

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0030.7
(504 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
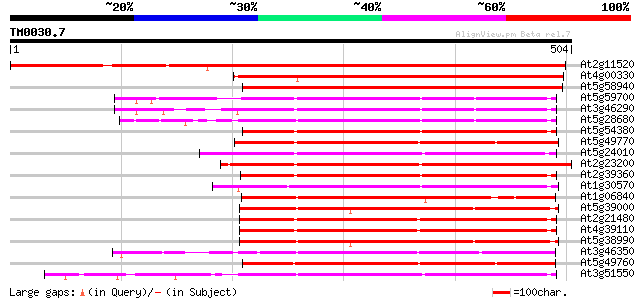
Score E
Sequences producing significant alignments: (bits) Value
At2g11520 putative protein kinase 560 e-160
At4g00330 unknown protein 337 9e-93
At5g58940 receptor-like protein kinase precursor - like 335 3e-92
At5g59700 receptor-like protein kinase precursor - like 244 8e-65
At3g46290 receptor protein kinase -like 244 1e-64
At5g28680 receptor-like protein kinase precursor - like 239 3e-63
At5g54380 receptor-protein kinase-like protein 236 2e-62
At5g49770 receptor protein kinase-like 236 3e-62
At5g24010 receptor-protein kinase-like protein 234 6e-62
At2g23200 putative protein kinase 233 2e-61
At2g39360 putative protein kinase 232 3e-61
At1g30570 putative serine/threonine protein kinase 232 3e-61
At1g06840 receptor protein kinase, putative 231 7e-61
At5g39000 protein kinase - like protein 229 2e-60
At2g21480 putative protein kinase 229 3e-60
At4g39110 putative receptor-like protein kinase 229 3e-60
At5g38990 protein kinase - like protein 228 5e-60
At3g46350 receptor-like protein kinase homolog 228 5e-60
At5g49760 receptor protein kinase-like 227 1e-59
At3g51550 receptor protein kinase like protein 226 2e-59
>At2g11520 putative protein kinase
Length = 510
Score = 560 bits (1444), Expect = e-160
Identities = 290/504 (57%), Positives = 368/504 (72%), Gaps = 13/504 (2%)
Query: 1 MAITVLY-LVILLQLSTFLGSSVVLQSKECGTDWLAHSYSSHGQELFYINGNVVNQVSFC 59
+ IT L+ L++LLQ+ S+ + S C +D L ++ LF INGN V ++ FC
Sbjct: 11 LVITALFGLLMLLQIKETSASTSFVSSSVCKSDHLTYTKPYQQGSLFTINGNPVEKLRFC 70
Query: 60 EALQLYIANGCDVKDYFGSSGCAVGGSFVNFPSKAGRKLLQKDSSKESTSQGDSKALPTN 119
EAL+ + ANGC +D F C + S GR+ L++ + K+S +
Sbjct: 71 EALRFHKANGCIFEDSFSDDFCTIH-------SLLGRRFLEEKTVKDSKNSKPKTEYSHV 123
Query: 120 KVGIFAGGALLACCAVLCPCFYGKKRKATAHAVLAKDPISMDSATSFDASVSASVSG--- 176
KV I G LL CCA+ CPCF+ K+RKA +H VL K+ S+ +SF+ S S+
Sbjct: 124 KVSIAGSGFLLLCCALCCPCFH-KERKANSHEVLPKESNSVHQVSSFEMSPSSEKIPQSP 182
Query: 177 -KIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKA 235
+ P SP RVP SPSR++MSP+ SRL LNL +SQ+ AT NF+++ QIG+GGFG V+K
Sbjct: 183 FRAPPSPSRVPQSPSRYAMSPRPSRLGPLNLTMSQINTATGNFADSHQIGEGGFGVVFKG 242
Query: 236 NLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYV 295
L+DG VVA+KRAK+EHFE+LRTEF SEV+LL+KI HRNLVKLLG++DKG+ER++ITEYV
Sbjct: 243 VLDDGQVVAIKRAKKEHFENLRTEFKSEVDLLSKIGHRNLVKLLGYVDKGDERLIITEYV 302
Query: 296 PNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESM 355
NGTLR+HLDG RG L+FNQRLEI IDV HGLTYLH YAE+QIIHRD+KSSNILLT+SM
Sbjct: 303 RNGTLRDHLDGARGTKLNFNQRLEIVIDVCHGLTYLHSYAERQIIHRDIKSSNILLTDSM 362
Query: 356 RAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEIL 415
RAKVADFGFAR GP + +QTHI T+VKGTVGYLDPEYMKT+ LT KSDVYSFGILL+EIL
Sbjct: 363 RAKVADFGFARGGPTDSNQTHILTQVKGTVGYLDPEYMKTYHLTAKSDVYSFGILLVEIL 422
Query: 416 TGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAP 475
TGRRPVE K+ DER+T+RWAF KYNEG V EL+DP E V+ +L KM L+FQCAAP
Sbjct: 423 TGRRPVEAKRLPDERITVRWAFDKYNEGRVFELVDPNARERVDEKILRKMFSLAFQCAAP 482
Query: 476 IRTDRPNMKSVGEQLWAIRADYLK 499
+ +RP+M++VG+QLWAIR+ YL+
Sbjct: 483 TKKERPDMEAVGKQLWAIRSSYLR 506
>At4g00330 unknown protein
Length = 411
Score = 337 bits (864), Expect = 9e-93
Identities = 167/301 (55%), Positives = 216/301 (71%), Gaps = 6/301 (1%)
Query: 202 HTLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLR---T 258
HT ++ AT+NFS + +IGQGGFGTVYK L DG AVKRAK+ + +
Sbjct: 104 HT-RFTFDEIYDATKNFSPSFRIGQGGFGTVYKVKLRDGKTFAVKRAKKSMHDDRQGADA 162
Query: 259 EFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRL 318
EF SE++ LA++ H +LVK GF+ +E+IL+ EYV NGTLR+HLD GK LD RL
Sbjct: 163 EFMSEIQTLAQVTHLSLVKYYGFVVHNDEKILVVEYVANGTLRDHLDCKEGKTLDMATRL 222
Query: 319 EIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGP-VNGDQTHI 377
+IA DVAH +TYLH+Y + IIHRD+KSSNILLTE+ RAKVADFGFARL P + TH+
Sbjct: 223 DIATDVAHAITYLHMYTQPPIIHRDIKSSNILLTENYRAKVADFGFARLAPDTDSGATHV 282
Query: 378 STKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAF 437
ST+VKGT GYLDPEY+ T+QLT KSDVYSFG+LL+E+LTGRRP+EL + ER+T+RWA
Sbjct: 283 STQVKGTAGYLDPEYLTTYQLTEKSDVYSFGVLLVELLTGRRPIELSRGQKERITIRWAI 342
Query: 438 RKYNEGSVVELMDPLMEE-AVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRAD 496
+K+ G + ++DP +E+ + N L K+L+++FQC AP R RP+MK E LW IR D
Sbjct: 343 KKFTSGDTISVLDPKLEQNSANNLALEKVLEMAFQCLAPHRRSRPSMKKCSEILWGIRKD 402
Query: 497 Y 497
Y
Sbjct: 403 Y 403
>At5g58940 receptor-like protein kinase precursor - like
Length = 466
Score = 335 bits (860), Expect = 3e-92
Identities = 163/289 (56%), Positives = 218/289 (75%), Gaps = 2/289 (0%)
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHF-ESLRTEFSSEVELLA 268
Q+ RAT NFS QIG+GGFGTV+K L+DG +VA+KRA++ ++ +S EF +E+ L+
Sbjct: 135 QLQRATANFSSVHQIGEGGFGTVFKGKLDDGTIVAIKRARKNNYGKSWLLEFKNEIYTLS 194
Query: 269 KIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGL 328
KI+H NLVKL GF++ G+E++++ EYV NG LREHLDGLRG L+ +RLEIAIDVAH L
Sbjct: 195 KIEHMNLVKLYGFLEHGDEKVIVVEYVANGNLREHLDGLRGNRLEMAERLEIAIDVAHAL 254
Query: 329 TYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYL 388
TYLH Y + IIHRD+K+SNIL+T +RAKVADFGFARL + THIST+VKG+ GY+
Sbjct: 255 TYLHTYTDSPIIHRDIKASNILITNKLRAKVADFGFARLVSEDLGATHISTQVKGSAGYV 314
Query: 389 DPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVEL 448
DP+Y++T QLT KSDVYSFG+LL+EILTGRRP+ELK+ +R+T++WA R+ + V +
Sbjct: 315 DPDYLRTFQLTDKSDVYSFGVLLVEILTGRRPIELKRPRKDRLTVKWALRRLKDDEAVLI 374
Query: 449 MDPLMEEAVNA-DVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAIRAD 496
MDP ++ A +V KML L+ +C P R RP MK + E+LWAIR +
Sbjct: 375 MDPFLKRNRAAIEVAEKMLRLASECVTPTRATRPAMKGIAEKLWAIRRE 423
>At5g59700 receptor-like protein kinase precursor - like
Length = 829
Score = 244 bits (623), Expect = 8e-65
Identities = 161/406 (39%), Positives = 220/406 (53%), Gaps = 37/406 (9%)
Query: 95 GRKLLQKDSSKESTSQGD-----SKALPTNKVGIFAG---GALLACCAVLCPCFYGKKRK 146
G ++++ ++SK S G S + VG+ G G+LLA VL F K++
Sbjct: 373 GLEIMKMNNSKSQLSIGTFLPSGSSSTTKKNVGMIIGLTIGSLLAL-VVLGGFFVLYKKR 431
Query: 147 ATAHAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNL 206
+K I + S + +S +++ S R+P
Sbjct: 432 GRDQDGNSKTWIPLSSNGTTSSSNGTTLASIASNSSYRIP-------------------- 471
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
L V AT +F E IG GGFG VYK L DG VAVKRA + + L EF +E+E+
Sbjct: 472 -LVAVKEATNSFDENRAIGVGGFGKVYKGELHDGTKVAVKRANPKSQQGL-AEFRTEIEM 529
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAH 326
L++ HR+LV L+G+ D+ NE IL+ EY+ NGTL+ HL G L + QRLEI I A
Sbjct: 530 LSQFRHRHLVSLIGYCDENNEMILVYEYMENGTLKSHLYGSGLLSLSWKQRLEICIGSAR 589
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL YLH K +IHRDVKS+NILL E++ AKVADFG ++ GP DQTH+ST VKG+ G
Sbjct: 590 GLHYLHTGDAKPVIHRDVKSANILLDENLMAKVADFGLSKTGP-EIDQTHVSTAVKGSFG 648
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYNEGSV 445
YLDPEY + QLT KSDVYSFG+++ E+L RPV E V L WA + +G +
Sbjct: 649 YLDPEYFRRQQLTEKSDVYSFGVVMFEVLCA-RPVIDPTLTREMVNLAEWAMKWQKKGQL 707
Query: 446 VELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
++DP + + D L K + +C A DRP+M G+ LW
Sbjct: 708 EHIIDPSLRGKIRPDSLRKFGETGEKCLADYGVDRPSM---GDVLW 750
>At3g46290 receptor protein kinase -like
Length = 830
Score = 244 bits (622), Expect = 1e-64
Identities = 158/409 (38%), Positives = 222/409 (53%), Gaps = 41/409 (10%)
Query: 95 GRKLLQKDSSKESTSQGD----SKALPTNKVGIFAGGALLACCAVL----CPCFYGKKRK 146
G ++++ ++SK S G S + + +G+ G A+ + AV+ C Y K+++
Sbjct: 374 GLEIMKMNNSKGQLSTGTFVPGSSSSSKSNLGLIVGSAIGSLLAVVFLGSCFVLYKKRKR 433
Query: 147 ATAHAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHT--- 203
S T S++ + G S++S L+ + T
Sbjct: 434 GQ----------DGHSKTWMPFSINGTSMG-------------SKYSNGTTLTSITTNAN 470
Query: 204 LNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSE 263
+ + V AT NF E+ IG GGFG VYK L DG VAVKR + + L EF +E
Sbjct: 471 YRIPFAAVKDATNNFDESRNIGVGGFGKVYKGELNDGTKVAVKRGNPKSQQGL-AEFRTE 529
Query: 264 VELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAID 323
+E+L++ HR+LV L+G+ D+ NE ILI EY+ NGT++ HL G L + QRLEI I
Sbjct: 530 IEMLSQFRHRHLVSLIGYCDENNEMILIYEYMENGTVKSHLYGSGLPSLTWKQRLEICIG 589
Query: 324 VAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKG 383
A GL YLH K +IHRDVKS+NILL E+ AKVADFG ++ GP DQTH+ST VKG
Sbjct: 590 AARGLHYLHTGDSKPVIHRDVKSANILLDENFMAKVADFGLSKTGP-ELDQTHVSTAVKG 648
Query: 384 TVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYNE 442
+ GYLDPEY + QLT KSDVYSFG++L E+L RPV E V L WA + +
Sbjct: 649 SFGYLDPEYFRRQQLTDKSDVYSFGVVLFEVLCA-RPVIDPTLPREMVNLAEWAMKWQKK 707
Query: 443 GSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
G + +++D + + D L K + +C A DRP+M G+ LW
Sbjct: 708 GQLDQIIDQSLRGNIRPDSLRKFAETGEKCLADYGVDRPSM---GDVLW 753
>At5g28680 receptor-like protein kinase precursor - like
Length = 858
Score = 239 bits (609), Expect = 3e-63
Identities = 155/397 (39%), Positives = 229/397 (57%), Gaps = 27/397 (6%)
Query: 99 LQKDSSKESTSQGDSKALPTNKVGIFAGGALLACCAVLCPCFYGKKRKATAHAVLAKD-- 156
+Q + + QGD K + +G G A + CA LC Y +KRK +
Sbjct: 416 MQANEDVKKDFQGD-KRITAFVIGSAGGVAAVLFCA-LCFTMYQRKRKFSGSDSHTSSWL 473
Query: 157 PISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFS-MSPKLSRLHTLNLNLSQVVRAT 215
PI +S TS + +++SGK ++ S S ++ L R +LS++ T
Sbjct: 474 PIYGNSHTS---ATKSTISGK--------SNNGSHLSNLAAGLCR----RFSLSEIKHGT 518
Query: 216 RNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNL 275
NF E+ IG GGFG VYK ++ G VA+K++ + L EF +E+ELL+++ H++L
Sbjct: 519 HNFDESNVIGVGGFGKVYKGVIDGGTKVAIKKSNPNSEQGLN-EFETEIELLSRLRHKHL 577
Query: 276 VKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYA 335
V L+G+ D+G E LI +Y+ GTLREHL + L + +RLEIAI A GL YLH A
Sbjct: 578 VSLIGYCDEGGEMCLIYDYMSLGTLREHLYNTKRPQLTWKRRLEIAIGAARGLHYLHTGA 637
Query: 336 EKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKT 395
+ IIHRDVK++NILL E+ AKV+DFG ++ GP N + H++T VKG+ GYLDPEY +
Sbjct: 638 KYTIIHRDVKTTNILLDENWVAKVSDFGLSKTGP-NMNGGHVTTVVKGSFGYLDPEYFRR 696
Query: 396 HQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYNEGSVVELMDPLME 454
QLT KSDVYSFG++L E+L RP + E+V+L WA +G++ +++DP ++
Sbjct: 697 QQLTEKSDVYSFGVVLFEVLCA-RPALNPSLSKEQVSLGDWAMNCKRKGTLEDIIDPNLK 755
Query: 455 EAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
+N + L K D + +C + DRP M G+ LW
Sbjct: 756 GKINPECLKKFADTAEKCLSDSGLDRPTM---GDVLW 789
>At5g54380 receptor-protein kinase-like protein
Length = 855
Score = 236 bits (603), Expect = 2e-62
Identities = 130/282 (46%), Positives = 178/282 (63%), Gaps = 5/282 (1%)
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAK 269
+++ AT F E+ +G GGFG VYK LEDG VAVKR + + EF +E+E+L+K
Sbjct: 502 EIMDATNKFDESSLLGVGGFGRVYKGTLEDGTKVAVKRGNPRSEQGM-AEFRTEIEMLSK 560
Query: 270 IDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLT 329
+ HR+LV L+G+ D+ +E IL+ EY+ NG LR HL G L + QRLEI I A GL
Sbjct: 561 LRHRHLVSLIGYCDERSEMILVYEYMANGPLRSHLYGADLPPLSWKQRLEICIGAARGLH 620
Query: 330 YLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLD 389
YLH A + IIHRDVK++NILL E++ AKVADFG ++ GP + DQTH+ST VKG+ GYLD
Sbjct: 621 YLHTGASQSIIHRDVKTTNILLDENLVAKVADFGLSKTGP-SLDQTHVSTAVKGSFGYLD 679
Query: 390 PEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELM 449
PEY + QLT KSDVYSFG++L+E+L R + ++ WA +G + ++M
Sbjct: 680 PEYFRRQQLTEKSDVYSFGVVLMEVLCCRPALNPVLPREQVNIAEWAMAWQKKGLLDQIM 739
Query: 450 DPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
D + VN L K + + +C A DRP+M G+ LW
Sbjct: 740 DSNLTGKVNPASLKKFGETAEKCLAEYGVDRPSM---GDVLW 778
>At5g49770 receptor protein kinase-like
Length = 946
Score = 236 bits (601), Expect = 3e-62
Identities = 125/292 (42%), Positives = 190/292 (64%), Gaps = 4/292 (1%)
Query: 203 TLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSS 262
T ++ + T NFS+ +G GG+G VYK L +G V+A+KRA++ + EF +
Sbjct: 619 TKAFTFEELSKCTNNFSDANDVGGGGYGQVYKGTLPNGQVIAIKRAQQGSMQGA-FEFKT 677
Query: 263 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAI 322
E+ELL+++ H+N+VKLLGF E++L+ EY+PNG+LR+ L G G LD+ +RL+IA+
Sbjct: 678 EIELLSRVHHKNVVKLLGFCFDQKEQMLVYEYIPNGSLRDGLSGKNGVKLDWTRRLKIAL 737
Query: 323 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVK 382
GL YLH A+ IIHRDVKS+NILL E + AKVADFG ++L + ++ H++T+VK
Sbjct: 738 GSGKGLAYLHELADPPIIHRDVKSNNILLDEHLTAKVADFGLSKL-VGDPEKAHVTTQVK 796
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNE 442
GT+GYLDPEY T+QLT KSDVY FG+++LE+LTG+ P++ + V + + N
Sbjct: 797 GTMGYLDPEYYMTNQLTEKSDVYGFGVVMLELLTGKSPIDRGSYVVKEVKKKMD-KSRNL 855
Query: 443 GSVVELMD-PLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAI 493
+ EL+D +++ + N K +D++ QC P +RP M V ++L +I
Sbjct: 856 YDLQELLDTTIIQNSGNLKGFEKYVDVALQCVEPEGVNRPTMSEVVQELESI 907
>At5g24010 receptor-protein kinase-like protein
Length = 824
Score = 234 bits (598), Expect = 6e-62
Identities = 130/321 (40%), Positives = 189/321 (58%), Gaps = 5/321 (1%)
Query: 171 SASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLSQVVRATRNFSETLQIGQGGFG 230
S+ +G P R S+ + S HTL ++ +++ T NF +L IG GGFG
Sbjct: 442 SSESTGWTPLRRFRGSSNSRTTERTVSSSGYHTLRISFAELQSGTNNFDRSLVIGVGGFG 501
Query: 231 TVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERIL 290
V++ +L+D VAVKR + L EF SE+ +L+KI HR+LV L+G+ ++ +E IL
Sbjct: 502 MVFRGSLKDNTKVAVKRGSPGSRQGL-PEFLSEITILSKIRHRHLVSLVGYCEEQSEMIL 560
Query: 291 ITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNIL 350
+ EY+ G L+ HL G L + QRLE+ I A GL YLH + + IIHRD+KS+NIL
Sbjct: 561 VYEYMDKGPLKSHLYGSTNPPLSWKQRLEVCIGAARGLHYLHTGSSQGIIHRDIKSTNIL 620
Query: 351 LTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGIL 410
L + AKVADFG +R GP D+TH+ST VKG+ GYLDPEY + QLT KSDVYSFG++
Sbjct: 621 LDNNYVAKVADFGLSRSGPCI-DETHVSTGVKGSFGYLDPEYFRRQQLTDKSDVYSFGVV 679
Query: 411 LLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSF 470
L E+L R V+ ++ WA +G + +++DP + + + L K + +
Sbjct: 680 LFEVLCARPAVDPLLVREQVNLAEWAIEWQRKGMLDQIVDPNIADEIKPCSLKKFAETAE 739
Query: 471 QCAAPIRTDRPNMKSVGEQLW 491
+C A DRP ++G+ LW
Sbjct: 740 KCCADYGVDRP---TIGDVLW 757
>At2g23200 putative protein kinase
Length = 834
Score = 233 bits (593), Expect = 2e-61
Identities = 133/317 (41%), Positives = 198/317 (61%), Gaps = 6/317 (1%)
Query: 190 SRFSMSPKLSRLHT-LNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRA 248
S++ SP L LH L + + ++ AT NF E L IG+GGFG VYKA L DG A+KR
Sbjct: 460 SQYHNSP-LRNLHLGLTIPFTDILSATNNFDEQLLIGKGGFGYVYKAILPDGTKAAIKRG 518
Query: 249 KREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLR 308
K + + EF +E+++L++I HR+LV L G+ ++ +E IL+ E++ GTL+EHL G
Sbjct: 519 KTGSGQGI-LEFQTEIQVLSRIRHRHLVSLTGYCEENSEMILVYEFMEKGTLKEHLYGSN 577
Query: 309 GKILDFNQRLEIAIDVAHGLTYLHLY-AEKQIIHRDVKSSNILLTESMRAKVADFGFARL 367
L + QRLEI I A GL YLH +E IIHRDVKS+NILL E AKVADFG +++
Sbjct: 578 LPSLTWKQRLEICIGAARGLDYLHSSGSEGAIIHRDVKSTNILLDEHNIAKVADFGLSKI 637
Query: 368 GPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAA 427
N D+++IS +KGT GYLDPEY++TH+LT KSDVY+FG++LLE+L R ++
Sbjct: 638 H--NQDESNISINIKGTFGYLDPEYLQTHKLTEKSDVYAFGVVLLEVLFARPAIDPYLPH 695
Query: 428 DERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVG 487
+E W ++G++ E++DP + + + L K ++++ +C +RP+M+ V
Sbjct: 696 EEVNLSEWVMFCKSKGTIDEILDPSLIGQIETNSLKKFMEIAEKCLKEYGDERPSMRDVI 755
Query: 488 EQLWAIRADYLKSARRE 504
L + + + RRE
Sbjct: 756 WDLEYVLQLQMMTNRRE 772
>At2g39360 putative protein kinase
Length = 815
Score = 232 bits (592), Expect = 3e-61
Identities = 125/285 (43%), Positives = 183/285 (63%), Gaps = 6/285 (2%)
Query: 208 LSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELL 267
L+ + AT +F E+L IG GGFG VYK L D VAVKR + + L EF +EVE+L
Sbjct: 477 LALIKEATDDFDESLVIGVGGFGKVYKGVLRDKTEVAVKRGAPQSRQGL-AEFKTEVEML 535
Query: 268 AKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKI-LDFNQRLEIAIDVAH 326
+ HR+LV L+G+ D+ +E I++ EY+ GTL++HL L K L + QRLEI + A
Sbjct: 536 TQFRHRHLVSLIGYCDENSEMIIVYEYMEKGTLKDHLYDLDDKPRLSWRQRLEICVGAAR 595
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL YLH + + IIHRDVKS+NILL ++ AKVADFG ++ GP + DQTH+ST VKG+ G
Sbjct: 596 GLHYLHTGSTRAIIHRDVKSANILLDDNFMAKVADFGLSKTGP-DLDQTHVSTAVKGSFG 654
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVV 446
YLDPEY+ QLT KSDVYSFG+++LE++ GR ++ ++ + WA + +G +
Sbjct: 655 YLDPEYLTRQQLTEKSDVYSFGVVMLEVVCGRPVIDPSLPREKVNLIEWAMKLVKKGKLE 714
Query: 447 ELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
+++DP + V + + K +++ +C + +RP M G+ LW
Sbjct: 715 DIIDPFLVGKVKLEEVKKYCEVTEKCLSQNGIERPAM---GDLLW 756
>At1g30570 putative serine/threonine protein kinase
Length = 849
Score = 232 bits (592), Expect = 3e-61
Identities = 132/318 (41%), Positives = 191/318 (59%), Gaps = 12/318 (3%)
Query: 183 LRVPSSPSRFSMSPKLSRLHTL-------NLNLSQVVRATRNFSETLQIGQGGFGTVYKA 235
L V +S + + RL+TL L+++ AT+NF + L IG GGFG VY+
Sbjct: 478 LHVNNSTANAKATGGSLRLNTLAASTMGRKFTLAEIRAATKNFDDGLAIGVGGFGKVYRG 537
Query: 236 NLEDGLVVAVKRAKREHFESLRTEFSSEVELLAKIDHRNLVKLLGFIDKGNERILITEYV 295
LEDG ++A+KRA H + EF +E+ +L+++ HR+LV L+GF D+ NE IL+ EY+
Sbjct: 538 ELEDGTLIAIKRAT-PHSQQGLAEFETEIVMLSRLRHRHLVSLIGFCDEHNEMILVYEYM 596
Query: 296 PNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESM 355
NGTLR HL G L + QRLE I A GL YLH +E+ IIHRDVK++NILL E+
Sbjct: 597 ANGTLRSHLFGSNLPPLSWKQRLEACIGSARGLHYLHTGSERGIIHRDVKTTNILLDENF 656
Query: 356 RAKVADFGFARLGPVNGDQTHISTKVKGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEIL 415
AK++DFG ++ GP + D TH+ST VKG+ GYLDPEY + QLT KSDVYSFG++L E +
Sbjct: 657 VAKMSDFGLSKAGP-SMDHTHVSTAVKGSFGYLDPEYFRRQQLTEKSDVYSFGVVLFEAV 715
Query: 416 TGRRPVELKKAADERVTLRWAFRKYNEGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAP 475
R + D+ WA + ++ ++D + + + L K +++ +C A
Sbjct: 716 CARAVINPTLPKDQINLAEWALSWQKQRNLESIIDSNLRGNYSPESLEKYGEIAEKCLAD 775
Query: 476 IRTDRPNMKSVGEQLWAI 493
+RP M GE LW++
Sbjct: 776 EGKNRPMM---GEVLWSL 790
>At1g06840 receptor protein kinase, putative
Length = 939
Score = 231 bits (589), Expect = 7e-61
Identities = 129/286 (45%), Positives = 175/286 (61%), Gaps = 11/286 (3%)
Query: 209 SQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLA 268
+++ AT NF+ + QIGQGG+G VYK L G VVA+KRA+ + + EF +E+ELL+
Sbjct: 602 AELALATDNFNSSTQIGQGGYGKVYKGTLGSGTVVAIKRAQEGSLQGEK-EFLTEIELLS 660
Query: 269 KIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGL 328
++ HRNLV LLGF D+ E++L+ EY+ NGTLR+++ + LDF RL IA+ A G+
Sbjct: 661 RLHHRNLVSLLGFCDEEGEQMLVYEYMENGTLRDNISVKLKEPLDFAMRLRIALGSAKGI 720
Query: 329 TYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNG----DQTHISTKVKGT 384
YLH A I HRD+K+SNILL AKVADFG +RL PV H+ST VKGT
Sbjct: 721 LYLHTEANPPIFHRDIKASNILLDSRFTAKVADFGLSRLAPVPDMEGISPQHVSTVVKGT 780
Query: 385 VGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGS 444
GYLDPEY THQLT KSDVYS G++LLE+ TG +P+ K + + Y GS
Sbjct: 781 PGYLDPEYFLTHQLTDKSDVYSLGVVLLELFTGMQPITHGKNIVREINI-----AYESGS 835
Query: 445 VVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
++ +D M +V + L K L+ +C RP+M V +L
Sbjct: 836 ILSTVDKRM-SSVPDECLEKFATLALRCCREETDARPSMAEVVREL 880
>At5g39000 protein kinase - like protein
Length = 873
Score = 229 bits (585), Expect = 2e-60
Identities = 124/292 (42%), Positives = 186/292 (63%), Gaps = 10/292 (3%)
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGL-VVAVKRAKREHFESLRTEFSSEVE 265
++ ++ AT +F + L IG GGFG+VYK ++ G +VAVKR + + + EF +E+E
Sbjct: 507 SIFEIKSATNDFEDKLIIGVGGFGSVYKGQIDGGATLVAVKRLEITSNQGAK-EFETELE 565
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHL---DGLRGKILDFNQRLEIAI 322
+L+K+ H +LV L+G+ D+ NE +L+ EY+P+GTL++HL D L + +RLEI I
Sbjct: 566 MLSKLRHVHLVSLIGYCDEDNEMVLVYEYMPHGTLKDHLFRRDKTSDPPLSWKRRLEICI 625
Query: 323 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVK 382
A GL YLH A+ IIHRD+K++NILL E+ KV+DFG +R+GP + QTH+ST VK
Sbjct: 626 GAARGLQYLHTGAKYTIIHRDIKTTNILLDENFVTKVSDFGLSRVGPTSASQTHVSTVVK 685
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYN 441
GT GYLDPEY + LT KSDVYSFG++LLE+L RP+ ++ E+ L RW Y
Sbjct: 686 GTFGYLDPEYYRRQVLTEKSDVYSFGVVLLEVLC-CRPIRMQSVPPEQADLIRWVKSNYR 744
Query: 442 EGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAI 493
G+V +++D + + + L K +++ +C +RP M V +WA+
Sbjct: 745 RGTVDQIIDSDLSADITSTSLEKFCEIAVRCVQDRGMERPPMNDV---VWAL 793
>At2g21480 putative protein kinase
Length = 871
Score = 229 bits (584), Expect = 3e-60
Identities = 125/285 (43%), Positives = 177/285 (61%), Gaps = 6/285 (2%)
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
+LS++ T+NF + IG GGFG VY ++DG VA+KR + + + TEF +E+++
Sbjct: 514 SLSELQEVTKNFDASEIIGVGGFGNVYIGTIDDGTQVAIKRGNPQSEQGI-TEFHTEIQM 572
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAH 326
L+K+ HR+LV L+G+ D+ E IL+ EY+ NG R+HL G L + QRLEI I A
Sbjct: 573 LSKLRHRHLVSLIGYCDENAEMILVYEYMSNGPFRDHLYGKNLSPLTWKQRLEICIGAAR 632
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL YLH + IIHRDVKS+NILL E++ AKVADFG ++ V Q H+ST VKG+ G
Sbjct: 633 GLHYLHTGTAQGIIHRDVKSTNILLDEALVAKVADFGLSK--DVAFGQNHVSTAVKGSFG 690
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVV 446
YLDPEY + QLT KSDVYSFG++LLE L R + + ++ WA +G +
Sbjct: 691 YLDPEYFRRQQLTDKSDVYSFGVVLLEALCARPAINPQLPREQVNLAEWAMLWKQKGLLE 750
Query: 447 ELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
+++DP + AVN + + K + + +C A DRP M G+ LW
Sbjct: 751 KIIDPHLVGAVNPESMKKFAEAAEKCLADYGVDRPTM---GDVLW 792
>At4g39110 putative receptor-like protein kinase
Length = 878
Score = 229 bits (583), Expect = 3e-60
Identities = 124/285 (43%), Positives = 178/285 (61%), Gaps = 6/285 (2%)
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVEL 266
+LS++ AT+NF + IG GGFG VY L+DG VAVKR + + + TEF +E+++
Sbjct: 515 SLSELQEATKNFEASQIIGVGGFGNVYIGTLDDGTKVAVKRGNPQSEQGI-TEFQTEIQM 573
Query: 267 LAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAH 326
L+K+ HR+LV L+G+ D+ +E IL+ E++ NG R+HL G L + QRLEI I A
Sbjct: 574 LSKLRHRHLVSLIGYCDENSEMILVYEFMSNGPFRDHLYGKNLAPLTWKQRLEICIGSAR 633
Query: 327 GLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVG 386
GL YLH + IIHRDVKS+NILL E++ AKVADFG ++ V Q H+ST VKG+ G
Sbjct: 634 GLHYLHTGTAQGIIHRDVKSTNILLDEALVAKVADFGLSK--DVAFGQNHVSTAVKGSFG 691
Query: 387 YLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVV 446
YLDPEY + QLT KSDVYSFG++LLE L R + + ++ WA + +G +
Sbjct: 692 YLDPEYFRRQQLTDKSDVYSFGVVLLEALCARPAINPQLPREQVNLAEWAMQWKRKGLLE 751
Query: 447 ELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
+++DP + +N + + K + + +C DRP M G+ LW
Sbjct: 752 KIIDPHLAGTINPESMKKFAEAAEKCLEDYGVDRPTM---GDVLW 793
>At5g38990 protein kinase - like protein
Length = 880
Score = 228 bits (582), Expect = 5e-60
Identities = 124/292 (42%), Positives = 187/292 (63%), Gaps = 10/292 (3%)
Query: 207 NLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGL-VVAVKRAKREHFESLRTEFSSEVE 265
++ ++ AT +F E L IG GGFG+VYK ++ G +VAVKR + + + EF +E+E
Sbjct: 514 SIYEIKSATNDFEEKLIIGVGGFGSVYKGRIDGGATLVAVKRLEITSNQGAK-EFDTELE 572
Query: 266 LLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHL---DGLRGKILDFNQRLEIAI 322
+L+K+ H +LV L+G+ D NE +L+ EY+P+GTL++HL D L + +RLEI I
Sbjct: 573 MLSKLRHVHLVSLIGYCDDDNEMVLVYEYMPHGTLKDHLFRRDKASDPPLSWKRRLEICI 632
Query: 323 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVK 382
A GL YLH A+ IIHRD+K++NILL E+ AKV+DFG +R+GP + QTH+ST VK
Sbjct: 633 GAARGLQYLHTGAKYTIIHRDIKTTNILLDENFVAKVSDFGLSRVGPTSASQTHVSTVVK 692
Query: 383 GTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTL-RWAFRKYN 441
GT GYLDPEY + LT KSDVYSFG++LLE+L RP+ ++ E+ L RW +N
Sbjct: 693 GTFGYLDPEYYRRQILTEKSDVYSFGVVLLEVLC-CRPIRMQSVPPEQADLIRWVKSNFN 751
Query: 442 EGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLWAI 493
+ +V +++D + + + + K +++ +C +RP M V +WA+
Sbjct: 752 KRTVDQIIDSDLTADITSTSMEKFCEIAIRCVQDRGMERPPMNDV---VWAL 800
>At3g46350 receptor-like protein kinase homolog
Length = 871
Score = 228 bits (582), Expect = 5e-60
Identities = 147/402 (36%), Positives = 214/402 (52%), Gaps = 34/402 (8%)
Query: 93 KAGRKLL---QKDSSKESTSQGDSKALPTNKVGIFAGGALLACCAVLCPCFYGKKRKATA 149
K G K+L K+ STS K V I A + L F+G ++K T+
Sbjct: 462 KKGLKILFDGDKNDPCLSTSCNPKKKFSVMIVAIVASTVVFVLVVSLA-LFFGLRKKKTS 520
Query: 150 HAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLHTLNLNLS 209
V A P P +PL S S S ++ R + S
Sbjct: 521 SHVKAIPPS--------------------PTTPLENVMSTSISETSIEMKRK---KFSYS 557
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAK 269
+V++ T NF L G+GGFGTVY +L+ VAVK + + + EF +EV+LL +
Sbjct: 558 EVMKMTNNFQRAL--GEGGFGTVYHGDLDSSQQVAVKLLSQSSTQGYK-EFKAEVDLLLR 614
Query: 270 IDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRG-KILDFNQRLEIAIDVAHGL 328
+ H NL+ L+G+ D+ + LI EY+ NG L+ HL G G +L +N RL IA+D A GL
Sbjct: 615 VHHINLLNLVGYCDERDHLALIYEYMSNGDLKHHLSGEHGGSVLSWNIRLRIAVDAALGL 674
Query: 329 TYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYL 388
YLH+ ++HRDVKS+NILL E+ AK+ADFG +R + G ++H+ST V G++GYL
Sbjct: 675 EYLHIGCRPSMVHRDVKSTNILLDENFMAKIADFGLSR-SFILGGESHVSTVVAGSLGYL 733
Query: 389 DPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVEL 448
DPEY +T +L SDVYSFGI+LLEI+T +R ++ K ++ W N G + +
Sbjct: 734 DPEYYRTSRLAEMSDVYSFGIVLLEIITNQRVID--KTREKPHITEWTAFMLNRGDITRI 791
Query: 449 MDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
MDP + N+ + + L+L+ CA P +RP+M V +L
Sbjct: 792 MDPNLNGDYNSHSVWRALELAMSCANPSSENRPSMSQVVAEL 833
>At5g49760 receptor protein kinase-like
Length = 953
Score = 227 bits (579), Expect = 1e-59
Identities = 121/282 (42%), Positives = 188/282 (65%), Gaps = 4/282 (1%)
Query: 210 QVVRATRNFSETLQIGQGGFGTVYKANLEDGLVVAVKRAKREHFESLRTEFSSEVELLAK 269
++ + T NFSE +G GG+G VY+ L +G ++A+KRA++ + EF +E+ELL++
Sbjct: 623 ELKKCTDNFSEANDVGGGGYGKVYRGILPNGQLIAIKRAQQGSLQG-GLEFKTEIELLSR 681
Query: 270 IDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAIDVAHGLT 329
+ H+N+V+LLGF NE++L+ EY+ NG+L++ L G G LD+ +RL+IA+ GL
Sbjct: 682 VHHKNVVRLLGFCFDRNEQMLVYEYISNGSLKDSLSGKSGIRLDWTRRLKIALGSGKGLA 741
Query: 330 YLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKVKGTVGYLD 389
YLH A+ IIHRD+KS+NILL E++ AKVADFG ++L + ++TH++T+VKGT+GYLD
Sbjct: 742 YLHELADPPIIHRDIKSNNILLDENLTAKVADFGLSKL-VGDPEKTHVTTQVKGTMGYLD 800
Query: 390 PEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYNEGSVVELM 449
PEY T+QLT KSDVY FG++LLE+LTGR P+E K V + + + + EL+
Sbjct: 801 PEYYMTNQLTEKSDVYGFGVVLLELLTGRSPIERGKYVVREVKTKMN-KSRSLYDLQELL 859
Query: 450 D-PLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQL 490
D ++ + N K +DL+ +C +RP+M V +++
Sbjct: 860 DTTIIASSGNLKGFEKYVDLALRCVEEEGVNRPSMGEVVKEI 901
>At3g51550 receptor protein kinase like protein
Length = 895
Score = 226 bits (576), Expect = 2e-59
Identities = 152/470 (32%), Positives = 245/470 (51%), Gaps = 42/470 (8%)
Query: 32 DWLAHSYSSHGQELFYI--NGNVVNQVSFCEALQLYIANGCDVKDYFGSSGCAVGGSFVN 89
D++ + +GQ+ ++ + N VN+ + ++L NG ++ S G G + +
Sbjct: 368 DYVVNPPEGNGQQDLWLALHPNPVNKPEYYDSL----LNGVEIFKMNTSDGNLAGTNPIP 423
Query: 90 FPSKAG--RKLLQKDSSKESTSQGDSKALPTNKVGIFAGGALLACCAVLCPCFYGKKRKA 147
P K+L+ + K SK+ G +G +LA C ++RK
Sbjct: 424 GPQVTADPSKVLRPTTRK-------SKSNTAIIAGAASGAVVLALIIGFCVFGAYRRRKR 476
Query: 148 -----TAHAVLAKDPISMDSATSFDASVSASVSGKIPASPLRVPSSPSRFSMSPKLSRLH 202
+ A P+S+ + S + +G +S +PS+ R
Sbjct: 477 GDYQPASDATSGWLPLSLYGNSHSAGSAKTNTTGSYASS---LPSNLCR----------- 522
Query: 203 TLNLNLSQVVRATRNFSETLQIGQGGFGTVYKANLEDGLV-VAVKRAKREHFESLRTEFS 261
+ + +++ AT+NF E+ +G GGFG VY+ ++ G VA+KR + + EF
Sbjct: 523 --HFSFAEIKAATKNFDESRVLGVGGFGKVYRGEIDGGTTKVAIKRGNPMSEQGVH-EFQ 579
Query: 262 SEVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIA 321
+E+E+L+K+ HR+LV L+G+ ++ E IL+ +Y+ +GT+REHL + L + QRLEI
Sbjct: 580 TEIEMLSKLRHRHLVSLIGYCEENCEMILVYDYMAHGTMREHLYKTQNPSLPWKQRLEIC 639
Query: 322 IDVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKV 381
I A GL YLH A+ IIHRDVK++NILL E AKV+DFG ++ GP D TH+ST V
Sbjct: 640 IGAARGLHYLHTGAKHTIIHRDVKTTNILLDEKWVAKVSDFGLSKTGPTL-DHTHVSTVV 698
Query: 382 KGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVELKKAADERVTLRWAFRKYN 441
KG+ GYLDPEY + QLT KSDVYSFG++L E L R + A ++ WA Y
Sbjct: 699 KGSFGYLDPEYFRRQQLTEKSDVYSFGVVLFEALCARPALNPTLAKEQVSLAEWAPYCYK 758
Query: 442 EGSVVELMDPLMEEAVNADVLMKMLDLSFQCAAPIRTDRPNMKSVGEQLW 491
+G + +++DP ++ + + K + + +C +RP+M G+ LW
Sbjct: 759 KGMLDQIVDPYLKGKITPECFKKFAETAMKCVLDQGIERPSM---GDVLW 805
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,607,954
Number of Sequences: 26719
Number of extensions: 444294
Number of successful extensions: 4650
Number of sequences better than 10.0: 1016
Number of HSP's better than 10.0 without gapping: 869
Number of HSP's successfully gapped in prelim test: 147
Number of HSP's that attempted gapping in prelim test: 1217
Number of HSP's gapped (non-prelim): 1118
length of query: 504
length of database: 11,318,596
effective HSP length: 104
effective length of query: 400
effective length of database: 8,539,820
effective search space: 3415928000
effective search space used: 3415928000
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0030.7


