

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0033b.5
(88 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
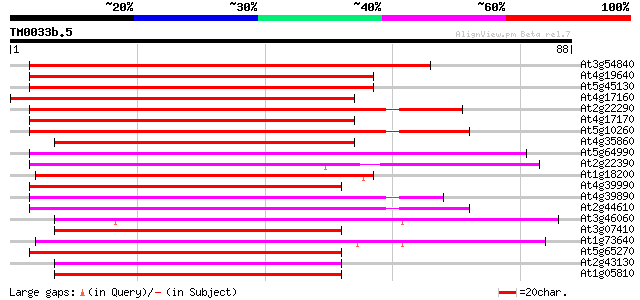
Score E
Sequences producing significant alignments: (bits) Value
At3g54840 small GTPase Ara6 (ARA6) 76 3e-15
At4g19640 small GTP-binding protein - like 66 3e-12
At5g45130 ras-related GTP-binding protein RHA1 (sp|P31582) 64 2e-11
At4g17160 GTP-binding RAB2A like protein 59 4e-10
At2g22290 putative GTP-binding protein 57 2e-09
At4g17170 GTP-binding RAB2A like protein 54 1e-08
At5g10260 GTP-binding protein 54 1e-08
At4g35860 GTP-binding protein GB2 53 2e-08
At5g64990 GTP binding protein-like 52 5e-08
At2g22390 putative GTP-binding protein 50 2e-07
At1g18200 hypothetical protein 50 2e-07
At4g39990 GTP-binding protein GB3 50 3e-07
At4g39890 small GTP-binding protein - like 50 3e-07
At2g44610 putative small GTP-binding protein 49 4e-07
At3g46060 GTP-binding protein ara-3 49 6e-07
At3g07410 GTP-binding protein like 49 6e-07
At1g73640 putative ras-related GTP-binding protein 49 6e-07
At5g65270 GTP-binding protein 46 3e-06
At2g43130 Ras-related GTP-binding protein (ARA-4) 45 5e-06
At1g05810 RAS-related protein ARA-1 45 5e-06
>At3g54840 small GTPase Ara6 (ARA6)
Length = 202
Score = 76.3 bits (186), Expect = 3e-15
Identities = 37/63 (58%), Positives = 47/63 (73%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+VMALV NK+DL KREV E+G + A++NGMF++ETSAKT +NIN LF EIG+ P
Sbjct: 140 IVMALVGNKADLHEKREVPTEDGMELAEKNGMFFIETSAKTADNINQLFEEIGKRLPRPA 199
Query: 64 RSS 66
SS
Sbjct: 200 PSS 202
>At4g19640 small GTP-binding protein - like
Length = 200
Score = 65.9 bits (159), Expect = 3e-12
Identities = 30/54 (55%), Positives = 40/54 (73%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NKSDL R+V E+ +AQENG+F+METSAKT N+ ++F+EI R
Sbjct: 116 MVMALAGNKSDLLDARKVTAEDAQTYAQENGLFFMETSAKTATNVKEIFYEIAR 169
>At5g45130 ras-related GTP-binding protein RHA1 (sp|P31582)
Length = 200
Score = 63.5 bits (153), Expect = 2e-11
Identities = 29/54 (53%), Positives = 39/54 (71%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGR 57
+VMAL NK+DL R+V EE +AQEN +F+METSAKT N+ D+F+EI +
Sbjct: 116 MVMALAGNKADLLDARKVSAEEAEIYAQENSLFFMETSAKTATNVKDIFYEIAK 169
>At4g17160 GTP-binding RAB2A like protein
Length = 205
Score = 58.9 bits (141), Expect = 4e-10
Identities = 27/54 (50%), Positives = 36/54 (66%)
Query: 1 SQILVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE 54
S+ + L+ NK DLE KR V EEG QFA+E+G+ +ME SAKT N+ + F E
Sbjct: 109 SENMTTMLIGNKCDLEDKRTVSTEEGEQFAREHGLIFMEASAKTAHNVEEAFVE 162
>At2g22290 putative GTP-binding protein
Length = 207
Score = 56.6 bits (135), Expect = 2e-09
Identities = 31/68 (45%), Positives = 42/68 (61%), Gaps = 2/68 (2%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V IEEG +E G+ ++ETSAK NI LF +I PG
Sbjct: 115 VIIVLVGNKTDLVEKRQVSIEEGDSKGREYGVMFIETSAKAGFNIKPLFRKIAAAL--PG 172
Query: 64 RSSYAPTK 71
SY+ TK
Sbjct: 173 MESYSNTK 180
>At4g17170 GTP-binding RAB2A like protein
Length = 211
Score = 54.3 bits (129), Expect = 1e-08
Identities = 23/51 (45%), Positives = 35/51 (68%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE 54
+ + L+ NK DL +R V EEG QFA+E+G+ +ME SAKT +N+ + F +
Sbjct: 112 MTIMLIGNKCDLAHRRAVSTEEGEQFAKEHGLIFMEASAKTAQNVEEAFIK 162
>At5g10260 GTP-binding protein
Length = 207
Score = 53.9 bits (128), Expect = 1e-08
Identities = 29/69 (42%), Positives = 43/69 (62%), Gaps = 2/69 (2%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V IEEG A++ G+ ++ETSAK NI LF +I PG
Sbjct: 115 VIIVLVGNKTDLVDKRQVSIEEGDNKARDYGVIFIETSAKAGFNIKPLFRKIAAAL--PG 172
Query: 64 RSSYAPTKE 72
+ + TK+
Sbjct: 173 METLSSTKQ 181
>At4g35860 GTP-binding protein GB2
Length = 211
Score = 53.1 bits (126), Expect = 2e-08
Identities = 23/47 (48%), Positives = 33/47 (69%)
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE 54
L+ NK DL KR V EEG QFA+E+G+ ++E SA+T +N+ + F E
Sbjct: 116 LIGNKCDLAHKRAVSKEEGQQFAKEHGLLFLEASARTAQNVEEAFIE 162
>At5g64990 GTP binding protein-like
Length = 206
Score = 52.0 bits (123), Expect = 5e-08
Identities = 29/78 (37%), Positives = 42/78 (53%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V IEEG A+E G +METSAK NI LF +I +
Sbjct: 113 VIIVLVGNKTDLVNKRQVSIEEGENKAREFGALFMETSAKAGFNIKPLFCKITSALQGNE 172
Query: 64 RSSYAPTKEWASAGLSSM 81
S+ ++ L +
Sbjct: 173 AVSWTKQEDLVDVNLKPL 190
>At2g22390 putative GTP-binding protein
Length = 176
Score = 50.4 bits (119), Expect = 2e-07
Identities = 29/84 (34%), Positives = 48/84 (56%), Gaps = 7/84 (8%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENI----NDLFFEIGRGF 59
+V+ L+ NK+DLE +R V +E+ +FA++ G+F++ETSA N+ N L EI F
Sbjct: 90 IVIILIGNKTDLENQRSVPVEDAKEFAEKEGLFFLETSALNSTNVENSFNTLLTEI---F 146
Query: 60 ESPGRSSYAPTKEWASAGLSSMPP 83
+ + A T S+ +S + P
Sbjct: 147 NKVNKKNLAKTTVSCSSQVSLLRP 170
>At1g18200 hypothetical protein
Length = 354
Score = 50.4 bits (119), Expect = 2e-07
Identities = 26/54 (48%), Positives = 35/54 (64%), Gaps = 1/54 (1%)
Query: 5 VMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFE-IGR 57
V+ LV NKSDL REVE EEG A+ G++++ETSA +N+ + F IGR
Sbjct: 120 VVVLVGNKSDLGQSREVEEEEGKTLAESEGLYFLETSALENQNVEEAFLSMIGR 173
>At4g39990 GTP-binding protein GB3
Length = 224
Score = 49.7 bits (117), Expect = 3e-07
Identities = 21/49 (42%), Positives = 34/49 (68%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+V+ L+ NKSDLE +R V E+ +FA++ G+F++ETSA N+ + F
Sbjct: 123 IVIILIGNKSDLEDQRAVPTEDAKEFAEKEGLFFLETSALNATNVENSF 171
>At4g39890 small GTP-binding protein - like
Length = 214
Score = 49.7 bits (117), Expect = 3e-07
Identities = 28/65 (43%), Positives = 39/65 (59%), Gaps = 2/65 (3%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V I EG +E G+ ++ETSAK NI LF +I PG
Sbjct: 116 VIIVLVGNKTDLVEKRQVSISEGEDKGKEYGVMFIETSAKENFNIKALFRKIAAAL--PG 173
Query: 64 RSSYA 68
SY+
Sbjct: 174 VDSYS 178
>At2g44610 putative small GTP-binding protein
Length = 208
Score = 48.9 bits (115), Expect = 4e-07
Identities = 28/69 (40%), Positives = 41/69 (58%), Gaps = 2/69 (2%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFESPG 63
+++ LV NK+DL KR+V IEE A+E + ++ETSAK NI LF +I PG
Sbjct: 115 VIVVLVGNKTDLVDKRQVSIEEAEAKARELNVMFIETSAKAGFNIKALFRKIAAAL--PG 172
Query: 64 RSSYAPTKE 72
+ + TK+
Sbjct: 173 METLSSTKQ 181
>At3g46060 GTP-binding protein ara-3
Length = 216
Score = 48.5 bits (114), Expect = 6e-07
Identities = 31/90 (34%), Positives = 44/90 (48%), Gaps = 11/90 (12%)
Query: 8 LVANKSDL-EPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFFEIGRGFE------ 60
LV NK+D+ E KR V +G A E G+ + ETSAKT N+ ++FF IGR +
Sbjct: 125 LVGNKADMDESKRAVPTAKGQALADEYGIKFFETSAKTNLNVEEVFFSIGRDIKQRLSDT 184
Query: 61 ----SPGRSSYAPTKEWASAGLSSMPPVSC 86
P + T + A AG ++ C
Sbjct: 185 DSRAEPATIKISQTDQAAGAGQATQKSACC 214
>At3g07410 GTP-binding protein like
Length = 217
Score = 48.5 bits (114), Expect = 6e-07
Identities = 23/45 (51%), Positives = 29/45 (64%)
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
LV NK DLE R V +EEG A+E G+F+METSA N++ F
Sbjct: 122 LVGNKCDLEDIRAVSVEEGKALAEEEGLFFMETSALDATNVDKAF 166
>At1g73640 putative ras-related GTP-binding protein
Length = 233
Score = 48.5 bits (114), Expect = 6e-07
Identities = 33/88 (37%), Positives = 46/88 (51%), Gaps = 8/88 (9%)
Query: 5 VMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLFF-EIGRGFE--- 60
V+ LV NKSDL REVE +EG A+ G++++ETSA N+ + F IGR E
Sbjct: 120 VVVLVGNKSDLRQSREVEEDEGKTLAESEGLYFLETSALENVNVEEAFLVMIGRIHEVVT 179
Query: 61 ----SPGRSSYAPTKEWASAGLSSMPPV 84
S +S+ A T G ++ PV
Sbjct: 180 QRIASENKSNGAATPHINGNGNGTVLPV 207
>At5g65270 GTP-binding protein
Length = 226
Score = 46.2 bits (108), Expect = 3e-06
Identities = 19/49 (38%), Positives = 32/49 (64%)
Query: 4 LVMALVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
+V+ L+ NKSDL +R + E+ +FA++ G+F++ETSA N+ F
Sbjct: 123 IVIILIGNKSDLVDQRAIPTEDAKEFAEKEGLFFLETSAFNATNVESAF 171
>At2g43130 Ras-related GTP-binding protein (ARA-4)
Length = 214
Score = 45.4 bits (106), Expect = 5e-06
Identities = 21/45 (46%), Positives = 27/45 (59%)
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
L+ NK DLE R V +EEG A+ G+F+METSA N+ F
Sbjct: 122 LIGNKCDLESIRAVSVEEGKSLAESEGLFFMETSALDSTNVKTAF 166
>At1g05810 RAS-related protein ARA-1
Length = 261
Score = 45.4 bits (106), Expect = 5e-06
Identities = 22/45 (48%), Positives = 28/45 (61%)
Query: 8 LVANKSDLEPKREVEIEEGGQFAQENGMFYMETSAKTVENINDLF 52
LV NK DLE R V +EEG A+E G+F++ETSA N+ F
Sbjct: 165 LVGNKCDLENIRAVSVEEGKALAEEEGLFFVETSALDSTNVKTAF 209
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,039,177
Number of Sequences: 26719
Number of extensions: 71651
Number of successful extensions: 195
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 136
Number of HSP's gapped (non-prelim): 62
length of query: 88
length of database: 11,318,596
effective HSP length: 64
effective length of query: 24
effective length of database: 9,608,580
effective search space: 230605920
effective search space used: 230605920
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0033b.5


