK 305
Query: 303 AMNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCD 362
AM++VGIS E+Q+AIFR +A+ILHLGNIEF+ G E DSS+ +DEKS FH++ AA L MC+
Sbjct: 306 AMDVVGISSEEQDAIFRVVASILHLGNIEFAKGTEIDSSIPRDEKSWFHLKTAAELLMCN 365
Query: 363 VDLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDI 422
L +LC R + TR+ +I K LD AA+ RD LAK +Y+RLFDWLV+KIN S+GQD
Sbjct: 366 EKSLEDSLCKRIMATRDETITKTLDPEAALLSRDALAKVMYSRLFDWLVEKINTSIGQDP 425
Query: 423 NSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYI 482
+S+ IGVLDIYGFE FK NSFEQFCIN NEKLQQHFN+HVFKMEQEEY +EEINWSYI
Sbjct: 426 DSKYLIGVLDIYGFESFKTNSFEQFCINLTNEKLQQHFNQHVFKMEQEEYKKEEINWSYI 485
Query: 483 EFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTD 542
EFVDNQD+L+LIEKKP GI+ALLDEACMFP+STHETF+ KL+Q + +H R K K +++D
Sbjct: 486 EFVDNQDILDLIEKKPGGIIALLDEACMFPRSTHETFAQKLYQTYKNHKRFTKPKLARSD 545
Query: 543 FAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFS 602
F I HYAG VTY T+ FLDKN+DYV+ EH LL++S C FV+ LFP ++S +S +FS
Sbjct: 546 FTICHYAGDVTYQTELFLDKNKDYVIAEHQALLNASTCSFVANLFPPVSDDSKQS--KFS 603
Query: 603 SVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRIS 662
S+ +RFKQQL +L+E LN+TEPHY+RC+KPN+L +P +FEN +V+ QLRCGGV+EA+RIS
Sbjct: 604 SIGTRFKQQLVSLLEILNTTEPHYIRCIKPNNLLKPGIFENQNVLQQLRCGGVMEAIRIS 663
Query: 663 LAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQ 722
AGYPTR+ + EF++RFG+IA + +D + ++ A +K+L K LE +Q+G++KVFLRAGQ
Sbjct: 664 CAGYPTRKHFDEFLNRFGIIAPQVLDKNSNEPAACKKLLDKAGLEGYQIGKSKVFLRAGQ 723
Query: 723 IGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRE 782
+ LD+RR E+L +A IQR++R+++A++ FI +R +A +QA CRGY+ + +Y R
Sbjct: 724 MADLDTRRTEILGRSASIIQRKVRSYLAQKTFIQLRISATQIQAVCRGYLARSIYEGMRR 783
Query: 783 TAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWR 842
AAA+ +Q+ +R +L RK Y +L S+ +IQ+ +RG ++R+ ++ +AAT IQ R
Sbjct: 784 EAAALKIQRDLRKFLARKAYTELFSATILIQAGMRGMVSRKELCLRRQTKAATIIQTRCR 843
Query: 843 MYKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELTW 902
+Y R + + + + + QC WR + A++EL+ LK A ETGAL+ AK+KLEKQ++ELTW
Sbjct: 844 VYLARLHYRKLKKAAITTQCAWRGKVARKELKNLKMAARETGALQEAKNKLEKQVEELTW 903
Query: 903 RLHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIK 962
RL LEK++R E+AK+ E +K + +E + + + I E + + IK
Sbjct: 904 RLQLEKRMRTDLEEAKKQENAKYESSLEEIQNKFKETEALLIKEREAAKTVSEVLPI-IK 962
Query: 963 EKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFEQK 1022
E + +EL ++++ EN LK + + E K EL + + +++ E K
Sbjct: 963 EVPVVDQEL--MEKLTNENEKLKGMVSSLEIKIDETAKELHETARISQDRLKQALAAESK 1020
Query: 1023 CSQLEQNVKSLEEKMLSLEDENHV-LRQKALSAPLKSNRQGFAKSLSERYSNAVASRTER 1081
++L+ ++ LEEK+ +E E + L+Q L+ P+KS VA
Sbjct: 1021 VAKLKTAMQRLEEKISDMETEKQIMLQQTILNTPVKS----------------VAGHPPT 1064
Query: 1082 KPI--FESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAA 1139
I E+ T L F + K AER +N + L C+KEN+GF NGKP+AA
Sbjct: 1065 ATIKNLENGHRTNLENQFNEVEVNGNAGKSAAERQLENVDTLIDCVKENIGFSNGKPIAA 1124
Query: 1140 PIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQRNLR 1199
IYKCLLHW FESE+T+ FD +IE I ++ DD+ L YWL+NTSALL LLQ++L+
Sbjct: 1125 FTIYKCLLHWKCFESEKTSAFDRLIEMIGSAIENEDDNGHLAYWLTNTSALLFLLQKSLK 1184
Query: 1200 SNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEARYP 1259
G T ++ + L R ++ +S N + ++ + + VEA+YP
Sbjct: 1185 PAGAGATASKKPPITTSLFGR---MALSFRSSPNLAAAAEAAALAV-----IRPVEAKYP 1236
Query: 1260 AILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQQPS 1319
A+LFKQQL A VEKIFG++RD+LKKELS L+ +CIQAP+ + G + RS L +
Sbjct: 1237 ALLFKQQLAAYVEKIFGMIRDNLKKELSALISMCIQAPRISK---GGIQRSARSLGKDSP 1293
Query: 1320 GGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFSNGE 1379
W +I++ L+SL++ L N+VP I+K+ TQ FSF+N+ LFNSLLLR+ECCTFSNGE
Sbjct: 1294 AIHWQSIIDGLNSLLAILKDNYVPLVLIQKIHTQTFSFVNVQLFNSLLLRKECCTFSNGE 1353
Query: 1380 YMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALT 1439
++KSGLAELE W EYAG SW EL +IRQAVGFLVIHQK + S D+I DLCP L+
Sbjct: 1354 FVKSGLAELELW-CGQVNEYAGPSWDELKHIRQAVGFLVIHQKYRVSYDDIVHDLCPILS 1412
Query: 1440 VRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDD 1490
V+Q+YRI T+YWDD Y T+SVS EV+S MR +++ + N + SNSFLLDD+
Sbjct: 1413 VQQLYRICTLYWDDCYNTRSVSQEVISSMRALMTEESN-DADSNSFLLDDN 1462
>At2g31900 putative unconventional myosin
Length = 1490
Score = 1446 bits (3744), Expect = 0.0
Identities = 760/1478 (51%), Positives = 1011/1478 (67%), Gaps = 60/1478 (4%)
Query: 66 MTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPLG 125
MT+LAYL+EPGVL+NL R+ LN+IYTYTG+ILIAVNPF +LPHLY HMMEQYKGA G
Sbjct: 1 MTKLAYLHEPGVLHNLDCRFALNEIYTYTGNILIAVNPFQRLPHLYSVHMMEQYKGAAFG 60
Query: 126 ELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDRS 185
ELSPH+FAVAD SYRAM+NE +SQSILVSGESGAGKTETTK++M+YL F+GGR+ + RS
Sbjct: 61 ELSPHLFAVADTSYRAMINEARSQSILVSGESGAGKTETTKMLMRYLAFMGGRSDTEGRS 120
Query: 186 VEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVVQ 245
VEQQVLESNP+LEAFGNA+TV+N+NSSRFGKFVEIQFD G+ISGAAIRTYLLERSRV Q
Sbjct: 121 VEQQVLESNPVLEAFGNAKTVKNNNSSRFGKFVEIQFDKRGKISGAAIRTYLLERSRVCQ 180
Query: 246 ITDPERNYHCFYQLCAFETD-AEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRAM 304
++DPERNYHCFY LCA + A+K+++G P FHYLNQ+ YE+ V + EY+ TR AM
Sbjct: 181 VSDPERNYHCFYMLCAAPPEEAKKFKVGDPRTFHYLNQTNCYEVSNVDDAREYLETRNAM 240
Query: 305 NIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDVD 364
+IVGI E Q+AIFR +AAILHLGN+ F G+E DSS ++D+KSR+H+Q AA L MC+
Sbjct: 241 DIVGIGQEAQDAIFRVVAAILHLGNVNFIKGEEADSSKLRDDKSRYHLQTAAELLMCNEK 300
Query: 365 LLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDINS 424
++ +LC R I T +G+I K LD +A + RD LAKTVY+RLFDW+VDKIN S+GQD ++
Sbjct: 301 MMEDSLCKRVIVTPDGNITKPLDPESAASNRDALAKTVYSRLFDWIVDKINSSIGQDPDA 360
Query: 425 QMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIEF 484
+ IGVLDIYGFE FK NSFEQ CIN NEKLQQHFN+HVFKMEQEEY REEINWSY+EF
Sbjct: 361 KSLIGVLDIYGFESFKINSFEQLCINLTNEKLQQHFNQHVFKMEQEEYTREEINWSYVEF 420
Query: 485 VDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFA 544
VDNQDVL+LIEKKP GI+ALLDEACMFPKSTHETF+ K++Q + H R K K +QT F
Sbjct: 421 VDNQDVLDLIEKKPGGIIALLDEACMFPKSTHETFAQKMYQTYKGHKRFSKPKLAQTAFT 480
Query: 545 ISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSSV 604
++HYAG VTY + FLDKN+DYVV EH LL +SKC FV+ LFP PE++S+ S +FSS+
Sbjct: 481 VNHYAGDVTYSAEQFLDKNKDYVVAEHQALLDASKCSFVANLFPPLPEDASKQS-KFSSI 539
Query: 605 ASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISLA 664
+RFKQQLQALMETLN+TEPHY+RCVKPN++ +P +FEN +V++QLRCGGVLEA+RIS A
Sbjct: 540 GTRFKQQLQALMETLNTTEPHYIRCVKPNAVLKPGIFENDNVLNQLRCGGVLEAIRISCA 599
Query: 665 GYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQIG 724
GYPT+R + EF+DRF ++A + +GS D+K+ I K+ L+ +Q+G+TK+FLRAGQ+
Sbjct: 600 GYPTKRAFDEFLDRFVMLATDVPEGS-DEKSACASICNKMGLKGYQIGKTKIFLRAGQMA 658
Query: 725 ILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRETA 784
LD+RR EVL A + IQRQ+RT++ R+ F+ + A + +Q R + +K+Y + R A
Sbjct: 659 ELDARRTEVLAGATKLIQRQIRTYLTRKEFLGQKRATIYMQKLWRAKLARKLYQNMRREA 718
Query: 785 AAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRMY 844
A+I +QK IR RK Y KL +SAT+IQ+ +R R + H + +AA IQ WR +
Sbjct: 719 ASICIQKNIRAHRARKNYTKLQASATVIQTGLRTMSARNKHRHRRRTKAAIIIQREWRRH 778
Query: 845 KFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELTWRL 904
+ A+ +H+ + +A+QCLWR + A++EL+ L+ A ETGAL+ AK KLEK+++ELTWRL
Sbjct: 779 QVHEAYKKHKKATLALQCLWRAKVARKELKNLRMAARETGALKEAKDKLEKRVEELTWRL 838
Query: 905 HLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIKEK 964
LEK + EDAK EI+KLQ + L +LD A A I + + +L+I++
Sbjct: 839 ELEKNQKADLEDAKAQEIAKLQNNLTELQEKLDEAYAAIIRD-------KEAAKLAIEQA 891
Query: 965 SALKRELVAVD------------EIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDET 1012
+ +E+ VD E+ E A LK + FE KC ALE +
Sbjct: 892 PPIIKEVPVVDNTQLELLNSQNNELEVEVAKLKGKIKEFEVKCFALE-------NDSRAS 944
Query: 1013 IEKLREFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERY- 1071
+ + + + K + ++ ++ L + +LE EN VLRQ+AL+A G SL ++
Sbjct: 945 VTEAEDAKSKAVEFQEIIERLHTNLSNLESENQVLRQQALAASTSVEEIGELNSLKDKVA 1004
Query: 1072 ---SNAVASRTERKPIFESPTPTKLIPTFTPGMSDSRRSKLTA----------------- 1111
S R + + ++ P ++ + ++ + ++ A
Sbjct: 1005 ILESENETLRRQTESAEKTMPPARVFASEKNLENEHQTKEIQATKEPRNPINVLAKQGSL 1064
Query: 1112 -ERHQDNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEV 1170
+R Q++ E L +C+ + F N K +AA I+YK LL W FE+E+T IFD I+ I
Sbjct: 1065 TDRQQESHEVLMKCLTDERRFDNEKSVAAWIVYKALLQWRLFEAEKTNIFDRIVHKIRSS 1124
Query: 1171 LKARDDDDVLPYWLSNTSALLCLLQRNLRSNGFLTTTGQRYTGS-AGLASRTVHVSYLVS 1229
++ +DD L YWL+ +S LL LLQ L+ + +R S A L R V S
Sbjct: 1125 IEGQDDTRELAYWLTTSSTLLYLLQSTLKFSNTNNAASRRNRSSHATLFGRLVQGMQPSS 1184
Query: 1230 TSINHSCGPKSPLKFIGYDDGVSHVEARYPAILFKQQLTACVEKIFGLLRDDLKKELSPL 1289
+ S G G + VEA+YPA+LFKQ L A VEK +G++RD LKKE++PL
Sbjct: 1185 VGLETSSGYSG---MAGIPNDQQMVEAKYPALLFKQHLAAYVEKTYGMIRDKLKKEINPL 1241
Query: 1290 LGLCIQAPKTGR----QHGGKLSRSPSGLPQQPSGGQWANIVNFLDSLMSKLHGNHVPSF 1345
L LCI AP+ R + K + QQ S QW NIVN L+ ++ + NHVPS
Sbjct: 1242 LNLCIHAPRPTRAKTLRDVTKSIHLTTIAKQQASYVQWQNIVNKLEHTLTFMAENHVPSM 1301
Query: 1346 FIRKLVTQVFSFINITLFNSLLLRRECCTFSNGEYMKSGLAELEKWIVNAKEEYAGTSWH 1405
RKL QVFS+IN+ LFNSLLLRRECC+ SNGEY+K GL ELE+W + A +E + W
Sbjct: 1302 ITRKLFHQVFSYINVQLFNSLLLRRECCSVSNGEYLKMGLHELEQWCLKADDEATRSPWD 1361
Query: 1406 ELNYIRQAVGFLVIHQKRKKSLDEIRQDLCPALTVRQIYRISTMYWDDKYGTQSVSNEVV 1465
EL +IRQAV FLV HQK +KSLDEI +++CP L++ Q+YRI TM+WDDKYGTQ +S EV+
Sbjct: 1362 ELQHIRQAVMFLVSHQKTQKSLDEIAKEICPVLSIPQVYRIGTMFWDDKYGTQGLSPEVI 1421
Query: 1466 SEMREIVSSKDNQNITSNSFLLDDDMSIPFSAEDIDMA 1503
++MR+++ ++D+ N+T SFLLD D SIPFS ED+ +
Sbjct: 1422 NQMRKLM-TEDSANMTYPSFLLDVDSSIPFSVEDVSQS 1458
>At4g28710 myosin heavy chain - like protein (fragment)
Length = 1446
Score = 1412 bits (3654), Expect = 0.0
Identities = 752/1531 (49%), Positives = 1013/1531 (66%), Gaps = 95/1531 (6%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VWV+D + AW+ EV+ G +V+ SGK V + P+D E GV+
Sbjct: 2 VGSCVWVEDPEVAWIDGEVIEVKGSDIKVKCT--SGKTVCFTISSAYPKDV-EAPASGVD 58
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMTRLAYL+EPGVL N+K R+ +N+IYTYTG+ILIAVNPF +LPHLY+NHMM+QYKGA
Sbjct: 59 DMTRLAYLHEPGVLQNMKSRFDINEIYTYTGNILIAVNPFRRLPHLYNNHMMQQYKGAGF 118
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDR 184
GELSPH FAVADA+YR M N+G SQSILVSGESGAGKTETTKL+MQYL +GGRA + R
Sbjct: 119 GELSPHPFAVADAAYRQMKNQGISQSILVSGESGAGKTETTKLLMQYLADMGGRAVSEGR 178
Query: 185 SVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVV 244
+VE++VLESNP+LEAFGNA+TVRN+NSSRFGKFVEIQFD GRISGAAIRTYLLERSRV
Sbjct: 179 TVEKKVLESNPVLEAFGNAKTVRNNNSSRFGKFVEIQFDQRGRISGAAIRTYLLERSRVC 238
Query: 245 QITDPERNYHCFYQLCAFET-DAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRA 303
Q++DPERNYHCFY LCA D +K++L P FHYLNQS+ EL+ + + +EY TR+A
Sbjct: 239 QVSDPERNYHCFYMLCAAPPEDIKKWKLADPRKFHYLNQSQCIELERMDDAKEYRETRKA 298
Query: 304 MNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDV 363
M++VGI+ E+QEAIF+ +AAILHLGN+EF GKE DSS KD+ S +H++ AA LFMCD
Sbjct: 299 MDVVGINSEEQEAIFQVVAAILHLGNVEFGKGKEADSSAPKDDTSNYHLKTAAELFMCDE 358
Query: 364 DLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDIN 423
L +LC R I TR +I K LD +A RD LAKTVY+RLFDW+V+KIN S+GQD +
Sbjct: 359 QALEDSLCKRVIVTRGETITKCLDQESAALSRDALAKTVYSRLFDWIVNKINDSIGQDPD 418
Query: 424 SQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIE 483
S+ IGVLDIYGFE FK NSFEQFCIN NEKLQQHFN+HVFKMEQ+EY +EEI+WSYIE
Sbjct: 419 SEYLIGVLDIYGFESFKTNSFEQFCINLTNEKLQQHFNQHVFKMEQDEYNKEEIDWSYIE 478
Query: 484 FVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDF 543
FVDNQ++L+LIEKK GI++LL+EACMFP++THETF+ K++Q F H K K S+TDF
Sbjct: 479 FVDNQEILDLIEKKAGGIISLLNEACMFPRATHETFAEKMYQTFKDHKHFSKPKLSRTDF 538
Query: 544 AISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSS 603
I HYAG VTY T+ FL+KN+DYVV EH LL++S+C FV+ LFPL E++++ S +FSS
Sbjct: 539 TICHYAGDVTYQTEQFLEKNKDYVVAEHQTLLNASRCAFVASLFPLLAEDANKKS-KFSS 597
Query: 604 VASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISL 663
++SRFKQQL L+ETL++TEPHY+RCVKPN+L +P +FEN +V+ QLRCGGV+EA+RIS
Sbjct: 598 ISSRFKQQLVTLLETLSTTEPHYIRCVKPNNLLKPLIFENQNVLQQLRCGGVMEAIRISC 657
Query: 664 AGYPTRRTYSEFVDRFGLIALEFMDGSYD-------DKAVAEKILQKLKLENFQLGRTKV 716
AG+PTR+ + EF++RF ++A E +D S D D +K+L+K+ L+ +Q+G+TKV
Sbjct: 658 AGFPTRKKFEEFLERFSVLAPEVLDKSTDGWPLSSTDDVACKKLLEKVALQGYQIGKTKV 717
Query: 717 FLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKM 776
FLRAGQ+ LD+RR EVL AA IQR+ R++++R+ F+ +R A +QA CRG + + +
Sbjct: 718 FLRAGQMADLDARRNEVLGRAASRIQRKFRSYLSRKTFLMLRKVATNMQAVCRGQLSRLI 777
Query: 777 YASKRETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATS 836
+ R AA + +Q+ IRM L RK+Y +L +A IQ +RG +R R ++ +AA
Sbjct: 778 FEGLRRDAAVLEIQRDIRMHLARKSYKELYFAAVSIQLGIRGMASRGRLRFQRQDKAAIM 837
Query: 837 IQACWRMYKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQ 896
IQ+ R + + + R + + + Q WR R A++ELR+LK A ETG L AKSKLEKQ
Sbjct: 838 IQSHCRKFLAQLHYQRLKKAAITTQSAWRARLARKELRKLKMAAKETGVLEAAKSKLEKQ 897
Query: 897 LDELTWRLHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQ 956
++ELTW+L LEK++R E++K E +KL+ +E + L+ K + E +
Sbjct: 898 VEELTWKLQLEKRMRTDMEESKTQENAKLRSALEEMQLQFKETKALHLQEVEAAKKMAET 957
Query: 957 FELSIKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKL 1016
+ ++E + ELV +++ EN LK + + ++K E + K +E +++
Sbjct: 958 VPV-LQEVPVVDTELV--EKLTSENEKLKSLVSSLDQKIDETEKKFEERSKINEERLKQA 1014
Query: 1017 REFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVA 1076
E E L+ V L+EK+L +E EN +LRQK+ +
Sbjct: 1015 IEAETTIVNLKTAVHELQEKILDVESENKILRQKS-----------------------LI 1051
Query: 1077 SRTERKPIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKP 1136
+ P PTP N L C+ N+GF GKP
Sbjct: 1052 QASGHLP----PTP--------------------------NIGALINCVVNNIGFNQGKP 1081
Query: 1137 LAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSALLCLLQR 1196
+AA IYKCLLHW +FE+ERT++FD +++ I +K D++ L YWLSNTS LL ++Q+
Sbjct: 1082 VAAFTIYKCLLHWKSFEAERTSVFDRLVQMIGSAIKDEGDNEHLAYWLSNTSTLLFMIQQ 1141
Query: 1197 NLRSNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEA 1256
+L+ T Q+ S L R +S S ++ + + V A
Sbjct: 1142 SLKPGA---TPQQKTPVSTSLFGRMAMGFRSAPSSAETSAAAEAAAAAV-----IRPVVA 1193
Query: 1257 RYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQ 1316
+ PA+LFKQQLTA VEKIFG++RD+LK EL LL LCIQAP+T + RS +
Sbjct: 1194 KDPALLFKQQLTAYVEKIFGMIRDNLKNELQTLLSLCIQAPRTSTGRSLRSFRSSKTMRN 1253
Query: 1317 QPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFS 1376
W I + L++++S L N VP I+ + Q FSFIN+ LFNSLLLRRECCTFS
Sbjct: 1254 NSPLDHWNGIYDGLNAILSTLQENFVPPVLIQNIFIQTFSFINVQLFNSLLLRRECCTFS 1313
Query: 1377 NGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCP 1436
NGE+ YAG+SW EL +IRQAVGF+VIH+K + S D+I DLCP
Sbjct: 1314 NGEF------------------YAGSSWDELKHIRQAVGFMVIHKKYRISYDDIAHDLCP 1355
Query: 1437 ALTVRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFS 1496
L+V+Q+YRI T+YWDD Y T+SVS +V++ MR ++ ++D+ N S++FLLD+D SIPFS
Sbjct: 1356 ILSVQQLYRICTLYWDDSYNTRSVSQDVIANMR-VLMTEDSNNADSSAFLLDEDSSIPFS 1414
Query: 1497 AEDIDMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
A+D+ ++ D E+ + E FL
Sbjct: 1415 ADDLSSSMKEKDFAEMKPAEELEENPAFSFL 1445
>At2g20290 putative myosin heavy chain
Length = 1502
Score = 1333 bits (3450), Expect = 0.0
Identities = 736/1538 (47%), Positives = 1017/1538 (65%), Gaps = 79/1538 (5%)
Query: 4 RMGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGV 63
++GS VWVQD ++AW+ EV+ +G +VQ SGK V+A P+D E GV
Sbjct: 18 KVGSIVWVQDPEEAWIDGEVVEVNGEDIKVQCT--SGKTVVAKGSNTYPKDM-EVPPSGV 74
Query: 64 EDMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAP 123
+DMT LAYL+EPGVL NLK RY +++IYTYTG+ILIAVNPF +LP+LY++HMM QYKGA
Sbjct: 75 DDMTTLAYLHEPGVLQNLKSRYYIDEIYTYTGNILIAVNPFKQLPNLYNDHMMAQYKGAA 134
Query: 124 LGELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDD 183
LGELSPH FAVADA+YR M+NEG SQSILVSGESGAGKTET K++M+YL +GGRA D
Sbjct: 135 LGELSPHPFAVADAAYRQMINEGISQSILVSGESGAGKTETAKMLMKYLAKMGGRAVSDR 194
Query: 184 RSVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRV 243
R+VE QVLESNP+LEAFGNA+TV+N+NSSRFGKFVEIQFD GRISGAAIRTYLLERSRV
Sbjct: 195 RTVEDQVLESNPVLEAFGNAKTVKNNNSSRFGKFVEIQFDQRGRISGAAIRTYLLERSRV 254
Query: 244 VQITDPERNYHCFYQLCAF-ETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRR 302
Q++DPERNYHCFY LCA D K +L P+ F YLNQS +LDGV + +EY +TR
Sbjct: 255 CQVSDPERNYHCFYMLCAAPPEDKRKLKLNDPTEFRYLNQSHCIKLDGVDDSKEYTKTRE 314
Query: 303 AMNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCD 362
AM IVGI+ E+QEAIFR +AAILHLGNIEF+ G+E DSSV DE S+ ++++AA LFMCD
Sbjct: 315 AMGIVGINLEEQEAIFRVVAAILHLGNIEFAIGEEPDSSVPTDE-SKKYLKIAAELFMCD 373
Query: 363 VDLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDI 422
L +LC R + T E +I + LD N+A RD LAK VY+RLFDW+V+KIN S+GQD
Sbjct: 374 EQALEDSLCKRIMVTPEETISRCLDPNSAALSRDALAKFVYSRLFDWIVNKINNSIGQDP 433
Query: 423 NSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYI 482
+S+ IGVLDIYGFE FK NSFEQFCIN NEKLQQHF +HV KMEQEEY +EEI WS I
Sbjct: 434 DSKDMIGVLDIYGFESFKTNSFEQFCINLTNEKLQQHFTQHVLKMEQEEYTKEEIEWSQI 493
Query: 483 EFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTD 542
F DN+ VLELIEKK GI+ALLDEACMFP+STH+TFS KL++ + K K S+TD
Sbjct: 494 TFPDNRYVLELIEKKRGGIIALLDEACMFPRSTHKTFSQKLYETLKDNKYFSKPKLSRTD 553
Query: 543 FAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFS 602
F I HYAG VTY T+ FL+KN+DYVV EH LL +S+C F++GLFP E++++ S +FS
Sbjct: 554 FTICHYAGDVTYQTEQFLEKNKDYVVAEHQALLGASRCTFIAGLFPPLVEDANKQS-KFS 612
Query: 603 SVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRIS 662
S+AS+FKQQL +L+E LN+TEPHY+RCVKPN+L +P +FEN + + QLRCGGV+E +R+
Sbjct: 613 SIASQFKQQLASLIEGLNTTEPHYIRCVKPNNLLKPSIFENQNSLQQLRCGGVMETIRVC 672
Query: 663 LAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQ 722
AGYPTR+ + EF+DRFG++ +D S D+KA +K+L+ + L FQ+G+TKVFL+AGQ
Sbjct: 673 RAGYPTRKHFDEFLDRFGILDSATLDKSSDEKAACKKLLETVGLNGFQIGKTKVFLKAGQ 732
Query: 723 IGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRE 782
+ LD RR EVL AA IQ + R+++ R+ FI +R AA+ +QA RG + + + + R
Sbjct: 733 MAELDDRRTEVLGRAACIIQWKFRSYLTRQSFIMLRNAAINIQAVYRGQVARYRFENLRR 792
Query: 783 TAAAISVQKYIRMWL-RRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACW 841
AAA+ +Q+ +R+ L R+++Y++ + +QS +RG R + ++ +A T IQ+
Sbjct: 793 EAAALKIQRALRIHLDRKRSYIE---AVVTVQSGLRGMAA--RVVLRRKTKATTVIQSHC 847
Query: 842 RMYKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELT 901
R + + + + + + Q WR R A++ELR+LK +A +T L+ AKS L ++++ELT
Sbjct: 848 RRLRAELHYKKLKKAAITTQSAWRARLARKELRKLKTDARDTVVLQAAKSMLAEKVEELT 907
Query: 902 WRLHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINE---CNKNAVLQNQFE 958
WRL LEK++RV E +K E +KLQ +E + L+ + K++ + E K A +
Sbjct: 908 WRLDLEKRMRVDMEVSKAQENAKLQLALEEIQLQFEETKVSLLKEVEAAKKTAAIVP--- 964
Query: 959 LSIKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLRE 1018
+KE + + V ++++ EN LK + + E K E + +K +E ++K +
Sbjct: 965 -VVKEVPVV--DTVLMEKLTSENEKLKSLVTSLELKIDETEKKFEETKKISEERLKKALD 1021
Query: 1019 FEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAV-AS 1077
E K L+ + +LEEK+ ++ EN+ L++ L+ P+K+ F + + N + S
Sbjct: 1022 AENKIDNLKTAMHNLEEKLKEVKLENNFLKESVLTTPVKTASGRFLSTPLKNLQNGLFTS 1081
Query: 1078 RTERKPIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSR--------CIKENL 1129
+ E TP ++ + G R +H+D FL + + +N+
Sbjct: 1082 EESQLSGAEFTTPPRIQES---GSDTKSRGSHIDPQHRDLLGFLEKEDVDALINSVTKNV 1138
Query: 1130 GFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARDDDDVLPYWLSNTSA 1189
GF GKP+AA IYKCLLHW +FE+ERT +FD +++ I +K D+D L YWLSNTS
Sbjct: 1139 GFSQGKPVAAFTIYKCLLHWKSFEAERTNVFDRLVQMIGSAIKDEDNDANLAYWLSNTST 1198
Query: 1190 LLCLLQRNLRSNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDD 1249
LL +LQ++L+S G TG+ L V ++ G +SP +
Sbjct: 1199 LLFMLQQSLKSGG---------TGATPLRQSPSLVRWMTK-------GFRSPAA-----E 1237
Query: 1250 GVSHVEARYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSR 1309
+ V+A+ PA+ FKQQL A VEKI G++ D+LKKEL+ +L LCIQAPKT +
Sbjct: 1238 AIRPVDAKDPALHFKQQLEAYVEKILGIIWDNLKKELNTVLALCIQAPKTFK-------- 1289
Query: 1310 SPSGLPQQPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLR 1369
+ L + W +I+ LD+L+S L + VP I+K+ +Q FS IN+ + NSL+ R
Sbjct: 1290 -GNALISITTANYWQDIIEGLDALLSTLKESFVPPVLIQKIFSQAFSLINVQVCNSLVTR 1348
Query: 1370 RECCTFSNGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDE 1429
+ C+F NGEY+KSGL +LEKW KEEYAG+SW EL + RQAVGFL+IH+K S DE
Sbjct: 1349 PDNCSFINGEYLKSGLEKLEKWCCETKEEYAGSSWDELKHTRQAVGFLLIHKKYNISYDE 1408
Query: 1430 IRQDLCPALTVRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDD 1489
I DLCP L ++Q +++ T+Y D+ Y T+SVS +V++ M +++ S+ FLL +
Sbjct: 1409 IANDLCPNLQIQQHFKLCTLYKDEIYNTKSVSQDVIASMTGVMTD-------SSDFLLKE 1461
Query: 1490 DMS--IPFSAEDI-----DMAIPAIDPDE--IDLPAFV 1518
D S I S +D+ D + P E ++ P+F+
Sbjct: 1462 DSSNIISLSIDDLCSSMQDKDFAQVKPAEELLENPSFI 1499
>At2g33240 putative myosin heavy chain
Length = 1611
Score = 1320 bits (3417), Expect = 0.0
Identities = 748/1632 (45%), Positives = 1030/1632 (62%), Gaps = 177/1632 (10%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGK-------------KVLASPEKLC 51
+GS+VWV+D D+AW+ EV+ ++G +V T + KV+A +
Sbjct: 8 VGSQVWVEDPDEAWLDGEVVEANGQEIKVNCQTKTVSPFSPKQRDNVLVLKVVAKVNAVH 67
Query: 52 PRDADEEEHGGVEDMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLY 111
P+D + E G V+DMT+LAYL+EPGVL NLK RY N+IYTYTG+ILIAVNPF +LPHLY
Sbjct: 68 PKDPEFPELG-VDDMTKLAYLHEPGVLLNLKARYNANEIYTYTGNILIAVNPFKRLPHLY 126
Query: 112 DNHMMEQYKGAPLGELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQY 171
N +MEQYKG GELSPH FAVAD++YR M+NEG SQ+ILVSGESGAGKTE+TK++MQY
Sbjct: 127 GNEIMEQYKGTDFGELSPHPFAVADSAYRKMINEGVSQAILVSGESGAGKTESTKMLMQY 186
Query: 172 LTFVGGRAAGDDRSVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGA 231
L ++GG+A + RSVEQQVLESNP+LEAFGNA+TVRN+NSSRFGKFVEIQF+ GRISGA
Sbjct: 187 LAYMGGKAESEGRSVEQQVLESNPVLEAFGNAKTVRNNNSSRFGKFVEIQFNHMGRISGA 246
Query: 232 AIRTYLLERSRVVQITDPERNYHCFYQLCAF-ETDAEKYELGHPSHFHYLNQSKIYELDG 290
AIRTYLLERSRV Q++DPERNYHCFY LCA E + E+Y+LG PS FHYLNQS + LD
Sbjct: 247 AIRTYLLERSRVCQVSDPERNYHCFYMLCAAPEQETERYQLGKPSTFHYLNQSNCHALDA 306
Query: 291 VSNVEEYVRTRRAMNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRF 350
+ + +EY+ TR+AM++VGIS E+Q+AIFR +AAILHLGNIEF+ +E D + KD+KSRF
Sbjct: 307 IDDSKEYLATRKAMDVVGISPEEQDAIFRVVAAILHLGNIEFAKSEESDGAEPKDDKSRF 366
Query: 351 HMQMAANLFMCDVDLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWL 410
H+++AA LFMCD L ++LC R + TR SI K LD +A RD LAK VY++LFDWL
Sbjct: 367 HLKVAAKLFMCDEKALENSLCNRVMVTRGESITKPLDPGSAALSRDALAKIVYSKLFDWL 426
Query: 411 VDKINRSVGQDINSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQE 470
V KIN S+GQD +S+ IGVLDIYGFE FK NSFEQFCIN NEKLQQHFN+HVFKMEQE
Sbjct: 427 VTKINNSIGQDSSSKYIIGVLDIYGFESFKTNSFEQFCINLTNEKLQQHFNQHVFKMEQE 486
Query: 471 EYGREEINWSYIEFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSH 530
EY +EEI+WSYIEF+DNQDVL+LIEKKP GI+ALLDEACMFP+STH+T + KL+Q F SH
Sbjct: 487 EYTKEEIDWSYIEFIDNQDVLDLIEKKPGGIIALLDEACMFPRSTHDTLAEKLYQTFGSH 546
Query: 531 PRLGKEKFSQTDFAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLP 590
R K K ++TDF I HYAG VTY T+ FLDKN+DYVV EH +L++SS C FVS LFP
Sbjct: 547 KRFTKPKLARTDFTICHYAGDVTYQTELFLDKNKDYVVGEHQSLMNSSDCSFVSSLFPKS 606
Query: 591 PEESSRSSYRFSSVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQL 650
EESS+SS +FSS+ S+FKQQLQ+L+ETLN+TEPHY+RCVKPN++ +P++FEN +V+HQL
Sbjct: 607 REESSKSS-KFSSIGSQFKQQLQSLLETLNTTEPHYIRCVKPNNVLKPEIFENVNVLHQL 665
Query: 651 RCGGVLEAVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQ 710
RCGGV+EA+RIS AGYPTR+ ++EF+ RF ++A E + S+D+ +K+L ++ L+ FQ
Sbjct: 666 RCGGVMEAIRISCAGYPTRKPFNEFLTRFRILAPEATERSFDEVDACKKLLARVDLKGFQ 725
Query: 711 LGRTKVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRG 770
+G+TKVFLRAGQ+ LD+ RAEVL ++AR IQR++ T+++R+ ++ +++A+ +QA CRG
Sbjct: 726 IGKTKVFLRAGQMAELDAHRAEVLGHSARIIQRKVITYLSRKKYLLLQSASTEIQAFCRG 785
Query: 771 YIGQKMYASKRETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKE 830
+I + + + R AA++ +QK R ++ + + KLC+SA IQS +R R F + +
Sbjct: 786 HIARVQFKATRREAASVRIQKQARTYICQTAFKKLCASAISIQSGLRAMAARVEFQYRTK 845
Query: 831 HRAATSIQACW---------------RMYKFRSAFHRHQASIVAIQCLWRRRQAKRELRR 875
+AA IQA R R + R + + + QC WR + A RELR+
Sbjct: 846 RKAAIIIQASLKPHIDDKDLSFFSQIRRCLCRRRYLRTKKAAITTQCGWRVKVAHRELRK 905
Query: 876 LKQEANETGALRLAKSKLEKQLDELTWRLHLEKKIRVSNED------------------- 916
LK A ETGAL+ AK+KLEK+++ELT L LEK++R+ E
Sbjct: 906 LKMAAKETGALQDAKTKLEKEVEELTSCLELEKQMRMELEQVKTQEVEDLRSALNDMKLQ 965
Query: 917 ------AKQIEISKLQKMIEALNLELD--------AAKLATINECNKNAV--LQNQF--- 957
K EI KLQ ++ + LE + LA NE K+ V LQ +
Sbjct: 966 LGETQVTKSEEILKLQSALQDMQLEFEELAKELEMTNDLAAENEQLKDLVSSLQRKIDES 1025
Query: 958 -----ELSIKEKSALKRELVAVDE-----IRKENAMLKVSLDAFEKKCTALE------VE 1001
E S + +K+E+ +D+ + EN LK + EKK +L+ V+
Sbjct: 1026 DSKYEETSKLSEERVKQEVPVIDQGVIIKLEAENQKLKALVSTLEKKIDSLDRKHDDLVD 1085
Query: 1002 LINAQKGRDETIEKLREFEQKCSQLEQNVKSLEEK-----MLSLEDENHVLRQKALSAPL 1056
L+ ++ DET +K E + C + + V E+K L E V+ + L
Sbjct: 1086 LL--ERKIDETEKKYEEASKLCEERLKQVVDTEKKYEEASRLCEERLKQVVDTETKLIEL 1143
Query: 1057 KSNRQGFAKSLS--ERYSNAVASRTERKPIFESPTPTKLI-----------------PTF 1097
K++ Q + +S E + + R +P K + +F
Sbjct: 1144 KTSMQRLEEKVSDMEAEDKILRQQALRNSASRKMSPQKSLDLFVFMYLFQPVENGHHESF 1203
Query: 1098 TP------GMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHA 1151
P G RRS++ + H + + L +C+ +N+GF +GKP+AA IYKCL+HW
Sbjct: 1204 APIPSRRFGAMSFRRSQIEQQPH-EFVDVLLKCVSKNVGFSHGKPVAAFTIYKCLIHWKL 1262
Query: 1152 FESERTAIFDYIIEGINEVLKAR---------------DDDDVLPYWLSNTSALLCLLQR 1196
FE+E+T++FD I+ ++ +DD L YWL+NTS LL LLQR
Sbjct: 1263 FEAEKTSVFDRIVPIFGSAIEVTWKRFNQYALIYFQNPEDDSNLAYWLTNTSTLLFLLQR 1322
Query: 1197 NLRSNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHVEA 1256
+L+S+ + ++ R G +SP D V V+A
Sbjct: 1323 SLKSHSTTGASPKKPPQPTSFFGRMTQ-------------GFRSPSSASLSGDVVQQVDA 1369
Query: 1257 RYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAPKTGRQHGGKLSRSPSGLPQ 1316
RYPA+LFKQQLTA +E I+G+ ++++K++L+P+L CIQ SP+
Sbjct: 1370 RYPALLFKQQLTAYIETIYGIFQENVKRKLAPVLSSCIQ------------ENSPT---- 1413
Query: 1317 QPSGGQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFINITLFNSLLLRRECCTFS 1376
W +++ L+ L+ L N+ K+ Q F IN+ LFNS LL+RECCTF
Sbjct: 1414 ----ETWQDVIGLLNQLLGTLKKNY-------KIFCQTFQDINVQLFNS-LLQRECCTFI 1461
Query: 1377 NGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCP 1436
G+ + L ELE W A E++ G+SW EL RQA+ LV QK + D++ +LCP
Sbjct: 1462 MGKKVNVWLNELESWCSQATEDFVGSSWDELKNTRQALVLLVTEQKSTITYDDLTTNLCP 1521
Query: 1437 ALTVRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFS 1496
AL+ +Q+YRI T+ D + Q+VS +V+S ++ +V+ +D S SFLLD++ SIPF+
Sbjct: 1522 ALSTQQLYRICTLCKIDDHEDQNVSPDVISNLKLLVTDEDED---SRSFLLDNNSSIPFA 1578
Query: 1497 AEDIDMAIPAID 1508
A++I ++ D
Sbjct: 1579 ADEISNSMQEKD 1590
>At1g04600 putative myosin heavy chain
Length = 1730
Score = 1110 bits (2871), Expect = 0.0
Identities = 604/1178 (51%), Positives = 801/1178 (67%), Gaps = 69/1178 (5%)
Query: 5 MGSKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVE 64
+GS VWV+D D AW+ EV + + V SGK V+A + P+D + E G V+
Sbjct: 9 VGSHVWVEDPDDAWIDGEV---EEVNSEEITVNCSGKTVVAKLNNVYPKDPEFPELG-VD 64
Query: 65 DMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPL 124
DMT+LAYL+EPGVL NLK RY N+IYTYTG+ILIAVNPF +LPHLY + M+QYKG
Sbjct: 65 DMTKLAYLHEPGVLLNLKCRYNANEIYTYTGNILIAVNPFKRLPHLYGSETMKQYKGTAF 124
Query: 125 GELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDR 184
GELSPH FAVAD++YR M+NEG SQ+ILVSGESGAGKTE+TK++MQYL ++GGRA + R
Sbjct: 125 GELSPHPFAVADSAYRKMINEGVSQAILVSGESGAGKTESTKMLMQYLAYMGGRAESEGR 184
Query: 185 SVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVV 244
SVEQQVLESNP+LEAFGNA+TVRN+NSSRFGKFVEIQFD GRISGAAIRTYLLERSRV
Sbjct: 185 SVEQQVLESNPVLEAFGNAKTVRNNNSSRFGKFVEIQFDQRGRISGAAIRTYLLERSRVC 244
Query: 245 QITDPERNYHCFYQLCAF-ETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRA 303
Q++DPERNYHCFY LCA E + E+Y+LG PS F YLNQS Y LDG+ + +EY+ TR+A
Sbjct: 245 QVSDPERNYHCFYMLCAAPEQETERYKLGKPSTFRYLNQSNCYALDGLDDSKEYLATRKA 304
Query: 304 MNIVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDV 363
M++VGI+ E+Q+ IFR +AAILHLGNIEF+ G+E ++S KDEKSRFH+++AA LFMCD
Sbjct: 305 MDVVGINSEEQDGIFRVVAAILHLGNIEFAKGEESEASEPKDEKSRFHLKVAAELFMCDG 364
Query: 364 DLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDIN 423
L +LC R + TR+ SI K+LD ++A GRD LAK VY++LFDWLV KIN S+GQD N
Sbjct: 365 KALEDSLCKRVMVTRDESITKSLDPDSAALGRDALAKIVYSKLFDWLVTKINNSIGQDPN 424
Query: 424 SQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIE 483
S+ IGVLDIYGFE FK NSFEQFCIN NEKLQQHFN+HVFKMEQEEY +EEI+WSYIE
Sbjct: 425 SKHIIGVLDIYGFESFKTNSFEQFCINLTNEKLQQHFNQHVFKMEQEEYTKEEIDWSYIE 484
Query: 484 FVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDF 543
F+DNQDVL+LIEKKP GI+ALLDEACMFP+STH+TF+ KL+Q F +H R GK K +QTDF
Sbjct: 485 FIDNQDVLDLIEKKPGGIIALLDEACMFPRSTHDTFAQKLYQTFKNHKRFGKPKLAQTDF 544
Query: 544 AISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSS 603
I HYAG VTY T+ FLDKN+DYVV EH LLSSS C FVS LFP PEESS++S +FSS
Sbjct: 545 TICHYAGDVTYQTELFLDKNKDYVVGEHQALLSSSDCSFVSSLFPPLPEESSKTS-KFSS 603
Query: 604 VASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISL 663
+ S+FKQQLQ+L+E+L++TEPHY+RCVKPN+L +P +FEN +++HQLRCGGV+EA+RIS
Sbjct: 604 IGSQFKQQLQSLLESLSTTEPHYIRCVKPNNLLKPDIFENINILHQLRCGGVMEAIRISC 663
Query: 664 AGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQI 723
AGYPTR+ ++EF+ RF ++A E SYD+ +K+L K+ L+ FQ+G+TKVFLRAGQ+
Sbjct: 664 AGYPTRKPFNEFLTRFRILAPETTKSSYDEVDACKKLLAKVDLKGFQIGKTKVFLRAGQM 723
Query: 724 GILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRET 783
+D+ RAEVL ++AR IQR + T+ +R+ F+ ++AA+ +QA CRG + + + + R
Sbjct: 724 AEMDAHRAEVLGHSARIIQRNVLTYQSRKKFLLLQAASTEIQALCRGQVARVWFETMRRE 783
Query: 784 AAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRM 843
AA++ +QK R ++ + Y LCSSA IQ+ +R R K+ RA IQ+ R
Sbjct: 784 AASLRIQKQARTYICQNAYKTLCSSACSIQTGMRAKAARIELQLRKKRRATIIIQSQIRR 843
Query: 844 YKFRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDELTWR 903
+ R + + + QC WR + A+RELR LK A ETGAL+ AK+KLE Q++ELT
Sbjct: 844 CLCHQRYVRTKKAAITTQCGWRVKVARRELRNLKMAAKETGALQDAKTKLENQVEELTSN 903
Query: 904 LHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNKNAVLQNQFELSIKE 963
L LEK++R+ E+AK EI LQ ++ + L+L + E + + +L +++
Sbjct: 904 LELEKQMRMEIEEAKSQEIEALQSVLTDIKLQLRDTQETKSKEISDLQSVLTDIKLQLRD 963
Query: 964 ------------KSALKRELVAVDEIRK----------ENAMLKVSLDAFEKKCTAL--- 998
+SAL+ + ++E+ K EN LK S+ + + K
Sbjct: 964 TQETKSKEISDLQSALQDMQLEIEELSKGLEMTNDLAAENEQLKESVSSLQNKIDESERK 1023
Query: 999 --EVELINAQKGRDE-------TIEKLREFEQKCSQLEQNVKSLEEKMLSLE-------- 1041
E+ I+ ++ +DE I KL QK L V S+EEK+ L+
Sbjct: 1024 YEEISKISEERIKDEVPVIDQSAIIKLETENQKLKAL---VSSMEEKIDELDRKHDETSP 1080
Query: 1042 ------------DENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVASRTERKPIFESPT 1089
D V +A + LK+ K ++E +N+ + E K I + +
Sbjct: 1081 NITEKLKEDVSFDYEIVSNLEAENERLKALVGSLEKKINESGNNSTDEQEEGKYILKEES 1140
Query: 1090 PTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKE 1127
T+ D+ R K A+ ++D + +S K+
Sbjct: 1141 LTE------DASIDNERVKKLADENKDLNDLVSSLEKK 1172
Score = 306 bits (785), Expect = 5e-83
Identities = 220/690 (31%), Positives = 344/690 (48%), Gaps = 96/690 (13%)
Query: 882 ETGALRLAKSKLEKQLDELTWRLHLEKKIRVSNEDAKQIEIS-KLQKMIEALNLELDA-- 938
E L+ S +E+++DEL R H E ++ + + + ++ +EA N L A
Sbjct: 1053 ENQKLKALVSSMEEKIDELD-RKHDETSPNITEKLKEDVSFDYEIVSNLEAENERLKALV 1111
Query: 939 -AKLATINECNKNAV-LQNQFELSIKEKSALKRELVAVDEIRK---ENAMLKVSLDAFEK 993
+ INE N+ Q + + +KE+S + + + ++K EN L + + EK
Sbjct: 1112 GSLEKKINESGNNSTDEQEEGKYILKEESLTEDASIDNERVKKLADENKDLNDLVSSLEK 1171
Query: 994 KCTALEVELINAQKGRDETIEKLREFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALS 1053
K E + A + +E +++ + E L+ +++ LEEK+ +E + RQ+AL
Sbjct: 1172 KIDETEKKYEEASRLCEERLKQALDAETGLIDLKTSMQRLEEKVSDMETAEQIRRQQAL- 1230
Query: 1054 APLKSNRQGFAKSLSERYSNAVASRTERKPIFES-PTPTKLIPTFTPGMSDSRRSKLTAE 1112
S S R S V S T P+ P IP+ G RRS++ +
Sbjct: 1231 ----------VNSASRRMSPQV-SFTGAPPLENGHQEPLAPIPSRRFGTESFRRSRIERQ 1279
Query: 1113 RHQDNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLK 1172
H+ + L +C+ +N+GF +GKP+AA IYKCL+ W FE+E+T+IFD I+ ++
Sbjct: 1280 PHEF-VDVLLKCVSKNIGFSHGKPVAALTIYKCLMRWKIFEAEKTSIFDRIVPVFGSAIE 1338
Query: 1173 ARDDDDVLPYWLSNTSALLCLLQRNLR---SNGFLTTTGQRYTGSAGLASRTVHVSYLVS 1229
++DD+ L YWL+NTS LL LLQR+LR S G T + T G ++ + +
Sbjct: 1339 NQEDDNHLAYWLTNTSTLLFLLQRSLRQQSSTGSSPTKPPQPTSFFGRMTQ----GFRST 1394
Query: 1230 TSINHSCGPKSPLKFIGYDDGVSHVEARYPAILFKQQLTACVEKIFGLLRDDLKKELSPL 1289
+S N S D V V+ARYPA+LFKQQLTA VE ++G++R+++K+E+S L
Sbjct: 1395 SSPNLS------------TDVVQQVDARYPALLFKQQLTAYVETMYGIIRENVKREVSSL 1442
Query: 1290 LGLCIQA----------------------PKTGRQHGGKLSRSPSGLPQQPSG------- 1320
L CIQ+ P + S P++ SG
Sbjct: 1443 LSSCIQSLKESSCDSSVVNSPSKSSEENLPAKSSEENSPKKSSEENSPKESSGDKSPQKL 1502
Query: 1321 ----------------------GQWANIVNFLDSLMSKLHGNHVPSFFIRKLVTQVFSFI 1358
W +I+ FL+ ++ N+VP F ++K+ +Q F +I
Sbjct: 1503 SDDNSPSKEGQAVKSSEENSPASSWQSIIEFLNYILITWKKNYVPLFLVQKMFSQTFQYI 1562
Query: 1359 NITLFNSLLLRRECCTFSNGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLV 1418
N+ LFNSLLL RE CT + G +K+GL ELE W A EE+ G+SW EL + RQAV LV
Sbjct: 1563 NVQLFNSLLLEREYCTVNMGIKVKAGLDELESWCSQATEEFVGSSWDELKHTRQAVVLLV 1622
Query: 1419 IHQKRKKSLDEIRQDLCPALTVRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQ 1478
K + D++ +LC L+ Q+YRI T+ D G +VS EV+S ++ +++++D
Sbjct: 1623 TEPKSTITYDDLTINLCSVLSTEQLYRICTLCKDKDDGDHNVSPEVISNLKLLLTNEDE- 1681
Query: 1479 NITSNSFLLDDDMSIPFSAEDIDMAIPAID 1508
S SFLLDDD SIPF ++I + D
Sbjct: 1682 --NSRSFLLDDDSSIPFDTDEISSCMQEKD 1709
>At3g58160 myosin heavy chain MYA3
Length = 1242
Score = 992 bits (2565), Expect = 0.0
Identities = 512/921 (55%), Positives = 667/921 (71%), Gaps = 9/921 (0%)
Query: 7 SKVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVEDM 66
S VWV+D ++AW+ VL G ++ T+ G+ V+A+ +L P+D + G VEDM
Sbjct: 9 SHVWVEDPERAWIDGVVLNIKG--EEAEIKTNDGRDVIANLSRLYPKDTEAPSEG-VEDM 65
Query: 67 TRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPLGE 126
TRL+YL+EP VL NL RY LN+IYTYTG+ILIAVNPF LPHLYD +ME+YK A E
Sbjct: 66 TRLSYLHEPAVLDNLATRYELNEIYTYTGNILIAVNPFQGLPHLYDAEVMEKYKEAYFKE 125
Query: 127 LSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDRSV 186
L+PHVFA+ +YR M+NEG+++ ILVSGESG+GKTETTK++M+YL + GG A + R+V
Sbjct: 126 LNPHVFAIGGIAYREMINEGRNKCILVSGESGSGKTETTKMLMRYLAYFGGHTAVEGRTV 185
Query: 187 EQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVVQI 246
E QVLESNP+LEAFGNA+TV+N+NSSRFGKFVEIQFD GRISGAAIRTYLLERSRV Q+
Sbjct: 186 ENQVLESNPVLEAFGNAKTVKNNNSSRFGKFVEIQFDDVGRISGAAIRTYLLERSRVCQV 245
Query: 247 TDPERNYHCFYQLCAFET-DAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRAMN 305
+DPERNYHCFY LCA D E+++LG P F YLNQS Y+LDGV++ EEY+ TRRAM+
Sbjct: 246 SDPERNYHCFYLLCAAPPEDVERFKLGDPKSFRYLNQSSCYKLDGVNDAEEYLATRRAMD 305
Query: 306 IVGISHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDVDL 365
+VGIS ++Q+AIFR +A+ILHLGNIEFS G++ DSS +KDE+S FH+QM + L MCD
Sbjct: 306 VVGISEKEQDAIFRVVASILHLGNIEFSKGEDADSSSVKDEQSMFHLQMTSELLMCDPHS 365
Query: 366 LLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRSVGQDINSQ 425
L LC R + T E I ++LD A RD LAKT+Y+RLFDWLV+KIN S+GQD +S+
Sbjct: 366 LEDALCKRMMVTPEEVIKRSLDPLGAAVSRDGLAKTIYSRLFDWLVNKINISIGQDSHSR 425
Query: 426 MQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIEFV 485
IGVLDIYGFE FK NSFEQFCIN+ NEKLQQHFN+HVFKMEQ EY +EEI+WSY+EFV
Sbjct: 426 RLIGVLDIYGFESFKTNSFEQFCINYTNEKLQQHFNQHVFKMEQGEYQKEEIDWSYVEFV 485
Query: 486 DNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFAI 545
DN+DV++LIEKKP GI+ALLDEACM PKST ETFS KL+ F H R K K +++DF +
Sbjct: 486 DNKDVVDLIEKKPGGIIALLDEACMLPKSTPETFSEKLYHTFKDHKRFMKPKLTRSDFTL 545
Query: 546 SHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFSSVA 605
HYAG V Y +D FLDKN+DYVV EH +LL++SKC FVSGLFP P+ESS+S +FSS+
Sbjct: 546 VHYAGDVQYQSDQFLDKNKDYVVAEHQDLLNASKCSFVSGLFPPLPKESSKS--KFSSIG 603
Query: 606 SRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLEAVRISLAG 665
+RFK QLQ LMETLNSTEPHY+RCVKPN+L +P +F+N +V+HQLR GGVLEA+R+ AG
Sbjct: 604 ARFKLQLQQLMETLNSTEPHYIRCVKPNNLLQPTVFDNANVLHQLRSGGVLEAIRVKCAG 663
Query: 666 YPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKLENFQLGRTKVFLRAGQIGI 725
YPT RT+ EF++RF ++A E + G Y+ + + IL+K L +Q+G++KVFLRAGQ+
Sbjct: 664 YPTNRTFIEFLNRFLILAPEILKGEYEAEVACKWILEKKGLTGYQIGKSKVFLRAGQMAE 723
Query: 726 LDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQKMYASKRETAA 785
LD+ R VL +AR IQ Q+RT + R F+ +R A+V +QA RG I +K+ R A
Sbjct: 724 LDAHRTRVLGESARMIQGQVRTRLTRERFVLMRRASVNIQANWRGNIARKISKEMRREEA 783
Query: 786 AISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHRAATSIQACWRMYK 845
AI +QK +R + +K Y K SSA +QS VR R F + RAAT IQA WR Y
Sbjct: 784 AIKIQKNLRRQIAKKDYGKTKSSALTLQSGVRTMAARHEFRYKLTTRAATVIQAYWRGYS 843
Query: 846 FRSAFHRHQASIVAIQCLWRRRQAKRELRRLKQEANETGALRLAKSKLEKQLDE---LTW 902
S + + + + + R R A+++L + KQ + + K +L + +E +++
Sbjct: 844 AISDYKKLKRVSLLCKSNLRGRIARKQLGQSKQADRKEETEKERKVELSNRAEEAVDMSF 903
Query: 903 RLHLEKKIRVSNEDAKQIEIS 923
LH E+ + ++ ++S
Sbjct: 904 VLHSEQSDDAESGHGRKAKLS 924
>At3g19960 myosin
Length = 1166
Score = 629 bits (1621), Expect = e-180
Identities = 374/947 (39%), Positives = 554/947 (58%), Gaps = 61/947 (6%)
Query: 10 WVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVEDMTRL 69
W+Q + W ++L++ G + + L GK + E L P + D + GV+D+ +L
Sbjct: 118 WIQLPNGNWELGKILSTSGEESVISL--PEGKVIKVISETLVPANPDILD--GVDDLMQL 173
Query: 70 AYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPLGELSP 129
+YLNEP VLYNL RY + IYT G +L+AVNPF ++P LY N +E Y+ SP
Sbjct: 174 SYLNEPSVLYNLNYRYNQDMIYTKAGPVLVAVNPFKEVP-LYGNRYIEAYRKK--SNESP 230
Query: 130 HVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDRSVEQQ 189
HV+A+AD + R M+ + +QSI++SGESGAGKTET K+ MQYL +GG + +E +
Sbjct: 231 HVYAIADTAIREMIRDEVNQSIIISGESGAGKTETAKIAMQYLAALGGGSG-----IEYE 285
Query: 190 VLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVVQITDP 249
+L++NP+LEAFGNA+T+RNDNSSRFGK +EI F +G+ISGA I+T+LLE+SRVVQ +
Sbjct: 286 ILKTNPILEAFGNAKTLRNDNSSRFGKLIEIHFSESGKISGAQIQTFLLEKSRVVQCAEG 345
Query: 250 ERNYHCFYQLCAFETDA--EKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRAMNIV 307
ER+YH FYQLCA + A EK L + YL QS Y ++GV + E + + A++IV
Sbjct: 346 ERSYHIFYQLCAGASPALREKLNLTSAHEYKYLGQSNCYSINGVDDAERFHTVKEALDIV 405
Query: 308 GISHEDQEAIFRTLAAILHLGNIEFSP-GKEYDSSVIKDEKSRFHMQMAANLFMCDVDLL 366
+S EDQE++F LAA+L LGN+ F+ E + DE + A L C+++ L
Sbjct: 406 HVSKEDQESVFAMLAAVLWLGNVSFTVIDNENHVEPVADES----LSTVAKLIGCNINEL 461
Query: 367 LSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRS--VGQDINS 424
TL R+++ R +IV+ L A+ RD LAK++Y+ LFDWLV++IN+S VG+
Sbjct: 462 TLTLSKRNMRVRNDTIVQKLTLPQAIDARDALAKSIYSCLFDWLVEQINKSLAVGKRRTG 521
Query: 425 QMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIEF 484
+ I +LDIYGFE F NSFEQFCIN+ANE+LQQHFN H+FK+EQEEY ++ I+W+ ++F
Sbjct: 522 R-SISILDIYGFESFDKNSFEQFCINYANERLQQHFNRHLFKLEQEEYIQDGIDWTRVDF 580
Query: 485 VDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFA 544
DNQ+ L L EKKP+G+++LLDE FP T T + KL QH S+ +K F
Sbjct: 581 EDNQNCLSLFEKKPLGLLSLLDEESTFPNGTDLTLANKLKQHLQSNSCFRGDKGKL--FT 638
Query: 545 ISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKC----------------PFVSGLFP 588
+ HYAG+VTY T FL+KNRD + + LLSS C P V L+
Sbjct: 639 VVHYAGEVTYETTGFLEKNRDLLHSDSIQLLSSCSCLLPQAFASSMLIQSEKPVVGPLYK 698
Query: 589 LPPEESSRSSYRFSSVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIH 648
+S R SVA++FK QL LM+ L +T PH++RC+KPN++ P ++E G V+
Sbjct: 699 AGGADSQR-----LSVATKFKSQLFQLMQRLGNTTPHFIRCIKPNNIQSPGVYEQGLVLQ 753
Query: 649 QLRCGGVLEAVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKL-- 706
QLRC GVLE VRIS +G+PTR ++ +F R+G + +E + D +V+ IL + +
Sbjct: 754 QLRCCGVLEVVRISRSGFPTRMSHQKFSRRYGFLLVENI-ADRDPLSVSVAILHQFNILP 812
Query: 707 ENFQLGRTKVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQA 766
E +Q+G TK+F R GQIG+L+ R L R +Q R + AR + ++ LQ+
Sbjct: 813 EMYQVGYTKLFFRTGQIGVLEDTRNRTLHGILR-VQSSFRGYQARCLLKELKRGISILQS 871
Query: 767 CCRGYIGQKMYAS-KRETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQ-- 823
RG +K +A +R AA ++Q ++ + R Y + ++ +IQS +RG++ R+
Sbjct: 872 FVRGEKIRKEFAELRRRHKAAATIQSQVKSKIARIQYKGIADASVVIQSAIRGWLVRRCS 931
Query: 824 -RFLHIKEHRAATSIQACWRMYKFRSAFHRHQASIVAIQCLWRRRQAKREL--RRLKQEA 880
+K A T+ + S Q ++ + R ++ + ++ +RL+Q
Sbjct: 932 GDIGWLKSGGAKTN--ELGEVLVKASVLSELQRRVLKAEAALREKEEENDILQQRLQQYE 989
Query: 881 NETGALRLAKSKLE----KQLDELTWRLHLEKKIRVSNEDAKQIEIS 923
N +E KQ+ L L + KK + A+ + S
Sbjct: 990 NRWSEYETKMKSMEEIWQKQMRSLQSSLSIAKKSLAVEDSARNSDAS 1036
>At5g54280 myosin heavy chain
Length = 1111
Score = 596 bits (1536), Expect = e-170
Identities = 353/820 (43%), Positives = 502/820 (61%), Gaps = 47/820 (5%)
Query: 8 KVWVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVEDMT 67
+VW + + W ++ ++ + V L T + KV S E+L P + D E GVED+
Sbjct: 56 RVWCRVSNGQWQLGKIQSTSADTSLVMLSTANVVKV--STEELFPANPDILE--GVEDLI 111
Query: 68 RLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPLGEL 127
+L+YLNEP VLYNL+ RY + IY+ G +LIAVNPF + +Y N ++ Y+ +
Sbjct: 112 QLSYLNEPSVLYNLRVRYLQDVIYSKAGPVLIAVNPFKNV-EIYGNDVISAYQKKVMD-- 168
Query: 128 SPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDRSVE 187
+PHV+AVADA+Y MM E K+QS+++SGESGAGKTET K MQYL +GG + G VE
Sbjct: 169 APHVYAVADAAYDEMMRE-KNQSLIISGESGAGKTETAKFAMQYLAALGGGSCG----VE 223
Query: 188 QQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVVQIT 247
++L++ +LEAFGNA+T RN NSSRFGK +EI F + G+I GA + T+L ++SRVVQ+
Sbjct: 224 YEILKTTCILEAFGNAKTSRNANSSRFGKLIEIHFSAMGKICGAKLETFLFDQSRVVQLF 283
Query: 248 DPERNYHCFYQLCAFETDA--EKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRAMN 305
+ ER+YH FY+LCA + E+ +L S + YL+QS + GV + +++ + A +
Sbjct: 284 NGERSYHIFYELCAGASPILKERLKLKTASEYTYLSQSDCLTIAGVDDAQKFHKLLEAFD 343
Query: 306 IVGISHEDQEAIFRTLAAILHLGNIEFS-PGKEYDSSVIKDEKSRFHMQMAANLFMCDVD 364
IV I E QE F LAA+L LGN+ F E V+ DE + AA L C+ +
Sbjct: 344 IVQIPKEHQERAFALLAAVLWLGNVSFRVTDNENHVEVVADEA----VANAAMLMGCNTE 399
Query: 365 LLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRS--VGQDI 422
L+ L TR +Q I K L A RD +AK +YA LFDWLV++IN + VG+
Sbjct: 400 ELMVVLSTRKLQAGTDCIAKKLTLRQATDMRDGIAKFIYANLFDWLVEQINIALEVGKSR 459
Query: 423 NSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYI 482
+ I +LDIYGFE FK+NSFEQFCIN+ANE+LQQHFN H+FK+EQEEY + I+W+ +
Sbjct: 460 TGR-SISILDIYGFESFKNNSFEQFCINYANERLQQHFNRHLFKLEQEEYEEDGIDWTKV 518
Query: 483 EFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTD 542
EFVDNQ+ L+LIEKKPIG+++LLDE FPK+T TF+ KL QH ++ E+
Sbjct: 519 EFVDNQECLDLIEKKPIGLLSLLDEESNFPKATDLTFANKLKQHLKTNSCFKGER--GRA 576
Query: 543 FAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPLPPEESSRSSYRFS 602
F ++HYAG+V Y T+ FL+KNRD + + NLLSS C + LF S+ S
Sbjct: 577 FRVNHYAGEVLYDTNGFLEKNRDPLPADLINLLSSCDCQLLK-LFSTKMRGKSQKPLMLS 635
Query: 603 -----SVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVLE 657
+V ++FK QL LM L +T PH++RC+KPNS P+++E V+ QLRC GVLE
Sbjct: 636 DSTNQTVGTKFKGQLFKLMNKLENTSPHFIRCIKPNSKQLPRVYEEDLVLQQLRCCGVLE 695
Query: 658 AVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKL--ENFQLGRTK 715
VRIS +GYPTR T+ EF R+G + L + D +V+ +L++ + E +Q+G TK
Sbjct: 696 VVRISRSGYPTRLTHQEFAGRYGFL-LSDKKVAQDPLSVSIAVLKQYDVHPEMYQVGYTK 754
Query: 716 VFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQK 775
++LR GQIGI + RR +VL +Q+ R ++R F +R + LQ+ RG ++
Sbjct: 755 LYLRTGQIGIFEDRRKKVLQGIVG-LQKHFRGHLSRAYFQNMRKVTLVLQSYIRGENARR 813
Query: 776 MY-------------ASKRETAAAISVQKYIRMWLRRKTY 802
++ AS E +A I +Q +R WL RK +
Sbjct: 814 LFDTEAKFHADSVSEASTDELSAVIHLQSAVRGWLARKHF 853
Score = 32.3 bits (72), Expect = 2.2
Identities = 29/117 (24%), Positives = 55/117 (46%), Gaps = 7/117 (5%)
Query: 953 LQNQFEL---SIKEKSALKRELVAVDEIRKENAMLK-VSLDAFEKKCTALEVELINAQKG 1008
+Q Q EL + K K R + +I E ++ S+ +K+ E L ++
Sbjct: 856 MQRQKELRNVATKSKRKAGRRISEDKDIPLEQPQVQPTSMSDLQKRILKSEAALSQKEEE 915
Query: 1009 RDETIEKLREFEQKCSQLEQNVKSLEE---KMLSLEDENHVLRQKALSAPLKSNRQG 1062
E+LR+FE++ S+ + +KS+EE K +S + +K+L+A + + G
Sbjct: 916 NTALREQLRQFEERWSEYDIKMKSMEETWQKQMSSLQMSLAAARKSLAAESITGQAG 972
>At4g27370 myosin heavy chain - like protein
Length = 1126
Score = 568 bits (1464), Expect = e-162
Identities = 342/769 (44%), Positives = 463/769 (59%), Gaps = 48/769 (6%)
Query: 55 ADEEEHGGVEDMTRLAYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNH 114
A+ E GVED+T+L+YLNEP +LYNL+ RY+ + IY+ G +LIAVNPF + +Y
Sbjct: 158 ANPEILEGVEDLTQLSYLNEPSLLYNLRVRYSQDLIYSKAGPVLIAVNPFKNV-QIYGEE 216
Query: 115 MMEQYKGAPLGELSPHVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTF 174
+ Y+ L +PHV+AVADA+Y MM GESGAGKTET K MQYL
Sbjct: 217 FLSAYQKNALD--APHVYAVADAAYDDMMRG--------DGESGAGKTETAKYAMQYLEA 266
Query: 175 VGGRAAGDDRSVEQQVLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIR 234
+GG + G VE ++L++N +LEAFGNA+T RNDNSSRFGK +EI F + G+I GA +
Sbjct: 267 LGGGSFG----VENEILKTNCILEAFGNAKTSRNDNSSRFGKLMEIHFSAKGKICGAKLE 322
Query: 235 TYLLERSRVVQITDPERNYHCFYQLCAFETDA--EKYELGHPSHFHYLNQSKIYELDGVS 292
T+ L++SRV Q+ + ER YH FYQLCA + E+ ++ S ++YLNQS +D
Sbjct: 323 TFSLDQSRVAQLCNGERCYHIFYQLCAGASPILKERLKIKAASEYNYLNQSNCLTIDRTD 382
Query: 293 NVEEYVRTRRAMNIVGISHEDQEAIFRTLAAILHLGNIEFSP-GKEYDSSVIKDEKSRFH 351
+ +++ + A NIV I E QE F LAA+L LGN+ F E V+ DE
Sbjct: 383 DAQKFHKLMEAFNIVQIPQEYQERTFALLAAVLWLGNVSFEVIDNENHVEVVADEA---- 438
Query: 352 MQMAANLFMCDVDLLLSTLCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLV 411
+ A L C+ L+ L T +Q I K L A RD+LAK +YA LF+WLV
Sbjct: 439 VTNVAMLMGCNSKKLMVVLSTCKLQAGRDCIAKRLTLRQATDMRDSLAKIIYASLFNWLV 498
Query: 412 DKINRS--VGQDINSQMQIGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQ 469
++IN S VG + I +LDIYGFE FKDNSFEQFCIN+ANE+LQQHFN H+FK+EQ
Sbjct: 499 EQINISLEVGNSRTGR-SISILDIYGFESFKDNSFEQFCINYANERLQQHFNRHLFKLEQ 557
Query: 470 EEYGREEINWSYIEFVDNQDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPS 529
EEY + I+W+ +EF+DNQ+ L LIEKKPIG+V+LL+E FPK+T TF+ KL QH +
Sbjct: 558 EEYEGDGIDWTKVEFIDNQECLNLIEKKPIGLVSLLNEESNFPKATDTTFANKLKQHLNA 617
Query: 530 HPRLGKEKFSQTDFAISHYAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKCPFVSGLFPL 589
+ E+ F I HYAG+V Y+T+ FL+KNRD + V+ LLS KC ++ LF
Sbjct: 618 NSCFKGER--GRGFRIKHYAGEVLYNTNGFLEKNRDPLHVDLIQLLSLCKCQLLN-LFST 674
Query: 590 PPEESSRSSYRFS-----SVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENG 644
FS SV ++FK QL LM L T PH++RC+KPNS P ++E
Sbjct: 675 KMHHDFLKPATFSDSMNQSVIAKFKGQLFKLMNKLEDTTPHFIRCIKPNSNQLPGLYEEN 734
Query: 645 SVIHQLRCGGVLEAVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKL 704
V+ QLRC GVLE VRIS +GYPTR T+ E R+G + L+ S D + ++ IL++
Sbjct: 735 HVLQQLRCCGVLEIVRISRSGYPTRLTHQELAVRYGCLLLDTRI-SQDPLSTSKAILKQC 793
Query: 705 KL--ENFQLGRTKVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAV 762
L E +Q+G TK++LR G I +L+ R+ VL +Q+Q R + R F +R AAV
Sbjct: 794 NLPPEMYQVGYTKIYLRTGVISVLEERKKYVL-RGILGLQKQFRGYQTREYFHNMRNAAV 852
Query: 763 CLQACCRGYIGQKMY-----------ASKRETAAAISVQKYIRMWLRRK 800
LQ+ RG ++ Y A +E AAI +Q +R WL RK
Sbjct: 853 ILQSYIRGENARRNYIVVGESAIVSTAITKELDAAIHLQYMVRKWLARK 901
>At1g50360 myosin, putative
Length = 1085
Score = 466 bits (1199), Expect = e-131
Identities = 318/962 (33%), Positives = 495/962 (51%), Gaps = 145/962 (15%)
Query: 10 WVQDRDQAWVAAEVLASDGGGNRVQLVTDSGKKVLASPEKLCPRDADEEEHGGVEDMTRL 69
WVQ + W +++++ G +V GK + E L P + D + GV+D+ +L
Sbjct: 120 WVQLPNGNWELGKIMSTSG--EESVIVVTEGKVLKVKSETLVPANPDILD--GVDDLMQL 175
Query: 70 AYLNEPGVLYNLKRRYTLNDIYTYTGSILIAVNPFTKLPHLYDNHMMEQYKGAPLGELSP 129
+YLNEP VLYNL+ RY + IYT G +L+AVNPF ++P LY N +E Y+ SP
Sbjct: 176 SYLNEPAVLYNLEYRYNQDMIYTKAGPVLVAVNPFKEVP-LYGNRNIEAYRKR--SNESP 232
Query: 130 HVFAVADASYRAMMNEGKSQSILVSGESGAGKTETTKLIMQYLTFVGGRAAGDDRSVEQQ 189
HV+A+AD + R M+ + +QSI++SGESGAGKTET K+ MQYL +GG + +E +
Sbjct: 233 HVYAIADTAIREMIRDEVNQSIIISGESGAGKTETAKIAMQYLAALGGGSG-----IEYE 287
Query: 190 VLESNPLLEAFGNARTVRNDNSSRFGKFVEIQFDSNGRISGAAIRTYLLERSRVVQITDP 249
+L++NP+LEA FG ++ D++
Sbjct: 288 ILKTNPILEA--------------FGNAKTLRNDNS------------------------ 309
Query: 250 ERNYHCFYQLCAFETDAEKYELGHPSHFHYLNQSKIYELDGVSNVEEYVRTRRAMNIVGI 309
+K L ++YL QS Y ++GV + E + + A++IV +
Sbjct: 310 -----------------KKLNLTSAKQYNYLKQSNCYSINGVDDAERFHAVKEALDIVHV 352
Query: 310 SHEDQEAIFRTLAAILHLGNIEFSPGKEYDSSVIKDEKSRFHMQMAANLFMCDVDLLLST 369
S EDQE +F LAA+L LGN+ F+ + + ++S + A L C+++ L
Sbjct: 353 SKEDQENVFAMLAAVLWLGNVSFTIIDNENHVEPEPDES---LSTVAKLIGCNINELKLA 409
Query: 370 LCTRSIQTREGSIVKALDCNAAVAGRDTLAKTVYARLFDWLVDKINRS--VGQDINSQMQ 427
L R+++ +IV+ L + A+ RD LAK++YA LFDWLV++IN+S VG+ +
Sbjct: 410 LSKRNMRVNNDTIVQKLTLSQAIDARDALAKSIYACLFDWLVEQINKSLAVGKRRTGR-S 468
Query: 428 IGVLDIYGFECFKDNSFEQFCINFANEKLQQHFNEHVFKMEQEEYGREEINWSYIEFVDN 487
I +LDIYGFE F NSFEQFCIN+ANE+LQQHFN H+FK+EQEEY ++ I+W+ ++F DN
Sbjct: 469 ISILDIYGFESFNKNSFEQFCINYANERLQQHFNRHLFKLEQEEYIQDGIDWTRVDFEDN 528
Query: 488 QDVLELIEKKPIGIVALLDEACMFPKSTHETFSTKLFQHFPSHPRLGKEKFSQTDFAISH 547
Q+ L L EKKP+G+++LLDE FP T T + KL QH + ++ F ++H
Sbjct: 529 QECLSLFEKKPLGLLSLLDEESTFPNGTDLTLANKLKQHLNDNSCFRGDRGKA--FTVAH 586
Query: 548 YAGKVTYHTDTFLDKNRDYVVVEHCNLLSSSKC----PFVSGLF-----PL--PPEESSR 596
YAG+VTY T FL+KNRD + + LLSS C F S + PL P ++
Sbjct: 587 YAGEVTYETTGFLEKNRDLLHSDSIQLLSSCSCHLPQAFASSMLIYSEKPLVGPLHKAGG 646
Query: 597 SSYRFSSVASRFKQQLQALMETLNSTEPHYVRCVKPNSLNRPQMFENGSVIHQLRCGGVL 656
+ + SVA++FK QL LM+ L +T PH++RC+KPN++ ++E G V+ QLRC GVL
Sbjct: 647 ADSQRLSVATKFKGQLFQLMQRLGNTTPHFIRCIKPNNVQSAGLYEQGLVLQQLRCCGVL 706
Query: 657 EAVRISLAGYPTRRTYSEFVDRFGLIALEFMDGSYDDKAVAEKILQKLKL--ENFQLGRT 714
E + + D +V+ IL + + E +Q+G T
Sbjct: 707 ENI-----------------------------AAKDPLSVSVAILHQFNILPEMYQVGYT 737
Query: 715 KVFLRAGQIGILDSRRAEVLDNAARCIQRQLRTFIARRVFIAVRAAAVCLQACCRGYIGQ 774
K+F R GQIG+L+ R L R +Q R AR ++ LQ+ RG +
Sbjct: 738 KLFFRTGQIGVLEDTRNRTLHGILR-LQSYFRGHQARCRLKELKTGITILQSFVRGEKMR 796
Query: 775 KMYAS--KRETAAAISVQKYIRMWLRRKTYMKLCSSATIIQSCVRGFMTRQRFLHIKEHR 832
K Y +R A+A ++Q +++ + + Y ++ +IQS +RG + R R
Sbjct: 797 KEYTELLQRHRASA-AIQSHVKRRIASQQYKATVDASAVIQSAIRGELVR---------R 846
Query: 833 AATSIQACWRMYKFRSAFHRHQASIVAIQCLWRRRQAKRELR---RLKQEANETGALRLA 889
A I + R+++ V ++ + +R LR L+++ E LR
Sbjct: 847 CAGDIG-----WLSSGGTKRNESDEVLVKASYLSDLQRRVLRTEAALREKEEENDILRQR 901
Query: 890 KSKLEKQLDELTWRLHLEKKIRVSNEDAKQIEISKLQKMIEALNLELDAAKLATINECNK 949
+ + + E E K++ S E+ Q ++ LQ + L+ A ++ +
Sbjct: 902 VQQYDNRWSE------YETKMK-SMEEIWQKQMKSLQSSLSIAKKSLEVEDSARNSDASV 954
Query: 950 NA 951
NA
Sbjct: 955 NA 956
>At5g20470 myosin-like protein
Length = 556
Score = 159 bits (401), Expect = 2e-38
Identities = 78/151 (51%), Positives = 112/151 (73%), Gaps = 2/151 (1%)
Query: 1377 NGEYMKSGLAELEKWIVNAKEEYAGTSWHELNYIRQAVGFLVIHQKRKKSLDEIRQDLCP 1436
NGEY+K+GLAELE+W + A +EYAG++W EL +IRQAVGFLV +QK K SL I P
Sbjct: 399 NGEYVKAGLAELEQWCIEATDEYAGSAWDELRHIRQAVGFLVTYQKPKMSLAVI-TSFFP 457
Query: 1437 ALTVRQIYRISTMYWDDKYGTQSVSNEVVSEMREIVSSKDNQNITSNSFLLDDDMSIPFS 1496
L+++Q+YRIST YWD+KYGT SVS++V++ MR ++ ++D+ N S+SFLLD+D SIPF+
Sbjct: 458 VLSIQQLYRISTNYWDEKYGTHSVSSDVIANMR-VMMTEDSNNAVSSSFLLDEDDSIPFT 516
Query: 1497 AEDIDMAIPAIDPDEIDLPAFVPEYSCAQFL 1527
DI ++ ++ ++I+LP + E S FL
Sbjct: 517 VGDITESMEQVNVNDIELPQLIRENSSFSFL 547
Score = 150 bits (380), Expect = 4e-36
Identities = 103/321 (32%), Positives = 152/321 (47%), Gaps = 59/321 (18%)
Query: 876 LKQEANETGALRLAKSKLEKQLDELTWRLHLEKKIRVSNEDAKQIEISKLQKMIEALNLE 935
LK A +TGALR AK KLEK+++ELT RL LE + R E+AK E +K Q+ ++A+ L+
Sbjct: 96 LKMAARDTGALREAKDKLEKRVEELTLRLQLETRQRTDLEEAKTQEYAKQQEALQAMWLQ 155
Query: 936 LDAAKLATINECNKNAVLQNQFELSIKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKC 995
++ A + E + IKE L + ++ + E LK A E
Sbjct: 156 VEEANAVVVREREAARKAIEEAPPVIKEIPVLVEDTEKINSLTSEVEALKAERQAAEH-- 213
Query: 996 TALEVELINAQKGRDETIEKLREFEQKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAP 1055
LE + E +L +K QL ++V+ LEEK+ + E E VLRQ+AL+
Sbjct: 214 --LEKAFSETEARNSELATELENATRKADQLHESVQRLEEKLSNSESEIQVLRQQALAI- 270
Query: 1056 LKSNRQGFAKSLSERYSNAVASRTERKPIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQ 1115
S +K T E
Sbjct: 271 ------------------------------------------------SGETKTTPE--- 279
Query: 1116 DNCEFLSRCIKENLGFKNGKPLAAPIIYKCLLHWHAFESERTAIFDYIIEGINEVLKARD 1175
+ L +CI +NLG+ P+AA +IYKCLLHW +FE ERT++FD IIE I ++ +
Sbjct: 280 ---DILVKCISQNLGYNGDMPVAACVIYKCLLHWRSFELERTSVFDRIIETIGSAVEVLE 336
Query: 1176 DDDVLPYWLSNTSALLCLLQR 1196
D++VL YWLSN ++L L++
Sbjct: 337 DNEVLAYWLSNLASLSLFLEQ 357
>At5g20450 putative protein
Length = 356
Score = 147 bits (371), Expect = 5e-35
Identities = 110/343 (32%), Positives = 173/343 (50%), Gaps = 64/343 (18%)
Query: 961 IKEKSALKRELVAVDEIRKENAMLKVSLDAFEKKCTALEVELINAQKGRDETIEKLREFE 1020
I+E+ ++ + ++ KEN+ ++ + AL+ L + ++ ++ E E
Sbjct: 35 IREQETARKAIEEAPQVIKENSEDTEKFNSLTSEVEALKASLQSERQAAEDLRNAFSEAE 94
Query: 1021 QKCSQLEQNVKSLEEKMLSLEDENHVLRQKALSAPLKSNRQGFAKSLSERYSNAVASRTE 1080
+ S+L N++++ ++ L + SA L+S +Q A+ L + S A A E
Sbjct: 95 ARNSELATNLENVTRRVDQLCE----------SASLQSEQQA-AEDLRKALSLAEARNLE 143
Query: 1081 RKPIFESPTPTKLIPTFTPGMSDSRRSKLTAERHQDNCEFLSRCIKENLGFKNGKPLAAP 1140
E+ T RR E ++ E L +CI +NLG+ GKP+AA
Sbjct: 144 LTTKLENVT---------------RRVDQLCE--SESQEVLVKCISQNLGYDGGKPVAAC 186
Query: 1141 IIYKCLLHWHAFESERTAIFDYIIE------GINEVLKARDDDDVLPYWLSNTSALLCLL 1194
+IYKCLLHW +FE ERT IFD I++ +++ K D+++VL YWLSN++ L+C+L
Sbjct: 187 VIYKCLLHWRSFEVERTNIFDRIVKIIASAIEVSDSYKVSDNNEVLAYWLSNSAMLVCVL 246
Query: 1195 QRNLRSNGFLTTTGQRYTGSAGLASRTVHVSYLVSTSINHSCGPKSPLKFIGYDDGVSHV 1254
QR RS+ L+ S GL S V VSYL D S V
Sbjct: 247 QRTFRSSTMLS--------SKGLMS-GVLVSYL---------------------DRQSQV 276
Query: 1255 EARYPAILFKQQLTACVEKIFGLLRDDLKKELSPLLGLCIQAP 1297
+ PA+LFK+QL ++KI+G++RD+LKKE+ P L C QAP
Sbjct: 277 VPKCPAMLFKKQLIDFLQKIYGMMRDNLKKEILPHLEYCKQAP 319
>At1g42680 myosin heavy chain ATM2, putative
Length = 162
Score = 101 bits (251), Expect = 4e-21
Identities = 48/80 (60%), Positives = 63/80 (78%), Gaps = 5/80 (6%)
Query: 157 SGAGKTETTKLIMQYLTFVGGRAAGDDRSVEQQVLESNPLLEAFGNARTVRNDNSSRFGK 216
SGAGKTETTK+ MQYL +GG + +E ++L++NP+LEAFGNA+T+RNDNSSRFGK
Sbjct: 51 SGAGKTETTKIAMQYLAALGGGSG-----IEYEILKTNPILEAFGNAKTLRNDNSSRFGK 105
Query: 217 FVEIQFDSNGRISGAAIRTY 236
+EI F G+ISGA I+T+
Sbjct: 106 LIEIHFSETGKISGAQIQTF 125
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.135 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,242,842
Number of Sequences: 26719
Number of extensions: 1455955
Number of successful extensions: 5093
Number of sequences better than 10.0: 188
Number of HSP's better than 10.0 without gapping: 42
Number of HSP's successfully gapped in prelim test: 151
Number of HSP's that attempted gapping in prelim test: 4506
Number of HSP's gapped (non-prelim): 455
length of query: 1532
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1420
effective length of database: 8,326,068
effective search space: 11823016560
effective search space used: 11823016560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0034a.6


