

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.7
(715 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
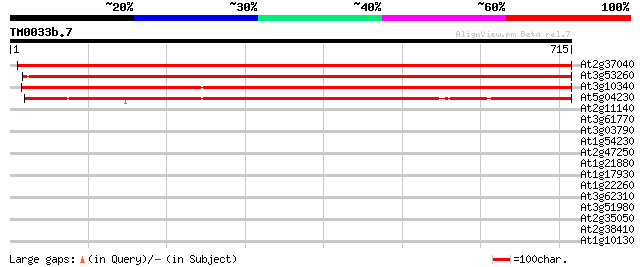
Score E
Sequences producing significant alignments: (bits) Value
At2g37040 phenylalanine ammonia lyase (PAL1) 1174 0.0
At3g53260 phenylalanine ammonia-lyase 1160 0.0
At3g10340 putative phenylalanine ammonia-lyase 1114 0.0
At5g04230 phenylalanine ammonia-lyase PAL3 985 0.0
At2g11140 putative retroelement pol polyprotein 33 0.73
At3g61770 unknown protein 32 1.6
At3g03790 unknown protein 32 1.6
At1g54230 hypothetical protein 32 1.6
At2g47250 putative pre-mRNA splicing factor RNA helicase 31 2.1
At1g21880 receptor-like GPI-anchored protein (lysM) 1 31 2.8
At1g17930 unknown protein 30 3.6
At1g22260 hypothetical protein 30 4.7
At3g62310 ATP-dependent RNA helicase-like protein 30 6.2
At3g51980 unknown protein 30 6.2
At2g35050 putative protein kinase 30 6.2
At2g38410 unknown protein 29 8.0
At1g10130 putative calcium ATPase 29 8.0
>At2g37040 phenylalanine ammonia lyase (PAL1)
Length = 725
Score = 1174 bits (3038), Expect = 0.0
Identities = 577/705 (81%), Positives = 645/705 (90%)
Query: 11 GSIFCNTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAI 70
G I + A DPL+WG AAE MKGSHLDEVKRMVAE+RK VV LGGETLTI QVAAI
Sbjct: 21 GDIKTKNMVINAEDPLNWGAAAEQMKGSHLDEVKRMVAEFRKPVVNLGGETLTIGQVAAI 80
Query: 71 AANHQGVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLE 130
+ V VEL E+ARAGV ASSDWVM SMN GTDSYGVTTGFGATSHRRTKNG ALQ E
Sbjct: 81 STIGNSVKVELSETARAGVNASSDWVMESMNKGTDSYGVTTGFGATSHRRTKNGVALQKE 140
Query: 131 LIRFLNAGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNIT 190
LIRFLNAGIFG+ E++HTLP ATRAAMLVRINTLLQG+SGIRFEILEAIT +NNNIT
Sbjct: 141 LIRFLNAGIFGSTKETSHTLPHSATRAAMLVRINTLLQGFSGIRFEILEAITSFLNNNIT 200
Query: 191 PCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQ 250
P LPLRGT+TASGDLVPLSYIAGLLTGRPNSKA GP+GE + A++AF+LAGI SGFF+LQ
Sbjct: 201 PSLPLRGTITASGDLVPLSYIAGLLTGRPNSKATGPNGEALTAEEAFKLAGISSGFFDLQ 260
Query: 251 PKEGLALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHH 310
PKEGLALVNGTAVGSG+AS+VLF+ N+L++LAE+LSA+FAEVM GKPEFTDHLTH+LKHH
Sbjct: 261 PKEGLALVNGTAVGSGMASMVLFETNVLSVLAEILSAVFAEVMSGKPEFTDHLTHRLKHH 320
Query: 311 PGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFST 370
PGQIEAAAIMEHILDGSSYMK A+KLHE+DPLQKPKQDRYALRTSPQWLGP IEVIR++T
Sbjct: 321 PGQIEAAAIMEHILDGSSYMKLAQKLHEMDPLQKPKQDRYALRTSPQWLGPQIEVIRYAT 380
Query: 371 KSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELV 430
KSIEREINSVNDNPLIDVSRNKA+HGGNFQGTPIGVSMDNTRLA+AAIGKLMFAQF+ELV
Sbjct: 381 KSIEREINSVNDNPLIDVSRNKAIHGGNFQGTPIGVSMDNTRLAIAAIGKLMFAQFSELV 440
Query: 431 DDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVN 490
+D YNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVT+HVQSAEQHNQDVN
Sbjct: 441 NDFYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTSHVQSAEQHNQDVN 500
Query: 491 SLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTTG 550
SLGLISSRKT+EA++ILKLMS+TFL+A+CQA+DLRHLEENL+ +VK+TVSQVAK+ LTTG
Sbjct: 501 SLGLISSRKTSEAVDILKLMSTTFLVAICQAVDLRHLEENLRQTVKNTVSQVAKKVLTTG 560
Query: 551 VNGELHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDS 610
VNGELHPSRFCEKDLLKVVDRE ++ Y DDPC ATYPL+QKLRQV+VDHAL+NGE EK++
Sbjct: 561 VNGELHPSRFCEKDLLKVVDREQVYTYADDPCSATYPLIQKLRQVIVDHALINGESEKNA 620
Query: 611 KTSIFQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTE 670
TSIF KI FE+ELK++LPKEVE+ARAAY++G A+PN+I ECRSYPLY+FVR+ELGTE
Sbjct: 621 VTSIFHKIGAFEEELKAVLPKEVEAARAAYDNGTSAIPNRIKECRSYPLYRFVREELGTE 680
Query: 671 LLTGEKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
LLTGEK SPGEE DK+FTAIC+GKIIDP++ECL EWNGAP+PIC
Sbjct: 681 LLTGEKVTSPGEEFDKVFTAICEGKIIDPMMECLNEWNGAPIPIC 725
>At3g53260 phenylalanine ammonia-lyase
Length = 717
Score = 1160 bits (3002), Expect = 0.0
Identities = 572/699 (81%), Positives = 641/699 (90%), Gaps = 1/699 (0%)
Query: 17 TVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQG 76
T KT A DPL+WG+AA+ MKGSHLDEVK+MV EYR+ VV LGGETLTI QVAAI+
Sbjct: 20 TTKTLA-DPLNWGLAADQMKGSHLDEVKKMVEEYRRPVVNLGGETLTIGQVAAISTVGGS 78
Query: 77 VSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLN 136
V VEL E++RAGVKASSDWVM SMN GTDSYGVTTGFGATSHRRTKNG ALQ ELIRFLN
Sbjct: 79 VKVELAETSRAGVKASSDWVMESMNKGTDSYGVTTGFGATSHRRTKNGTALQTELIRFLN 138
Query: 137 AGIFGNGTESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPLR 196
AGIFGN E+ HTLPQ ATRAAMLVR+NTLLQGYSGIRFEILEAIT L+N+NI+P LPLR
Sbjct: 139 AGIFGNTKETCHTLPQSATRAAMLVRVNTLLQGYSGIRFEILEAITSLLNHNISPSLPLR 198
Query: 197 GTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGLA 256
GT+TASGDLVPLSYIAGLLTGRPNSKA GP GE + AK+AF+ AGI +GFF+LQPKEGLA
Sbjct: 199 GTITASGDLVPLSYIAGLLTGRPNSKATGPDGESLTAKEAFEKAGISTGFFDLQPKEGLA 258
Query: 257 LVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEA 316
LVNGTAVGSG+AS+VLF+AN+ A+LAEVLSAIFAEVM GKPEFTDHLTH+LKHHPGQIEA
Sbjct: 259 LVNGTAVGSGMASMVLFEANVQAVLAEVLSAIFAEVMSGKPEFTDHLTHRLKHHPGQIEA 318
Query: 317 AAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIERE 376
AAIMEHILDGSSYMK A+K+HE+DPLQKPKQDRYALRTSPQWLGP IEVIR +TKSIERE
Sbjct: 319 AAIMEHILDGSSYMKLAQKVHEMDPLQKPKQDRYALRTSPQWLGPQIEVIRQATKSIERE 378
Query: 377 INSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYNN 436
INSVNDNPLIDVSRNKA+HGGNFQGTPIGVSMDNTRLA+AAIGKLMFAQF+ELV+D YNN
Sbjct: 379 INSVNDNPLIDVSRNKAIHGGNFQGTPIGVSMDNTRLAIAAIGKLMFAQFSELVNDFYNN 438
Query: 437 GLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLIS 496
GLPSNLTAS NPSLDYGFKGAEIAMASYCSELQYLANPVT+HVQSAEQHNQDVNSLGLIS
Sbjct: 439 GLPSNLTASSNPSLDYGFKGAEIAMASYCSELQYLANPVTSHVQSAEQHNQDVNSLGLIS 498
Query: 497 SRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTTGVNGELH 556
SRKT+EA++ILKLMS+TFL+ +CQA+DLRHLEENL+ +VK+TVSQVAK+ LTTG+NGELH
Sbjct: 499 SRKTSEAVDILKLMSTTFLVGICQAVDLRHLEENLRQTVKNTVSQVAKKVLTTGINGELH 558
Query: 557 PSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSIFQ 616
PSRFCEKDLLKVVDRE +F Y+DDPC ATYPLMQ+LRQV+VDHAL NGE EK++ TSIFQ
Sbjct: 559 PSRFCEKDLLKVVDREQVFTYVDDPCSATYPLMQRLRQVIVDHALSNGETEKNAVTSIFQ 618
Query: 617 KIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGEK 676
KI FE+ELK++LPKEVE+ARAAY +G +PN+I ECRSYPLY+FVR+ELGT+LLTGEK
Sbjct: 619 KIGAFEEELKAVLPKEVEAARAAYGNGTAPIPNRIKECRSYPLYRFVREELGTKLLTGEK 678
Query: 677 TRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
SPGEE DK+FTA+C+GK+IDPL++CL EWNGAP+PIC
Sbjct: 679 VVSPGEEFDKVFTAMCEGKLIDPLMDCLKEWNGAPIPIC 717
>At3g10340 putative phenylalanine ammonia-lyase
Length = 707
Score = 1114 bits (2881), Expect = 0.0
Identities = 558/701 (79%), Positives = 616/701 (87%), Gaps = 2/701 (0%)
Query: 16 NTVKTAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQ 75
N + + DPL+W AE++KGSHLDEVKRMV EYRK VKLGGETLTI QVAA+A
Sbjct: 8 NHITAVSGDPLNWNATAEALKGSHLDEVKRMVKEYRKEAVKLGGETLTIGQVAAVARGGG 67
Query: 76 GVSVELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFL 135
G +VEL E ARAGVKASS+WVM SMN GTDSYGVTTGFGATSHRRTK G ALQ ELIRFL
Sbjct: 68 GSTVELAEEARAGVKASSEWVMESMNRGTDSYGVTTGFGATSHRRTKQGGALQNELIRFL 127
Query: 136 NAGIFGNGT-ESTHTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLP 194
NAGIFG G +++HTLP+P TRAAMLVR+NTLLQGYSGIRFEILEAITKL+N+ ITPCLP
Sbjct: 128 NAGIFGPGAGDTSHTLPKPTTRAAMLVRVNTLLQGYSGIRFEILEAITKLLNHEITPCLP 187
Query: 195 LRGTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEG 254
LRGT+TASGDLVPLSYIAGLLTGRPNSKAVGPSGE + A +AF+LAG+ S FFELQPKEG
Sbjct: 188 LRGTITASGDLVPLSYIAGLLTGRPNSKAVGPSGETLTASEAFKLAGVSS-FFELQPKEG 246
Query: 255 LALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQI 314
LALVNGTAVGSGLAS VLFDANILA+L+EV+SA+FAEVMQGKPEFTDHLTHKLKHHPGQI
Sbjct: 247 LALVNGTAVGSGLASTVLFDANILAVLSEVMSAMFAEVMQGKPEFTDHLTHKLKHHPGQI 306
Query: 315 EAAAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIE 374
EAAAIMEHILDGSSY+K A+ LHE+DPLQKPKQDRYALRTSPQWLGP IEVIR +TK IE
Sbjct: 307 EAAAIMEHILDGSSYVKEAQLLHEMDPLQKPKQDRYALRTSPQWLGPQIEVIRAATKMIE 366
Query: 375 REINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHY 434
REINSVNDNPLIDVSRNKALHGGNFQGTPIGV+MDN+RLA+A+IGKLMFAQF+ELV+D Y
Sbjct: 367 REINSVNDNPLIDVSRNKALHGGNFQGTPIGVAMDNSRLAIASIGKLMFAQFSELVNDFY 426
Query: 435 NNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGL 494
NNGLPSNL+ RNPSLDYGFKGAEIAMASYCSELQ+LANPVT HVQSAEQHNQDVNSLGL
Sbjct: 427 NNGLPSNLSGGRNPSLDYGFKGAEIAMASYCSELQFLANPVTNHVQSAEQHNQDVNSLGL 486
Query: 495 ISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTTGVNGE 554
ISSRKT EA++ILKLMS+T+L+ALCQA+DLRHLEENLK +VKS VSQVAKR LT G NGE
Sbjct: 487 ISSRKTAEAVDILKLMSTTYLVALCQAVDLRHLEENLKKAVKSAVSQVAKRVLTVGANGE 546
Query: 555 LHPSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSI 614
LHPSRF E+D+L+VVDRE +F+Y DDPC TYPLMQKLR +LVDHAL + E E +S TS+
Sbjct: 547 LHPSRFTERDVLQVVDREYVFSYADDPCSLTYPLMQKLRHILVDHALADPEREANSATSV 606
Query: 615 FQKIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTG 674
F KI FE ELK LLPKEVE R YE G A+ N+I ECRSYPLY+FVR EL TELLTG
Sbjct: 607 FHKIGAFEAELKLLLPKEVERVRVEYEEGTSAIANRIKECRSYPLYRFVRDELNTELLTG 666
Query: 675 EKTRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
E RSPGEE DK+F AI GK+IDPLLECL EWNGAP+ IC
Sbjct: 667 ENVRSPGEEFDKVFLAISDGKLIDPLLECLKEWNGAPVSIC 707
>At5g04230 phenylalanine ammonia-lyase PAL3
Length = 694
Score = 985 bits (2547), Expect = 0.0
Identities = 507/699 (72%), Positives = 577/699 (82%), Gaps = 16/699 (2%)
Query: 20 TAATDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLGGETLTIAQVAAIAANHQGVSV 79
TA +DPL+W VAAE++KGSHL+EVK+MV +YRK V+LGGETLTI QVAA+A+ G +V
Sbjct: 9 TALSDPLNWNVAAEALKGSHLEEVKKMVKDYRKGTVQLGGETLTIGQVAAVASG--GPTV 66
Query: 80 ELCESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLNAGI 139
EL E AR GVKASSDWVM SMN TD+YG+TTGFG++S RRT G ALQ ELIR+LNAGI
Sbjct: 67 ELSEEARGGVKASSDWVMESMNRDTDTYGITTGFGSSSRRRTDQGAALQKELIRYLNAGI 126
Query: 140 FGNGTES---THTLPQPATRAAMLVRINTLLQGYSGIRFEILEAITKLINNNITPCLPLR 196
F G E ++TLP+PATRAAML+R+NTLLQGYSGIRFEILEAIT L+N ITP LPLR
Sbjct: 127 FATGNEDDDRSNTLPRPATRAAMLIRVNTLLQGYSGIRFEILEAITTLLNCKITPLLPLR 186
Query: 197 GTVTASGDLVPLSYIAGLLTGRPNSKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGLA 256
GT+TASGDLVPLSYIAG L GRPNS++VGPSGE++ A +AF+LAG+ S FFEL+PKEGLA
Sbjct: 187 GTITASGDLVPLSYIAGFLIGRPNSRSVGPSGEILTALEAFKLAGVSS-FFELRPKEGLA 245
Query: 257 LVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEA 316
LVNGTAVGS LAS VL+DANIL + +EV SA+FAEVMQGKPEFTDHLTHKLKHHPGQIEA
Sbjct: 246 LVNGTAVGSALASTVLYDANILVVFSEVASAMFAEVMQGKPEFTDHLTHKLKHHPGQIEA 305
Query: 317 AAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIERE 376
AAIMEHILDGSSY+K A LH++DPLQKPKQDRYALRTSPQWLGP IEVIR +TK IERE
Sbjct: 306 AAIMEHILDGSSYVKEALHLHKIDPLQKPKQDRYALRTSPQWLGPQIEVIRAATKMIERE 365
Query: 377 INSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYNN 436
INSVNDNPLIDVSRNKA+HGGNFQGTPIGV+MDNTRLALA+IGKLMFAQFTELV+D YNN
Sbjct: 366 INSVNDNPLIDVSRNKAIHGGNFQGTPIGVAMDNTRLALASIGKLMFAQFTELVNDFYNN 425
Query: 437 GLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGLIS 496
GLPSNL+ RNPSLDYG KGAE+AMASYCSELQ+LANPVT HV+SA QHNQDVNSLGLIS
Sbjct: 426 GLPSNLSGGRNPSLDYGLKGAEVAMASYCSELQFLANPVTNHVESASQHNQDVNSLGLIS 485
Query: 497 SRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYSVKSTVSQVAKRTLTTGVNGELH 556
SR T EA+ ILKLMS+T+L+ALCQA DLRHLEE LK +V VS AK L +
Sbjct: 486 SRTTAEAVVILKLMSTTYLVALCQAFDLRHLEEILKKAVNEVVSHTAKSVLA------IE 539
Query: 557 PSRFCEKDLLKVVDRETLFAYIDDPCLATYPLMQKLRQVLVDHALVNGEYEKDSKTSIFQ 616
P R D+L VV+RE +F+Y+DDP T PLMQKLR VL D AL E E D ++F+
Sbjct: 540 PFR-KHDDILGVVNREYVFSYVDDPSSLTNPLMQKLRHVLFDKALAEPEGETD---TVFR 595
Query: 617 KIATFEDELKSLLPKEVESARAAYESGNPAMPNKINECRSYPLYKFVRKELGTELLTGEK 676
KI FE ELK LLPKEVE R YE+G + N+I +CRSYPLY+FVR EL T LLTGE
Sbjct: 596 KIGAFEAELKFLLPKEVERVRTEYENGTFNVANRIKKCRSYPLYRFVRNELETRLLTGED 655
Query: 677 TRSPGEECDKLFTAICQGKIIDPLLECLGEWNGAPLPIC 715
RSPGE+ DK+F AI QGK+IDPL ECL EWNGAP+ IC
Sbjct: 656 VRSPGEDFDKVFRAISQGKLIDPLFECLKEWNGAPISIC 694
>At2g11140 putative retroelement pol polyprotein
Length = 411
Score = 32.7 bits (73), Expect = 0.73
Identities = 20/71 (28%), Positives = 36/71 (50%), Gaps = 1/71 (1%)
Query: 260 GTAVGSGLASIVLFDANILAILAEVLSAIFAEVMQGK-PEFTDHLTHKLKHHPGQIEAAA 318
G+AV S + +L D ++A+ ++ A G P+ D++ + L H +
Sbjct: 19 GSAVRSAREATILEDQKLMALKQQLTQRSDAYGGNGSNPKVPDNVDNPLILHSSDHPGLS 78
Query: 319 IMEHILDGSSY 329
I+ H+LDGS+Y
Sbjct: 79 IVAHVLDGSNY 89
>At3g61770 unknown protein
Length = 284
Score = 31.6 bits (70), Expect = 1.6
Identities = 24/75 (32%), Positives = 37/75 (49%), Gaps = 9/75 (12%)
Query: 255 LALVNGTA-----VGSGLASIVLFDA----NILAILAEVLSAIFAEVMQGKPEFTDHLTH 305
+AL +G A V G + IV++DA + AEVL+ I ++ +G P L
Sbjct: 194 VALCHGVADSLFPVCLGFSLIVMYDAIGVRRHAGMQAEVLNLIIRDLFEGHPISQRKLKE 253
Query: 306 KLKHHPGQIEAAAIM 320
L H P Q+ A A++
Sbjct: 254 LLGHTPSQVLAGALV 268
>At3g03790 unknown protein
Length = 1078
Score = 31.6 bits (70), Expect = 1.6
Identities = 27/123 (21%), Positives = 49/123 (38%), Gaps = 1/123 (0%)
Query: 415 LAAIGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSELQYLANP 474
L +G L + +V N LD GF ++ + +QY NP
Sbjct: 500 LLVVGSLYHPAYAPIVLKKSQTLQADKCREEENEELDEGFMFDDVESVNVLQSVQY-DNP 558
Query: 475 VTTHVQSAEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHLEENLKYS 534
V S + + V + ++ R + +EI + + L C+ I +R+L+ L +S
Sbjct: 559 KERIVPSLKSLCEKVAAECIVEPRNAIQLLEIADSLGAEDLKKYCEDIVIRNLDFILTFS 618
Query: 535 VKS 537
+S
Sbjct: 619 PQS 621
>At1g54230 hypothetical protein
Length = 266
Score = 31.6 bits (70), Expect = 1.6
Identities = 32/123 (26%), Positives = 57/123 (46%), Gaps = 5/123 (4%)
Query: 409 DNTRLALAA-IGKLMFAQFTELVDDHYNNGLPSNLTASRNPSLDYGFKGAEIAMASYCSE 467
DN+R L + KL+ + E V + Y +P + A+R S A+A +E
Sbjct: 68 DNSRERLRGEVEKLVAERKIEKVGNRYMMMMPQRVPATREDSTTPQRDPQAEAVAKLVAE 127
Query: 468 LQYLANPVTTHVQSAEQHNQDVNSLGLISSRKTNEAIEILKLMSSTFLIALCQAIDLRHL 527
++L + A++++ +NS +I A+EIL ++ +I LC ++ R
Sbjct: 128 TEHLEFQAKEAQELADRYSLMLNSECIILEL----AVEILNRCANGQMIFLCTSLVPRAP 183
Query: 528 EEN 530
EEN
Sbjct: 184 EEN 186
>At2g47250 putative pre-mRNA splicing factor RNA helicase
Length = 729
Score = 31.2 bits (69), Expect = 2.1
Identities = 32/128 (25%), Positives = 60/128 (46%), Gaps = 22/128 (17%)
Query: 270 IVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSY 329
I+L +A+ + +VL + EV++ +P+ KL +EA E+ G+
Sbjct: 188 IILDEAHERTLATDVLFGLLKEVLRNRPDL------KLVVMSATLEAEKFQEY-FSGAPL 240
Query: 330 MKAAKKLHEVDPL--QKPKQD--RYALRTSPQ--WLGPLIEVIRFST---------KSIE 374
MK +LH V+ Q+P++D A+RT Q P +++ F T + I
Sbjct: 241 MKVPGRLHPVEIFYTQEPERDYLEAAIRTVVQIHMCEPPGDILVFLTGEEEIEDACRKIN 300
Query: 375 REINSVND 382
+E++++ D
Sbjct: 301 KEVSNLGD 308
>At1g21880 receptor-like GPI-anchored protein (lysM) 1
Length = 416
Score = 30.8 bits (68), Expect = 2.8
Identities = 23/72 (31%), Positives = 33/72 (44%), Gaps = 3/72 (4%)
Query: 319 IMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIER--E 376
+ HIL ++K VD ++K Y R S LG + + + S E+ E
Sbjct: 81 VENHILPSKLFLKIPITCSCVDGIRKSVSTHYKTRPSDN-LGSIADSVYGGLVSAEQIQE 139
Query: 377 INSVNDNPLIDV 388
NSVND L+DV
Sbjct: 140 ANSVNDPSLLDV 151
>At1g17930 unknown protein
Length = 478
Score = 30.4 bits (67), Expect = 3.6
Identities = 18/52 (34%), Positives = 29/52 (55%), Gaps = 5/52 (9%)
Query: 175 FEILEAITKLINNNITPCLPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVGP 226
F+ +A+ L +N++ C P G + D+VP S+ LL GR +K +GP
Sbjct: 273 FKYRKALDDLDPSNVSWC-PFEGDL----DIVPQSFKDNLLLGRSRTKLIGP 319
>At1g22260 hypothetical protein
Length = 851
Score = 30.0 bits (66), Expect = 4.7
Identities = 30/118 (25%), Positives = 47/118 (39%), Gaps = 18/118 (15%)
Query: 293 MQGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSYMKAAKKLHEVDPLQKPKQDRYAL 352
+Q + E + +T H Q++A Y KKL E LQ+ K++R
Sbjct: 643 IQAENELKERITALKSEHDAQLKAFKCQ--------YEDDCKKLQEELDLQRKKEERQRA 694
Query: 353 RTSPQWLGPLIEVIRFSTKSIEREINSVNDNPLIDVSRNKALHGGNFQGTPIGVSMDN 410
QW + E+E+NS N +S++ L GG+ + I V DN
Sbjct: 695 LVQLQW------KVMSDNPPEEQEVNS---NKNYSISKDSRL-GGSKRSEHIRVRSDN 742
>At3g62310 ATP-dependent RNA helicase-like protein
Length = 726
Score = 29.6 bits (65), Expect = 6.2
Identities = 25/92 (27%), Positives = 45/92 (48%), Gaps = 11/92 (11%)
Query: 270 IVLFDANILAILAEVLSAIFAEVMQGKPEFTDHLTHKLKHHPGQIEAAAIMEHILDGSSY 329
I+L +A+ + +VL + EV++ +P+ KL +EA ++ G+
Sbjct: 184 IILDEAHERTLATDVLFGLLKEVLKNRPDL------KLVVMSATLEAEKFQDY-FSGAPL 236
Query: 330 MKAAKKLHEVDPL--QKPKQD--RYALRTSPQ 357
MK +LH V+ Q+P++D A+RT Q
Sbjct: 237 MKVPGRLHPVEIFYTQEPERDYLEAAIRTVVQ 268
>At3g51980 unknown protein
Length = 382
Score = 29.6 bits (65), Expect = 6.2
Identities = 37/169 (21%), Positives = 73/169 (42%), Gaps = 12/169 (7%)
Query: 23 TDPLSWGVAAESMKGSHLDEVKRMVAEYRKAVVKLG-GETLTIAQVAAIAANHQGVSVEL 81
+DP + AA+ + LDE+++ E ++ V KL + Q+A N+ +S+E
Sbjct: 76 SDPATLKEAAKDAEKMSLDELQKRQLELKELVEKLKMPSNAKLMQIAIDDLNNSSLSLED 135
Query: 82 CESARAGVKASSDWVMNSMNNGTDSYGVTTGFGATSHRRTKNGNALQLELIRFLNAGIFG 141
A + + + N+ N+ + S G+ G +H T+ +R L A + G
Sbjct: 136 RHRALQELLILVEPIDNA-NDLSKSGGLRVVAGELNHDDTE---------VRKLAAWVLG 185
Query: 142 NGTESTHTLPQPATRAAMLVRINTLLQGYSGIR-FEILEAITKLINNNI 189
+++ + + L + ++ S + L A++ LI NNI
Sbjct: 186 KASQNNPFVQEQVLELGALTTLIKMVNSSSTEEAVKALFAVSALIRNNI 234
>At2g35050 putative protein kinase
Length = 1257
Score = 29.6 bits (65), Expect = 6.2
Identities = 34/135 (25%), Positives = 58/135 (42%), Gaps = 12/135 (8%)
Query: 182 TKLINNNITPC----LPLRGTVTASGDLVPLSYIAGLLTGRPNSKAVG-PSGEVVNAKDA 236
T + N N P PL+ AS + P + G + +P+ + P V+ ++
Sbjct: 874 TSMTNTNGVPIDYSYPPLQSEKVASSQIHPQIHFDGNI--KPDVSTITIPDLNTVDTQED 931
Query: 237 F---QLAGIDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAILAEVLSAIFAEVM 293
+ Q+ G +S L G+ L++ A SG+ S+ + + L L E+ S F V
Sbjct: 932 YSQSQIKGAESTDATLNA--GVPLIDFMAADSGMRSLQVIKNDDLEELKELGSGTFGTVY 989
Query: 294 QGKPEFTDHLTHKLK 308
GK TD ++K
Sbjct: 990 HGKWRGTDVAIKRIK 1004
>At2g38410 unknown protein
Length = 671
Score = 29.3 bits (64), Expect = 8.0
Identities = 42/199 (21%), Positives = 81/199 (40%), Gaps = 29/199 (14%)
Query: 336 LHEVDPLQKPKQDRYALRTSPQWLGPLIEVIRFSTKSIEREINSVNDNPLID--VSRNKA 393
L VDP DR A++ + + L+E R + K + + + S D+ L+ + N +
Sbjct: 252 LQAVDP-----SDREAVKD--EVIVDLVERCRSNQKKLMQMLTSTGDDELLGRGLDLNDS 304
Query: 394 L------HGGNFQGTPIGVSMDNTRLALAAIGKLMFAQFTELVDDHYN-----NGLPSNL 442
L H G+P+ V + L++ A + + D + + +P+ +
Sbjct: 305 LQILLAKHDAIASGSPLPVQASGSPLSVQASKPADSSPKSSEAKDSSSIAGSSSPIPATV 364
Query: 443 TASRNPSLDYGFKGAEIAMASYCSELQYLANPVTTHVQSAEQHNQDVNSLGL-------- 494
+ ++P +D ++ E A VTT S E HN N+L L
Sbjct: 365 STGKSP-IDEEYEEEEDEFAQLARRHSKPPASVTTDPTSLESHNAASNALALALPDPPPP 423
Query: 495 ISSRKTNEAIEILKLMSST 513
+++ K + I++L + T
Sbjct: 424 VNTTKEQDMIDLLSITLCT 442
>At1g10130 putative calcium ATPase
Length = 992
Score = 29.3 bits (64), Expect = 8.0
Identities = 20/64 (31%), Positives = 32/64 (49%), Gaps = 11/64 (17%)
Query: 221 SKAVGPSGEVVNAKDAFQLAGIDSGFFELQPKEGLALVNGTAVGSGLASIVLFDANILAI 280
++ V +G+ VN A + A I G+A+ +GTAV + +VL D N +I
Sbjct: 678 NEVVAMTGDGVNDAPALKKADI-----------GIAMGSGTAVAKSASDMVLADDNFASI 726
Query: 281 LAEV 284
+A V
Sbjct: 727 VAAV 730
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,184,175
Number of Sequences: 26719
Number of extensions: 629344
Number of successful extensions: 1558
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 1545
Number of HSP's gapped (non-prelim): 18
length of query: 715
length of database: 11,318,596
effective HSP length: 106
effective length of query: 609
effective length of database: 8,486,382
effective search space: 5168206638
effective search space used: 5168206638
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0033b.7


