

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0026.15
(145 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
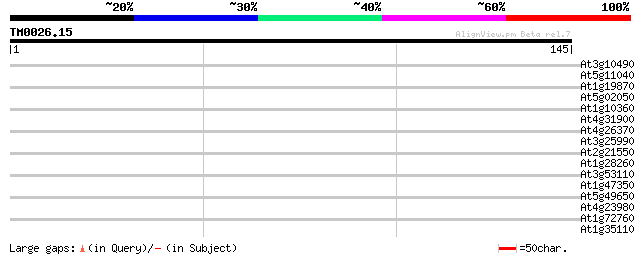
Score E
Sequences producing significant alignments: (bits) Value
At3g10490 unknown protein 29 0.76
At5g11040 putative protein 28 2.2
At1g19870 unknown protein 27 2.9
At5g02050 unknown protein 27 3.8
At1g10360 putative glutathione S-transferase TSI-1 27 3.8
At4g31900 putative protein 27 4.9
At4g26370 unknown protein 27 4.9
At3g25990 putative DNA-binding protein, GT-1 27 4.9
At2g21550 putative dihydrofolate reductase-thymidylate synthase 27 4.9
At1g28260 unknown protein 27 4.9
At3g53110 RNA helicase -like protein 26 6.5
At1g47350 hypothetical protein 26 6.5
At5g49650 xylulose kinase 26 8.4
At4g23980 auxin response factor 9 (ARF9) 26 8.4
At1g72760 putative protein kinase 26 8.4
At1g35110 unknown protein 26 8.4
>At3g10490 unknown protein
Length = 451
Score = 29.3 bits (64), Expect = 0.76
Identities = 23/72 (31%), Positives = 30/72 (40%), Gaps = 17/72 (23%)
Query: 77 SHQLLNMKSEP-EKDPLELVPDDENEHYVDSK----------------LMSKPMEEEELP 119
S L+N+ P E PL++ D +N H D K L + MEEEE P
Sbjct: 234 SKSLININEPPRETAPLDIESDQQNHHENDLKPEEHNNNNNYDENEETLKREQMEEEERP 293
Query: 120 PLEGELWNWEDP 131
P + N E P
Sbjct: 294 PRPVCVLNKEAP 305
>At5g11040 putative protein
Length = 1186
Score = 27.7 bits (60), Expect = 2.2
Identities = 20/76 (26%), Positives = 37/76 (48%), Gaps = 7/76 (9%)
Query: 69 FELPIVVGSHQLLNMKSEPEKDPLELVPDDEN-EHYVDSKLMSK---PMEEEELPPLEGE 124
FE+ + V QL N E + P++ P+ E + +D ++ P+E +LP L+G
Sbjct: 905 FEISVFV---QLENSAKEDDSSPVQDSPEYEYPKTRIDRDYSARVLIPLEHFKLPVLDGS 961
Query: 125 LWNWEDPPDDELRARS 140
+ + PP +R+
Sbjct: 962 FFTKDPPPGSPSSSRN 977
>At1g19870 unknown protein
Length = 805
Score = 27.3 bits (59), Expect = 2.9
Identities = 13/45 (28%), Positives = 22/45 (48%)
Query: 84 KSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNW 128
KSEP +L+ + +H ++S +KP+ + P WNW
Sbjct: 291 KSEPNAAAQKLLENKFAKHLMESTPKTKPINIKCDPTKPSSAWNW 335
>At5g02050 unknown protein
Length = 267
Score = 26.9 bits (58), Expect = 3.8
Identities = 26/114 (22%), Positives = 50/114 (43%), Gaps = 11/114 (9%)
Query: 25 VNDEHQVRRIYVGSAAVRSFDQRGRWCETYLMNGRVCLPYAPRSFELPIVVGS---HQLL 81
+ HQ+RR V S +V SF +T V + + E+ V H
Sbjct: 39 IRGSHQIRRGSVSSFSVSSFSTESAISKTTADENLVSVIES----EIECAVAEEAPHDTD 94
Query: 82 NMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDPPDDE 135
++ +PE P E++ D+ E V L+++ E+E + + +++D ++E
Sbjct: 95 FLEDKPEGFPFEII-DNSGERTV---LLTRKFEDETIQVEVDSVASYDDEDEEE 144
>At1g10360 putative glutathione S-transferase TSI-1
Length = 227
Score = 26.9 bits (58), Expect = 3.8
Identities = 22/67 (32%), Positives = 31/67 (45%), Gaps = 9/67 (13%)
Query: 48 GRWCETYLMNGRVCLPYAPRSFE-LPIVVGSHQLLNMKSEP--EKDPLELVPD----DEN 100
G W Y+M R+ L S+E L GS L +KS P +K P+ + D + N
Sbjct: 10 GSWASVYVMRARIALHLKSISYEFLQETYGSKSELLLKSNPVHKKMPVLIHADKPVCESN 69
Query: 101 --EHYVD 105
HY+D
Sbjct: 70 IIVHYID 76
>At4g31900 putative protein
Length = 1067
Score = 26.6 bits (57), Expect = 4.9
Identities = 13/38 (34%), Positives = 22/38 (57%)
Query: 83 MKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPP 120
M +E + D LE + D+E+E+ +D ++ EEE P
Sbjct: 701 MYAEDDLDGLEEISDEEDEYCLDDLKVTSDEEEEADEP 738
>At4g26370 unknown protein
Length = 301
Score = 26.6 bits (57), Expect = 4.9
Identities = 23/78 (29%), Positives = 37/78 (46%), Gaps = 11/78 (14%)
Query: 37 GSAAVRSFDQR--GRWCETYLMNGRVCLPYAPRSFELPIVVGSHQLLNMKSEPEKDPLEL 94
GS +R F++R R Y + L Y SF P V K+E +++ EL
Sbjct: 115 GSDPIRLFEKRINARREPGYEFDKSSLLEYNHMSFGGPPV---------KTETKEEEDEL 165
Query: 95 VPDDENEHYVDSKLMSKP 112
V DE E ++++++S P
Sbjct: 166 VRHDEKESKIEAEVLSAP 183
>At3g25990 putative DNA-binding protein, GT-1
Length = 372
Score = 26.6 bits (57), Expect = 4.9
Identities = 16/63 (25%), Positives = 27/63 (42%), Gaps = 17/63 (26%)
Query: 81 LNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDPPDDELRARS 140
LN+++E + D L L + + P+ +PP WNW D P + + +
Sbjct: 194 LNLETELDHDGLPL------------PIAADPITANGVPP-----WNWRDTPGNGVDGQP 236
Query: 141 FVG 143
F G
Sbjct: 237 FAG 239
>At2g21550 putative dihydrofolate reductase-thymidylate synthase
Length = 492
Score = 26.6 bits (57), Expect = 4.9
Identities = 16/58 (27%), Positives = 30/58 (51%)
Query: 69 FELPIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELW 126
F L +V QL++ + P+++ + DE + Y+DS ++ E+ + P L G W
Sbjct: 281 FWLGVVEEILQLISGSNNPKENGSHIWDTDEAKEYLDSFGVNATEEDGDNPFLHGLHW 338
>At1g28260 unknown protein
Length = 880
Score = 26.6 bits (57), Expect = 4.9
Identities = 18/75 (24%), Positives = 35/75 (46%), Gaps = 2/75 (2%)
Query: 53 TYLMNGRVCLPYAPRSFEL--PIVVGSHQLLNMKSEPEKDPLELVPDDENEHYVDSKLMS 110
TY+ + + L + RS + P S+ LL + + PL+ + D +Y++
Sbjct: 190 TYVSDELLALYHCVRSLAVKEPFPGASNNLLLLFEKNRSSPLQSLSTDAEFNYLNPSEKK 249
Query: 111 KPMEEEELPPLEGEL 125
++E +L +GEL
Sbjct: 250 VSVKERDLSKAKGEL 264
>At3g53110 RNA helicase -like protein
Length = 496
Score = 26.2 bits (56), Expect = 6.5
Identities = 20/57 (35%), Positives = 29/57 (50%), Gaps = 7/57 (12%)
Query: 84 KSEP--EKDPLELVPDDENEHYVDSKLMSKPMEEEELPPLEGELWNWEDPPDDELRA 138
K+EP EK V DD++E S+L S ++EEE P E+P D ++A
Sbjct: 28 KTEPTTEKKKWGDVEDDDDEEEAVSELNSLSIKEEEKPDS-----ILEEPEDSNIKA 79
>At1g47350 hypothetical protein
Length = 528
Score = 26.2 bits (56), Expect = 6.5
Identities = 15/44 (34%), Positives = 25/44 (56%), Gaps = 1/44 (2%)
Query: 99 ENEHYVDSKLMSKPMEEEELPPLEG-ELWNWEDPPDDELRARSF 141
E+ + +DSKL+ + E+ P LE E + ++ DDE RS+
Sbjct: 455 EDINVIDSKLLKSKIYEDTYPNLEDYEQSDHDETSDDEESLRSY 498
>At5g49650 xylulose kinase
Length = 558
Score = 25.8 bits (55), Expect = 8.4
Identities = 19/78 (24%), Positives = 33/78 (41%), Gaps = 11/78 (14%)
Query: 55 LMNGRVCLPYAPRSFELPIVVGSHQLL-------NMKSEPEKDPLELVPDDENEHYVDSK 107
L NG++ Y P+ VGSH+ + +++ E++ E P E ++ +
Sbjct: 372 LNNGKLGFYYTENEILPPLPVGSHRYILENFSGESLEGVKEQEVGEFDPPSEVRALIEGQ 431
Query: 108 LMSKPMEEEEL----PPL 121
+SK E PPL
Sbjct: 432 FLSKRAHTERFGMPSPPL 449
>At4g23980 auxin response factor 9 (ARF9)
Length = 638
Score = 25.8 bits (55), Expect = 8.4
Identities = 12/39 (30%), Positives = 22/39 (55%), Gaps = 6/39 (15%)
Query: 6 RTGHSWEIILHDEDRDRAWVNDE------HQVRRIYVGS 38
R+ + WEI+ D++ D V D+ + V+RI++ S
Sbjct: 565 RSRNQWEIVFTDDEGDMMLVGDDPWPEFCNMVKRIFIWS 603
>At1g72760 putative protein kinase
Length = 697
Score = 25.8 bits (55), Expect = 8.4
Identities = 14/36 (38%), Positives = 21/36 (57%), Gaps = 3/36 (8%)
Query: 12 EIILHDEDRDRAWVN---DEHQVRRIYVGSAAVRSF 44
E+ILHD D A VN + + + + VG++A SF
Sbjct: 87 EVILHDIDISNAIVNYITNNYYIANLVVGASARNSF 122
>At1g35110 unknown protein
Length = 1311
Score = 25.8 bits (55), Expect = 8.4
Identities = 19/62 (30%), Positives = 26/62 (41%), Gaps = 10/62 (16%)
Query: 89 KDPLELVPDDENEHYVDSKL-----MSKPMEEEELPPLEGELWN-----WEDPPDDELRA 138
KDP + PDDE + + DS P+EE + + E E PDD + A
Sbjct: 715 KDPNDNEPDDEGKIHFDSSANVSNKKQSPLEEHNIMEVNLEAGEASQSVVEPHPDDPMEA 774
Query: 139 RS 140
S
Sbjct: 775 NS 776
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.137 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,972,012
Number of Sequences: 26719
Number of extensions: 177158
Number of successful extensions: 480
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 476
Number of HSP's gapped (non-prelim): 18
length of query: 145
length of database: 11,318,596
effective HSP length: 90
effective length of query: 55
effective length of database: 8,913,886
effective search space: 490263730
effective search space used: 490263730
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0026.15


