

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0032.12
(921 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
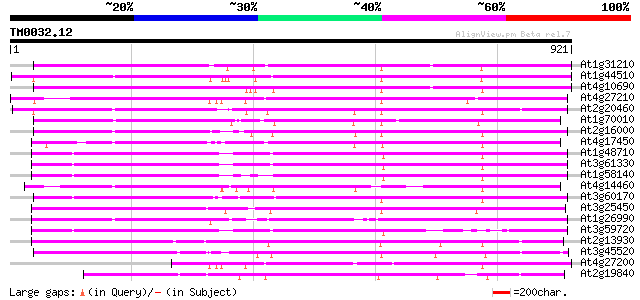
At1g31210 putative reverse transcriptase 663 0.0
At1g44510 polyprotein, putative 638 0.0
At4g10690 retrotransposon like protein 620 e-178
At4g27210 putative protein 561 e-160
At2g20460 putative retroelement pol polyprotein 551 e-157
At1g70010 hypothetical protein 531 e-151
At2g16000 putative retroelement pol polyprotein 514 e-146
At4g17450 retrotransposon like protein 509 e-144
At1g48710 hypothetical protein 493 e-139
At3g61330 copia-type polyprotein 491 e-138
At1g58140 hypothetical protein 490 e-138
At4g14460 retrovirus-related like polyprotein 481 e-136
At3g60170 putative protein 457 e-128
At3g25450 hypothetical protein 447 e-125
At1g26990 polyprotein, putative 437 e-122
At3g59720 copia-type reverse transcriptase-like protein 432 e-121
At2g13930 putative retroelement pol polyprotein 428 e-120
At3g45520 copia-like polyprotein 413 e-115
At4g27200 putative protein 409 e-114
At2g19840 copia-like retroelement pol polyprotein 367 e-101
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 663 bits (1710), Expect = 0.0
Identities = 367/923 (39%), Positives = 506/923 (54%), Gaps = 52/923 (5%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
VWH+RLGH S AL HL+N+K I + SR+S VC+ C +GK RLPF S++ L D
Sbjct: 455 VWHHRLGHANSKALQHLQNSKAIQINKSRTSPVCEPCQMGKSSRLPFLISDSRVLHPLDR 514
Query: 99 LHSDLW-TSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
+H DLW SP++S G +YY + +DD++ + W +P+ KS+ +F + L++ Q +
Sbjct: 515 IHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSWFYPLHNKSEFLSVFISFQKLVENQLNTK 574
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
IK Q D G E+ ++ + + +G+ R SCP+T QNG +ERK R + + ++L HS
Sbjct: 575 IKVFQSDGGGEFVSNKLKTHLSEHGIHHRISCPYTPQQNGLAERKHRHLVELGLSMLFHS 634
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSAT 277
P FW + A Y++N LP L+N SP + L+ P YS LRVFG C+P
Sbjct: 635 HTPQKFWVESFFTANYIINRLPSSVLKNLSPYEALFGEKPDYSSLRVFGSACYPCLRPLA 694
Query: 278 INKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYD 337
NK PRS CVFLGY ++GY+C+ K+ ISR+VIF+ES PF + Y
Sbjct: 695 QNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGKVYISRNVIFNESELPFKE-------KYQ 747
Query: 338 CFTEDLPPSLIHHWQTTS--------------TRPPDLSIPPSSPTDSTMPSPAPTSSPT 383
L+ WQ ++P DL+ S + P PTS+
Sbjct: 748 SLVPQYSTPLLQAWQHNKISEISVPAAPVQLFSKPIDLNTYAGSQVTEQLTDPEPTSNNE 807
Query: 384 SSAPLPLPPVPPTPP-------TRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPK 436
S P + MTT S GI KP ++L I + P
Sbjct: 808 GSDEEVNPVAEEIAANQEQVINSHAMTTRSKAGIQKPNTRYAL---ITSRMNTAEPKTLA 864
Query: 437 QALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARL 496
A+ P W A+ E N + +TW LVP D+N++ +F+ K +G + KARL
Sbjct: 865 SAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNILSSKWVFKTKLHPDGSIDKLKARL 924
Query: 497 VGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYM 556
V G+ Q GVD ETF+ VV+ ATIR VL ++ S+ WPI LDV NAFLHG+L E V+M
Sbjct: 925 VAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSKGWPIKQLDVSNAFLHGELQEPVFM 984
Query: 557 HQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS------- 609
+QP GF D Q P +VCR K++YGLKQAPRAW+ F++++ GF S SD S
Sbjct: 985 YQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGFVCSKSDPSLFVCHQD 1044
Query: 610 ----YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLS 665
Y+LLYVDDI+L S L + + L F+MKDLGP YFLGI + Y GLFL
Sbjct: 1045 GKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYFLGIQIEDYANGLFLH 1104
Query: 666 QSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPN 725
Q+ YA++I+ +AGM+ CNP TP+ +L + + + T ++SLAG LQYLT TRP+
Sbjct: 1105 QTAYATDILQQAGMSDCNPMPTPLPQ--QLDNLNSELFAEPTYFRSLAGKLQYLTITRPD 1162
Query: 726 ISYVVQQVCLHMHAPHTEH--MLAMFVAL*LTVFTCIPPLLRNLFLIRML-----TGGMS 778
I + V +C MH+P T +L + P+ RN L G
Sbjct: 1163 IQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGMGLPIKRNSTLTLSAYSDSDHAGCK 1222
Query: 779 CTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFP 838
TRRST+G+C+ LG NLISWS+KRQPT+S+SS EAEYR + E WI LL +L P
Sbjct: 1223 NTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYAAREITWISFLLRDLGIP 1282
Query: 839 LSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADI 898
T V+CDN+S++YLS NP H R+KH + D H++RE+VA G H+ + Q+AD+
Sbjct: 1283 QYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGLIETQHISATFQLADV 1342
Query: 899 FTKGLPRVLFDDFRSSLSFGEPP 921
FTK LPR F D RS L P
Sbjct: 1343 FTKSLPRRAFVDLRSKLGVSGSP 1365
>At1g44510 polyprotein, putative
Length = 1459
Score = 638 bits (1645), Expect = 0.0
Identities = 390/1034 (37%), Positives = 537/1034 (51%), Gaps = 118/1034 (11%)
Query: 3 FSVSDFQTGMPLLRCNSLGDLY--PVTCSSHFAGLAS-------GVWHNRLGHPTSSALN 53
F V D TG LL+ + DLY PVT A S WH+RLGHP++S LN
Sbjct: 427 FQVKDLNTGTLLLQGRTKDDLYEWPVTNPPATALFTSPSPKTTLSSWHSRLGHPSASILN 486
Query: 54 HLRNN-KLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTA 112
L + L S + T C C++ K +LPF++S + + + +D+WTSPI+S
Sbjct: 487 TLLSKFSLPVSVASSNKTSCSDCLINKSHKLPFATSSIHSSSPLEYIFTDVWTSPIISHD 546
Query: 113 GHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDND 172
++YY++++D +T + W++P+ +KSQV F L++ +F A I+ L DNG E+
Sbjct: 547 NYKYYLVLVDHYTRYTWLYPLQQKSQVKATFIAFKALVENRFQAKIRTLYSDNGGEFI-- 604
Query: 173 SFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMAT 232
+ R + +NG+ S PHT NG SERK R I TLL +SVP +W +A A
Sbjct: 605 ALRDFLVSNGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLTQASVPREYWTYAFATAV 664
Query: 233 YLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLG 292
YL+N +P L SP Q+L+ P+Y LRVFGCLCFP+ T NKL+ RS CVFLG
Sbjct: 665 YLINRMPTPVLCLQSPFQKLFGSSPNYQRLRVFGCLCFPWLRPYTRNKLEERSKRCVFLG 724
Query: 293 YPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFAD-----------LPLESTSSYDCFTE 341
Y + Y C D+ N +L SRHV+FDES +PFA P ES+SS
Sbjct: 725 YSLTQTAYLCLDVDNNRLYTSRHVMFDESTYPFAASIREQSQSSLVTPPESSSSSSPANS 784
Query: 342 DLPPSLI-----------------------------------HHWQ----TTSTRP---- 358
P S++ HH Q T S P
Sbjct: 785 GFPCSVLRLQSPPASSPETPSPPQQQNDSPVSPRQTGSPTPSHHSQVRDSTLSPSPSVSN 844
Query: 359 ------------PDLSIPPSSPTDSTMPSPAPTSSPTSSA---PLPLPPVPPTPPTRT-- 401
P+ P+SP +P+P P ++P+SS P+ PP +T
Sbjct: 845 SEPTAPHENGPEPEAQSNPNSPFIGPLPNPNPETNPSSSIEQRPVDKSTTTALPPNQTTI 904
Query: 402 ----------------MTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWK 445
M T S + I+KPK SL+V++ P +S P QAL D W+
Sbjct: 905 AATSNSRSQPPKNNHQMKTRSKNNITKPKTKTSLTVALTQPHLSE-PNTVTQALKDKKWR 963
Query: 446 FAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIA 505
FAM EF+A RN+TW+LVP +++ +F+ K NGL + YKARLV G++Q
Sbjct: 964 FAMSDEFDAQQRNHTWDLVPPNPTQHLVGCRWVFKLKYLPNGLIDKYKARLVAKGFNQQY 1023
Query: 506 GVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDS 565
GVD ETF+ V+K TIR VL +A+ ++WP+ LDV+NAFL G L E VYM QP GF D
Sbjct: 1024 GVDYAETFSPVIKATTIRVVLDVAVKKNWPLKQLDVNNAFLQGTLTEEVYMAQPPGFVDK 1083
Query: 566 QHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS-----------YILLY 614
P +VCR +K++YGLKQAPRAWY ++ +IGF +S +D S Y+L+Y
Sbjct: 1084 DRPSHVCRLRKAIYGLKQAPRAWYMELKQHLLNIGFVNSLADTSLFIYSHGTTLLYLLVY 1143
Query: 615 VDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEII 674
VDDII+ S H + ++ LA F++KD L YFLGI TR GL L Q Y ++++
Sbjct: 1144 VDDIIVTGSDHKSVSAVLSSLAERFSIKDPTDLHYFLGIEATRTNTGLHLMQRKYMTDLL 1203
Query: 675 ARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVC 734
A+ M P ATP+ T KL+ GT D + Y+S+ G+LQYL FTRP+I++ V ++
Sbjct: 1204 AKHNMLDAKPVATPLPTSPKLTLHGGTKLNDASEYRSVVGSLQYLAFTRPDIAFAVNRLS 1263
Query: 735 LHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRML-------TGGMSCTRRSTSGY 787
MH P ++H A L T + N L G S ST+ Y
Sbjct: 1264 QFMHQPTSDHWQAAKRVLRYLAGTTTHGIFLNSSSPIHLHAFSDADWAGDSADYVSTNAY 1323
Query: 788 CVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHC 847
++LG N ISWSSK+Q +S SS E+EYR VAN SE W+ +LL ELH L + C
Sbjct: 1324 VIYLGRNPISWSSKKQRGVSRSSTESEYRAVANAASEIRWLCSLLTELHIRLPHGPTIFC 1383
Query: 848 DNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVL 907
DN+ + Y+ NPV H R KHI +D HFVR + R+ HV + Q+AD TK L R
Sbjct: 1384 DNIGATYICANPVFHSRMKHIALDYHFVRGMIQSRALRVSHVSTNDQLADALTKSLSRPH 1443
Query: 908 FDDFRSSLSFGEPP 921
F RS + + P
Sbjct: 1444 FLSARSKIGVRQLP 1457
>At4g10690 retrotransposon like protein
Length = 1515
Score = 620 bits (1600), Expect = e-178
Identities = 364/972 (37%), Positives = 518/972 (52%), Gaps = 92/972 (9%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
VWH RLGHP L HL K I + + SS +C++C +GK RLPF +SE ++ R +
Sbjct: 457 VWHQRLGHPNKEVLQHLIKTKAIVVNKT-SSNMCEACQMGKVCRLPFVASEFVSSRPLER 515
Query: 99 LHSDLW-TSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
+H DLW +P+ S G +YYV+ +D+++ F W +P+ KS + +F L++ Q+
Sbjct: 516 IHCDLWGPAPVTSAQGFQYYVIFIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHK 575
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
I QCD G E+ + F + + G+ SCPHT QNG +ER+ R + + +L+ HS
Sbjct: 576 IAMFQCDGGGEFVSYKFVAHLASCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHS 635
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLR-NDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSA 276
VP W A + +L N+LP TL N SP + L+ P Y+ LRVFG C+P+
Sbjct: 636 KVPHKLWVEAFFTSNFLSNLLPSSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPY 695
Query: 277 TINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADL--PLESTS 334
NK P+S CVFLGY ++GY+C K+ I RHV+FDE +FP++D+ ++ S
Sbjct: 696 AKNKFDPKSLLCVFLGYNNKYKGYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTIS 755
Query: 335 SYDCFT--EDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAP----- 387
FT + S +T ST D+ P ++ + S AP + T++AP
Sbjct: 756 GSPLFTAWQKGFSSTALSRETPSTNVEDIIFPSATVSSSVPTGCAPNIAETATAPDVDVA 815
Query: 388 ----LPLPPVP------PTPPTRT------MTTHSMHGISKPKKPFSLSVSI-DDPSISP 430
+ +PP P PT P + +T S IS P S++VS+ +D P
Sbjct: 816 AAHDMVVPPSPITSTSLPTQPEESTSDQNHYSTDSETAISSAMTPQSINVSLFEDSDFPP 875
Query: 431 L-------------------------------------------PCNPKQALSDPNWKFA 447
L P + K+AL D W A
Sbjct: 876 LQSVISSTTAAPETSHPMITRAKSGITKPNPKYALFSVKSNYPEPKSVKEALKDEGWTNA 935
Query: 448 MQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGV 507
M E + +TW+LVP ++ +F+ K S+G + KARLV G+ Q GV
Sbjct: 936 MGEEMGTMHETDTWDLVPPEMVDRLLGCKWVFKTKLNSDGSLDRLKARLVARGYEQEEGV 995
Query: 508 DCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQH 567
D ET++ VV+ AT+R++L +A W + LDV NAFLH +L ETV+M QP GF D
Sbjct: 996 DYVETYSPVVRSATVRSILHVATINKWSLKQLDVKNAFLHDELKETVFMTQPPGFEDPSR 1055
Query: 568 PDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS-----------YILLYVD 616
PDYVC+ KK++Y LKQAPRAW+ +F+ Y+ GF S SD S ++LLYVD
Sbjct: 1056 PDYVCKLKKAIYDLKQAPRAWFDKFSSYLLKYGFICSFSDPSLFVYLKGRDVMFLLLYVD 1115
Query: 617 DIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIAR 676
D+IL ++ L + + +L+ EF MKD+G L YFLGI + GLFLSQ Y S+++
Sbjct: 1116 DMILTGNNDVLLQQLLNILSTEFRMKDMGALHYFLGIQAHYHNDGLFLSQEKYTSDLLVN 1175
Query: 677 AGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLH 736
AGM+ C+ TP+ L + P + T ++ LAG LQYLT TRP+I + V VC
Sbjct: 1176 AGMSDCSSMPTPLQL--DLLQGNNKPFPEPTYFRRLAGKLQYLTLTRPDIQFAVNFVCQK 1233
Query: 737 MHAPHTE--HMLAMFVAL*LTVFTCIPPLLRNL-FLIRMLT----GGMSCTRRSTSGYCV 789
MHAP H+L + T L N ++R + G TRRST G+C
Sbjct: 1234 MHAPTMSDFHLLKRILHYLKGTMTMGINLSSNTDSVLRCYSDSDWAGCKDTRRSTGGFCT 1293
Query: 790 FLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDN 849
FLG N+ISWS+KR PT+S SS EAEYR ++ SE WI LL E+ P Q ++CDN
Sbjct: 1294 FLGYNIISWSAKRHPTVSKSSTEAEYRTLSFAASEVSWIGFLLQEIGLPQQQIPEMYCDN 1353
Query: 850 VSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFD 909
+S++YLS NP H R+KH ++D ++VRE+VA G + H+P+ Q+ADIFTK LP+ F
Sbjct: 1354 LSAVYLSANPALHSRSKHFQVDYYYVRERVALGALTVKHIPASQQLADIFTKSLPQAPFC 1413
Query: 910 DFRSSLSFGEPP 921
D R L PP
Sbjct: 1414 DLRFKLGVVLPP 1425
>At4g27210 putative protein
Length = 1318
Score = 561 bits (1447), Expect = e-160
Identities = 355/996 (35%), Positives = 479/996 (47%), Gaps = 128/996 (12%)
Query: 2 GFSVSDFQTGMPLLRCNSLGDLYPVTCSSHFAGLASG--------VWHNRLGHPTSSALN 53
G V+D T LL ++ LY + F S VWH RLGHP L
Sbjct: 266 GVRVNDKATKKLLLMGSNRDGLYCLKDDKQFQAFFSTRQRSASDEVWHRRLGHPHPQILQ 325
Query: 54 HLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLW-TSPILSTA 112
L +H DLW + I S
Sbjct: 326 PLER-----------------------------------------VHCDLWGPTTITSVQ 344
Query: 113 GHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDND 172
G RYY + +D ++ F W++P+ KS Y+IF L++ Q S I QCD G E+ +
Sbjct: 345 GFRYYAVFIDHYSRFSWIYPLKLKSDFYNIFLAFHKLVENQLSQKISVFQCDGGGEFVSH 404
Query: 173 SFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMAT 232
F ++ ++G+ + SCPHT QNG +ERK R + + ++L S VP FW A A
Sbjct: 405 KFLQHLQSHGIQQQLSCPHTPQQNGLAERKHRHLVELGLSMLFQSHVPHKFWVEAFFTAN 464
Query: 233 YLLNILPRKTLRND-SPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFL 291
+L+N+LP L+ SP ++LY + P Y+ LR FG CFP NK P S CVFL
Sbjct: 465 FLINLLPTSALKESISPYEKLYDKKPDYTSLRSFGSACFPTLRDYAENKFNPCSLKCVFL 524
Query: 292 GYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFA----------------------DLP 329
GY ++GY+C +L ISRHVIFDES +PF+ D P
Sbjct: 525 GYNEKYKGYRCLYPPTGRLYISRHVIFDESVYPFSHTYKHLHPQPRTPLLAAWLRSSDSP 584
Query: 330 LESTSSYDC-----FTE-DLPP----------------SLIHHWQTTSTRPPDLSIPPSS 367
STS+ FT D PP S+ H T+ + PD ++
Sbjct: 585 APSTSTSPSSRSPLFTSADFPPLPQRKTPLLPTLVPISSVSHASNITTQQSPDFDSERTT 644
Query: 368 PTDSTMPSPAPTSSPTSS--------APLPLPPVPPTPPTRTMTTHSMHGISKPKKPFSL 419
DS + SS S A + + + M T + GISKP +
Sbjct: 645 DFDSASIGDSSHSSQAGSDSEETIQQASVNVHQTHASTNVHPMVTRAKVGISKPNPRY-- 702
Query: 420 SVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IF 479
V + P P AL P W AM E TW LVP D++V+ +F
Sbjct: 703 -VFLSHKVSYPEPKTVTAALKHPGWTGAMTEEIGNCSETQTWSLVPYKSDMHVLGSKWVF 761
Query: 480 RHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*L 539
R K ++G KAR+V G+ Q G+D ET++ VV+ T+R VL +A + +W I +
Sbjct: 762 RTKLHADGTLNKLKARIVAKGFLQEEGIDYLETYSPVVRTPTVRLVLHLATALNWDIKQM 821
Query: 540 DVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSI 599
DV NAFLHGDL ETVYM QP GF D PD+VC KS+YGLKQ+PRAW+ +F+ ++
Sbjct: 822 DVKNAFLHGDLKETVYMTQPAGFVDPSKPDHVCLLHKSIYGLKQSPRAWFDKFSTFLLEF 881
Query: 600 GFQHSSSDHS-----------YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLS 648
GF S SD S +LLYVDD+++ +S S +A L EF M D+G L
Sbjct: 882 GFFCSKSDPSLFIYAHNNNLILLLLYVDDMVITGNSSQTLTSLLAALNKEFRMTDMGQLH 941
Query: 649 YFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTL 708
YFLGI V R GLF+SQ YA +++ A M C P TP+ + D T
Sbjct: 942 YFLGIQVQRQQNGLFMSQQKYAEDLLIAASMEHCTPLPTPLPVQLDRVPHQEELFSDPTY 1001
Query: 709 YQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTE--HMLAMF-------VAL*LTVFTC 759
++S+AG LQYLT TRP+I + V VC MH P H+L + + ++
Sbjct: 1002 FRSIAGKLQYLTLTRPDIQFAVNFVCQKMHQPTISDFHLLKRILRYIKGTITMGISYSRD 1061
Query: 760 IPPLLRNLFLIRMLTGGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVA 819
P LL+ G TRRS G C F+G NL+SWSSK+ PT+S SS EAEY+ ++
Sbjct: 1062 SPTLLQ--AYSDSDWGNCKQTRRSVGGLCTFMGTNLVSWSSKKHPTVSRSSTEAEYKSLS 1119
Query: 820 NVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKV 879
+ SE W+ LL EL PL + CDN+S++YL+ NP H RTKH ++D HFVRE+V
Sbjct: 1120 DAASEILWLSTLLRELRIPLPDTPELFCDNLSAVYLTANPAFHARTKHFDIDFHFVRERV 1179
Query: 880 ARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSL 915
A + H+P QIADIFTK LP F R L
Sbjct: 1180 ALKALVVKHIPGSEQIADIFTKSLPYEAFIHLRGKL 1215
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 551 bits (1420), Expect = e-157
Identities = 329/947 (34%), Positives = 498/947 (51%), Gaps = 59/947 (6%)
Query: 5 VSDFQTGMPLLRCNSLGDLYPVTCSSHFAGLAS----GVWHNRLGHPTSSALNHLRNNKL 60
+ D G+ L +G+LY + S + + VWH RLGHP+ S L+ L
Sbjct: 530 IQDLTKGLTLGEGKRIGNLYVLDTQSPAISVNAVVDVSVWHKRLGHPSFSRLDSLSEVLG 589
Query: 61 IYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDILHSDLWTSPILSTA-GHRYYVL 119
++ S C C L K +L F S+ I +F++LH D+W + T G++Y++
Sbjct: 590 TTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLHIDVWGPFSVETVEGYKYFLT 649
Query: 120 VLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSFRRYCH 179
++DDH+ W++ + KS V +F L++ Q+ +K ++ DN +E +F +
Sbjct: 650 IVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAKEL---AFTEFYK 706
Query: 180 ANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILP 239
A G++ SCP T QN ERK + I N+ R L+ S++ +W + A +L+N P
Sbjct: 707 AKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFLINRTP 766
Query: 240 RKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRG 299
L N +P + L + P YS L+ FGCLC+ S +K PRS CVFLGYP +G
Sbjct: 767 SALLSNKTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYPFGFKG 826
Query: 300 YKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYDCFTEDLPPSLIHHWQTTSTRPP 359
YK DL + + ISR+V F E FP A +T++ D FT P
Sbjct: 827 YKLLDLESNVVHISRNVEFHEELFPLASSQQSATTASDVFT-----------------PM 869
Query: 360 DLSIPPSSPTDSTMPSPAPTSSPTSSAP----LPLPPVPPTPPTRTMTTHSMHGISKPKK 415
D P SS T P+P SP++ P + H IS
Sbjct: 870 D---PLSSGNSITSHLPSPQISPSTQISKRRITKFPAHLQDYHCYFVNKDDSHPISSSLS 926
Query: 416 PFSLSVS----IDDPSISPLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVN 471
+S S I++ S P+P + +A W A+ E A+ R +TWE+ P
Sbjct: 927 YSQISPSHMLYINNISKIPIPQSYHEAKDSKEWCGAIDQEIGAMERTDTWEITSLPPGKK 986
Query: 472 VIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALS 531
+ +F K ++G E +KAR+V G++Q G+D ETF+ V K AT++ +L ++ S
Sbjct: 987 AVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETFSPVAKMATVKLLLKVSAS 1046
Query: 532 RSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRD----SQHPDYVCRRKKSLYGLKQAPRA 587
+ W ++ LD+ NAFL+GDL ET+YM P G+ D S P+ VCR KKS+YGLKQA R
Sbjct: 1047 KKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLKKSIYGLKQASRQ 1106
Query: 588 WYQRFADYVSSIGFQHSSSDHS-----------YILLYVDDIILVASSHDLRKSFMALLA 636
W+ +F++ + ++GF+ DH+ +L+YVDDI++ +++ +S L
Sbjct: 1107 WFLKFSNSLLALGFEKQHGDHTLFVRCIGSEFIVLLVYVDDIVIASTTEQAAQSLTEALK 1166
Query: 637 FEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLS 696
F +++LGPL YFLG+ V R G+ LSQ YA E++ A M C PS+ P+ +LS
Sbjct: 1167 ASFKLRELGPLKYFLGLEVARTSEGISLSQRKYALELLTSADMLDCKPSSIPMTPNIRLS 1226
Query: 697 SSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTV 756
+ G ED +Y+ L G L YLT TRP+I++ V ++C AP T H+ A++ L
Sbjct: 1227 KNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPRTAHLAAVYKVLQYIK 1286
Query: 757 FTCIPPLLRNL---FLIRMLT----GGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHS 809
T L + ++ T G +RRST+G+ +F+G +LISW SK+QPT+S S
Sbjct: 1287 GTVGQGLFYSAEDDLTLKGYTDADWGTCPDSRRSTTGFTMFVGSSLISWRSKKQPTVSRS 1346
Query: 810 SAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIE 869
SAEAEYR +A E W+ LLL L S +++ D+ +++Y++ NPV H+RTKHIE
Sbjct: 1347 SAEAEYRALALASCEMAWLSTLLLALRVH-SGVPILYSDSTAAVYIATNPVFHERTKHIE 1405
Query: 870 MDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSLS 916
+D H VREK+ GQ ++LHV ++ Q+ADI TK L F S +S
Sbjct: 1406 IDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQFAHLLSKMS 1452
>At1g70010 hypothetical protein
Length = 1315
Score = 531 bits (1367), Expect = e-151
Identities = 321/904 (35%), Positives = 474/904 (51%), Gaps = 54/904 (5%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
+WH RLGHP+ L + + + + C C + K LPF S + R FD+
Sbjct: 406 LWHKRLGHPSVQKLQPMSSLLSFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDL 465
Query: 99 LHSDLWTSPILSTA-GHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
+H D W + T G+RY++ ++DD++ W++ + KS V + T T+++ QF
Sbjct: 466 IHIDTWGPFSVQTHDGYRYFLTIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETT 525
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
IK ++ DN E + F ++ H+ G++ SCP T QN ERK + I N+ R+L S
Sbjct: 526 IKGVRSDNAPELN---FTQFYHSKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQS 582
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSAT 277
+P S+W + A YL+N LP L + P + L P+Y H++VFGCLC+
Sbjct: 583 HIPISYWGDCILTAVYLINRLPAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKD 642
Query: 278 INKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYD 337
+K PR+ C F+GYP +GYK DL +I+SRHV+F E FPF L S +
Sbjct: 643 RHKFSPRAKACAFIGYPSGFKGYKLLDLETHSIIVSRHVVFHEELFPFLGSDL-SQEEQN 701
Query: 338 CFTEDLPPSLIHHWQTTSTRPPDLS-----IPPSSPTDSTMPSPAPTSSPTSSAPLPLPP 392
F + P + + P D S +P ++PT++ +P P S TS P
Sbjct: 702 FFPDLNPTPPMQRQSSDHVNPSDSSSSVEILPSANPTNN-VPEP---SVQTSHRKAKKPA 757
Query: 393 VPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPC--------NPKQALSDPNW 444
++ + + H I K F I+DP ++ L C N +A W
Sbjct: 758 YLQDYYCHSVVSSTPHEIRK----FLSYDRINDPYLTFLACLDKTKEPSNYTEAEKLQVW 813
Query: 445 KFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQI 504
+ AM EF+ L +TWE+ P D I IF+ K S+G E YKARLV G++Q
Sbjct: 814 RDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQK 873
Query: 505 AGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFR- 563
G+D +ETF+ V K +++ +L +A + LD+ NAFL+GDL E +YM P G+
Sbjct: 874 EGIDYNETFSPVAKLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYAS 933
Query: 564 ---DSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS----------- 609
DS P+ VCR KKSLYGLKQA R WY +F+ + +GF S DH+
Sbjct: 934 RQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKISDGIFL 993
Query: 610 YILLYVDDIILVASSH---DLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQ 666
+L+Y+DDII+ +++ D+ KS M F ++DLG L YFLG+ + R G+ +SQ
Sbjct: 994 CVLVYIDDIIIASNNDAAVDILKSQMKSF---FKLRDLGELKYFLGLEIVRSDKGIHISQ 1050
Query: 667 STYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNI 726
YA +++ G C PS+ P+D + SG +V Y+ L G L YL TRP+I
Sbjct: 1051 RKYALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDI 1110
Query: 727 SYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNL-----FLIRMLTGGMSC-- 779
++ V ++ AP H+ A++ L T L + + SC
Sbjct: 1111 TFAVNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQGLFYSATSELQLKVYANADYNSCRD 1170
Query: 780 TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPL 839
+RRSTSGYC+FLGD+LI W S++Q +S SSAEAEYR ++ E W+ N L EL PL
Sbjct: 1171 SRRSTSGYCMFLGDSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPL 1230
Query: 840 SQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIF 899
S+ TL+ CDN ++I+++ N V H+RTKHIE D H VRE++ +G + H+ + QIAD F
Sbjct: 1231 SKPTLLFCDNEAAIHIANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPF 1290
Query: 900 TKGL 903
TK L
Sbjct: 1291 TKPL 1294
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 514 bits (1325), Expect = e-146
Identities = 312/920 (33%), Positives = 468/920 (49%), Gaps = 73/920 (7%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
+WH RLGH + L+ + ++ ++ S C C L K +L F +S + FD+
Sbjct: 557 MWHRRLGHASLQRLDAISDSLGTTRHKNKGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDL 616
Query: 99 LHSDLWTSPILSTA-GHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSAN 157
LH D+W + T G++Y++ ++DDH+ WM+ + KS+V +F ++ Q+
Sbjct: 617 LHIDVWGPFSVETVEGYKYFLTIVDDHSRATWMYLLKTKSEVLTVFPAFIQQVENQYKVK 676
Query: 158 IKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHS 217
+K ++ DN E F + G++ SCP T QN ERK + I N+ R L+ S
Sbjct: 677 VKAVRSDNAPEL---KFTSFYAEKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQS 733
Query: 218 SVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSAT 277
VP S W + A +L+N P + L N +P + L P Y LR FGCLC+
Sbjct: 734 QVPLSLWGDCVLTAVFLINRTPSQLLMNKTPYEILTGTAPVYEQLRTFGCLCYSSTSPKQ 793
Query: 278 INKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYD 337
+K QPRS C+FLGYP ++GYK DL + + ISR+V F E FP A P S SS
Sbjct: 794 RHKFQPRSRACLFLGYPSGYKGYKLMDLESNTVFISRNVQFHEEVFPLAKNP-GSESSLK 852
Query: 338 CFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLPPVPP-- 395
FT +P S S T SP SS P + +PP
Sbjct: 853 LFTPMVPVS-------------------SGIISDTTHSP-------SSLPSQISDLPPQI 886
Query: 396 ------TPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSIS-----------PLPCNPKQA 438
PP H S K P S ++S S S P+P N +A
Sbjct: 887 SSQRVRKPPAHLNDYHCNTMQSDHKYPISSTISYSKISPSHMCYINNITKIPIPTNYAEA 946
Query: 439 LSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVG 498
W A+ E A+ + NTWE+ P + +F K ++G E YKARLV
Sbjct: 947 QDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGCKWVFTLKFLADGNLERYKARLVA 1006
Query: 499 DGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQ 558
G++Q G+D +TF+ V K TI+ +L ++ S+ W + LDV NAFL+G+L E ++M
Sbjct: 1007 KGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKKWFLKQLDVSNAFLNGELEEEIFMKI 1066
Query: 559 PLGFRDSQ----HPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS----- 609
P G+ + + + V R K+S+YGLKQA R W+++F+ + S+GF+ + DH+
Sbjct: 1067 PEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWFKKFSSSLLSLGFKKTHGDHTLFLKM 1126
Query: 610 ------YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLF 663
+L+YVDDI++ ++S L F ++DLG L YFLG+ V R G+
Sbjct: 1127 YDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQRFKLRDLGDLKYFLGLEVARTTAGIS 1186
Query: 664 LSQSTYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTR 723
+ Q YA E++ GM +C P + P+ K+ G ED+ Y+ + G L YLT TR
Sbjct: 1187 ICQRKYALELLQSTGMLACKPVSVPMIPNLKMRKDDGDLIEDIEQYRRIVGKLMYLTITR 1246
Query: 724 PNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLTG-----GMS 778
P+I++ V ++C AP T H+ A + L T L + L G S
Sbjct: 1247 PDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTVGQGLFYSASSDLTLKGFADSDWAS 1306
Query: 779 C--TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELH 836
C +RRST+ + +F+GD+LISW SK+Q T+S SSAEAEYR +A E W+ LL+ L
Sbjct: 1307 CQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEAEYRALALATCEMVWLFTLLVSLQ 1366
Query: 837 FPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIA 896
+++ D+ ++IY++ NPV H+RTKHI++D H VRE++ G+ ++LHV + Q+A
Sbjct: 1367 -ASPPVPILYSDSTAAIYIATNPVFHERTKHIKLDCHTVRERLDNGELKLLHVRTEDQVA 1425
Query: 897 DIFTKGLPRVLFDDFRSSLS 916
DI TK L F+ +S +S
Sbjct: 1426 DILTKPLFPYQFEHLKSKMS 1445
>At4g17450 retrotransposon like protein
Length = 1433
Score = 509 bits (1310), Expect = e-144
Identities = 307/894 (34%), Positives = 454/894 (50%), Gaps = 60/894 (6%)
Query: 37 SGVWHNRLGHPTSSALNHLRNN-----KLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETI 91
S WH RLGHP S ++ L + K I + S VC C L K L F S + +
Sbjct: 550 SVTWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHLSFQSRQNM 609
Query: 92 TLRSFDILHSDLWTSPILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
+FD++H D W + T D W++ + KS V +F ++
Sbjct: 610 CSAAFDLVHIDTWGPFSVPT-------------NDATWIYLLKNKSDVLHVFPAFINMVH 656
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
TQ+ +K ++ DN E F A+G++ SCP T QN ERK + I N+ R
Sbjct: 657 TQYQTKLKSVRSDNAHEL---KFTDLFAAHGIVAYHSCPETPEQNSVVERKHQHILNVAR 713
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
LL S++P FW + A +L+N LP L N SP ++L + P+Y L+ FGCLC+
Sbjct: 714 ALLFQSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYESLKTFGCLCYS 773
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLE 331
+K +PR+ CVFLGYP+ ++GYK D+ + ISRHVIF E FPF +
Sbjct: 774 STSPKQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHEDIFPF----IS 829
Query: 332 STSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP 391
ST D +D P L R DL + +S D T P +SS PL
Sbjct: 830 STIKDD--IKDFFPLL-----QFPARTDDLPLEQTSIID-THPHQDVSSSKALVPFDPLS 881
Query: 392 PVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPE 451
PP H + ++P F I++ + + +P +A W AM+ E
Sbjct: 882 KRQKKPPKHLQDFHCYNNTTEPFHAF-----INNITNAVIPQRYSEAKDFKAWCDAMKEE 936
Query: 452 FNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDE 511
A++R NTW +V P + I +F K ++G E YKARLV G++Q G+D +E
Sbjct: 937 IGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGYTQEEGLDYEE 996
Query: 512 TFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRD----SQH 567
TF+ V K ++R +L +A W +H LD+ NAFL+GDL E +YM P G+ D +
Sbjct: 997 TFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVGEALP 1056
Query: 568 PDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSY-----------ILLYVD 616
P +CR KS+YGLKQA R WY + ++ + +GFQ S++DH+ +L+YVD
Sbjct: 1057 PHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKYANGVLMGVLVYVD 1116
Query: 617 DIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIAR 676
DI++V++S D F A L F ++DLG YFLGI + R G+ + Q Y E+++
Sbjct: 1117 DIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKGISICQRKYILELLST 1176
Query: 677 AGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLH 736
G PS+ P+D KL+ G P D T Y+ L G L YL TRP+I+Y V +C
Sbjct: 1177 TGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVNTLCQF 1236
Query: 737 MHAPHTEHMLAMFVAL*LTVFTCIPPLLRNL---FLIRMLT----GGMSCTRRSTSGYCV 789
HAP + H+ A+ L T L + F +R T G + +RR + YC+
Sbjct: 1237 SHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDSDFGSCTDSRRCVAAYCM 1296
Query: 790 FLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDN 849
F+GD L+SW SK+Q T+S S+AEAE+R ++ E W+ L + P ++CDN
Sbjct: 1297 FIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDDFKVPFIPPAYLYCDN 1356
Query: 850 VSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGL 903
+++++ N V H+RTK +E+D + RE V G + + V + Q+AD TK +
Sbjct: 1357 TAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETGEQVADPLTKAI 1410
>At1g48710 hypothetical protein
Length = 1352
Score = 493 bits (1269), Expect = e-139
Identities = 302/902 (33%), Positives = 456/902 (50%), Gaps = 50/902 (5%)
Query: 37 SGVWHNRLGHPTSSALNHLRNNKLIYCDP--SRSSTVCDSCVLGKHVRLPF-SSSETITL 93
S +WH R GH L L +++ P + + VC+ C+LGK ++ F S +
Sbjct: 465 SWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQFKMSFPKESSSRAQ 524
Query: 94 RSFDILHSDLWTSPIL--STAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
+S +++H+D+ PI S Y++L +DD + W++ + +KS+V++IF ++
Sbjct: 525 KSLELIHTDV-CGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSEVFEIFKKFKAHVE 583
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+ IK ++ D G E+ + F +YC NG+ + + P + QNG +ERK RTI M R
Sbjct: 584 KESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGVAERKNRTILEMAR 643
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
++L +P W A+ A YLLN P K++ +P + R SHLRVFG +
Sbjct: 644 SMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKSGVSHLRVFGSIAHA 703
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLE 331
P +KL +S +F+GY N +GYK Y+ +K IISR+++FDE + E
Sbjct: 704 HVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVFDEEGEWDWNSNEE 763
Query: 332 STSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP 391
+ + F ED P PT PS PT+ PTS +
Sbjct: 764 DYNFFPHFEEDEP----------------------EPTREEPPSEEPTTPPTSPTSSQIE 801
Query: 392 PVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPE 451
R + ++ +++ ++ +L + P + ++A+ W+ AM E
Sbjct: 802 ESSSERTPRFRSIQELYEVTENQENLTLFCLFAECE----PMDFQEAIEKKTWRNAMDEE 857
Query: 452 FNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDE 511
++ +N+TWEL P I +++ KK S G E YKARLV G+ Q AG+D DE
Sbjct: 858 IKSIQKNDTWELTSLPNGHKTIGVKWVYKAKKNSKGEVERYKARLVAKGYIQRAGIDYDE 917
Query: 512 TFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYV 571
F V + T+R ++S+A W IH +DV +AFL+GDL E VY+ QP G+ D V
Sbjct: 918 VFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKV 977
Query: 572 CRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSYIL-----------LYVDDIIL 620
R KK+LYGLKQAPRAW R Y F +H+ + LYVDD+I
Sbjct: 978 LRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIF 1037
Query: 621 VASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMA 680
++ + + F + EF M D+G +SY+LGI V + G+F++Q YA E++ + M
Sbjct: 1038 TGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMD 1097
Query: 681 SCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAP 740
NP TP++ KLS D T ++SL G+L+YLT TRP+I Y V V +M P
Sbjct: 1098 DSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHP 1157
Query: 741 HTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLT-------GGMSCTRRSTSGYCVFLGD 793
T H A L T L + L GG R+STSG+ ++GD
Sbjct: 1158 TTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGD 1217
Query: 794 NLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSI 853
+W SK+QP + S+ EAEY + V + W+RNLL EL P + T + DN S+I
Sbjct: 1218 TAFTWMSKKQPIVVLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAI 1277
Query: 854 YLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRS 913
L+ NPV H R+KHI+ H++RE V++ ++ +V + Q+ADIFTK L R F RS
Sbjct: 1278 ALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIFTKPLKREDFIKMRS 1337
Query: 914 SL 915
L
Sbjct: 1338 LL 1339
>At3g61330 copia-type polyprotein
Length = 1352
Score = 491 bits (1263), Expect = e-138
Identities = 300/902 (33%), Positives = 454/902 (50%), Gaps = 50/902 (5%)
Query: 37 SGVWHNRLGHPTSSALNHLRNNKLIYCDP--SRSSTVCDSCVLGKHVRLPF-SSSETITL 93
S +WH R GH L L +++ P + + VC+ C+LGK ++ F S +
Sbjct: 465 SWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQFKMSFPKESSSRAQ 524
Query: 94 RSFDILHSDLWTSPIL--STAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
+ +++H+D+ PI S Y++L +DD + W++ + +KS+V++IF ++
Sbjct: 525 KPLELIHTDV-CGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSEVFEIFKKFKAHVE 583
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+ IK ++ D G E+ + F +YC NG+ + + P + QNG ERK RTI M R
Sbjct: 584 KESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGVVERKNRTILEMAR 643
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
++L +P W A+ A YLLN P K++ +P + R P SHLRVFG +
Sbjct: 644 SMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKPGVSHLRVFGSIAHA 703
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLE 331
P +KL +S +F+GY N +GYK Y+ +K IISR+++FDE + E
Sbjct: 704 HVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVFDEEGEWDWNSNEE 763
Query: 332 STSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP 391
+ + F ED P PT PS PT+ PTS +
Sbjct: 764 DYNFFPHFEEDEP----------------------EPTREEPPSEEPTTPPTSPTSSQIE 801
Query: 392 PVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPE 451
R + ++ +++ ++ +L + P + ++A+ W+ AM E
Sbjct: 802 ESSSERTPRFRSIQELYEVTENQENLTLFCLFAECE----PMDFQKAIEKKTWRNAMDEE 857
Query: 452 FNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDE 511
++ +N+TWEL P I +++ KK S G E YKARLV G+SQ G+D DE
Sbjct: 858 IKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRVGIDYDE 917
Query: 512 TFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYV 571
F V + T+R ++S+A W IH +DV +AFL+GDL E VY+ QP G+ D V
Sbjct: 918 VFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKV 977
Query: 572 CRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSYIL-----------LYVDDIIL 620
R KK LYGLKQAPRAW R Y F +H+ + LYVDD+I
Sbjct: 978 LRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIF 1037
Query: 621 VASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMA 680
++ + + F + EF M D+G +SY+LGI V + G+F++Q YA E++ + +
Sbjct: 1038 TGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKID 1097
Query: 681 SCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAP 740
NP TP++ KLS D T ++SL G+L+YLT TRP+I Y V V +M P
Sbjct: 1098 DSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHP 1157
Query: 741 HTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLT-------GGMSCTRRSTSGYCVFLGD 793
T H A L T L + L GG R+STSG+ ++GD
Sbjct: 1158 TTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGD 1217
Query: 794 NLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSI 853
+W SK+QP ++ S+ EAEY + V + W+RNLL EL P + T + DN S+I
Sbjct: 1218 TAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAI 1277
Query: 854 YLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRS 913
L+ NPV H R+KHI+ H++RE V++ ++ +V + Q+AD FTK L R F RS
Sbjct: 1278 ALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADFFTKPLKRENFIKMRS 1337
Query: 914 SL 915
L
Sbjct: 1338 LL 1339
>At1g58140 hypothetical protein
Length = 1320
Score = 490 bits (1261), Expect = e-138
Identities = 303/902 (33%), Positives = 448/902 (49%), Gaps = 82/902 (9%)
Query: 37 SGVWHNRLGHPTSSALNHLRNNKLIYCDP--SRSSTVCDSCVLGKHVRLPF-SSSETITL 93
S +WH R GH L L +++ P + + VC+ C+LGK ++ F S +
Sbjct: 465 SWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQFKMSFPKESSSRAQ 524
Query: 94 RSFDILHSDLWTSPIL--STAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
+ +++H+D+ PI S Y++L +DD + W++ + +KS+V++IF ++
Sbjct: 525 KPLELIHTDV-CGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSEVFEIFKKFKAHVE 583
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+ IK ++ D G E+ + F +YC NG+ + + P + QNG +ERK RTI M R
Sbjct: 584 KESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGVAERKNRTILEMAR 643
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
++L +P W A+ A YLLN P K++ +P + R P SHLRVFG +
Sbjct: 644 SMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKPGVSHLRVFGSIAHA 703
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLE 331
P +KL +S +F+GY N +GYK Y+ +K IISR+++FDE + E
Sbjct: 704 HVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVFDEEGEWDWNSNEE 763
Query: 332 STSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP 391
+ + F ED P PT PS PT+
Sbjct: 764 DYNFFPHFEEDKP----------------------EPTREEPPSEEPTT----------- 790
Query: 392 PVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPE 451
PPT PT + P + ++A+ W+ AM E
Sbjct: 791 --PPTSPTSSQIEEKCE-----------------------PMDFQEAIEKKTWRNAMDEE 825
Query: 452 FNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDE 511
++ +N+TWEL P I +++ KK S G E YKARLV G+SQ AG+D DE
Sbjct: 826 IKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRAGIDYDE 885
Query: 512 TFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYV 571
F V + T+R ++S+A W IH +DV +AFL+GDL E VY+ QP G+ D V
Sbjct: 886 VFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKV 945
Query: 572 CRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSYIL-----------LYVDDIIL 620
R KK+LYGLKQAPRAW R Y F +H+ + LYVDD+I
Sbjct: 946 LRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIF 1005
Query: 621 VASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMA 680
++ + + F + EF M D+G +SY+LGI V + G+F++Q YA E++ + M
Sbjct: 1006 TGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMD 1065
Query: 681 SCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAP 740
NP TP++ KLS D T ++SL G+L+YLT TRP+I Y V V +M P
Sbjct: 1066 DSNPVCTPMECGIKLSKKEEGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHP 1125
Query: 741 HTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLT-------GGMSCTRRSTSGYCVFLGD 793
T H A L T L + L GG R+STSG+ ++GD
Sbjct: 1126 TTTHFKAAKRILRYIKGTVNFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGD 1185
Query: 794 NLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSI 853
+W SK+QP ++ S+ EAEY + V + W+RNLL EL P + T + DN S+I
Sbjct: 1186 TAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAI 1245
Query: 854 YLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRS 913
L+ NPV H R+KHI+ H++RE V++ ++ +V + Q+ADIFTK L R F RS
Sbjct: 1246 ALAKNPVFHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIFTKPLKREDFIKMRS 1305
Query: 914 SL 915
L
Sbjct: 1306 LL 1307
>At4g14460 retrovirus-related like polyprotein
Length = 1489
Score = 481 bits (1239), Expect = e-136
Identities = 319/950 (33%), Positives = 455/950 (47%), Gaps = 143/950 (15%)
Query: 25 PVTCSSHFAGLASG-VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRL 83
P C + L G +WH RLGHP+S L L+ RL
Sbjct: 586 PAACLFTGSVLNDGHLWHQRLGHPSSVVLQKLK-------------------------RL 620
Query: 84 PFSSSETITLRSFDILHSDLWTS-PILSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDI 142
+ S + FD++H D+W I S G RY++ V+DD T W++ + K V +
Sbjct: 621 AYISHNNLASNPFDLVHLDIWGPFSIESIEGFRYFLTVVDDCTRTTWVYMLRNKKDVSSV 680
Query: 143 FTTLATLIKTQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERK 202
F L+ TQF+A IK ++ DN E F +G++ FSC +T QN ERK
Sbjct: 681 FPEFIKLVSTQFNAKIKAIRSDNAPEL---GFTEIVKEHGMLHHFSCAYTPQQNSVVERK 737
Query: 203 IRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHL 262
+ I N+ R LL S++P +W + A +L+N LP L N SP + + ++ P YS L
Sbjct: 738 HQHILNVARALLFQSNIPMQYWSDCVTTAVFLINRLPSPLLNNKSPYELILNKQPDYSLL 797
Query: 263 RVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESR 322
+ FGCLCF + K PR+ CVFLGYP ++GYK DL + + +SR+V+F E
Sbjct: 798 KNFGCLCFVSTNAHERTKFTPRARACVFLGYPSGYKGYKVLDLESHSVTVSRNVVFKEHV 857
Query: 323 FPFADLPLESTSSYDCFTEDLPP-----------------SLI----------HHWQTTS 355
FPF L + + D F + P SLI +H ++S
Sbjct: 858 FPFKTSELLN-KAVDMFPNSILPLPAPLHFVETMPLIDEDSLIPTTTDSRTADNHASSSS 916
Query: 356 TRPPDLSIPPSSPTDS----TMPSPAPTSSPTSSAPLPL--------PPVPPTPPT-RTM 402
+ P + IPPSS T++ + P S T+ AP L P + PPT ++
Sbjct: 917 SALPSI-IPPSSNTETQDIDSNAVPITRSKRTTRAPSYLSEYHCSLVPSISTLPPTDSSI 975
Query: 403 TTHSMHGI---SKPKK--PFSLSVSIDDPSISPL-------------PCNPKQALSDPNW 444
H + I S PKK P+ +S + +PL P QA+ W
Sbjct: 976 PIHPLPEIFTASSPKKTTPYPISTVVSYDKYTPLCQSYIFAYNTETEPKTFSQAMKSEKW 1035
Query: 445 KFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQI 504
E A+ N TW + P D NV+ +F K +G E YKARLV G++Q
Sbjct: 1036 IRVAVEELQAMELNKTWSVESLPPDKNVVGCKWVFTIKYNPDGTVERYKARLVAQGFTQQ 1095
Query: 505 AGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRD 564
G+D +TF+ V K + + +L +A W + +DV +AFLHGDL E ++M P G+
Sbjct: 1096 EGIDFLDTFSPVAKLTSAKMMLGLAAITGWTLTQMDVSDAFLHGDLDEEIFMSLPQGYTP 1155
Query: 565 SQ----HPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSYILLYVDDIIL 620
P+ VCR KS+YGLKQA R WY+RF L+Y+DDI++
Sbjct: 1156 PAGTILPPNPVCRLLKSIYGLKQASRQWYKRFV----------------AALVYIDDIMI 1199
Query: 621 VASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMA 680
+++ ++ ALL EF +KDLGP +FL G+
Sbjct: 1200 ASNNDAEVENLKALLRSEFKIKDLGPARFFL--------------------------GLL 1233
Query: 681 SCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAP 740
C PS+ P+D L GTP + T Y+ L G L YLT TRP+I+Y V Q+ + AP
Sbjct: 1234 GCKPSSIPMDPTLHLVRDMGTPLPNPTAYRKLIGRLLYLTITRPDITYAVHQLSQFISAP 1293
Query: 741 HTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLTG-------GMSCTRRSTSGYCVFLGD 793
H+ A L L+ + L G TRRS SG+C++LG
Sbjct: 1294 SDIHLQAAHKVLRYIKANPGQGLMYSADYEICLNGFSDADWAACKDTRRSISGFCIYLGT 1353
Query: 794 NLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSI 853
+LISW SK+Q S SS E+EYR +A E W++ LL +LH PL+ + CDN S++
Sbjct: 1354 SLISWKSKKQAVASRSSTESEYRSMAQATCEIIWLQQLLKDLHIPLTCPAKLFCDNKSAL 1413
Query: 854 YLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGL 903
+ S NPV H+RTKHIE+D H VR+++ G + LHVP+ +Q ADI TK L
Sbjct: 1414 HSSLNPVFHERTKHIEIDCHTVRDQIKAGNLKALHVPTENQHADILTKAL 1463
>At3g60170 putative protein
Length = 1339
Score = 457 bits (1176), Expect = e-128
Identities = 287/907 (31%), Positives = 458/907 (49%), Gaps = 41/907 (4%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSST--VCDSCVLGKHVRLPFSSSETI-TLRS 95
+WH R GH L L + K++ P +T +C C+ GK R S + +
Sbjct: 435 LWHCRFGHLNQEGLKLLAHKKMVIGLPILKATKEICAICLTGKQHRESMSKKTSWKSSTQ 494
Query: 96 FDILHSDLWTSPI--LSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQ 153
++HSD+ PI +S +G RY + +DD T W++ + +KS+ + F ++ +
Sbjct: 495 LQLVHSDI-CGPITPISHSGKRYILSFIDDFTRKTWVYFLHEKSEAFATFKIFKASVEKE 553
Query: 154 FSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTL 213
A + CL+ D G E+ ++ F +C ++G+ + + T QNG +ERK RTI N +R++
Sbjct: 554 IGAFLTCLRTDRGGEFTSNEFGEFCRSHGISRQLTAAFTPQQNGVAERKNRTIMNAVRSM 613
Query: 214 LAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFF 273
L+ VP FW A + + ++ N P + +P + R P + RVFGC+ +
Sbjct: 614 LSERQVPKMFWSEATKWSVHIQNRSPTAAVEGMTPEEAWSGRKPVVEYFRVFGCIGYVHI 673
Query: 274 PSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESR---FPFADLPL 330
P +KL +S CVFLG + ++ YD +K++IS+ V+FDE + + AD+
Sbjct: 674 PDQKRSKLDDKSKKCVFLGVSEESKAWRLYDPVMKKIVISKDVVFDEDKSWDWDQADVEA 733
Query: 331 ESTSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPL 390
+ + +C ED + P ++ P +D+ + S +P +P+S AP P+
Sbjct: 734 KEVT-LECGDED------DEKNSEVVEPIAVASPNHVGSDNNVSS-SPILAPSSPAPSPV 785
Query: 391 PP--VPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAM 448
P M + + ++ S+ + + P+ + A+ D W+ AM
Sbjct: 786 AAKVTRERRPPGWMADYETGEGEEIEENLSVMLLMMMTEADPIQFD--DAVKDKIWREAM 843
Query: 449 QPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVD 508
+ E ++++NNTWEL P I +++ K +G + YKARLV G++Q G+D
Sbjct: 844 EHEIESIVKNNTWELTTLPKGFTPIGVKWVYKTKLNEDGEVDKYKARLVAKGYAQCYGID 903
Query: 509 CDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHP 568
E F V + T+RT+L+I+ +W I LDV +AFLHG+L E VY+ QP GF
Sbjct: 904 YTEVFAPVARLDTVRTILAISSQFNWEIFQLDVKSAFLHGELKEEVYVRQPEGFIREGEE 963
Query: 569 DYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS-----------YILLYVDD 617
+ V + +K+LYGLKQAPRAWY R Y F+ S+H+ + LYVDD
Sbjct: 964 EKVYKLRKALYGLKQAPRAWYSRIEAYFLKEEFERCPSEHTLFTKTRVGNILIVSLYVDD 1023
Query: 618 IILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARA 677
+I S + F + EF M DLG + +FLGI V + GG+F+ Q YA E++AR
Sbjct: 1024 LIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHFLGIEVKQSDGGIFICQRRYAREVLARF 1083
Query: 678 GMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHM 737
GM N P+ KL+ D T+++ L G+L YLT TRP++ Y V + M
Sbjct: 1084 GMDESNAVKNPIVPGTKLTKDENGEKVDETMFKQLVGSLMYLTVTRPDLMYGVCLISRFM 1143
Query: 738 HAPHTEHMLA---MFVAL*LTVFTCIPPLLRNLFLIRMLT------GGMSCTRRSTSGYC 788
P H LA + L TV I R ++++ G RRSTSG+
Sbjct: 1144 SNPRMSHWLAAKRILRYLKGTVELGIFYRRRKNRSLKLMAFTDSDYAGDLNDRRSTSGFV 1203
Query: 789 VFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCD 848
+ I W+SK+QP ++ S+ EAEY A + W+R +L +L T+++CD
Sbjct: 1204 FLMASGAICWASKKQPVVALSTTEAEYIAAAFCACQCVWLRKVLEKLGAEEKSATVINCD 1263
Query: 849 NVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLF 908
N S+I LS +PV H ++KHIE+ H++R+ V ++ + P+ Q+ADIFTK L F
Sbjct: 1264 NSSTIQLSKHPVLHGKSKHIEVRFHYLRDLVNGDVVKLEYCPTEDQVADIFTKPLKLEQF 1323
Query: 909 DDFRSSL 915
+ R+ L
Sbjct: 1324 EKLRALL 1330
>At3g25450 hypothetical protein
Length = 1343
Score = 447 bits (1151), Expect = e-125
Identities = 284/911 (31%), Positives = 447/911 (48%), Gaps = 49/911 (5%)
Query: 37 SGVWHNRLGHPTSSALNHLRNNKLIYCDPS---RSSTVCDSCVLGKHVRLPFSSSETI-T 92
S +WH RLGH + + + +L+ S + C SC+ GK R F + +
Sbjct: 422 STIWHARLGHISFETIKAMIKKELVIGISSSVPQEKETCGSCLFGKQARHSFPKATSYRA 481
Query: 93 LRSFDILHSDLWTSPILSTAGHRYYVLVL-DDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
+ +++H DL STA + YV VL DDH+ ++W + +KS+ + F L++
Sbjct: 482 AQVLELIHGDLCGPISPSTAAKKRYVFVLIDDHSRYMWSILLKEKSEAFGKFKEFKALVE 541
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+ A IK + D G E+ + F+ +C G+ + P+T QNG ER+ RT+ M R
Sbjct: 542 QECGAIIKTFRTDRGGEFLSHEFQEFCAKEGINRHLTAPYTPQQNGVVERRNRTLLGMTR 601
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
++L H ++P W A++ +TYL+N + ++L N +P + H+ P+ HLRVFGC+ +
Sbjct: 602 SILKHMNMPNYLWGEAVRHSTYLINRVGTRSLSNQTPYEVFKHKKPNVEHLRVFGCVSYA 661
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLE 331
+ KL RS V+LG + Y+ D + R++ +SR V+FDE+R + + E
Sbjct: 662 KVEVPNLKKLDDRSRMLVYLGTEPGSKAYRLLDPTKRRIFVSRDVVFDENR---SWMWQE 718
Query: 332 STSSYDCFTEDLPPSLIHHWQT------TSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSS 385
S+S D + +L ST P + + D + A T S
Sbjct: 719 SSSETDKESGTFTITLSEFGNNGVTENDISTEPEETEEAEINGEDENIIEEAETEEHDQS 778
Query: 386 APLPLPPVPPTPPTRTMTTHSMHGISKPK--KPFSLSVSIDDP----SISPLPCNPKQAL 439
P P S + +P K + L I+ +++ P + K+A
Sbjct: 779 QEEPQP-----------VRRSQRQVIRPNYLKDYVLCAEIEAEHLLLAVNDEPWDFKEAN 827
Query: 440 SDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGD 499
W+ A + E ++ +N TW LV P I +F+ K S+G YKARLV
Sbjct: 828 KSKEWRDACKEEIQSIEKNRTWSLVDLPVGSKAIGVKWVFKLKHNSDGSINKYKARLVAK 887
Query: 500 GWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQP 559
G+ Q GVD +E F V + T+R ++++A S W IH LDV AFLHG+L E VY+ QP
Sbjct: 888 GYVQRHGVDFEEVFAPVARIETVRLIIALAASNGWEIHHLDVKTAFLHGELREDVYVSQP 947
Query: 560 LGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS---------- 609
GF + + + V + K+LYGL+QAPRAW + + + + F+ + S
Sbjct: 948 EGFTNKESKEKVYKLHKALYGLRQAPRAWNTKLNEILKELKFEKCHKEPSLYRKQEGENI 1007
Query: 610 -YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQST 668
+ +YVDD+++ S+ D+ +F + +F M DLG L+Y+LGI V + G+ L Q
Sbjct: 1008 LVVAVYVDDLLVTGSNLDIILNFKKGMVGKFEMSDLGKLTYYLGIEVLQSKDGITLKQER 1067
Query: 669 YASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISY 728
YA +I+ AGM+ CN TP+ +LS + D T Y+ G L+YL TRP++SY
Sbjct: 1068 YAKKILEEAGMSKCNTVNTPMIASLELSKAQDEKRIDETDYRRNIGCLRYLLHTRPDLSY 1127
Query: 729 VVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLL----RNLFLIRMLTGGMSC---TR 781
V + ++ P H A+ L T L N LI +
Sbjct: 1128 NVGILSRYLQEPRESHGAALKQILRYLQGTTSHGLYFKKGENAGLIGYSDSSHNVDLDDG 1187
Query: 782 RSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQ 841
+ST G+ +L D I+W S++Q ++ SS EAE+ ++ W++ LL E+ +
Sbjct: 1188 KSTGGHIFYLNDCPITWCSQKQQVVTLSSCEAEFMAATEAAKQAIWLQELLAEVIGTECE 1247
Query: 842 ETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTK 901
+ + DN S+I L+ NPV H R+KHI HF+RE V GQ + HVP Q ADI TK
Sbjct: 1248 KVTIRVDNKSAIALTKNPVFHGRSKHIHRRYHFIRECVENGQIEVEHVPGVRQKADILTK 1307
Query: 902 GLPRVLFDDFR 912
L ++ F + R
Sbjct: 1308 ALGKIKFLEMR 1318
>At1g26990 polyprotein, putative
Length = 1436
Score = 437 bits (1123), Expect = e-122
Identities = 283/913 (30%), Positives = 447/913 (47%), Gaps = 75/913 (8%)
Query: 37 SGVWHNRLGHPTSSALNHLRNNKLIYCDP-SRSSTVCDSCVLGKHVRLPFSSSETITLRS 95
S +WH RLGHP+ + ++ L + ++ ++ S+ C C L K LPF S I ++
Sbjct: 555 SEMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLSKQKHLPFKSVNHIREKA 614
Query: 96 FDILHSDLWTSPILSTA-GHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQF 154
F+++H D W + T +RY++ ++DD + W++ + +KS V +F + +++TQ+
Sbjct: 615 FELVHIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLLKQKSDVLTVFPSFLKMVETQY 674
Query: 155 SANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLL 214
+ ++ DN E F G+ CP T QN ERK + + N+ R L+
Sbjct: 675 HTKVCSVRSDNAHEL---KFNELFAKEGIKADHPCPETPEQNFVVERKHQHLLNVARALM 731
Query: 215 AHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFP 274
S +P +W + A +L+N L + N++P +RL P YS L+ FGCLC+
Sbjct: 732 FQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKGKPDYSSLKAFGCLCYCSTS 791
Query: 275 SATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFAD------- 327
+ K PR+ C+FLGYPM ++GYK D+ + ISRHVIF E FPFA
Sbjct: 792 PKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRHVIFYEDIFPFASSNITDAA 851
Query: 328 -----------------LPLESTSSYDCFTEDLPPSLIH-HWQTTSTRPPDLSIPPSSPT 369
LPL +SS D S+I + STR L PS
Sbjct: 852 KDFFPHIYLPAPNNDEHLPLVQSSSDAPHNHDESSSMIFVPSEPKSTRQRKL---PSHLQ 908
Query: 370 DSTMPSPAPTSSPTSSAPLPLPPVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSIS 429
D + PT++ TS PL + S +PF ++I + +
Sbjct: 909 DFHCYNNTPTTTKTSPYPLT----------------NYISYSYLSEPFGAFINI--ITAT 950
Query: 430 PLPCNPKQALSDPNWKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLF 489
LP +A D W AM E +A +R TW + P + I K ++G
Sbjct: 951 KLPQKYSEARLDKVWNDAMGKEISAFVRTGTWSICDLPAGKVAVGCKWIITIKFLADGSI 1010
Query: 490 EHYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGD 549
E +KARLV G++Q G+D TF+ V K T++ +LS+A W +H LD+ NA L+GD
Sbjct: 1011 ERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLAPKMKWYLHQLDISNALLNGD 1070
Query: 550 LHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS 609
L E +YM P G+ + Q G + +P A + D+ + Q
Sbjct: 1071 LEEEIYMKLPPGYSEIQ-------------GQEVSPNA--KCHGDHTLFVKAQ--DGFFL 1113
Query: 610 YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTY 669
+L+YVDDI++ +++ + L+ F ++DLG +FLGI + R G+ L Q Y
Sbjct: 1114 VVLVYVDDILIASTTEAASAELTSQLSSFFQLRDLGEPKFFLGIEIARNADGISLCQRKY 1173
Query: 670 ASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYV 729
+++A + + C PS+ P++ KLS +GT ED Y+ + G LQYL TRP+I++
Sbjct: 1174 VLDLLASSDFSDCKPSSIPMEPNQKLSKDTGTLLEDGKQYRRILGKLQYLCLTRPDINFA 1233
Query: 730 VQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNL---FLIRMLTGG--MSC--TRR 782
V ++ + AP H+ A+ L T L F +R + +C TRR
Sbjct: 1234 VSKLAQYSSAPTDIHLQALHKILRYLKGTIGQGLFYGADTNFDLRGFSDSDWQTCPDTRR 1293
Query: 783 STSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQE 842
+G+ +F+G++L+SW SK+Q +S SSAEAEYR ++ E W+ +L P +
Sbjct: 1294 CVTGFAIFVGNSLVSWRSKKQDVVSMSSAEAEYRAMSVATKELIWLGYILTAFKIPFTHP 1353
Query: 843 TLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKG 902
++CDN ++++++ N V H+RTKHIE D H VRE + G + + V + +Q+AD TK
Sbjct: 1354 AYLYCDNEAALHIANNSVFHERTKHIENDCHKVRECIEAGILKTIFVRTDNQLADTLTKP 1413
Query: 903 LPRVLFDDFRSSL 915
L F + S L
Sbjct: 1414 LYPKPFRENNSKL 1426
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 432 bits (1112), Expect = e-121
Identities = 280/895 (31%), Positives = 427/895 (47%), Gaps = 116/895 (12%)
Query: 37 SGVWHNRLGHPTSSALNHLRNNKLIYCDP--SRSSTVCDSCVLGKHVRLPF-SSSETITL 93
S +WH R GH L L +++ P + + VC+ C+LG ++ F S +
Sbjct: 465 SWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGNQFKMSFPKESSSRAQ 524
Query: 94 RSFDILHSDLWTSPIL--STAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIK 151
+ +++H+D+ PI S Y++L +DD + W++ + +KS+V++IF ++
Sbjct: 525 KPLELIHTDV-CGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSEVFEIFKKFKAHVE 583
Query: 152 TQFSANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIR 211
+ IK ++ D+G E+ + F +YC NG+ + + P + QNG +ERK RTI M R
Sbjct: 584 KESGLVIKTMRSDSGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGVAERKNRTILEMAR 643
Query: 212 TLLAHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFP 271
++L +P W A+ A YLLN P K++ +P + R P SHLRVFG +
Sbjct: 644 SMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKPGVSHLRVFGSIAHA 703
Query: 272 FFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLE 331
P NKL +S +F+GY N +GYK Y+ +K IISR+++FDE + E
Sbjct: 704 HVPDEKRNKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVFDEEGEWDWNSNEE 763
Query: 332 STSSYDCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPLP 391
+ + F ED P PT PS PT+ PTS +
Sbjct: 764 DYNFFPHFEEDKP----------------------EPTREEPPSEEPTTPPTSPTSSQIE 801
Query: 392 PVPPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPE 451
R + ++ +++ ++ +L + P + ++A+ W+ AM E
Sbjct: 802 ESSSERTPRFRSIQELYEVTENQENLTLFCLFAECE----PMDFQEAIEKKTWRNAMDEE 857
Query: 452 FNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDE 511
++ +N+TWEL P I +++ KK S G E YKARLV G+SQ AG+D DE
Sbjct: 858 IKSIQKNDTWELTSLPNGHKAIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRAGIDYDE 917
Query: 512 TFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYV 571
F V + T+R ++S+A W IH +DV +AFL+GDL E VY+ QP G+ D V
Sbjct: 918 IFAPVARLETVRLIISLAAQNKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKV 977
Query: 572 CRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHSYIL-----------LYVDDIIL 620
R KK LYGLKQAPRAW R Y F +H+ + LYVDD+I
Sbjct: 978 LRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILIACLYVDDLIF 1037
Query: 621 VASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVTRYVGGLFLSQSTYASEIIARAGMA 680
++ + + F + EF M D+G +SY+LGI V + G+F++Q YA E++ + M
Sbjct: 1038 TGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMD 1097
Query: 681 SCNPSATPVDTK*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAP 740
NP SL G+L+YLT TRP+I Y V V +M P
Sbjct: 1098 DSNP--------------------------SLVGSLRYLTCTRPDILYAVGVVSRYMEHP 1131
Query: 741 HTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRMLTGGMSCTRRSTSGYCVFLGDNLISWSS 800
T H F +LR + + T + +
Sbjct: 1132 TTTH------------FKAAKRILRYI--------------KGTVNFGL----------- 1154
Query: 801 KRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPV 860
H S ++Y+ VV + W+RNLL EL P + T + DN S+I L+ NPV
Sbjct: 1155 -------HYSTTSDYK---LVVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNPV 1204
Query: 861 HHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSL 915
H R+KHI+ H++RE V++ ++ +V + Q+ADIFTK L R F RS L
Sbjct: 1205 FHDRSKHIDTRYHYIRECVSKKDVQLEYVKTHDQVADIFTKPLKREDFIKMRSLL 1259
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 428 bits (1100), Expect = e-120
Identities = 287/917 (31%), Positives = 461/917 (49%), Gaps = 50/917 (5%)
Query: 37 SGVWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSF 96
+ +WH+RLGH + + L + + + C+ CV GK R+ F+ ++ +T
Sbjct: 414 TALWHSRLGHMSQKGMEILVKKGCLRREVIKELEFCEDCVYGKQHRVSFAPAQHVTKEKL 473
Query: 97 DILHSDLWTSPI--LSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQF 154
+HSDLW SP S +Y++ +DD++ +W++ + KK + ++ F +++ Q
Sbjct: 474 AYVHSDLWGSPHNPASLGNSQYFISFVDDYSRKVWIYFLRKKDEAFEKFVEWKKMVENQS 533
Query: 155 SANIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLL 214
+K L+ DNG EY N F ++C G++ +C +T QNG +ER RTI + +R++L
Sbjct: 534 DRKVKKLRTDNGLEYCNHYFEKFCKEEGIVRHKTCAYTPQQNGIAERLNRTIMDKVRSML 593
Query: 215 AHSSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFP 274
+ S + FW A A YL+N P + D P ++ P S LR FGCL +
Sbjct: 594 SRSGMEKKFWAEAASTAVYLINRSPSTAINFDLPEEKWTGALPDLSSLRKFGCLA---YI 650
Query: 275 SATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTS 334
A KL PRS +F YP +GYK + L ++K +ISR+VIF E + F DL +S +
Sbjct: 651 HADQGKLNPRSKKGIFTSYPEGVKGYKVWVLEDKKCVISRNVIFRE-QVMFKDLKGDSQN 709
Query: 335 SY-DCFTEDL---PPSLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSSAPLPL 390
+ + EDL P + D S P + T +PT S +
Sbjct: 710 TISESDLEDLRVNPDMNDQEFTDQGGATQDNSNPSEATTSHNPVLNSPTHSQDEESEEED 769
Query: 391 PPVPPTPPTRTMTTHSMHGISKPKKPFSLSVSI-----DDPSISPLPCNPKQALSDPNWK 445
T + + K ++ S + + P P + ++AL DP+W+
Sbjct: 770 SDAVEDLSTYQLVRDRVRRTIKANPKYNESNMVGFAYYSEDDGKPEPKSYQEALLDPDWE 829
Query: 446 ---FAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGL-FEHYKARLVGDGW 501
AM+ E ++ +N+TW+LV +P V +I +F K G+ + ARLV G+
Sbjct: 830 KWNAAMKEEMVSMSKNHTWDLVTKPEKVKLIGCRWVFTRKAGIPGVEAPRFIARLVAKGF 889
Query: 502 SQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLG 561
+Q GVD +E F+ VVK +IR +LS+ + + + +DV AFLHG L E +YM QP G
Sbjct: 890 TQKEGVDYNEIFSPVVKHVSIRYLLSMVVHYNMELQQMDVKTAFLHGFLEEEIYMAQPEG 949
Query: 562 FRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSD------------HS 609
F + + VC K+SLYGLKQ+PR W RF +++ I + S+ D +
Sbjct: 950 FEIKRGSNKVCLLKRSLYGLKQSPRQWNLRFDEFMRGIKYTRSAYDSCVYFKKCNGDTYI 1009
Query: 610 YILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLGIAVT--RYVGGLFLSQS 667
Y+LLYVDD+++ +++ LL+ EF MKDLG LG+ ++ R G L LSQ
Sbjct: 1010 YLLLYVDDMLIASANKSEVNELKQLLSREFEMKDLGDAKKILGMEISRDRDAGLLTLSQE 1069
Query: 668 TYASEIIARAGMASCNPSATPVDTK*KLSSSSGTPCED------VTLYQSLAGALQY-LT 720
Y +++ M + P +TP+ KL +++ E+ + Y + G++ Y +
Sbjct: 1070 GYVKKVLRSFQMDNAKPVSTPLGIHFKLKAATDKEYEEQFERMKIVPYANTIGSIMYSMI 1129
Query: 721 FTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLL---RNLFLIRMLT--- 774
TRP+++Y + + M P +H A+ L T L + FL+R
Sbjct: 1130 GTRPDLAYSLGVISRFMSKPLKDHWQAVKWVLRYMRGTEKKKLCFRKQEDFLLRGYCDSD 1189
Query: 775 -GGMSCTRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLL 833
G TRRS +GY +G N ISW SK Q ++ SS EAEY + V E+ W++
Sbjct: 1190 YGSNFDTRRSITGYVFTVGGNTISWKSKLQKVVAISSTEAEYMALTEAVKEALWLKGFAA 1249
Query: 834 ELHFPLSQETL-VHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARILHVPSR 892
EL SQ+ + VH D+ S+I L+ N VHH+RTKHI++ +HF+R+ + G +++ + +
Sbjct: 1250 ELGH--SQDYVEVHSDSQSAITLAKNSVHHERTKHIDIRLHFIRDIICAGLIKVVKIATE 1307
Query: 893 HQIADIFTKGLPRVLFD 909
A+IFTK +P F+
Sbjct: 1308 CNPANIFTKTVPLAKFE 1324
>At3g45520 copia-like polyprotein
Length = 1363
Score = 413 bits (1061), Expect = e-115
Identities = 284/931 (30%), Positives = 441/931 (46%), Gaps = 72/931 (7%)
Query: 39 VWHNRLGHPTSSALNHLRNNKLIYCDPSRSSTVCDSCVLGKHVRLPFSSSETITLRSFDI 98
+WH RL H + ++ L + C+ C+ G+ ++ F+ ++ T + +
Sbjct: 447 LWHRRLCHMSQKNMSLLIKKGFLDKKKVSMLDTCEDCIYGRAKKIGFNLAQHDTKKKLEY 506
Query: 99 LHSDLWTSPI--LSTAGHRYYVLVLDDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSA 156
+HSDLW +P +S +Y++ +DD+T +W++ + K + ++ F + +L++ Q
Sbjct: 507 VHSDLWGAPTVPMSLGNCQYFISFIDDYTRKVWVYFLKTKDEAFEKFVSWISLVENQSGE 566
Query: 157 NIKCLQCDNGREYDNDSFRRYCHANGLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAH 216
+K L+ DNG E+ N F +C G +C +T QNG ER RTI +R++L
Sbjct: 567 RVKTLRTDNGLEFCNRMFDGFCEEKGFQRHRTCAYTPQQNGVVERMNRTIMEKVRSMLCD 626
Query: 217 SSVPPSFWHHALQMATYLLNILPRKTLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSA 276
S +P FW A A L+N P + + P +R + P YS+LR +GC+ F
Sbjct: 627 SGLPKRFWAEATHTAVLLINKTPCSAINFEFPDKRWSGKAPIYSYLRRYGCVTFVHTDG- 685
Query: 277 TINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPFADLPLESTSSY 336
KL R+ V +GYP +GYK + + +K ++SR+V F E+ + DL ++
Sbjct: 686 --GKLNLRAKKGVLIGYPSGVKGYKVWLIEEKKCVVSRNVSFQENAV-YKDL-MQRKEQV 741
Query: 337 DCFTEDLPPSLIHHWQTTSTRPPDLSIPPSSPTDS-TMPSPAPTSSPTSSAPLPLPPVPP 395
C +D S I DL + S S A + T A PP
Sbjct: 742 SCEEDDHAGSYI-----------DLDLEADKDNSSGGEQSQAQVTPATRGAVTSTPPRYE 790
Query: 396 TPPTRTMTTH----SMHGI-SKPKKPFSLSVSIDDPSI------------SPLPCNPKQA 438
T H S H + + ++ DD + P + K+A
Sbjct: 791 TDDIEETDVHQSPLSYHLVRDRERREIRAPRRFDDEDYYAEALYTTEDGDAVEPADYKEA 850
Query: 439 LSDPN---WKFAMQPEFNALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFE-HYKA 494
+ D N W+ AM E + ++N+TW V RP +I I+++K+ G+ E +KA
Sbjct: 851 VRDENWDKWRLAMNEEIESQLKNDTWTTVTRPEKQRIIGSRWIYKYKQGIPGVEEPRFKA 910
Query: 495 RLVGDGWSQIAGVDCDETFNLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETV 554
RLV G++Q GVD E F VVK +IR +LSI + + LDV AFLHG+L E +
Sbjct: 911 RLVAKGYAQREGVDYHEIFAPVVKHVSIRILLSIVAQENLELEQLDVKTAFLHGELKEKI 970
Query: 555 YMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDH------ 608
YM P G + VC KSLYGLKQAPR W ++F Y++ IGF+ S D
Sbjct: 971 YMMPPEGCESLFKENEVCLLNKSLYGLKQAPRQWNEKFNHYMTEIGFKRSDYDSCAYTKK 1030
Query: 609 ------SYILLYVDDIILVASSHDLRKSFMALLAFEFAMKDLGPLSYFLG--IAVTRYVG 660
Y+L YVDD+++ A++ + L+ +F MKDLG LG I + R G
Sbjct: 1031 LSDDSTMYLLFYVDDMLVAANNMQAIDALKKELSIKFEMKDLGAAKKILGIEIIIDREAG 1090
Query: 661 GLFLSQSTYASEIIARAGMASCNPSATPVDTK*KLSS------SSGTPCEDVTLYQSLAG 714
L+LSQ +Y ++++ M P+ TP+ K+ S S+ + Y S G
Sbjct: 1091 VLWLSQESYLNKVLKTFNMLESKPALTPLGAHLKMKSATEEKLSTEEEYMNSVPYSSAVG 1150
Query: 715 ALQY-LTFTRPNISYVVQQVCLHMHAPHTEHMLAMFVAL*LTVFTCIPPLLRNLFLIRML 773
++ Y + TRP+++Y V V M P EH L + L T L +
Sbjct: 1151 SIMYAMIGTRPDLAYPVGVVSRFMSQPAKEHWLGVKWVLRYIKGTVDTRLCYKRNSDFSI 1210
Query: 774 TGGMSC-------TRRSTSGYCVFLGDNLISWSSKRQPTLSHSSAEAEYRGVANVVSESC 826
G RRS +G LG N ISW S Q ++ SS E EY + V E+
Sbjct: 1211 CGYCDADYAADLDKRRSITGLVFTLGGNTISWKSGLQRVVAQSSTECEYMSLTEAVKEAI 1270
Query: 827 WIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQRTKHIEMDIHFVREKVARGQARI 886
W++ LL + + + + CD+ S+I LS N VHH+RTKHI++ HF+RE +A G+ +
Sbjct: 1271 WLKGLLKDFGYE-QKNVEIFCDSQSAIALSKNNVHHERTKHIDVKFHFIREIIADGKVEV 1329
Query: 887 LHVPSRHQIADIFTKGLPRVLFDDFRSSLSF 917
+ + ADIFTK LP + F+++L F
Sbjct: 1330 SKISTEKNPADIFTKVLP---VNKFQTALDF 1357
>At4g27200 putative protein
Length = 819
Score = 409 bits (1051), Expect = e-114
Identities = 267/717 (37%), Positives = 355/717 (49%), Gaps = 70/717 (9%)
Query: 266 GCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRGYKCYDLSNRKLIISRHVIFDESRFPF 325
G CFP NK P S CVFLGY ++GY+C +L ISRHVIFDES +PF
Sbjct: 15 GSACFPTLRDYAENKFNPCSLKCVFLGYNEKYKGYRCLYPPTGRLYISRHVIFDESVYPF 74
Query: 326 A----------------------DLPLESTSSYDC-----FTE-DLPP------------ 345
+ D P STS+ FT D PP
Sbjct: 75 SHTYKHLHPQPRTPLLAAWLRSSDSPAPSTSTSPSSRSPLFTSADFPPLPQRKTPLLPTL 134
Query: 346 ----SLIHHWQTTSTRPPDLSIPPSSPTDSTMPSPAPTSSPTSS--------APLPLPPV 393
S+ H T+ + PD ++ DS + SS S A + +
Sbjct: 135 VPISSVSHASNITTQQSPDFDSERTTDFDSASIGDSSHSSQAGSDSEETIQQASVNVHQT 194
Query: 394 PPTPPTRTMTTHSMHGISKPKKPFSLSVSIDDPSISPLPCNPKQALSDPNWKFAMQPEFN 453
P + M T + GISKP + V + P P AL P W AM E
Sbjct: 195 PASTNVHPMVTRAKVGISKPNPRY---VFLSHKVSYPEPKTVTAALKHPGWTGAMTEEIG 251
Query: 454 ALIRNNTWELVPRPCDVNVIRYM*IFRHKKQSNGLFEHYKARLVGDGWSQIAGVDCDETF 513
TW LVP D++V+ +FR K ++G KAR+V G+ Q G+D ET+
Sbjct: 252 NCSETQTWSLVPYKSDMHVLGSKWVFRTKLHADGTLNKLKARIVAKGFLQEEGIDYLETY 311
Query: 514 NLVVKPATIRTVLSIALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCR 573
+ VV+ T+R VL +A + +W I +DV NAFLHGDL ETVYM QP RD +VC
Sbjct: 312 SPVVRTPTVRLVLHLATALNWDIKQMDVKNAFLHGDLKETVYMTQPAANRD-----HVCL 366
Query: 574 RKKSLYGLKQAPRAWYQRFADYVSSIGFQHSSSDHS-YILLYVDDIILVASSHDLRKSFM 632
KS+YGLKQ+PRAW+ +F+ ++ GF SD S +I + +++IL+ S L S +
Sbjct: 367 LHKSIYGLKQSPRAWFDKFSTFLLEFGFFCRKSDPSLFIYAHNNNLILLLLSQTLT-SLL 425
Query: 633 ALLAFEFAMKDLGPLSY-FLGIAVTRYVGGLFLSQSTYASEIIARAGMASCNPSATPVDT 691
A L EF M D+G S FLGI V R GLF+SQ YA +++ A M C P TP+
Sbjct: 426 AALNKEFRMTDMGQHSLTFLGIQVQRQQNGLFMSQQKYAEDLLIAASMEHCTPLPTPLPV 485
Query: 692 K*KLSSSSGTPCEDVTLYQSLAGALQYLTFTRPNISYVVQQVCLHMHAPHTE--HMLAMF 749
+ D T ++S+AG LQYLT TRP+I + V VC MH P H+L
Sbjct: 486 QLDRVPHQEELFSDPTYFRSIAGKLQYLTLTRPDIQFAVNFVCQKMHQPTISDFHLLKRI 545
Query: 750 VAL*LTVFTCIPPLLRNL-FLIRMLT----GGMSCTRRSTSGYCVFLGDNLISWSSKRQP 804
+ T R+ L++ + G TRRS G C F+G NL+SWSSK+ P
Sbjct: 546 LRYIKGTITMGISYSRDSPTLLQAYSDSDWGNCKQTRRSVGGLCTFMGTNLVSWSSKKHP 605
Query: 805 TLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSGNPVHHQR 864
T+S SS EAEY+ +++ SE W+ LL EL PL + CDN+S++YL+ NP H R
Sbjct: 606 TVSRSSTEAEYKSLSDAASEILWLSTLLRELRIPLPDTPELFCDNLSAVYLTANPAFHAR 665
Query: 865 TKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDDFRSSLSFGEPP 921
TKH ++D HFVRE+VA + H+P QIADIFTK LP F R L PP
Sbjct: 666 TKHFDIDFHFVRERVALKALVVKHIPGSEQIADIFTKSLPYEAFIHLRGKLGVTLPP 722
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 367 bits (943), Expect = e-101
Identities = 255/833 (30%), Positives = 407/833 (48%), Gaps = 68/833 (8%)
Query: 122 DDHTDFLWMFPISKKSQVYDIFTTLATLIKTQFSANIKCLQCDNGREYDNDSFRRYCHAN 181
+D + +W++ + K + + FT +++TQ +K L+ DNG E+ N F C
Sbjct: 319 NDWSRKVWVYFLKTKDEAFASFTEWKKMVETQSERKLKHLRTDNGLEFCNHKFDEVCKKE 378
Query: 182 GLIFRFSCPHTSSQNGKSERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRK 241
G++ +C +T QNG +ER RTI N +R++L+ S + FW A A YL+N P
Sbjct: 379 GIVRHRTCTYTPQQNGVAERLNRTIMNKVRSMLSESGLDKKFWAKAASTAVYLINRSPSS 438
Query: 242 TLRNDSPTQRLYHRDPSYSHLRVFGCLCFPFFPSATINKLQPRSSPCVFLGYPMNHRGYK 301
++ N P + P++S L+ FGC+ + + KL PR+ VF+GYP +G++
Sbjct: 439 SIENKIPEELWTSAVPNFSGLKRFGCVVYVYSQEG---KLDPRAKKGVFVGYPNGVKGFR 495
Query: 302 CYDLSNRKLIISRHVIFDESRFPFADLPLESTSSYDCFTEDLPPSLIHHWQTTSTRPPD- 360
+ + + ISR+V+F E + D+ +STS F L + I ++ R D
Sbjct: 496 VWMIEEERCSISRNVVFREDVM-YKDILNQSTSGMS-FDFPLATNRIPSFECAGNRKEDE 553
Query: 361 LSIPPSSPTDSTMPS----PAPT-SSPTSSAPLPLPPVPPTPPTRTMTTHSMHGISKPKK 415
+S+ D T S P T SS +S P +T + ++
Sbjct: 554 ISVQGGVSDDDTKQSSEESPISTGSSGQNSGQRTYQIARDKPKRQTKIPDKLRDYELNEE 613
Query: 416 PFS----LSVSIDDPSISPLPCNPKQALSDPNWKF---AMQPEFNALIRNNTWELVPRPC 468
+ I + +P P + ++AL D ++K A+ E +L++NNTW LV R
Sbjct: 614 VLDEIAGYAYMITEDGGNPEPNDYQKALQDSDYKMWLKAVDEEIESLLKNNTWVLVNRDQ 673
Query: 469 DVNVIRYM*IFRHKKQSNGLFE-HYKARLVGDGWSQIAGVDCDETFNLVVKPATIRTVLS 527
I +F+ K G+ + +KARLV G+SQ G+D E F+ VVK +IR +LS
Sbjct: 674 FQKPIGCKWVFKRKSGIVGVEKPRFKARLVVKGYSQKEGIDYQEIFSPVVKHVSIRLLLS 733
Query: 528 IALSRSWPIH*LDVHNAFLHGDLHETVYMHQPLGFRDSQHPDYVCRRKKSLYGLKQAPRA 587
+ + +DV AFLHG L ET+Y+ QP G+ ++PD VC K+SLYGL+Q+PR
Sbjct: 734 MVTHCDMELQQMDVKTAFLHGYLDETIYIEQPEGYVHKRYPDKVCLLKRSLYGLRQSPRQ 793
Query: 588 WYQRFADYVSSIGFQHS------------SSDHSYILLYVDDIILVASSHDLRKSFMALL 635
W RF +++ IG++ S S ++ Y+LLYVDDI++ + ALL
Sbjct: 794 WNNRFNEFMQKIGYERSKYDSCVYFKELQSGEYIYLLLYVDDILIASRDKRTVCDLKALL 853
Query: 636 AFEFAMKDLGPLSYFLGIAVT--RYVGGLFLSQSTYASEIIARAGMASCNPSATPVDTK* 693
EF MKDLG LG+ + R G + +SQ Y +++ GM P TP+
Sbjct: 854 NSEFEMKDLGDAKKILGMEIVRDRKAGTMSISQEGYLLKVLGNFGMDQAKPVFTPMGAHF 913
Query: 694 KLSSSSG------TPCEDVTLYQSLAGALQY-LTFTRPNISYVVQQVCLHMHAPHTEHML 746
KL ++ + YQS G+L Y + TRP++++ V VC M P EH
Sbjct: 914 KLKPATDEEVMRQSEVMRAVPYQSAVGSLMYSMIGTRPDLAHSVGLVCRFMSKPLKEHWQ 973
Query: 747 AMFVAL*LTVFTCIPPLLRNLFLIRMLTGGMS---CTRRS----TSGYC--VFLGDNLIS 797
A+ +++R + G + C + GYC + D
Sbjct: 974 AV------------------KWILRYIRGSIDRKLCYKNEGELILEGYCDSDYAADKEGR 1015
Query: 798 WSSKRQPTLSHSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQETLVHCDNVSSIYLSG 857
S+ ++ SS EAEY + + E+ W++ + EL F + + +HCD+ S+I L+
Sbjct: 1016 RSTSGVKVVALSSTEAEYMALTDGAKEAIWLKGHVSELGF-VQKTVNIHCDSQSAIALAK 1074
Query: 858 NPVHHQRTKHIEMDIHFVREKVARGQARILHVPSRHQIADIFTKGLPRVLFDD 910
N V+H+RTKHI++ HF+R+ V G+ ++L + + ADIFTK LP F D
Sbjct: 1075 NAVYHERTKHIDVKYHFIRDLVNNGEVQVLKIDTEDNPADIFTKVLPVSKFQD 1127
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,772,692
Number of Sequences: 26719
Number of extensions: 1007542
Number of successful extensions: 12068
Number of sequences better than 10.0: 390
Number of HSP's better than 10.0 without gapping: 247
Number of HSP's successfully gapped in prelim test: 155
Number of HSP's that attempted gapping in prelim test: 5561
Number of HSP's gapped (non-prelim): 2056
length of query: 921
length of database: 11,318,596
effective HSP length: 108
effective length of query: 813
effective length of database: 8,432,944
effective search space: 6855983472
effective search space used: 6855983472
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0032.12


