

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0029a.5
(1393 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
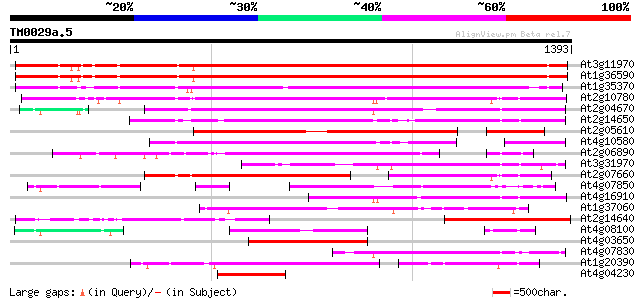
Sequences producing significant alignments: (bits) Value
At3g11970 hypothetical protein 1101 0.0
At1g36590 hypothetical protein 1099 0.0
At1g35370 hypothetical protein 986 0.0
At2g10780 pseudogene 711 0.0
At2g04670 putative retroelement pol polyprotein 645 0.0
At2g14650 putative retroelement pol polyprotein 634 0.0
At2g05610 putative retroelement pol polyprotein 624 e-179
At4g10580 putative reverse-transcriptase -like protein 516 e-146
At2g06890 putative retroelement integrase 480 e-135
At3g31970 hypothetical protein 452 e-127
At2g07660 putative retroelement pol polyprotein 384 e-106
At4g07850 putative polyprotein 382 e-106
At4g16910 retrotransposon like protein 378 e-104
At1g37060 Athila retroelment ORF 1, putative 311 1e-84
At2g14640 putative retroelement pol polyprotein 292 8e-79
At4g08100 putative polyprotein 288 2e-77
At4g03650 putative reverse transcriptase 285 1e-76
At4g07830 putative reverse transcriptase 276 5e-74
At1g20390 hypothetical protein 224 4e-58
At4g04230 putative transposon protein 198 2e-50
>At3g11970 hypothetical protein
Length = 1499
Score = 1101 bits (2848), Expect = 0.0
Identities = 591/1406 (42%), Positives = 859/1406 (61%), Gaps = 54/1406 (3%)
Query: 15 RVDIPMFNGNDAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLERA 74
++D P F+G W+ KVE FF + E K++M I + A W Q + + +
Sbjct: 111 KIDFPRFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTWHQSFIQSGVGLE 170
Query: 75 ----WEPFKQALFRRFQPALLQNPFGPLLSVKQKGSVMEYRENFELLAAPMRNADREVLK 130
W+ + + L RF+ +P L +++ +++Y + FEL+ + N E L
Sbjct: 171 VLYDWKGYVKLLKERFEDDC-DDPMAELKHLQETDGIIDYHQKFELIKTRV-NLSEEYLV 228
Query: 131 GVFLNGLQEEIKAEMKLYPADD------LAELMDRALLLEEKNT-------AMRGGKPKE 177
V+L GL+ + + ++++ L + ++A + NT A GG K
Sbjct: 229 SVYLAGLRTDTQMHVRMFQPQTVRHCLFLGKTYEKAHPKKPANTTWSTNRSAPTGGYNKY 288
Query: 178 EDKRGWKDLQNKGGTGNQDTEGKQPEKKWNGGQRLTQTELQERSRKGLCFKCGDKWGKEH 237
+ K G + G GN +QP+K ++Q E+ +R KGLC+ C +K+ EH
Sbjct: 289 Q-KEGESKTDHYGNKGNFKPVSQQPKK-------MSQQEMSDRRSKGLCYFCDEKYTPEH 340
Query: 238 ICSMKNYQLILMEVEEDEEEEEIFEEAEDGEFVLEGKVLQLSLNSKEGLTSNRSFKVKGK 297
K QL M+V DEE E+ EE + + + + Q+S+N+ G+ ++ +VKG
Sbjct: 341 YLVHKKTQLFRMDV--DEEFEDAREELVNDD---DEHMPQISVNAVSGIAGYKTMRVKGT 395
Query: 298 IGNREVLILIDCGATSNFISQDLVVELEIPVIATSEYVVEVGNGAKERNSGVCKNLKLEV 357
+ + ILID G+T NF+ + +L V V V +G K R G + ++
Sbjct: 396 YDKKIIFILIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWKL 455
Query: 358 QGISIMQHFFILGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCR 417
Q + ++ L G ++VLG+ WL +LG I F++L +++ QK++L G S
Sbjct: 456 QTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGS- 514
Query: 418 VTANWKSIKITEQQEAEGYYLSYEYQKEEEKTEAEV------------PEGMRKILEEYP 465
K+ K+ + QE + Q+ E TE E+ + ++L EYP
Sbjct: 515 -VREVKAQKLQKLQEDQVQLAMLCVQEVSESTEGELCTINALTSELGEESVVEEVLNEYP 573
Query: 466 EVFQEPKGLPP-RRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHS 524
++F EP LPP R +H I+L EG++ N RPYRY +QKNEI+KLV+++L +G ++ S
Sbjct: 574 DIFIEPTALPPFREKHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQAS 633
Query: 525 TSPFSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLK 584
+SP++SP +LVKKKDG WR CVDYR LN T+ D FPIP+I++L+DE+G AV+FSK+DL+
Sbjct: 634 SSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLR 693
Query: 585 SGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVF 644
+GYHQ+RM +DI KTAF+TH GH+EYLV+PFGLTNAP+TFQ LMN + +P+LRKFVLVF
Sbjct: 694 AGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVF 753
Query: 645 FDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADP 704
FDDIL+YS + E H+ HL+ V +V++ N L A KC+F P++ YLGH IS G+ DP
Sbjct: 754 FDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDP 813
Query: 705 SKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNSFQWTEGATQA 764
+KIK + +WP P +K LRGFLGL GYYRRFV+++ +A PL+ L K ++F+WT A QA
Sbjct: 814 AKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIAGPLHALTKTDAFEWTAVAQQA 873
Query: 765 FVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKS 824
F LK + PVL P FDK F++ETDA G+G+GAVLMQEG P+AY+S+ L + S
Sbjct: 874 FEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLS 933
Query: 825 VYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFE 884
+YE+EL+AV+ AV+KWRHYLL S F+I TDQRSL++L +QR+ QQ+W+ KL+ +D+E
Sbjct: 934 IYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYE 993
Query: 885 IKYKPGIENKAADALSR----KLQFSAISSVQCAEWADLEAEILEDERYRKVLQELATQG 940
I+Y+ G EN ADALSR ++ A++ V+C D++A D + + ++ L
Sbjct: 994 IQYRQGKENVVADALSRVEGSEVLHMAMTVVECDLLKDIQAGYANDSQLQDIITALQRDP 1053
Query: 941 NSAVGYQLKRGRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWE 1000
+S + + L K +IV+P T+L H + +GGH+G T++R+ LFYW+
Sbjct: 1054 DSKKYFSWSQNILRRKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKGLFYWK 1113
Query: 1001 GMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTI 1060
GM DIQ Y++ C CQ+ K + G LQPLPIP W+++SMDFI GLP + GK I
Sbjct: 1114 GMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPLPIPDTIWSEVSMDFIEGLPVSGGKTVI 1173
Query: 1061 LVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMF 1120
+VVVDR +K AHFIALSHPY+A +A ++ V +LHG PTSIVSDRD VF S FW E F
Sbjct: 1174 MVVVDRLSKAAHFIALSHPYSALTVAHAYLDNVFKLHGCPTSIVSDRDVVFTSEFWREFF 1233
Query: 1121 KLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHS 1180
L G LK +SAYHPQ+DGQTEVVNRC+ETYLRC+ +P+ W KWL+ AE+WYNTNYHS
Sbjct: 1234 TLQGVALKLTSAYHPQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAEYWYNTNYHS 1293
Query: 1181 AIKTTPFKALYGREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQA 1240
+ + TPF+ +YG+ PPV + V V + ER +L LK +L +AQ+RM+Q A
Sbjct: 1294 SSRMTPFEIVYGQVPPVHLPYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQHRMKQFA 1353
Query: 1241 NKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPE 1300
++HR + ++E+GD VY+K+QPY+ +S+ R+NQKLSP+Y+GPY II + AYKL LP
Sbjct: 1354 DQHRTEREFEIGDYVYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPS 1413
Query: 1301 GSQVHPVFHISLLKKAVNAGVQSQPLPAALTEEWELKVEPEAIMDTRENRDGD--LEVLI 1358
SQVHPVFH+S LK V + LP+ + + +E KV + + NR G +VL+
Sbjct: 1414 YSQVHPVFHVSQLKVLVGNVSTTVHLPSVMQDVFE-KVPEKVVERKMVNRQGKAVTKVLV 1472
Query: 1359 RWKDLPTFEDSWEDFSKLLDQFPNHQ 1384
+W + P E +WE L FP +
Sbjct: 1473 KWSNEPLEEATWEFLFDLQKTFPEFE 1498
>At1g36590 hypothetical protein
Length = 1499
Score = 1099 bits (2843), Expect = 0.0
Identities = 591/1406 (42%), Positives = 860/1406 (61%), Gaps = 54/1406 (3%)
Query: 15 RVDIPMFNGNDAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLERA 74
++D P F+G W+ KVE FF + E K++M I + A W Q + + +
Sbjct: 111 KIDFPRFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFDSHASTWHQSFIQSGVGLE 170
Query: 75 ----WEPFKQALFRRFQPALLQNPFGPLLSVKQKGSVMEYRENFELLAAPMRNADREVLK 130
W+ + + L RF+ +P L +++ +++Y + FEL+ + N E L
Sbjct: 171 VLYDWKGYVKLLKERFEDDC-DDPMAELKHLQETDGIIDYHQKFELIKTRV-NLSEEYLV 228
Query: 131 GVFLNGLQEEIKAEMKLYPADD------LAELMDRALLLEEKNT-------AMRGGKPKE 177
V+L GL+ + + ++++ L + ++A L + NT A GG K
Sbjct: 229 SVYLAGLRTDTQMHVRMFQPQTVRHCLFLGKTYEKAHLKKPANTTWSTNRSAPTGGYNKY 288
Query: 178 EDKRGWKDLQNKGGTGNQDTEGKQPEKKWNGGQRLTQTELQERSRKGLCFKCGDKWGKEH 237
+ K G + G GN +QP+K ++Q E+ +R KGLC+ C +K+ EH
Sbjct: 289 Q-KEGESKTDHYGNKGNFKPVSQQPKK-------MSQQEMSDRRSKGLCYFCDEKYTPEH 340
Query: 238 ICSMKNYQLILMEVEEDEEEEEIFEEAEDGEFVLEGKVLQLSLNSKEGLTSNRSFKVKGK 297
K QL M+V DEE E+ EE + + + + Q+S+N+ G+ ++ +VKG
Sbjct: 341 YLVHKKTQLFRMDV--DEEFEDAREELVNDD---DEHMPQISVNAVSGIAGYKTMRVKGT 395
Query: 298 IGNREVLILIDCGATSNFISQDLVVELEIPVIATSEYVVEVGNGAKERNSGVCKNLKLEV 357
+ + ILID G+T NF+ + +L V V V +G K R G + ++
Sbjct: 396 YDKKIIFILIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWKL 455
Query: 358 QGISIMQHFFILGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCR 417
Q + ++ L G ++VLG+ WL +LG I F++L +++ QK++L G S
Sbjct: 456 QTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGS- 514
Query: 418 VTANWKSIKITEQQEAEGYYLSYEYQKEEEKTEAEV------------PEGMRKILEEYP 465
K+ K+ + QE + Q+ E TE E+ + ++L EYP
Sbjct: 515 -VREVKAQKLQKLQEDQVQLAMLCVQEVSESTEGELCTINALTSELGEESVVEEVLNEYP 573
Query: 466 EVFQEPKGLPP-RRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHS 524
++F EP LPP R +H I+L EG++ N RPYRY +QKNEI+KLV+++L +G ++ S
Sbjct: 574 DIFIEPTALPPFREKHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQAS 633
Query: 525 TSPFSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLK 584
+SP++SP +LVKKKDG WR CVDYR LN T+ D FPIP+I++L+DE+G AV+FSK+DL+
Sbjct: 634 SSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLR 693
Query: 585 SGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVF 644
+GYHQ+RM +DI KTAF+TH GH+EYLV+PFGLTNAP+TFQ LMN + +P+LRKFVLVF
Sbjct: 694 AGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVF 753
Query: 645 FDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADP 704
FDDIL+YS + E H+ HL+ V +V++ N L A KC+F P++ YLGH IS G+ DP
Sbjct: 754 FDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDP 813
Query: 705 SKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNSFQWTEGATQA 764
+KIK + +WP P +K LRGFLGL GYYRRFV+++ +A PL+ L K ++F+WT A QA
Sbjct: 814 AKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIAGPLHALTKTDAFEWTAVAQQA 873
Query: 765 FVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKS 824
F LK + PVL P FDK F++ETDA G+G+GAVLMQEG P+AY+S+ L + S
Sbjct: 874 FEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLS 933
Query: 825 VYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFE 884
+YE+EL+AV+ AV+KWRHYLL S F+I TDQRSL++L +QR+ QQ+W+ KL+ +D+E
Sbjct: 934 IYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQRLNTPIQQQWLPKLLEFDYE 993
Query: 885 IKYKPGIENKAADALSR----KLQFSAISSVQCAEWADLEAEILEDERYRKVLQELATQG 940
I+Y+ G EN ADALSR ++ A++ V+C D++A D + + ++ L
Sbjct: 994 IQYRQGKENVVADALSRVEGSEVLHMAMTVVECDLLKDIQAGYANDSQLQDIITALQRDP 1053
Query: 941 NSAVGYQLKRGRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWE 1000
+S + + L K +IV+P T+L H + +GGH+G T++R+ LFY +
Sbjct: 1054 DSKKYFSWSQNILRRKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKGLFYSK 1113
Query: 1001 GMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTI 1060
GM DIQ Y++ C CQ+ K + G LQPLPIP W+++SMDFI GLP + GK I
Sbjct: 1114 GMIKDIQAYIRSCGTCQQCKSDPAASPGLLQPLPIPDTIWSEVSMDFIEGLPVSGGKTVI 1173
Query: 1061 LVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMF 1120
+VVVDR +K AHFIALSHPY+A +A+ ++ V +LHG PTSIVSDRD VF S FW E F
Sbjct: 1174 MVVVDRLSKAAHFIALSHPYSALTVAQAYLDNVFKLHGCPTSIVSDRDVVFTSEFWREFF 1233
Query: 1121 KLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHS 1180
L G LK +SAYHPQ+DGQTEVVNRC+ETYLRC+ +P+ W KWL+ AE+WYNTNYHS
Sbjct: 1234 TLQGVALKLTSAYHPQSDGQTEVVNRCLETYLRCMCHDRPQLWSKWLALAEYWYNTNYHS 1293
Query: 1181 AIKTTPFKALYGREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQA 1240
+ + TPF+ +YG+ PPV + V V + ER +L LK +L +AQ+RM+Q A
Sbjct: 1294 SSRMTPFEIVYGQVPPVHLPYLPGESKVAVVARSLQEREDMLLFLKFHLMRAQHRMKQFA 1353
Query: 1241 NKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPE 1300
++HR + ++E+GD VY+K+QPY+ +S+ R+NQKLSP+Y+GPY II + AYKL LP
Sbjct: 1354 DQHRTEREFEIGDYVYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPS 1413
Query: 1301 GSQVHPVFHISLLKKAVNAGVQSQPLPAALTEEWELKVEPEAIMDTRENRDGD--LEVLI 1358
SQVHPVFH+S LK V + LP+ + + +E KV + + NR G +VL+
Sbjct: 1414 YSQVHPVFHVSQLKVLVGNVSTTVHLPSVMQDVFE-KVPEKVVERKMVNRQGKAVTKVLV 1472
Query: 1359 RWKDLPTFEDSWEDFSKLLDQFPNHQ 1384
+W + P E +WE L FP +
Sbjct: 1473 KWSNEPLEEATWEFLFDLQKTFPEFE 1498
>At1g35370 hypothetical protein
Length = 1447
Score = 986 bits (2550), Expect = 0.0
Identities = 550/1385 (39%), Positives = 812/1385 (57%), Gaps = 95/1385 (6%)
Query: 15 RVDIPMFNGNDAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLERA 74
++D P F+G+ W+ KVE FF + E K++MV I + A W + + +
Sbjct: 116 KIDFPRFDGSRINEWLFKVEEFFGVDFTPEEMKVKMVAIHFDSHAATWHHSFIQSGIGLD 175
Query: 75 ----WEPFKQALFRRFQPALLQNPFGPLLSVKQKGSVMEYRENFELLAAPMRNADREVLK 130
W + + L RF+ A +P L +++ ++EY + FEL+ + N E L
Sbjct: 176 VFFNWPEYVKLLKDRFEDAC-DDPMAELKKLQETDGIVEYHQQFELIKVRL-NLSEEYLV 233
Query: 131 GVFLNGLQEEIKAEMKLYPADDLAELMDRALLLEEKNTAMRGGKPKEEDKRGWKDLQNKG 190
V+L GL+ + + ++++ + + + E + PK+ W +
Sbjct: 234 SVYLAGLRTDTQMHVRMFEPKTVRDCLRLGKYYERAH-------PKKTVSSTWSQKGTRS 286
Query: 191 GTGNQDTEGKQPEKKWNGGQRLTQTELQERSRKGLCFKCGDKWGKEHICSMKNYQLILME 250
G G R + Q+ GLC+ C +K+ EH K QL M+
Sbjct: 287 G----------------GSYRPVKEVEQKSDHLGLCYFCDEKFTPEHYLVHKKTQLFRMD 330
Query: 251 VEEDEEEEEIFEEAEDGEFVLEGKVLQLSLNSKEGLTSNRSFKVKGKIGNREVLILIDCG 310
V DEE E+ E D + + + Q+S+N+ G++ ++ VKG + R++ ILID G
Sbjct: 331 V--DEEFEDAVEVLSDDDHE-QKPMPQISVNAVSGISGYKTMGVKGTVDKRDLFILIDSG 387
Query: 311 ATSNFISQDLVVELEIPVIATSEYVVEVGNGAKERNSGVCKNLKLEVQGISIMQHFFILG 370
+T NFI + +L V + V V +G K G K ++Q + ++
Sbjct: 388 STHNFIDSTVAAKLGCHVESAGLTKVAVADGRKLNVDGQIKGFTWKLQSTTFQSDILLIP 447
Query: 371 LGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCRVTANWKSIKITEQ 430
L G ++VLG+ WL +LG I F++L +Q+ + Q++ L G +T + + IK +
Sbjct: 448 LQGVDMVLGVQWLETLGRISWEFKKLEMQFFYKNQRVWLHGI-----ITGSVRDIKAHKL 502
Query: 431 QEAEGYYLSY------EYQKEEEK--------TEAEVPEGM-RKILEEYPEVFQEPKGLP 475
Q+ + + E +EE+ T V E + + I+EE+P+VF EP LP
Sbjct: 503 QKTQADQIQLAMVCVREVVSDEEQEIGSISALTSDVVEESVVQNIVEEFPDVFAEPTDLP 562
Query: 476 P-RRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHSTSPFSSPAIL 534
P R DH I+L EGA+ N RPYRY +QK+EI+K+V++M+ SG I+ S+SPF+SP +L
Sbjct: 563 PFREKHDHKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIKSGTIQVSSSPFASPVVL 622
Query: 535 VKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKE 594
VKKKDG WR CVDY LN T+ D+F IP+I++L+DE+G +VVFSK+DL++GYHQ+RM
Sbjct: 623 VKKKDGTWRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQVRMDP 682
Query: 595 EDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKN 654
+DI KTAF+TH GH+EYLV+ FGLTNAP+TFQ+LMN V R +LRKFVLVFFDDILIYS +
Sbjct: 683 DDIQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILIYSSS 742
Query: 655 EELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWP 714
E HK+HLR+V +V++ + L A K +LGH IS + DP+KI+ + +WP
Sbjct: 743 IEEHKEHLRLVFEVMRLHKLFAKGSK--------EHLGHFISAREIETDPAKIQAVKEWP 794
Query: 715 IPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNSFQWTEGATQAFVKLKEVMTT 774
P VK +RGFLG GYYRRFV+N+ +A PL+ L K + F W+ A AF LK V+
Sbjct: 795 TPTTVKQVRGFLGFAGYYRRFVRNFGVIAGPLHALTKTDGFCWSLEAQSAFDTLKAVLCN 854
Query: 775 VPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELMAVV 834
PVL P FDK F++ETDA G+G+ AVLMQ+G P+AY+S+ L + S+YE+EL+A +
Sbjct: 855 APVLALPVFDKQFMVETDACGQGIRAVLMQKGHPLAYISRQLKGKQLHLSIYEKELLAFI 914
Query: 835 LAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIKYKPGIENK 894
AV+KWRHYLL S F+I TDQRSL++L +QR+ QQ+W+ KL+ +D+EI+Y+ G EN
Sbjct: 915 FAVRKWRHYLLPSHFIIKTDQRSLKYLLEQRLNTPVQQQWLPKLLEFDYEIQYRQGKENL 974
Query: 895 AADALSR----KLQFSAISSVQCAEWADLEAEILEDERYRKVLQELATQGNSAVGYQLKR 950
ADALSR ++ A+S V+C +++ D + ++ L ++ Y +
Sbjct: 975 VADALSRVEGSEVLHMALSIVECDFLKEIQVAYESDGVLKDIISALQQHPDAKKHYSWSQ 1034
Query: 951 GRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYV 1010
L K +IV+P +L+ H + +GG +G +++R+ +LFYW+GM DIQ ++
Sbjct: 1035 DILRRKSKIVVPNDVEITNKLLQWLHCSGMGGRSGRDASHQRVKSLFYWKGMVKDIQAFI 1094
Query: 1011 QKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKY 1070
+ C CQ+ K + G LQPLPIP + W D+SMDFI GLP + GK I+VVVDR +K
Sbjct: 1095 RSCGTCQQCKSDNAAYPGLLQPLPIPDKIWCDVSMDFIEGLPNSGGKSVIMVVVDRLSKA 1154
Query: 1071 AHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFS 1130
AHF+AL+HPY+A +A+ F+ V + HG PTSIVSDRD +F S FW E FKL G +L+ S
Sbjct: 1155 AHFVALAHPYSALTVAQAFLDNVYKHHGCPTSIVSDRDVLFTSDFWKEFFKLQGVELRMS 1214
Query: 1131 SAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKAL 1190
SAYHPQ+DGQTEVVNRC+E YLRC+ ++P W KWL AE+WYNTNYHS+ + TPF+ +
Sbjct: 1215 SAYHPQSDGQTEVVNRCLENYLRCMCHARPHLWNKWLPLAEYWYNTNYHSSSQMTPFELV 1274
Query: 1191 YGREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYE 1250
YG+ PP+ + V V + ER +L LK +L +AQ+RM+Q A++HR + ++
Sbjct: 1275 YGQAPPIHLPYLPGKSKVAVVARSLQERENMLLFLKFHLMRAQHRMKQFADQHRTERTFD 1334
Query: 1251 VGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQVHPVFHI 1310
+GD VY+K+QPY+ +S+ R NQKLSP+Y+GPY II K
Sbjct: 1335 IGDFVYVKLQPYRQQSVVLRVNQKLSPKYFGPYKIIEKCG-------------------- 1374
Query: 1311 SLLKKAVNAGVQSQPLPAALTEEWELKVEPEAIMD----TRENRDGDLEVLIRWKDLPTF 1366
+ V S LP+ L + +E PE I++ R+ R + VL++W P
Sbjct: 1375 ---EVMVGNVTTSTQLPSVLPDIFE--KAPEYILERKLVKRQGRAATM-VLVKWIGEPVE 1428
Query: 1367 EDSWE 1371
E +W+
Sbjct: 1429 EATWK 1433
>At2g10780 pseudogene
Length = 1611
Score = 711 bits (1836), Expect = 0.0
Identities = 474/1419 (33%), Positives = 754/1419 (52%), Gaps = 99/1419 (6%)
Query: 29 WVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLE--RAWEPFKQALFRRF 86
W ++++R F+ +R E + ++ + +E A W+ E++ + R++ F+ +++
Sbjct: 194 WRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVEKRRGDEVRSFADFEDEFNKKY 253
Query: 87 QP--ALLQNPFGPLLSVKQKGSVMEYRENFELLAAPM-RNADREVLK-GVFLNGLQEEIK 142
P A + L V+ +V EY E F L + R + E + F+ GL+ EI+
Sbjct: 254 FPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVGRELEEEQAQLRRFIRGLRIEIR 313
Query: 143 AEMKLYPADDLAELMDRALLLEEKNTAMRGGKPKEEDKRGWKDLQNKGGTGNQDTEGKQP 202
+ + ++EL++RA ++EE G + + R ++N NQ T+
Sbjct: 314 NHCLVRTFNSVSELVERAAMIEE------GIEEERYLNREKAPIRN-----NQSTKPADK 362
Query: 203 EKKWNGGQRLTQTELQE-------RSRKGLCFKCGDKWGKEHICSMKNYQLILMEVEEDE 255
++K++ T+++ + ++ G C+K G+ C K++ + E
Sbjct: 363 KRKFDKVDN-TKSDAKTGECVTCGKNHSGTCWKAIGACGR---CGSKDHAIQSCPKMEPG 418
Query: 256 EEEEIFEEAEDGEFV-----LEGKVLQLSLNSKEGLTSNRSFKVKGKIGNREVLI--LID 308
+ + + EE + L+ + +L+ + G NR R+ + + +
Sbjct: 419 QSKVLGEETRTCFYCGKTGHLKRECPKLTAEKQAGQRDNRGGNGLPPPPKRQAVASRVYE 478
Query: 309 CGATSNFISQDLVVELEIPVIATSEY-VVEVGNGAKERNSGVCKNLKLEVQGISIMQHFF 367
+N + ++Y +V G +G+ + + + V G+++
Sbjct: 479 LSEEANDAGNFRAITGGFRKEPNTDYGMVRAAGGQAMYPTGLVRGISVVVNGVNMPADLI 538
Query: 368 ILGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCRVTANWKSIKI 427
I+ L +V+LGMDWL + IQ+ + QG R T+ I
Sbjct: 539 IVPLKKHDVILGMDWLGKY-KAHIDCHRGRIQFERDEGMLKFQG----IRTTSGSLVISA 593
Query: 428 TEQQEAEGY----YLSYEYQKEEEKTEAEVPEGMRKILEEYPEVFQEPKGLPPRRTTDHA 483
+ + G YL+ KE + AE+ + I+ E+ +VF G+PP R+
Sbjct: 594 IQAERMLGKGCEAYLATITTKEVGAS-AELKD--IPIVNEFSDVFAAVSGVPPDRSDPFT 650
Query: 484 IQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHSTSPFSSPAILVKKKDGGWR 543
I+L+ G + + PYR + +++K ++E+L+ G IR S+SP+ +P + VKKKDG +R
Sbjct: 651 IELEPGTTPISKAPYRMAPAEMAKLKKQLEELLDKGFIRPSSSPWGAPVLFVKKKDGSFR 710
Query: 544 FCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFR 603
C+DYR LNK T+ +K+P+P IDEL+D++G A FSK+DL SGYHQI ++ D+ KTAFR
Sbjct: 711 LCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFR 770
Query: 604 THEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEELHKDHLR 663
T H+E++V+PFGLTNAP+ F +MN V R +L +FV++F +DIL+YSK+ E H++HLR
Sbjct: 771 TRYDHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFINDILVYSKSWEAHQEHLR 830
Query: 664 IVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWPIPKEVKGLR 723
VL+ L+E+ L A KCSF Q + +LGHVIS GV+ DP KI+ + +WP P+ +R
Sbjct: 831 AVLERLREHELFAKLSKCSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIR 890
Query: 724 GFLGLTGYYRRFVKNYSKLAQPLNQLL-KKNSFQWTEGATQAFVKLKEVMTTVPVLVPPN 782
FLGL GYYRRFV +++ +AQPL +L K +F W++ ++F++LK ++T PVLV P
Sbjct: 891 SFLGLAGYYRRFVMSFASMAQPLTRLTGKDTAFNWSDECEKSFLELKAMLTNAPVLVLPE 950
Query: 783 FDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELMAVVLAVQKWRH 842
+P+ + TDAS GLG VLMQ+G +AY S+ L + ++ E+ AVV ++ WR
Sbjct: 951 EGEPYTVYTDASIVGLGCVLMQKGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRS 1010
Query: 843 YLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIKYKPGIENKAADALSRK 902
YL G+K I+TD +SL+++ Q + Q++WM + Y+ +I Y PG N+ ADALSR+
Sbjct: 1011 YLYGAKVQIYTDHKSLKYIFTQPELNLRQRRWMELVADYNLDIAYHPGKANQVADALSRR 1070
Query: 903 -------------------LQFSAIS------SVQCAEWADLEAEI-LEDERYRKVLQEL 936
L +A+S + A+ ADL + I L ER ++ +
Sbjct: 1071 RSEVEAERSQVDLVNMMGTLHVNALSKEVEPLGLGAADQADLLSRIRLAQERDEEI--KG 1128
Query: 937 ATQGNSAVGYQLKRGRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISAL 996
Q N G ++ R+ +P +L+E H + H G + Y+ +
Sbjct: 1129 WAQNNKTEYQTSNNGTIVVNGRVCVPNDRALKEEILREAHQSKFSIHPGSNKMYRDLKRY 1188
Query: 997 FYWEGMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMG 1056
++W GMK D+ +V KC CQ K E P+G LQ LPIP W I+MDF+ GLP +
Sbjct: 1189 YHWVGMKKDVARWVAKCPTCQLVKAEHQVPSGLLQNLPIPEWKWDHITMDFVTGLPTGIK 1248
Query: 1057 K--DTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLST 1114
+ + VVVDR TK AHF+A+S A+ IAE +I E+VRLHG P SIVSDRD F S
Sbjct: 1249 SKHNAVWVVVDRLTKSAHFMAISDKDGAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSK 1308
Query: 1115 FWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWY 1174
FW K GT++ S+AYHPQTD Q+E + +E LR W K+L EF Y
Sbjct: 1309 FWKAFQKALGTRVNLSTAYHPQTDEQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAY 1368
Query: 1175 NTNYHSAIKTTPFKALYGRE-------PPVIFKGNDSLTSVDEVEKLTAERNLILEELKS 1227
N ++ ++I +P++ALYGR PV + T VDE T ER ++ LK
Sbjct: 1369 NNSFQASIGMSPYEALYGRACRTPLCWTPVGERRLFGPTIVDE----TTER---MKFLKI 1421
Query: 1228 NLEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIA 1287
L++AQ+R + ANK R++++++VGDLVYLK YK + S +KLSPRY GPY +I
Sbjct: 1422 KLKEAQDRQKSYANKRRKELEFQVGDLVYLKAMTYK-GAGRFTSRKKLSPRYVGPYKVIE 1480
Query: 1288 KINPAAYKLQL-PEGSQVHPVFHISLLKKAVNAGVQS-QPLPAALTEEWELKVEPEAIMD 1345
++ AYKL L P+ + H VFH+S L+K ++ +S + +P L E ++ P IMD
Sbjct: 1481 RVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVEDIPPGLKENMTVEAWPVRIMD 1540
Query: 1346 --TRENRDGDLEVL-IRWKDLPTFEDSWEDFSKLLDQFP 1381
T+ R ++L + W E +WE +K+ FP
Sbjct: 1541 RMTKGTRGKARDLLKVLWNCRGREEYTWETENKMKANFP 1579
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 645 bits (1664), Expect = 0.0
Identities = 383/1078 (35%), Positives = 583/1078 (53%), Gaps = 74/1078 (6%)
Query: 336 VEVGNGAKERNSGVCKNLKLEVQGISIMQHFFILGLGGTEVVLGMDWLASLGNIEANFQE 395
V+V G G K + +++ G S+ I + +V+LGMDWL + +
Sbjct: 365 VKVAGGQFLAVLGRAKGVDIQIAGESMPADLIISPVELYDVILGMDWLDHY-RVHLDCHR 423
Query: 396 LIIQWVSQGQKMVLQG-EPSVCRVTANWKSIKITEQQEAEGYYLSYEYQKEEEKTEAEVP 454
+ + ++V QG P+ + + K ++ E Y ++ + + +V
Sbjct: 424 GRVSFERPEGRLVYQGVRPTSGSLVISAVQAKKMIEKGCEAYLVTISMPE----SLGQVA 479
Query: 455 EGMRKILEEYPEVFQEPKGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKE 514
++++E+ +VFQ +GLPP R+ I+L+ G + + PYR + E++K +++
Sbjct: 480 VSDIRVIQEFEDVFQSLQGLPPSRSDPFTIELEPGTAPLSKAPYRMAPAEMTELKKQLED 539
Query: 515 MLNSGIIRHSTSPFSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGA 574
+L G IR STSP+ +P + VKKKDG +R C+DYR LN T+ +K+P+P IDELLD++
Sbjct: 540 LLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRG 599
Query: 575 AVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLR 634
A FSK+DL SGYHQI + E D+ KTAFRT GH+E++V+PF LTNAP+ F LMN V +
Sbjct: 600 ATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFALTNAPAAFMRLMNSVFQ 659
Query: 635 PYLRKFVLVFFDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHV 694
+L +FV++F DDIL+YSK+ E H+ HLR V++ L+E L A KCSF Q EI +LGH+
Sbjct: 660 EFLDEFVIIFIDDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREIGFLGHI 719
Query: 695 ISQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKN- 753
+S GV+ DP KI+ + DWP P +R FL LTGYYRRFVK ++ +AQP+ +L K+
Sbjct: 720 VSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLRLTGYYRRFVKGFASMAQPMTKLTGKDV 779
Query: 754 SFQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMS 813
F W+ + FV LKE++T+ PVL P +P+++ TDAS GLG VLMQ G+ +AY S
Sbjct: 780 PFVWSPECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQRGKVIAYAS 839
Query: 814 KTLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQK 873
+ L ++ E+ V+ A++ WR YL G K + TD +SL+++ +Q + Q +
Sbjct: 840 RQLRKHEGNYPTHDLEMAVVIFALKIWRSYLYGGKVQVFTDHKSLKYIFNQPELNLRQMR 899
Query: 874 WMSKLMGYDFEIKYKPGIENKAADALSRK-------------------LQFSAISSVQCA 914
WM + YD EI Y PG N ADALSRK L+ ++
Sbjct: 900 WMELVADYDLEIAYHPGKANVVADALSRKRVGAAPGQSVEALVSEIGALRLCVVAREPLG 959
Query: 915 EWADLEAEILEDERYRKVLQELATQGNSAVG--YQL-KRGRLLYKDRIVLPKGSTKILTV 971
A A++L R + E + A G YQ G + R+ +PK +
Sbjct: 960 LEAVDRADLLTRARLAQEKDEGLIAASKAEGSEYQFAANGTIFVYGRVCVPKDEELRREI 1019
Query: 972 LKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQKCEVCQRNKYEALNPAGFLQ 1031
L E H + H G + Y+ + + W GMK D+ N+V +C+VCQ K E P
Sbjct: 1020 LSEAHASMFSIHPGATKMYRDLKRYYQWVGMKRDVANWVAECDVCQLVKAEHQVP----- 1074
Query: 1032 PLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIK 1091
D I V++DR TK AHF+A+ A +A+ ++
Sbjct: 1075 --------------------------DAIWVIMDRLTKSAHFLAIRKTDGAAVLAKKYVS 1108
Query: 1092 EVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETY 1151
E+V+LHG P SIVSDRD F FW GTK++ S+AYHPQTDGQ+E + +E
Sbjct: 1109 EIVKLHGVPVSIVSDRDSKFTFAFWRAFQAKMGTKVQMSTAYHPQTDGQSERTIQTLEDM 1168
Query: 1152 LRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGREPPVIFKGND----SLTS 1207
LR W LS EF YN +Y ++I PF+ALYGR + S+
Sbjct: 1169 LRMCVLDWGGHWADHLSLVEFAYNNSYQASIGMAPFEALYGRPCWTPLRWTQVEERSIYG 1228
Query: 1208 VDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSL 1267
D V++ T ER + LK N+++AQ R R A+K RR++++EVGD VYLK+ + +
Sbjct: 1229 ADYVQE-TTER---IRVLKLNMKEAQARQRSYADKRRRELEFEVGDRVYLKMAMLRGPNR 1284
Query: 1268 AKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQV-HPVFHISLLKKAVNAGVQ-SQP 1325
+ KLSPRY GP+ I+ ++ P AY+L+LP+ + H VFH+ +L+K ++ +
Sbjct: 1285 SILET-KLSPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVLMLRKCLHKDDEVLVK 1343
Query: 1326 LPAALTEEWELKVEPEAIMDTR--ENRDGDLEVL-IRWKDLPTFEDSWEDFSKLLDQF 1380
+P L L+ P +++ R E R + ++ + W E++WE +++ +F
Sbjct: 1344 IPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLWDCDGVTEETWEPEARMKARF 1401
Score = 43.5 bits (101), Expect = 8e-04
Identities = 49/200 (24%), Positives = 80/200 (39%), Gaps = 33/200 (16%)
Query: 25 DAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLER-----AWEPFK 79
+A W ++V+R F SR ++++ + +E A WW T R +W F
Sbjct: 146 EADSWRSRVQRNFGSSRCPAEYRVDLAVHFLEGDA---HLWWRSVTARRRQADMSWADFV 202
Query: 80 QALFRRFQPA-LLQNPFGPLLSVKQ-KGSVMEYRENFELLAAPMRNA--DREVLKGVFLN 135
++ P L L + Q + SV EY F L A D + FL
Sbjct: 203 AEFNAKYFPQEALDRMEARFLELTQGEWSVREYDREFNRLLAYAGRGMEDDQAQMRRFLR 262
Query: 136 GLQEEIKAEMKLYPADDLAELMDRALLLEE---------------KNTAM-----RGGKP 175
GL+ +++ + ++ A L++ A +EE KNT +GGKP
Sbjct: 263 GLRPDLRVQCRVSQYATKAALVETAAEVEEDLQRQVVGVSPAVQTKNTQQQVTPSKGGKP 322
Query: 176 KEEDKRGWKDLQNKGGTGNQ 195
+ KR W D ++ G G +
Sbjct: 323 AQGQKRKW-DHPSRAGQGGR 341
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 634 bits (1635), Expect = 0.0
Identities = 398/1094 (36%), Positives = 586/1094 (53%), Gaps = 91/1094 (8%)
Query: 298 IGNREVLILIDCGATSNFISQDLVVELEIPVIATSEY-VVEVGNGAKERNSGVCKNLKLE 356
+G E +L D GA+ FI+ + I + V+V G G K + ++
Sbjct: 308 VGGVEAHVLFDSGASHCFITPESASRGNIRGDPGEQLGAVKVAGGQFVAVLGRTKGVDIQ 367
Query: 357 VQGISIMQHFFILGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVC 416
+ G S+ I + +V+LGMDWL + + + + +V QG
Sbjct: 368 IAGESMPADLIISPVELYDVILGMDWLDHY-RVHLDCHRGRVSFERPEGSLVYQG----- 421
Query: 417 RVTANWKSIKITEQQEAEGYYLSYEYQKEEEKTEAEVPEGMRKILEEYPEVFQEPKGLPP 476
V S+ I+ Q E + +G LE + +VFQ +GLPP
Sbjct: 422 -VRPTSGSLVISAVQ-----------------AEKMIEKGCEAYLE-FEDVFQSLQGLPP 462
Query: 477 RRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHSTSPFSSPAILVK 536
R+ I+L+ G + + PYR + E++K ++++L + P + VK
Sbjct: 463 SRSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLEDLLGA------------PVLFVK 510
Query: 537 KKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEED 596
KKDG +R C+DYR LN T+ +K+P+P IDELLD++ A FSK+DL SGYH I + E D
Sbjct: 511 KKDGSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEAD 570
Query: 597 IPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEE 656
+ KTAFRT GH+E++V+PFGLTNAP+ F LMN V + L +FV++F DDIL+YSK+ E
Sbjct: 571 VRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLE 630
Query: 657 LHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWPIP 716
H+ HLR V++ L+E L A KCSF Q E+ +LGH++S GV+ DP KI+ + DW P
Sbjct: 631 EHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTP 690
Query: 717 KEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKN-SFQWTEGATQAFVKLKEVMTTV 775
+R FLGL GYYRRFVK ++ +AQP+ +L K+ F W+ + FV LKE++T+
Sbjct: 691 TNATEIRSFLGLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTST 750
Query: 776 PVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELMAVVL 835
PVL P +P+++ TDASG GLG VLMQ G+ +AY S+ L ++ E+ AV+
Sbjct: 751 PVLALPEHGEPYMVYTDASGVGLGCVLMQRGKVIAYASRQLRKHEGNYPTHDLEMAAVIF 810
Query: 836 AVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIKYKPGIENKA 895
A++ WR YL G + TD +SL+++ Q + Q++WM + YD EI Y PG N
Sbjct: 811 ALKIWRSYLYGGNVQVFTDHKSLKYIFTQPELNLRQRQWMELVADYDLEIAYHPGKANVV 870
Query: 896 ADALSRKLQFSAISSVQCAEWADLEAEILEDERYRKVLQELATQGNSAVGYQLKRGRLLY 955
ADALS K V A +EA + E R G AV R LL
Sbjct: 871 ADALSHK-------RVGAAPGQSVEALVSEIGALRLCAVAREPLGLEAV----DRADLLT 919
Query: 956 KDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQKCEV 1015
+ R+ K G+ YK EG + D+ N+V +C+V
Sbjct: 920 RVRLAQEKDE-------------------GLIAAYKA-------EGSE-DVANWVAECDV 952
Query: 1016 CQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIA 1075
CQ K E P G LQ LPIP W I++DF+ GLP + KD I V+VDR TK AHF+A
Sbjct: 953 CQLVKAEHQVPGGMLQSLPIPEWKWDFITIDFVVGLPVSRTKDAIWVIVDRLTKSAHFLA 1012
Query: 1076 LSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHP 1135
+ A +A+ ++ E+V+LHG P SIVSDRD F S FW GTK++ S+AYHP
Sbjct: 1013 IRKTDGAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGTKVQMSTAYHP 1072
Query: 1136 QTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGR-- 1193
QT GQ+E + +E LR W LS EF YN +Y ++I PF+ALY R
Sbjct: 1073 QTYGQSERTIQTLEDMLRMCVLDWGGHWADHLSLVEFAYNNSYPASIGMAPFEALYERPC 1132
Query: 1194 EPPVIFK--GNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEV 1251
P+ G S+ D V++ T ER + LK N+++AQ+R R A+K RR++++EV
Sbjct: 1133 RTPLCLTQVGERSIYGADYVQE-TTER---IRVLKLNMKEAQDRQRSYADKRRRELEFEV 1188
Query: 1252 GDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQV-HPVFHI 1310
GD VYLK+ + + S KLSPRY GP+ I+ ++ P AY+L+LP+ + H VFH+
Sbjct: 1189 GDRVYLKMAMLRGPN-RSISETKLSPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHV 1247
Query: 1311 SLLKKAVNAGVQ-SQPLPAALTEEWELKVEPEAIMDTR--ENRDGDLEVL-IRWKDLPTF 1366
S+L+K ++ + +P L L+ P +++ R E R + ++ + W
Sbjct: 1248 SMLRKCLHKDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLWDCDGVT 1307
Query: 1367 EDSWEDFSKLLDQF 1380
+++WE +++ +F
Sbjct: 1308 KETWEPEARMKARF 1321
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 624 bits (1610), Expect = e-179
Identities = 314/661 (47%), Positives = 435/661 (65%), Gaps = 56/661 (8%)
Query: 457 MRKILEEYPEVFQEPKGLPPRRTT-DHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEM 515
+ K++ E+P++F EP LPP R DH I+L E ++ N RPYRY +QKNEI+K+V +M
Sbjct: 7 VEKVVTEFPDIFVEPTDLPPFRAPHDHKIELLEDSNPVNQRPYRYVVHQKNEIDKIVDDM 66
Query: 516 LNSGIIRHSTSPFSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAA 575
L SG I+ S+SP++SP +LVKKKDG WR CVDYR LN T+ D+FPIP+I++L+DE+G +
Sbjct: 67 LASGTIQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELGGS 126
Query: 576 VVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRP 635
V+SK+DL++GYHQ+RM DI KTAF+TH GHYEYLV+PFGLTNAP++FQ+LMN +P
Sbjct: 127 NVYSKIDLRAGYHQVRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQSLMNSFFKP 186
Query: 636 YLRKFVLVFFDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVI 695
+LRKFVLVFFDDILIYS + E HK HL V +V++ ++L A KC+F P + YLGH I
Sbjct: 187 FLRKFVLVFFDDILIYSTSMEEHKKHLEAVFEVMRVHHLFAKMSKCAFAVPRVEYLGHFI 246
Query: 696 SQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNSF 755
S G+A DP+KIK + DWP+P +K L GFLGLTGYYRRF
Sbjct: 247 SGEGIATDPAKIKAVQDWPVPVNLKQLCGFLGLTGYYRRF-------------------- 286
Query: 756 QWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKT 815
F++E DA G G+GAVLMQEG P+A++S+
Sbjct: 287 -------------------------------FVVEMDACGHGIGAVLMQEGHPLAFISRQ 315
Query: 816 LSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWM 875
L + S+YE+EL+AV+ V+KWRHYL+ S F+I TDQRSL++L +QR+ QQ+W+
Sbjct: 316 LKGKQLHLSIYEKELLAVIFVVRKWRHYLISSHFIIKTDQRSLKYLLEQRLNTPIQQQWL 375
Query: 876 SKLMGYDFEIKYKPGIENKAADALSR----KLQFSAISSVQCAEWADLEAEILEDERYRK 931
KL+ +D+EI+YK G EN ADALSR ++ A+S V+C +++A + D +
Sbjct: 376 PKLLEFDYEIQYKQGKENLVADALSRVEGSEVLHMALSVVECDLLKEIQAGYVTDGDIQG 435
Query: 932 VLQELATQGNSAVGYQLKRGRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYK 991
++ L Q +S Y +G L K++IV+P S T+L+ H + +GGH+G T++
Sbjct: 436 IITILQQQADSKKHYTWSQGVLRRKNKIVVPNNSGIKDTILRWLHCSGMGGHSGKEVTHQ 495
Query: 992 RISALFYWEGMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGL 1051
R+ LFYW+ M DIQ +++ C CQ+ K + G LQPLPIP + W+D+SMDFI GL
Sbjct: 496 RVKGLFYWKSMVKDIQAFIRSCGTCQQCKSDNAASPGLLQPLPIPDRIWSDVSMDFIDGL 555
Query: 1052 PKAMGKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVF 1111
P + GK I+VVVDR +K AHFIAL+HPY+A +A+ ++ V +LHG P+SIV RV
Sbjct: 556 PLSNGKTVIMVVVDRLSKAAHFIALAHPYSAMTVAQAYLDNVFKLHGCPSSIVVSSRRVL 615
Query: 1112 L 1112
+
Sbjct: 616 V 616
Score = 126 bits (316), Expect = 1e-28
Identities = 63/142 (44%), Positives = 93/142 (65%)
Query: 1185 TPFKALYGREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHR 1244
TP++A+YG+ PP+ + V V + ER ++ LK +L +AQ+RM+Q A++H
Sbjct: 626 TPYEAVYGQPPPLHLPYLPGESKVAVVARSMQERESMILFLKFHLMRAQHRMKQLADQHI 685
Query: 1245 RDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQV 1304
+ ++EVGD V++K+QPY+ +S+ RS QKLSP+Y+GPY +I + AYKLQLP SQV
Sbjct: 686 TEREFEVGDYVFVKLQPYRQQSVVMRSTQKLSPKYFGPYKVIDRCGEVAYKLQLPANSQV 745
Query: 1305 HPVFHISLLKKAVNAGVQSQPL 1326
HPVFH+S L+ V S L
Sbjct: 746 HPVFHVSQLRVLVGTVTTSTHL 767
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 516 bits (1328), Expect = e-146
Identities = 286/763 (37%), Positives = 433/763 (56%), Gaps = 33/763 (4%)
Query: 348 GVCKNLKLEVQGISIMQHFFILGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKM 407
G K + +++ G S+ I + +V+LGMDWL + ++ + + ++
Sbjct: 351 GRAKGVDIQIAGESMPADLIISPVELYDVILGMDWL-DYYRVHLDWHRGRVFFERPEGRL 409
Query: 408 VLQGEPSVCRVTANWKSIKITEQQEAEGYYLSYEYQKEEEKTEAEVPEGMRKILEEYPEV 467
V QG V ++ + + ++ E +Y ++ +V ++++E+ +V
Sbjct: 410 VYQG---VRPISGSLVISAVQAEKMIEKGCEAYLVTISMPESVGQVAVSDIRVVQEFQDV 466
Query: 468 FQEPKGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHSTSP 527
FQ +GLPP ++ I+L+ G + + PYR + E++K +K++L G IR STSP
Sbjct: 467 FQSLQGLPPSQSDPFTIELEPGTAPLSKAPYRMAPAEMAELKKQLKDLLGKGFIRPSTSP 526
Query: 528 FSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGY 587
+ +P + VKKKDG +R C+DYR LN+ T+ +++P+P IDELLD++ A FSK+DL SGY
Sbjct: 527 WGAPVLFVKKKDGSFRLCIDYRELNRVTVKNRYPLPRIDELLDQLRGATCFSKIDLTSGY 586
Query: 588 HQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDD 647
HQI + E D+ KTAFRT GH+E++V+PFGLTNAP+ F LMN V + +L +FV++F DD
Sbjct: 587 HQIPIAEADVRKTAFRTRYGHFEFVVMPFGLTNAPAVFMRLMNSVFQEFLDEFVIIFIDD 646
Query: 648 ILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKI 707
IL+YSK+ E + HLR V++ L+E L A KCSF Q E+ +LGH++S GV+ DP KI
Sbjct: 647 ILVYSKSPEEQEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKI 706
Query: 708 KDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKN-SFQWTEGATQAFV 766
+ + DWP P +R FLG GYYRRFVK ++ +AQP+ +L K+ F W++ + FV
Sbjct: 707 EAIRDWPRPTNATEIRSFLGWAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSQECEEGFV 766
Query: 767 KLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVY 826
LKE++T+ PVL P +P+++ TDAS GLG VLMQ G+ +AY S+ L +
Sbjct: 767 SLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQHGKVIAYASRQLMKHEGNYPTH 826
Query: 827 ERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIK 886
+ E+ AV+ A++ WR YL G K + TD +SL+++ Q + Q++WM + YD EI
Sbjct: 827 DLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLEIA 886
Query: 887 YKPGIENKAADALSRKLQFSAISSVQCAEWADLEAEILEDERYRKVLQELATQGNSAVGY 946
Y PG N DALSRK +A+ +E + E R G AV
Sbjct: 887 YHPGKANVVVDALSRKRVGAALGQ-------SVEVLVSEIGALRLCAVAREPLGLEAV-- 937
Query: 947 QLKRGRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDI 1006
R LL + R+ K T + Y+ + + W GMK+D+
Sbjct: 938 --DRADLLTRVRLAQKKDEGLRAT-----------------KMYRDLKRYYQWVGMKMDV 978
Query: 1007 QNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDR 1066
N+V +C+VCQ K E G LQ LPIP W I+MD + GL + KD I V+VDR
Sbjct: 979 ANWVAECDVCQLVKAEHQVLGGMLQSLPIPEWKWDFITMDLVVGLRVSRTKDAIWVIVDR 1038
Query: 1067 FTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDR 1109
TK AHF+A+ A +A+ F+ E+V+LHG P ++ +DR
Sbjct: 1039 LTKSAHFLAIRKTDGAAVLAKKFVSEIVKLHGVPLNMKEAQDR 1081
Score = 92.0 bits (227), Expect = 2e-18
Identities = 52/158 (32%), Positives = 95/158 (59%), Gaps = 6/158 (3%)
Query: 1228 NLEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIA 1287
N+++AQ+R R A+K RR++++EVGD VYLK+ + + + S KLSPRY GP+ I+
Sbjct: 1074 NMKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPNRSI-SETKLSPRYMGPFKIVE 1132
Query: 1288 KINPAAYKLQLPEGSQV-HPVFHISLLKKAVNAGVQS-QPLPAALTEEWELKVEPEAIMD 1345
++ P AY+L+LP+ + H VFH+S+L+K ++ ++ +P L L+ P +++
Sbjct: 1133 RVEPVAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEALAKIPEDLQPNMTLEARPVRVLE 1192
Query: 1346 TR--ENRDGDLEVL-IRWKDLPTFEDSWEDFSKLLDQF 1380
R E R + ++ + W E++WE +++ +F
Sbjct: 1193 RRIKELRQKKIPLIKVLWDCDGVTEETWEPEARMKARF 1230
Score = 37.7 bits (86), Expect = 0.047
Identities = 47/211 (22%), Positives = 80/211 (37%), Gaps = 56/211 (26%)
Query: 14 RRVDIPMFNGN----DAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQ 69
+R+D F+G +A W ++V R F SR ++++ + +E A WW
Sbjct: 145 QRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVDLAVHFLEGDA---HLWWRSV 201
Query: 70 TLER-----AWEPFKQALFRRFQPALLQNPFGPLLSVKQKGSVMEYRENFELLAAPMRNA 124
T R +W F ++ P +P+ G ME + A MR
Sbjct: 202 TARRRQTDMSWADFVAEFKAKYFPQEALDPYA--------GQGMEDDQ------AQMRR- 246
Query: 125 DREVLKGVFLNGLQEEIKAEMKLYPADDLAELMDRALLLEE------------------- 165
FL GL+ +++ ++ A L++ A +EE
Sbjct: 247 --------FLRGLRPDLRVRCRVSQYATKAALVETAAEVEEDFQRQVVGVSPVVQPKKTQ 298
Query: 166 -KNTAMRGGKPKEEDKRGWKDLQNKGGTGNQ 195
+ T +GGKP + KR W D ++ G G +
Sbjct: 299 QQVTPSKGGKPAQGQKRKW-DHPSRAGQGGR 328
>At2g06890 putative retroelement integrase
Length = 1215
Score = 480 bits (1235), Expect = e-135
Identities = 333/1002 (33%), Positives = 515/1002 (51%), Gaps = 82/1002 (8%)
Query: 106 SVMEYRENFE-LLAAPMRNADREVLKGVFLNGLQEEIKAEMKLYPADDLAELMDRALLLE 164
SV EY + E L+ + DRE FL L +I+ ++ + E++ +A+L E
Sbjct: 31 SVEEYYKEMETLMLRADISEDREATLSRFLGDLNRDIQDRLETQYYVQIEEMLHKAILFE 90
Query: 165 E---KNTAMRGG-------KPKEEDKRGWKDLQNKGGTGNQDTEGKQPEKKWNGGQRLTQ 214
+ + ++ R KP + + NK + + + G+
Sbjct: 91 QQVKRKSSSRSSYGSGTIAKPTYQREERTSSYHNKPIVSPRAESKPYAAVQDHKGKAEIS 150
Query: 215 TELQERSRKGLCFKCGDKWGKEHICSMKNYQLILMEVEEDEEEEEI------FEEAED-- 266
T R R C+KC K + C K +IL++ E E EEEI +E E+
Sbjct: 151 TS---RVRDVRCYKCQGKGHYANECPNKRV-MILLDNGEIEPEEEIPDSPSSLKENEELP 206
Query: 267 --GEFVLEGKVLQLSLNSKEGLTSNRSFKVKGKIGNREVLILIDCGATSNFISQDLVVEL 324
GE ++ + L + + E F + + + ++ID G+ +N S+ +V +L
Sbjct: 207 AQGELLVARRTLSVQTKTDEQEQRKNLFHTRCHVHGKVCSLIIDGGSCTNVASETMVKKL 266
Query: 325 EIPVIATS-------EYVVEVGNGAKERNSGVCKNLKLEVQGISI---------MQHFFI 368
+ + S + VV + G K + +C L +E I + + H
Sbjct: 267 GLKWLNDSGKMRVKNQVVVPIVIG-KYEDEILCDVLPMEAGHILLGRPWQSDRKVMHDGF 325
Query: 369 LGLGGTEVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCRVTANWKSIKIT 428
E G L S+ E +Q+ I + Q ++ V++ +P+ + KS +
Sbjct: 326 TNRHSFEFKGGKTILVSMTPHEV-YQDQI--HLKQKKEQVVK-QPNFFAKSGEVKSAYSS 381
Query: 429 EQQEAEGYYLSYEYQKEEEKTEAEVPEGMRKILEEYPEVFQE--PKGLPPRRTTDHAIQL 486
+Q ++ E +P M +L++Y +VF E PKGLPP R +H I
Sbjct: 382 KQPML--LFVFKEALTSLTNFAPVLPSEMTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDF 439
Query: 487 QEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSGIIRHSTSPFSSPAILVKKKDGGWRFCV 546
GAS+PN P Y+ N +E KE L ++KDG WR C
Sbjct: 440 VPGASLPN-----RPAYRTNPVE--TKE-------------------LQRQKDGSWRMCF 473
Query: 547 DYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHE 606
D RA+N T+ PIP +D++LDE+ + +FSK+DLKSGYHQIRM E D KTAF+T
Sbjct: 474 DCRAINNVTVKYCHPIPRLDDMLDELHGSSIFSKIDLKSGYHQIRMNEGDEWKTAFKTKH 533
Query: 607 GHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEELHKDHLRIVL 666
G YE+LV+PFGLT+APSTF LMN VLR ++ FV+V+FDDIL+YS++ H +HL VL
Sbjct: 534 GLYEWLVMPFGLTHAPSTFMRLMNHVLRAFIGIFVIVYFDDILVYSESLREHIEHLDSVL 593
Query: 667 QVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWPIPKEVKGLRGFL 726
VL++ L AN KKC+F +++LG V+S GV D K+K + DWP PK V +R F
Sbjct: 594 NVLRKEELYANLKKCTFCTDNLVFLGFVVSADGVKVDEEKVKAIRDWPSPKTVGEVRSFH 653
Query: 727 GLTGYYRRFVKNYSKLAQPLNQLLKKN-SFQWTEGATQAFVKLKEVMTTVPVLVPPNFDK 785
GL G+YRRF K++S + PL +++KK+ F+W + +AF LK+ +T PVL+ F K
Sbjct: 654 GLAGFYRRFFKDFSTIVAPLTEVMKKDVGFKWEKAQEEAFQSLKDKLTNAPVLILSEFLK 713
Query: 786 PFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLL 845
F +E DASG G+GAVLMQ+ + +A+ S+ L Y++EL A+V A+Q+W+HYL
Sbjct: 714 TFEIECDASGIGIGAVLMQDQKLIAFFSEKLGGATLNYPTYDKELYALVRALQRWQHYLW 773
Query: 846 GSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIKYKPGIENKAADALSRKLQF 905
FVIHTD SL+ L Q+ + + +W+ + + + IKYK G +N ADALS++
Sbjct: 774 PKVFVIHTDHESLKHLKGQQKLNKRHARWVEFIETFAYVIKYKKGKDNVVADALSQRYTL 833
Query: 906 SAISSVQCAEWADLEAEILEDERYRKVLQELATQGNSAVGYQLKRGRLLYKDRIVLPKGS 965
+ +V+ + ++ D +++V + A + ++ Y + L Y++R+ +P S
Sbjct: 834 LSTLNVKLMGFEQIKEVYETDHDFQEVYK--ACEKFASGRYFRQDKFLFYENRLCVPNCS 891
Query: 966 TKILTVLKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQKCEVCQRNKYEALN 1025
+ L V +E H L GH GI +T + ++ F W MK D++ +C C++ K +
Sbjct: 892 LRDLFV-REAHGGGLMGHFGIAKTLEVMTEHFRWPHMKCDVKRICGRCNTCKQAK-SKIQ 949
Query: 1026 PAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRF 1067
P G PLPIP W DISMDF+ GLP+ GKD+I VV F
Sbjct: 950 PNGLYTPLPIPKHPWNDISMDFVMGLPRT-GKDSIFVVYSPF 990
Score = 57.8 bits (138), Expect = 4e-08
Identities = 39/120 (32%), Positives = 63/120 (52%), Gaps = 8/120 (6%)
Query: 1185 TPFKALYGREP----PVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQA 1240
+PF+ +YG P +I S+D +K + I E + N+E+ +QA
Sbjct: 988 SPFQIVYGFNPISPFDLIPLPLSERVSIDGKKKAELVQQ-IHENARRNIEEKTKLYAKQA 1046
Query: 1241 NKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPE 1300
NK RR+ +EVGD+V++ ++ + + K KL PR GP+ II +IN AY+L L +
Sbjct: 1047 NKGRREQIFEVGDMVWIHLRKERFPAQRK---SKLMPRIDGPFKIIKRINDNAYQLDLQD 1103
>At3g31970 hypothetical protein
Length = 1329
Score = 452 bits (1164), Expect = e-127
Identities = 288/835 (34%), Positives = 428/835 (50%), Gaps = 104/835 (12%)
Query: 576 VVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRP 635
V S + + + I + E ++ KTAFRT GH+E++V+PFGLTN P+ F LMN V +
Sbjct: 559 VAVSDIRVVHEFEDIPIAEANVRKTAFRTRYGHFEFVVMPFGLTNIPTAFMRLMNSVFQE 618
Query: 636 YLRKFVLVFFDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVI 695
+L +FV++F DDIL+YSK+ E H+ +Q + + A
Sbjct: 619 FLDEFVIIFIDDILVYSKSPEEHE------VQSEESDGEAAG------------------ 654
Query: 696 SQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKN-S 754
++ GV+ D KI+ + DWP P +R FLGL GYY+RFVK ++ +AQP+ +L K+
Sbjct: 655 AEVGVSVDLEKIEAIRDWPRPTNATEIRSFLGLAGYYKRFVKVFASMAQPMTKLTGKDVP 714
Query: 755 FQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSK 814
F W+ + F+ LKE++T+ PVL P +P+++ TDAS GL VLMQ G+
Sbjct: 715 FVWSPECDEGFMSLKEMLTSTPVLALPEHGEPYMVYTDASRVGLDCVLMQRGK------- 767
Query: 815 TLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKW 874
+ TD +SL+++ Q + Q++W
Sbjct: 768 ------------------------------------VFTDHKSLKYIFTQPELNLRQRRW 791
Query: 875 MSKLMGYDFEIKYKPGIENKAADALSRKLQFSAISS-----------VQCAEWADLEAEI 923
M + YD EI Y G N ADALSRK ++ + V E LEA
Sbjct: 792 MELVADYDLEIAYHSGKANVVADALSRKRVGGSVEALVSEIGALRLCVMAQEPLGLEAVD 851
Query: 924 LEDERYRKVLQELATQGNSAVG------YQLK-RGRLLYKDRIVLPKGSTKILTVLKEFH 976
D R L + +G A YQ G +L R+ +PK +L E H
Sbjct: 852 RADLLTRVRLAQEKDEGLIAASKADGSEYQFAANGTILVHGRVCVPKDEELRREILSEAH 911
Query: 977 DTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIP 1036
+ H + Y+ + + W GMK D+ N+V +C+VCQ K E P G LQ LPI
Sbjct: 912 ASMFSIHPRATKMYRDLKRYYQWVGMKRDVANWVTECDVCQLVKAEHQVPGGLLQSLPIL 971
Query: 1037 SQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRL 1096
W I+MDF+ GLP + KD I V+VDR TK AHF+A+ A +A+ ++ E+V L
Sbjct: 972 EWKWDFITMDFVVGLPVSRTKDAIWVIVDRLTKSAHFLAIRKTDGAVLLAKKYVSEIVEL 1031
Query: 1097 HGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVT 1156
HG P SIVSDRD F S FW GTK++ S+AYHPQTDGQ+E + +E LR
Sbjct: 1032 HGVPVSIVSDRDSKFTSAFWRAFQGEMGTKVQMSTAYHPQTDGQSERTIQTLEDMLRMCV 1091
Query: 1157 GSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGR--EPPVIFK--GNDSLTSVDEVE 1212
+ W LS EF YN +Y ++I+ PF+ALYGR P+ + G S+ D V
Sbjct: 1092 LDRGGHWADHLSLVEFAYNNSYQASIRMAPFEALYGRPCRTPLCWTQVGERSIYGADYVL 1151
Query: 1213 KLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSN 1272
+ T ER + LK N+++AQ+R R A+K RR++++EVGD VYLK+ + + S
Sbjct: 1152 E-TTER---IRVLKLNMKEAQDRQRSYADKRRRELEFEVGDRVYLKMAMLRGPN-RSISE 1206
Query: 1273 QKLSPRYYGPYPIIAKINPAAYKLQLPEGSQV-HPVFHISLLKKAVN------AGVQSQP 1325
KL+PRY GP+ I+ ++ P AY+L+LP+ + H VFH+S+L+K ++ A +
Sbjct: 1207 TKLTPRYMGPFRIVERVGPVAYRLELPDVMRAFHKVFHVSMLRKCLHKDDEVLAKILEDL 1266
Query: 1326 LPAALTEEWELKVEPEAIMDTRENRDGDLEVLIRWKDLPTFEDSWEDFSKLLDQF 1380
P E +++ I + R + ++VL W E++WE +++ F
Sbjct: 1267 QPNMTLEARPVRILERRIKELRRKKIPLIKVL--WNCDGVTEETWEPEARMKASF 1319
Score = 38.1 bits (87), Expect = 0.036
Identities = 77/371 (20%), Positives = 138/371 (36%), Gaps = 55/371 (14%)
Query: 25 DAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLER-----AWEPFK 79
+A W ++VE F SR ++++ + +E A WW T R +W F
Sbjct: 166 EADSWKSRVEHNFGSSRCPAEYRVDLAVHFLEGDA---HLWWRSVTARRRQAHMSWADFV 222
Query: 80 QALFRRFQP-ALLQNPFGPLLSVKQ-KGSVMEYRENFE--LLAAPMRNADREVLKGVFLN 135
++ P L L + Q + SV EY F L+ A D + FL
Sbjct: 223 AEFNAKYFPQEALDRMEARFLELTQGERSVREYEREFNRLLVYAGRGMQDDQAQMRRFLR 282
Query: 136 GLQEEIKAEMKLYPADDLAELMDRALLLEEK-NTAMRGGKPKEEDKRGWKDL-QNKGGTG 193
GL+ +++ ++ A L++ A +EE + G P + K+ + + +KGG
Sbjct: 283 GLRPDLRVRCRVLQYATKAALVETAAEVEEDLQRQVVGVSPAVKPKKTQQQVAPSKGGKP 342
Query: 194 NQDTEGKQPEKKWNGGQRLTQTELQERSRKGLCFKCGDKWGKEHICSMKNYQLILMEVEE 253
Q + K+ G R +LQ + ++ V+
Sbjct: 343 AQGQKRKE-----RGHLRPNCPKLQRMA----------------------VAVVQPAVQH 375
Query: 254 DEEEEEIFEEAEDGEFVLEGKVLQLSLNSKEGLTSNRSFKVKGKIGNREVLILIDCGATS 313
+ ++ ++ +G + + G TS R+ E +L D GA+
Sbjct: 376 GAQVQQRVQQLAHIAAAPQGYTTR-----EIGSTSKRAI--------TEAHVLFDSGASH 422
Query: 314 NFISQDLVVELEIPVIATSEY-VVEVGNGAKERNSGVCKNLKLEVQGISIMQHFFILGLG 372
FI+ + I + V+V G G K + +++ G S+ I +
Sbjct: 423 CFITPESASRGNIRGDPGEQLGAVKVAGGQFLAVLGRAKGVDIQIAGESMPVDLIISPVE 482
Query: 373 GTEVVLGMDWL 383
+V+LGMDWL
Sbjct: 483 LYDVILGMDWL 493
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 384 bits (987), Expect = e-106
Identities = 208/517 (40%), Positives = 319/517 (61%), Gaps = 15/517 (2%)
Query: 335 VVEVGNGAKERNSGVCKNLKLEVQGISIMQHFFILGLGGTEVVLGMDWLASL-GNIEANF 393
+V G + + + + + V G+++ I+ L +V+LGMDWL G+I+ +
Sbjct: 1 MVRAAGGQAMYPTRLVRGISVVVNGVNMPADLIIVPLKKHDVILGMDWLGKYKGHIDCHR 60
Query: 394 QELIIQWVSQGQKMVLQGEPSVCRVTANWKSIKITEQQEAEGY----YLSYEYQKEEEKT 449
+ IQ+ + QG R T+ I + + G YL+ KE +
Sbjct: 61 ER--IQFERDEGMLKFQG----IRTTSGSLVISAIQAERMLGKGCEAYLATITTKEVGAS 114
Query: 450 EAEVPEGMRKILEEYPEVFQEPKGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIE 509
AE+ + + I+ E+ +VF G+PP R+ I+L+ G + + PYR + E++
Sbjct: 115 -AELKDIL--IVNEFSDVFAAVSGVPPDRSDPFTIELEPGTTPISKAPYRMAPAEMAELK 171
Query: 510 KLVKEMLNSGIIRHSTSPFSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELL 569
K ++E+L G IR S+SP+ +P + VKKKDG +R C+DYR LNK T+ +K+P+P IDEL+
Sbjct: 172 KQLEELLAKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDELM 231
Query: 570 DEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALM 629
D++G A FSK+DL SGYHQI ++ D+ KTAFRT GH+E++V+PFGLTNAP+ F +M
Sbjct: 232 DQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMKMM 291
Query: 630 NQVLRPYLRKFVLVFFDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEII 689
N V R +L +FV++F DDIL++SK+ E H++HLR VL+ L+E+ L A K SF Q +
Sbjct: 292 NGVFRDFLDEFVIIFIDDILVHSKSWEAHQEHLRAVLERLREHELFAKLSKFSFWQRSVG 351
Query: 690 YLGHVISQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQL 749
+LGHVIS GV+ DP KI+ + +WP P+ +R FLGL GYYRRFV +++ +AQPL +L
Sbjct: 352 FLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRL 411
Query: 750 L-KKNSFQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRP 808
K +F W++ ++F++LK ++ PVLV P +P+ + TDAS GLG VLMQ+G
Sbjct: 412 TGKDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSV 471
Query: 809 VAYMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLL 845
+AY S+ L + ++ E+ AVV A++ WR YL+
Sbjct: 472 IAYASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYLI 508
Score = 274 bits (700), Expect = 3e-73
Identities = 164/417 (39%), Positives = 234/417 (55%), Gaps = 20/417 (4%)
Query: 941 NSAVGYQLKR-GRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYW 999
N+ YQ G ++ R+ +P +L+E H + H G + Y+ + ++W
Sbjct: 524 NNKTEYQTSNNGTIVVNGRVCVPNDRALKEEILREAHQSKFSIHPGSNKMYRDLKRYYHW 583
Query: 1000 EGMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGK-- 1057
GM+ D+ +V KC CQ K E P+G LQ LPI W I+MDF+ LP +
Sbjct: 584 VGMRKDVARWVAKCPTCQLVKAEHQVPSGLLQNLPISEWKWDHITMDFVTRLPTGIKSKH 643
Query: 1058 DTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWS 1117
+ + VVVDR TK AHF+A+S A+ IAE +I E++RLHG P SIVSDRD F S FW+
Sbjct: 644 NAVWVVVDRLTKSAHFMAISDKDGAEIIAEKYIDEIMRLHGIPVSIVSDRDTRFTSKFWN 703
Query: 1118 EMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTN 1177
K GT++ S+AYHPQTDGQ+E + +E LR W K+L EF YN +
Sbjct: 704 AFQKALGTRVNLSTAYHPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLIEFAYNNS 763
Query: 1178 YHSAIKTTPFKALYGRE-------PPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLE 1230
+ ++I +P++ALYGR PV + T VDE T ER ++ LK L+
Sbjct: 764 FQASIGMSPYEALYGRACRTPLCWTPVGERRLFGPTIVDE----TTER---MKFLKIKLK 816
Query: 1231 KAQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKIN 1290
+AQ+R + ANK R++++++VGDLVYLK YK + S +KLSPRY GPY +I ++
Sbjct: 817 EAQDRQKSYANKRRKELEFQVGDLVYLKAMTYK-GAGRFTSRKKLSPRYVGPYKVIERVG 875
Query: 1291 PAAYKLQLPEGSQV-HPVFHISLLKKAVNAGVQS-QPLPAALTEEWELKVEPEAIMD 1345
AYKL LP V H VFH+S L+K ++ +S + +P L E ++ P IMD
Sbjct: 876 AVAYKLDLPPKLNVFHNVFHVSQLRKYLSDQEESVEDIPPGLKENMTVEAWPVRIMD 932
>At4g07850 putative polyprotein
Length = 1138
Score = 382 bits (981), Expect = e-106
Identities = 240/663 (36%), Positives = 335/663 (50%), Gaps = 110/663 (16%)
Query: 694 VISQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKN 753
V S GV D K+K + +WP PK V +R F GL G+YRRFV+++S LA PL +++KKN
Sbjct: 531 VASTDGVKVDEEKVKAIREWPSPKSVGKVRSFHGLAGFYRRFVRDFSTLAAPLTEVIKKN 590
Query: 754 -SFQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYM 812
F+W + AF LKE +T PVL P+F K F +E DA G G+GAVLMQ+ +P+AY
Sbjct: 591 VGFKWEQAPEDAFQALKEKLTHAPVLSLPDFLKTFEIECDAPGVGIGAVLMQDKKPIAYF 650
Query: 813 SKTLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQ 872
S+ L Y++EL A+V A+Q W+HYL +FVIHTD SL+ L Q+ + +
Sbjct: 651 SEKLGGATLNYPTYDKELYALVRALQTWQHYLWPKEFVIHTDHESLKHLKGQQKLNKRHA 710
Query: 873 KWMSKLMGYDFEIKYKPGIENKAADALSRKLQFSAISSVQCAEWADLEAEILEDERYRKV 932
+W+ + + + IKYK G +N ADALSR+ +
Sbjct: 711 RWVEFIETFPYVIKYKKGKDNVVADALSRRHE---------------------------- 742
Query: 933 LQELATQGNSAVGYQLKRGRLLYKDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKR 992
G L Y +R+ +P S + L + +E H L GH G+ +T K
Sbjct: 743 ------------------GFLFYDNRLCIPNSSLRELFI-REAHGGGLMGHFGVSKTLKV 783
Query: 993 ISALFYWEGMKLDIQNYVQKCEVCQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLP 1052
+ F+W MK D++ ++C C++ K ++ P G PLPIP W DISMDF+ GLP
Sbjct: 784 MQDHFHWPHMKRDVERMCERCTTCKQAKAKS-QPHGLCTPLPIPLHPWNDISMDFVVGLP 842
Query: 1053 KAM-GKDTILVVVDRFTKYAHFIALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVF 1111
+ GKD+I VVVDRF+K AHFI +A IA +F +EVVRLHG P +IVSDRD F
Sbjct: 843 RTRTGKDSIFVVVDRFSKMAHFIPCHKTDDAMHIANLFFREVVRLHGMPKTIVSDRDTKF 902
Query: 1112 LSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAE 1171
LS FW ++ GTKL FS+ HPQTDGQTEVVNR + T LR + K W L E
Sbjct: 903 LSYFWKTLWSKLGTKLLFSTTCHPQTDGQTEVVNRTLSTLLRALIKKNLKTWEDCLPHVE 962
Query: 1172 FWYNTNYHSAIKTTPFKALYGREPPVIFKGNDSLTSVDEVEKLTAERNLILEELKSNLEK 1231
F YN + HSA K +PF+ +YG P +T +D + L + +L K
Sbjct: 963 FAYNHSVHSATKFSPFQIVYGFNP---------ITPLDLMP---------LPLISIHLRK 1004
Query: 1232 AQNRMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINP 1291
+ P K KS KL PR G + ++ +IN
Sbjct: 1005 ER--------------------------FPNKRKS-------KLMPRMDGSFKVLKRIND 1031
Query: 1292 AAYKLQLPEGSQVHPVFHISLLKKAVNAGVQSQPLPAALTEEWELKVEPEAIMDTRENRD 1351
AY L L + L G + + +L + ++EPE + ++ D
Sbjct: 1032 NAYSLDLQD--------ETDLRSNPFQVG-EDDVIMGSLDHGAKEQLEPEEFGERKQLED 1082
Query: 1352 GDL 1354
G+L
Sbjct: 1083 GEL 1085
Score = 74.7 bits (182), Expect = 3e-13
Identities = 39/85 (45%), Positives = 51/85 (59%), Gaps = 2/85 (2%)
Query: 462 EEYPEVFQE--PKGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNSG 519
++Y +VF E P+GLPP R +H I GAS+PN YR + E+EK V E++ G
Sbjct: 446 KDYQDVFPEENPEGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVETKELEKQVNELMERG 505
Query: 520 IIRHSTSPFSSPAILVKKKDGGWRF 544
I S SP + P +LV KKDG WRF
Sbjct: 506 HICESMSPCAVPVLLVPKKDGSWRF 530
Score = 70.9 bits (172), Expect = 5e-12
Identities = 74/299 (24%), Positives = 122/299 (40%), Gaps = 26/299 (8%)
Query: 44 EAEKIEMVMIAMEDRALGWFQWWEEQTLER--------AWEPFKQALFRRFQPALLQNPF 95
EA K+++ D AL W W + T +R W K + RRF P+
Sbjct: 92 EANKVKVAATEFYDYALSW--WDQVVTTKRRLGDDSIETWNQLKNIMKRRFVPSHYHREL 149
Query: 96 GPLLS--VKQKGSVMEYRENFELLAAPMRNADR----EVLKGVFLNGLQEEIKAEMKLYP 149
L V+ +V EY + E L M AD E F+ GL +I ++
Sbjct: 150 HQRLRNLVQGNRTVEEYFKEMETL---MLRADVQEECEATMSRFMGGLNRDILDRFEVIH 206
Query: 150 ADDLAELMDRALLLEEKNTAMRGGKPKEEDKRGWKDLQNKGGTGNQDTEGKQPEKKWNGG 209
++L EL +A++ E K R KP + + K G + +P+ +
Sbjct: 207 YENLEELFHKAVMFE-KQIKRRSAKPSYNSSKPSYQREEKSGFQKEYKPFVKPKVEEISS 265
Query: 210 QRLTQTELQERSRKGLCFKCGDKWGKEHICSMKNYQLIL----MEVEEDEEEEEIFEEAE 265
+ + + R K CFKC CS K +I +E E+++ EE EEA
Sbjct: 266 KGKEKEVTRTRDLK--CFKCHGLGHYASECSNKRIMIIRDSGEVESEDEKPEESDVEEAP 323
Query: 266 DGEFVLEGKVLQLSLNSKEGLTSNRSFKVKGKIGNREVLILIDCGATSNFISQDLVVEL 324
GE ++ +VL + ++E F + I + ++ID G+ +N S+ +V +L
Sbjct: 324 KGELLVTMRVLSVLNKAEEQAQRENLFHTRCLIKGKVCSLIIDGGSCTNVASETMVQKL 382
>At4g16910 retrotransposon like protein
Length = 687
Score = 378 bits (970), Expect = e-104
Identities = 242/677 (35%), Positives = 361/677 (52%), Gaps = 51/677 (7%)
Query: 742 LAQPLNQLLKKNS-FQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGA 800
+AQP+ QL K++ F W+E ++F++LK ++T PVLV P +P+ + TDAS GLG
Sbjct: 1 MAQPMTQLTGKDTAFNWSEECERSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGC 60
Query: 801 VLMQEGRPVAYMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRF 860
VL+Q+G +AY S+ L + + E+ AVV A++ WR YL G+K I TD +SL++
Sbjct: 61 VLIQKGSVIAYASRQLRKHEKNYPTNDLEMAAVVFALKIWRSYLYGAKVQIFTDHKSLKY 120
Query: 861 LADQRIMGEEQQKWMSKLMGYDFEIKYKPGIENKAADALSR------------------- 901
+ Q + Q++W+ + YD I Y PG N+ DALSR
Sbjct: 121 IFTQPELNLRQRRWIKLVADYDLNIAYHPGKANQVVDALSRLRTVVEAERNQVNLVNMMG 180
Query: 902 KLQFSAISS------VQCAEWADLEAEILEDERYRKVLQELATQGNSAVGYQLKRGRLLY 955
L +A+S ++ A ADL + I + + ++ A Q N G ++
Sbjct: 181 TLHLNALSKEVEPLGLRAANQADLLSRIRSAQERDEEIKGWA-QNNKTEYQSSNNGTIVV 239
Query: 956 KDRIVLPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQKCEV 1015
R+ P +LKE H + H G + Y+ + ++W GMK D+ +V K
Sbjct: 240 NGRVCGPNDKALKEEILKEAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWVAK--- 296
Query: 1016 CQRNKYEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGK--DTILVVVDRFTKYAHF 1073
E P+G LQ LPIP W I MDF+ GLP + + + VVVDR TK AHF
Sbjct: 297 -----EEHQVPSGMLQNLPIPEWKWDHIMMDFVTGLPTGIKSKHNAVWVVVDRLTKSAHF 351
Query: 1074 IALSHPYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAY 1133
+A+S A+ IAE +I E+VRLHG P SIVSDRD F S FW K+ GT++ S+AY
Sbjct: 352 MAISDKDAAEIIAEKYIDEIVRLHGIPVSIVSDRDTRFTSKFWKPFQKVLGTRVNLSTAY 411
Query: 1134 HPQTDGQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGR 1193
HPQTDGQ+E + +E LR W K+L EF YN ++ ++I +P++ALYGR
Sbjct: 412 HPQTDGQSERTIQTLEDMLRACVLDWGGNWEKYLRLVEFAYNNSFQASIGMSPYEALYGR 471
Query: 1194 --EPPVIFK--GNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQY 1249
P+ + G L V++ T + ++ LK L++AQ+R + ANK R+++++
Sbjct: 472 AGRTPLCWTPVGERRLFGPAVVDETTKK----MKFLKIKLKEAQDRQKSYANKRRKELEF 527
Query: 1250 EVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQL-PEGSQVHPVF 1308
+VGDLVYLK YK + S +KL PRY GPY +I ++ AYKL L P+ H VF
Sbjct: 528 QVGDLVYLKAMTYK-GAGRFTSRKKLRPRYVGPYKVIERVGAVAYKLDLPPKLDAFHNVF 586
Query: 1309 HISLLKKAVNAGVQS-QPLPAALTEEWELKVEPEAIMD--TRENRDGDLEVL-IRWKDLP 1364
H+S L+K ++ +S + +P L E ++ P IMD + R +++L I W
Sbjct: 587 HVSQLRKCLSEQEESMEDVPPGLKENMTVEAWPVRIMDQMKKGTRGKSMDLLKILWNCGG 646
Query: 1365 TFEDSWEDFSKLLDQFP 1381
E +WE +K+ FP
Sbjct: 647 REEYTWETETKMKANFP 663
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 311 bits (798), Expect = 1e-84
Identities = 261/871 (29%), Positives = 404/871 (45%), Gaps = 117/871 (13%)
Query: 472 KGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNEI-EKLVKEMLNSGIIRH-STSPFS 529
KG+ P T H I L E S +I P R E+ +K + ++L++G+I S S +
Sbjct: 880 KGISPTLCT-HRIHL-ENESYSSIEPQRRLNPNLKEVVKKEILKLLDAGVIYPISDSTWV 937
Query: 530 SPAILVKKKDG------------------GWRFCVDYRALNKATIPDKFPIPIIDELLDE 571
SP V KK G G R C++YR LN A+ + FP+P ID +L+
Sbjct: 938 SPVHCVPKKGGMTVVKNSKDELIPTRTITGHRMCIEYRKLNVASRKEHFPLPFIDHMLER 997
Query: 572 IGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQ 631
+ + LD SG+ QI + D KT F G + Y +PFGL NAP+TFQ M
Sbjct: 998 LANHPYYCFLDSYSGFFQIPIHPNDQGKTTFTCPYGTFAYKRMPFGLCNAPATFQRCMTS 1057
Query: 632 VLRPYLRKFVLVFFDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYL 691
+ + + V VF DD +Y + +L VL+ +E NLV N +KC F E I L
Sbjct: 1058 IFSDLIEEMVEVFMDDFSVYGSSFSSCLLNLCRVLKRCEETNLVLNWEKCHFMVREGIVL 1117
Query: 692 GHVISQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLL- 750
G IS+ G+ D +KI M+ PK VK +R FLG G+YR F+K++SKLA+PL +LL
Sbjct: 1118 GRKISEEGIEVDKAKIDVMMQLQPPKTVKDIRSFLGHAGFYRIFIKDFSKLARPLTRLLC 1177
Query: 751 KKNSFQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVA 810
K+ F + + AF +KE + T P++ PN+D PF +
Sbjct: 1178 KETEFAFDDECLTAFKLIKEALITAPIVQAPNWDFPFEI--------------------- 1216
Query: 811 YMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEE 870
T+ D + E+EL+AVV A +K+R YL+GSK I+TD +LR + ++
Sbjct: 1217 ---ITMDDAQVRYATTEKELLAVVFAFEKFRSYLVGSKVTIYTDHAALRHIYAKKDTKPR 1273
Query: 871 QQKWMSKLMGYDFEIKYKPGIENKAADALSR-KLQFSAISSVQCAEWADLEAEILEDER- 928
+W+ L +D EI K GIEN AD LSR +++ + E + + L +++
Sbjct: 1274 LLRWILLLQEFDMEIVDKKGIENGVADHLSRMRIEDEVLIDDSMPEEQLMAIQQLNEKKL 1333
Query: 929 --YRKVLQELAT--QGNSAVGYQLKRG--------------RLLYKDRIVLP-KGSTKIL 969
Y + L + + + Y+ K+ L KD+I +I
Sbjct: 1334 PWYADHVNYLVSGEEPPNLSSYEKKKFFKDINHFYWDEPYLYTLCKDKIYRTCVSEDEIE 1393
Query: 970 TVLKEFHDTALGGHAGIFRTYKRI-SALFYWEGMKLDIQNYVQKCEVCQRNKYEALNPAG 1028
+L H A GGH F+T +I A F+W M D Q ++ KC+ +E + G
Sbjct: 1394 GILLHCHGFAYGGHFATFKTMSKILQAGFWWPSMFKDAQEFISKCD-----SFENFDVWG 1448
Query: 1029 FLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIALSHPYNAKEIAEV 1088
+DF+G P + G ILV +D +K+ IA SH +A+ + ++
Sbjct: 1449 ----------------IDFMGPFPSSYGNKYILVAIDYVSKWVEAIA-SHTNDARVVLKL 1491
Query: 1089 FIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTEVVNRCV 1148
F + G P ++SD + F++ + + K G K K T GQ E+ NR +
Sbjct: 1492 FKTIIFPRFGVPRIVISDGGKHFINKGFENLLKKHGVKHK--------TSGQVEISNREI 1543
Query: 1149 ETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGRE----PPVIFKGNDS 1204
+ L GS K W L+ + Y T + + I TTPF LYG+ + +K +
Sbjct: 1544 KAILEKTVGSTRKDWSAKLNDTLWAYRTAFKTPIGTTPFNLLYGKSCHLPVELEYKAMWA 1603
Query: 1205 LTSVDEVEKLTAERNLI-LEEL-KSNLEKAQ-NRMRQQANKHRRDVQ-----YEVGDLVY 1256
+ ++ K E+ LI L +L K LE + +++ ++ K D + ++VGD V
Sbjct: 1604 VKLLNFDIKTAEEKRLIQLNDLNKIRLEAYESSKIYKERTKSFHDKKIVSRDFKVGDQVL 1663
Query: 1257 LKIQPYKLKSLAKRSNQKLSPRYYGPYPIIA 1287
L S + KL R+ GP+ + A
Sbjct: 1664 L------FNSRLRLFPGKLKSRWSGPFSVTA 1688
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 292 bits (748), Expect = 8e-79
Identities = 145/318 (45%), Positives = 206/318 (64%), Gaps = 5/318 (1%)
Query: 1079 PYNAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTD 1138
P ++ +A F+ VV+LHG P SIVSD D +F+S FW E +KL+ TKL S+AYHPQTD
Sbjct: 628 PLLSRSVAAKFVSHVVKLHGIPRSIVSDCDPIFMSLFWQEFWKLSRTKLWMSTAYHPQTD 687
Query: 1139 GQTEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGREPPVI 1198
GQTEVVNRC+E +LRC PKQW ++ WAE+WYNT +H++ TPF+ALYGR P I
Sbjct: 688 GQTEVVNRCIEQFLRCFVHYHPKQWSSFIPWAEYWYNTTFHASTGMTPFQALYGRPPSPI 747
Query: 1199 FKGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEVGDLVYLK 1258
E+ + A R+ +L ELK +L A N M+QQA+ RDV ++VGD V L+
Sbjct: 748 PAYELGSVVCGELNEQMAARDELLAELKQHLVTANNCMKQQADSRLRDVSFQVGDWVLLR 807
Query: 1259 IQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQVHPVFHISLLKKAVN 1318
IQPY+ K+L +RS+QKLS R+YGP+ + +K AY+L LPEG+++HPVFH+SLLK V
Sbjct: 808 IQPYRQKTLFRRSSQKLSHRFYGPFQVASKHGEVAYRLTLPEGTRIHPVFHVSLLKPWVG 867
Query: 1319 AGVQSQPLPAALTEEWELKVEPEAIMDTR---ENRDGDLEVLIRWKDLPTFEDSWEDFSK 1375
G L ELK++P A+++ R +++ ++L++W+ L + +WE++ +
Sbjct: 868 DGEPDMGQLPPLRNNGELKLQPTAVLEVRWRSQDKKRVADLLVQWEGLHIEDATWEEYDQ 927
Query: 1376 LLDQFPNH--QLEDKLNL 1391
L FP LEDK+ L
Sbjct: 928 LAASFPEFVLNLEDKVRL 945
Score = 255 bits (651), Expect = 1e-67
Identities = 181/632 (28%), Positives = 299/632 (46%), Gaps = 99/632 (15%)
Query: 15 RVDIPMFNGNDAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQTLERA 74
++D P F+G D W++K +++F + + ++ A GW+Q TL R+
Sbjct: 105 KLDFPRFHGGDLTKWLSKAKQYFEYHEGPVEQAVRFTTFHLDGVANGWWQ----ATLRRS 160
Query: 75 WEPFKQALFRRFQPALLQNPFGPLLSVKQKGSVMEYRENFELLAAPMRNADREVLKGVFL 134
++ Q GS+ EY+ FE L + +EVL G F+
Sbjct: 161 ------------------------ATIGQTGSLSEYQHEFEWLQNKVYGWTQEVLVGAFM 196
Query: 135 NGLQEEIKAEMKLYPADDLAELMDRALLLEEKNTAMRGGKPKEEDKRGWKDLQNKGGTGN 194
NGL I ++++ L E+++ + G K K +D+R + + +
Sbjct: 197 NGLYYSISNGIRMFQPRSLREVINLRV----------GWKGKSKDRRRKRATHHLHAVLS 246
Query: 195 QDTEGKQPEKKWNGGQRLTQTELQERSRKGLCFKCGDKWGKEHICSMKNYQLILMEVEED 254
W +R + ++ RK ++ W Y +M+ E D
Sbjct: 247 LTV--------WWCAKRQSNRRQRDSVRKN--YERKGVWA---------YVSAVMKGEND 287
Query: 255 EEEEEIFEEAEDGEFVLEGKVLQLSLNSKEGLTSNRSFKVKGKIGNREVLILIDCGATSN 314
+E E EE+ + E GK +++L + EG+ + + +V+ I ++ LI+ G+T N
Sbjct: 288 DEAVE--EESTEAE----GKP-EITLYALEGVDTTSTIRVRATIHRNRLIALINSGSTHN 340
Query: 315 FISQDLVVELEIPVIATSEYVVEVGNGAKERNSGVCKNLKLEVQGISIMQHFFILGLGGT 374
FI + V + + T + V V NG + + + + G+ + L L G
Sbjct: 341 FIGEKAVRGMNLKATTTKPFTVRVVNGMPLVCRSRYEAIPVVMGGVVFPVTLYALPLMGL 400
Query: 375 EVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGEPSVCRVTANWKSIKITEQQEAE 434
++ +G+ WL++LG N++E +Q+ G ++ L G K+I ++A
Sbjct: 401 DLAMGVQWLSTLGPTLCNWKEQTLQFHWAGDEVRLMGIKPTGLRGVEHKTIT----KKAR 456
Query: 435 GYYLSYEYQKEEEKTEAEVPEG-MRKILEEYPEVFQEPKGLPPRRTTDHAIQLQEGASIP 493
+ + ++ P+G ++K++EE+ +F+ P LP R H I L++G
Sbjct: 457 MGHTIFAITMAHNSSDPNTPDGEIKKLIEEFAGLFETPTQLPSERPIVHRIALKKGTDPV 516
Query: 494 NIRPYRYPFYQKNEIEKLVKEMLNSGIIRHSTSPFSSPAILVKKKDGGWRFCVDYRALNK 553
N+RPYRY FYQK+E+ + +GIIR S+SPF SP +L
Sbjct: 517 NVRPYRYAFYQKDEMSR-------AGIIRPSSSPFLSPVLL------------------- 550
Query: 554 ATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLV 613
++FPIP +D++LDE+ AV F+KLDL +GY Q+RM DIPK AF+TH GHYEYLV
Sbjct: 551 ----ERFPIPTVDDMLDELNGAVYFTKLDLTAGYQQVRMHSPDIPKIAFQTHNGHYEYLV 606
Query: 614 LPFGLTNAPSTFQALMNQVLRPYLRKFVLVFF 645
+PFGL NAPSTFQALMN++ P L + V F
Sbjct: 607 MPFGLCNAPSTFQALMNEIFWPLLSRSVAAKF 638
>At4g08100 putative polyprotein
Length = 1054
Score = 288 bits (737), Expect = 2e-77
Identities = 149/342 (43%), Positives = 207/342 (59%), Gaps = 29/342 (8%)
Query: 547 DYRALNKATIPDKFPIPIIDELLDEIGAAVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHE 606
D RA+N T+ + PIP +D+ LD++ + +FSK+DLKSGYHQ RMKE D KTA +T +
Sbjct: 527 DCRAINNITVKYRHPIPRLDDTLDKLHGSSIFSKIDLKSGYHQTRMKEGDEWKTAIKTKQ 586
Query: 607 GHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIYSKNEELHKDHLRIVL 666
YE+LV+PFGLTNAP+TF LMN VLR ++ FV+V+FDDIL+YSKN E H HL++VL
Sbjct: 587 RLYEWLVMPFGLTNAPNTFMRLMNHVLRKHIGVFVIVYFDDILVYSKNLEYHVMHLKLVL 646
Query: 667 QVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDMLDWPIPKEVKGLRGFL 726
+L++ L AN KKC+F +++LG V+S G+ D K+K + +WP PK V
Sbjct: 647 DLLRKEKLYANLKKCTFCTDNLVFLGFVVSADGIKVDEEKVKAIREWPNPKNVSE----- 701
Query: 727 GLTGYYRRFVKNYSKLAQPLNQLLKKNSFQWTEGATQAFVKLKEVMTTVPVLVPPNFDKP 786
K F+W + AF LKE +T V + PNF K
Sbjct: 702 ------------------------KDIGFKWEDAQENAFQALKEKLTNSSVPILPNFMKS 737
Query: 787 FILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLLG 846
F +E DASG G+GAVLMQ+ +P+AY S+ L Y++EL A+V A+Q W+HYL
Sbjct: 738 FEIECDASGLGIGAVLMQDHKPIAYFSEKLGGATLNYPTYDKELYALVRALQTWQHYLWP 797
Query: 847 SKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIKYK 888
KFVIHTD SL+ L Q+ + + +W+ + + + IKYK
Sbjct: 798 KKFVIHTDHESLKHLKGQQKLNKRHARWVEFIETFPYVIKYK 839
Score = 58.2 bits (139), Expect = 3e-08
Identities = 40/130 (30%), Positives = 65/130 (49%), Gaps = 13/130 (10%)
Query: 1179 HSAIKTTPFKALYGREPPVIFKGNDSLTSV----DEVEKLTAERNLILEELKSNLEKAQN 1234
HSA K +PF+ +YG FK L + E L ++ L + NLE
Sbjct: 851 HSATKFSPFEIVYG------FKPTSPLDLIPLPLSERSSLDGKKKDDLVQQVLNLEARTK 904
Query: 1235 RMRQQANKHRRDVQYEVGDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAY 1294
+ ++ ANK R++V + GD V++ ++ K + + KL PR GP+ ++ +IN AY
Sbjct: 905 QYKKYANKGRKEVIFNEGDQVWVHLRK---KRFPEVRSSKLMPRIDGPFKVLKRINNNAY 961
Query: 1295 KLQLPEGSQV 1304
KL L + S +
Sbjct: 962 KLDLQDTSDL 971
Score = 56.6 bits (135), Expect = 1e-07
Identities = 71/294 (24%), Positives = 117/294 (39%), Gaps = 32/294 (10%)
Query: 13 GRRVDIPMFNGNDA----YGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEE 68
G ++ IP F+G D W K+E F ++ K+++ + AL WW++
Sbjct: 91 GLKLKIPAFHGTDNPDTYLEWEQKIELVFLCQECLQSNKVKIAATKFYNYAL---SWWDQ 147
Query: 69 QTLER---------AWEPFKQALFRRFQPALLQNPFGPLLSVKQKGS--VMEYRENFE-- 115
R W K + +RF P+ L +GS V EY E
Sbjct: 148 LVTSRRRTRDYPIKTWNQLKFVMRKRFVPSYYHRELHQRLRNLVQGSKTVEEYFLEMETL 207
Query: 116 LLAAPMRNADREVLKGVFLNGLQEEIKAEMKLYPADDLAELMDRALLLEE--KNTAMRGG 173
+L A ++ D E + F+ GL EI+ ++ +L E++ +A++ E+ K R
Sbjct: 208 MLRADLQE-DGEAVMSRFMGGLNREIQDRLETQHYVELEEMLHKAVMFEQQIKRKNARSS 266
Query: 174 KPKEEDKRGWKDLQNKGGTGNQ-DTEGKQPEKKWNGGQRLTQTELQERSRKGLCFKCGDK 232
K G Q + G Q D++ K + + E+ R+R CFK
Sbjct: 267 HTKTNYSSGKPSYQKEEKFGYQNDSKPFVKPKPVDLDPKGKGKEVITRARDIRCFK---S 323
Query: 233 WGKEHICSMKNYQLILM-----EVEEDEEEEEIFEEAEDGEFVLEGKVLQLSLN 281
G H S + + I++ EVE +E E ED E G+ + SL+
Sbjct: 324 QGLRHYASKCSNKRIMVHRDNGEVESKDESAEEDSMEEDVESPARGEFARRSLS 377
>At4g03650 putative reverse transcriptase
Length = 839
Score = 285 bits (729), Expect = 1e-76
Identities = 135/297 (45%), Positives = 195/297 (65%), Gaps = 1/297 (0%)
Query: 592 MKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRPYLRKFVLVFFDDILIY 651
+ E D+ KTAFRT GH+E++V+PFGLTNAP+ F LMN V + +L +FV++F DDIL+Y
Sbjct: 516 LAEADVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVY 575
Query: 652 SKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHVISQAGVAADPSKIKDML 711
SK+ E H+ HLR V++ L+E L A KCSF Q E+ +LGH++S GV+ DP KI+ +
Sbjct: 576 SKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIR 635
Query: 712 DWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKN-SFQWTEGATQAFVKLKE 770
DWP P +R FLGL GYYRRF+K ++ +AQP+ +L K+ F W+ + FV LKE
Sbjct: 636 DWPRPTNATEIRSFLGLAGYYRRFIKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKE 695
Query: 771 VMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEGRPVAYMSKTLSDRAQAKSVYEREL 830
++T+ PVL P +P+++ TDASG GLG LMQ G+ +AY S+ L ++ E+
Sbjct: 696 MLTSTPVLALPEHGEPYMVYTDASGVGLGCALMQRGKVIAYASRQLRKHEGNYPTHDLEM 755
Query: 831 MAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGEEQQKWMSKLMGYDFEIKY 887
AV+ A++ WR YL G K + TD +SL+++ Q + Q++WM + YD EI Y
Sbjct: 756 AAVIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLEIAY 812
Score = 38.5 bits (88), Expect = 0.027
Identities = 41/185 (22%), Positives = 80/185 (43%), Gaps = 24/185 (12%)
Query: 459 KILEEYPEVFQEPKGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNEIEKLVKEMLNS 518
++++E+ VFQ +GLPP R+ I+L+ + + PYR + E++K ++++L
Sbjct: 459 RVVQEFQYVFQSLQGLPPSRSDPFTIELEPKTAPLSKAPYRMAPAKMAELKKQLEDLLAE 518
Query: 519 GIIRHSTSPFSSPAILVKKKDGGWRFCV-DYRALNKATIPDKFPIPIIDELLDE-----I 572
+R + + + G + F V + N + + E LDE I
Sbjct: 519 ADVRKTA---------FRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEFLDEFVIIFI 569
Query: 573 GAAVVFSKLDLKSGYHQIRMKEE--------DIPKTAFRTHE-GHYEYLVLPFGLTNAPS 623
+V+SK + H R+ E+ + K +F E G ++V G++ P
Sbjct: 570 DDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPE 629
Query: 624 TFQAL 628
+A+
Sbjct: 630 KIEAI 634
Score = 34.7 bits (78), Expect = 0.39
Identities = 40/165 (24%), Positives = 66/165 (39%), Gaps = 16/165 (9%)
Query: 14 RRVDIPMFNGN----DAYGWVTKVERFFRLSRVEEAEKIEMVMIAMEDRALGWFQWWEEQ 69
+R+ F+G +A W ++VER F SR ++++ + +E A WW
Sbjct: 149 QRIGTEYFSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLEGDA---HLWWRSV 205
Query: 70 TLER-----AWEPFKQALFRRFQP-ALLQNPFGPLLSVKQ-KGSVMEYRENFE--LLAAP 120
T R +W F ++ P L L + Q SV EY F L+ A
Sbjct: 206 TARRRQADMSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRLLVYAG 265
Query: 121 MRNADREVLKGVFLNGLQEEIKAEMKLYPADDLAELMDRALLLEE 165
D + FL GL+ ++ ++ A L++ A +EE
Sbjct: 266 RGMEDDQAQMRRFLRGLRPNLRVRCRVSQYATKAALVETAAEVEE 310
Score = 34.7 bits (78), Expect = 0.39
Identities = 13/19 (68%), Positives = 17/19 (89%)
Query: 1124 GTKLKFSSAYHPQTDGQTE 1142
GTK++ S+ YHPQTDGQ+E
Sbjct: 818 GTKVQMSTTYHPQTDGQSE 836
>At4g07830 putative reverse transcriptase
Length = 611
Score = 276 bits (707), Expect = 5e-74
Identities = 189/609 (31%), Positives = 292/609 (47%), Gaps = 75/609 (12%)
Query: 803 MQEGRPVAYMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLA 862
MQ G+ +AY S+ L ++ E+ AV TD +SL+++
Sbjct: 1 MQRGKVIAYGSRQLKKHEGNYPTHDLEMAAVF------------------TDHKSLKYIF 42
Query: 863 DQRIMGEEQQKWMSKLMGYDFEIKYKPGIENKAADALSRK-------------------L 903
Q + ++WM + YD EI Y PG N DALSRK L
Sbjct: 43 TQLELNLRLRRWMKLVADYDLEIAYHPGKANVVTDALSRKRVGAAPGQSVETLVIEIGAL 102
Query: 904 QFSAISSVQCAEWADLEAEILEDERYRKVLQELATQGNSAVGYQLK---RGRLLYKDRIV 960
+ A++ A + ++L + + E + A G++ + G +L R+
Sbjct: 103 RLCAVAREPLGLEAVDQTDLLSRVQLAQEKDEGLIAASKAEGFEYQFAANGTILVHGRVC 162
Query: 961 LPKGSTKILTVLKEFHDTALGGHAGIFRTYKRISALFYWEGMKLDIQNYVQKCEVCQRNK 1020
+PK +L E H + H G + Y+ + + W GMK D+ N+V++C+VCQ K
Sbjct: 163 VPKDKELRQEILSEAHASMFSIHPGATKMYRDLKRYYQWVGMKRDVGNWVEECDVCQLVK 222
Query: 1021 YEALNPAGFLQPLPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIALSHPY 1080
E LQ LPIP W I+MDF+ GLP + KD I V+VDR TK AHF+A+
Sbjct: 223 IEHQVSGSLLQSLPIPEWKWDFITMDFVVGLPVSRTKDAIWVIVDRLTKSAHFLAIRKTD 282
Query: 1081 NAKEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQ 1140
A +A+ ++ E+V+LHG P SIVSDRD F S FW GTK++ S+AYHPQTDGQ
Sbjct: 283 GAAVLAKKYVSEIVKLHGVPVSIVSDRDSKFTSAFWRAFQAEMGTKVQMSTAYHPQTDGQ 342
Query: 1141 TEVVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGREPPVIF- 1199
+E + +E LR W LS EF YN +Y ++I PF+ LYGR +
Sbjct: 343 SERTIQTLEDMLRMCVLDWRGHWADHLSLVEFAYNNSYQASIGMAPFEVLYGRPCRTLCW 402
Query: 1200 --KGNDSLTSVDEVEKLTAERNLILEELKSNLEKAQNRMRQQANKHRRDVQYEVGDLVYL 1257
G S+ D V+++T + LK N+++AQNR R A+K R+++++EVGD V
Sbjct: 403 TQVGERSIYGADYVQEITER----IRVLKLNMKEAQNRQRSYADKRRKELEFEVGDSV-- 456
Query: 1258 KIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQV-HPVFHISLLKKA 1316
RS Q ++ P A++L+L + + H VFH+S+L+K
Sbjct: 457 -----SQDGHVARSEQ--------------RVGPVAFRLELSDVMRAFHKVFHVSMLRKC 497
Query: 1317 VNAGVQ-SQPLPAALTEEWELKVEPEAIMDTR----ENRDGDLEVLIRWKDLPTFEDSWE 1371
++ + +P L L+ P +++ R + L ++R D T E++WE
Sbjct: 498 LHKDDEVLAKIPEDLQPNMTLEARPVRVLERRIKELRRKKIPLIKVLRNCDGVT-EETWE 556
Query: 1372 DFSKLLDQF 1380
++L +F
Sbjct: 557 PEARLKARF 565
>At1g20390 hypothetical protein
Length = 1791
Score = 224 bits (570), Expect = 4e-58
Identities = 179/652 (27%), Positives = 306/652 (46%), Gaps = 54/652 (8%)
Query: 300 NREVLILIDCGATSNFISQDLVVELEIPVIATSEYVVEVG------NGAKERNSGVCKNL 353
+R +LID G++ + I +D++ + I T + V +G G K L
Sbjct: 580 SRVTRVLIDTGSSVDLIFKDVLTAMNI----TDRQIKPVSKPLAGFDGDFVMTIGTIK-L 634
Query: 354 KLEVQGISIMQHFFILGLGGT-EVVLGMDWLASLGNIEANFQELIIQWVSQGQKMVLQGE 412
+ V G+ F ++G V+LG W+ + I + + + + ++ + L+
Sbjct: 635 PIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIPSTYHQCV-KFPTHNGIFTLRA- 692
Query: 413 PSVCRVTANWKSIKITEQQEAEGYYLSYEYQKEEEKTEAEVPEGMR----KILEEYPEVF 468
P + + +S + +E E + AE+ +R +L+ + F
Sbjct: 693 PKEAKTPS--RSYEESELCRTEMVNIDESDPTRCVGVGAEISPSIRLELIALLKRNSKTF 750
Query: 469 ----QEPKGLPPRRTTDHAIQLQEGASIPNIRPYRYPFYQKNE---------IEKLVKEM 515
++ KG+ P T N+ P P QK + + V+++
Sbjct: 751 AWSIEDMKGIDPAITAHEL----------NVDPTFKPVKQKRRKLGPERARAVNEEVEKL 800
Query: 516 LNSG-IIRHSTSPFSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGA 574
L +G II + + ++VKKK+G WR CVDY LNKA D +P+P ID L++
Sbjct: 801 LKAGQIIEVKYPEWLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPHIDRLVEATSG 860
Query: 575 AVVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLR 634
+ S +D SGY+QI M ++D KT+F T G Y Y V+ FGL NA +T+Q +N++L
Sbjct: 861 NGLLSFMDAFSGYNQILMHKDDQEKTSFVTDRGTYCYKVMSFGLKNAGATYQRFVNKMLA 920
Query: 635 PYLRKFVLVFFDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSFGQPEIIYLGHV 694
+ + V V+ DD+L+ S E H +HL VL + N KC+FG +LG+V
Sbjct: 921 DQIGRTVEVYIDDMLVKSLKPEDHVEHLSKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYV 980
Query: 695 ISQAGVAADPSKIKDMLDWPIPKEVKGLRGFLGLTGYYRRFVKNYSKLAQPLNQLLKKNS 754
+++ G+ A+P +I+ +L+ P P+ + ++ G RF+ + P LLK+ +
Sbjct: 981 VTKRGIEANPKQIRAILELPSPRNAREVQRLTGRIAALNRFISRSTDKCLPFYNLLKRRA 1040
Query: 755 -FQWTEGATQAFVKLKEVMTTVPVLVPPNFDKPFILETDASGKGLGAVLMQEG----RPV 809
F W + + +AF KLK+ ++T P+LV P + L S + +VL++E RP+
Sbjct: 1041 QFDWDKDSEEAFEKLKDYLSTPPILVKPEVGETLYLYIAVSDHAVSSVLVREDRGEQRPI 1100
Query: 810 AYMSKTLSDRAQAKSVYERELMAVVLAVQKWRHYLLGSKFVIHTDQRSLRFLADQRIMGE 869
Y SK+L + V E+ +AVV + +K R Y + TDQ LR
Sbjct: 1101 FYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTDQ-PLRVALHSPSQSG 1159
Query: 870 EQQKWMSKLMGYDFEIKYKPGIENKA-ADAL-SRKLQFS--AISSVQCAEWA 917
KW +L YD + + +P ++++ AD L LQ + A+S + EW+
Sbjct: 1160 RMTKWAVELSEYDIDFRPRPAMKSQVLADFLIELPLQSAERAVSGNRGEEWS 1211
Score = 118 bits (295), Expect = 3e-26
Identities = 100/364 (27%), Positives = 162/364 (44%), Gaps = 22/364 (6%)
Query: 966 TKILTVLKEFHDTALGGHAGIFRTYKRISAL-FYWEGMKLDIQNYVQKCEVCQRNKYEAL 1024
T+I ++KE H+ A G H+G ++ L FYW M D + + KCE CQR+
Sbjct: 1433 TEINEIMKETHEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIH 1492
Query: 1025 NPAGFLQP--LPIPSQGWTDISMDFIGGLPKAMGKDTILVVVDRFTKYAHFIALSHPYNA 1082
P L+ P P W +MD +G +P + K ILV+ D FTK+ + + A
Sbjct: 1493 QPTELLRAGVAPYPFMRW---AMDIVGPMPASRQKRFILVMTDYFTKWVEAESYA-TIRA 1548
Query: 1083 KEIAEVFIKEVVRLHGFPTSIVSDRDRVFLSTFWSEMFKLAGTKLKFSSAYHPQTDGQTE 1142
++ K ++ HG P I++D F+S + +L S+ +PQ +GQ E
Sbjct: 1549 NDVQNFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAE 1608
Query: 1143 VVNRCVETYLRCVTGSKPKQWPKWLSWAEFWYNTNYHSAIKTTPFKALYGREPPVIFK-G 1201
N+ + + L+ K W L + Y T SA TPF YG E + G
Sbjct: 1609 ATNKTILSGLKKRLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEVG 1668
Query: 1202 NDSLTSVDEV------EKLTAERNLILEELK-SNLEKAQNRMRQQANKHRRDV---QYEV 1251
SL V +++ +R LEE++ + L + QN A + + V ++V
Sbjct: 1669 YSSLRRSMMVKNPELNDRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDV 1728
Query: 1252 GDLVYLKIQPYKLKSLAKRSNQKLSPRYYGPYPIIAKINPAAYKLQLPEGSQVHPVFHIS 1311
GDLV K+ ++ A+ + KL + G Y + + P Y+L G+ V ++
Sbjct: 1729 GDLVLRKV----FENTAEINAGKLGANWEGSYQVSKIVRPGDYELLTMSGTAVPRTWNSM 1784
Query: 1312 LLKK 1315
LK+
Sbjct: 1785 HLKR 1788
>At4g04230 putative transposon protein
Length = 315
Score = 198 bits (503), Expect = 2e-50
Identities = 94/168 (55%), Positives = 126/168 (74%)
Query: 516 LNSGIIRHSTSPFSSPAILVKKKDGGWRFCVDYRALNKATIPDKFPIPIIDELLDEIGAA 575
++ G IR S SP + P +LV KKDG WR C+D RA+N TI + PIP + ++LDE+ A
Sbjct: 125 IHKGYIRESLSPCAVPVLLVPKKDGTWRMCLDCRAINNITIKYRHPIPRLYDMLDELSGA 184
Query: 576 VVFSKLDLKSGYHQIRMKEEDIPKTAFRTHEGHYEYLVLPFGLTNAPSTFQALMNQVLRP 635
++FSK+DL+SGYHQ+RM+E D KTAF+T +G YE LV+PFGLTNAPSTF LMNQVLR
Sbjct: 185 IIFSKVDLRSGYHQVRMREGDEWKTAFKTKQGLYECLVMPFGLTNAPSTFMRLMNQVLRS 244
Query: 636 YLRKFVLVFFDDILIYSKNEELHKDHLRIVLQVLKENNLVANQKKCSF 683
++ KFV+V+FDDILIY+K+ H L ++L+ L++ L AN KKC+F
Sbjct: 245 FIGKFVVVYFDDILIYNKSYSDHIQPLELILKTLRKEGLYANLKKCTF 292
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,247,758
Number of Sequences: 26719
Number of extensions: 1474190
Number of successful extensions: 5845
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 81
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 5384
Number of HSP's gapped (non-prelim): 282
length of query: 1393
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1281
effective length of database: 8,326,068
effective search space: 10665693108
effective search space used: 10665693108
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0029a.5


