

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0023.5
(266 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
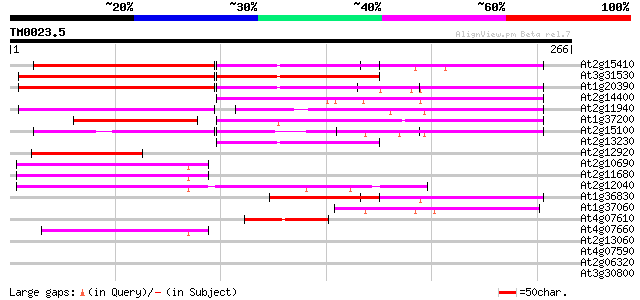
Score E
Sequences producing significant alignments: (bits) Value
At2g15410 putative retroelement pol polyprotein 81 7e-16
At3g31530 hypothetical protein 80 9e-16
At1g20390 hypothetical protein 80 1e-15
At2g14400 putative retroelement pol polyprotein 76 2e-14
At2g11940 putative retroelement gag/pol polyprotein 74 1e-13
At1g37200 hypothetical protein 64 1e-10
At2g15100 putative retroelement pol polyprotein 61 5e-10
At2g13230 pseudogene 59 2e-09
At2g12920 pseudogene 54 7e-08
At2g10690 putative retroelement pol polyprotein 51 7e-07
At2g11680 putative retroelement pol polyprotein 49 3e-06
At2g12040 T10J7.2 49 4e-06
At1g36830 hypothetical protein 47 8e-06
At1g37060 Athila retroelment ORF 1, putative 42 3e-04
At4g07610 42 4e-04
At4g07660 putative athila transposon protein 41 8e-04
At2g13060 pseudogene 40 0.002
At4g07590 39 0.003
At2g06320 putative retroelement pol polyprotein 39 0.003
At3g30800 hypothetical protein 38 0.006
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 80.9 bits (198), Expect = 7e-16
Identities = 39/86 (45%), Positives = 62/86 (71%)
Query: 12 LILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRP 71
L LY+AV+++AVS +L++E+ + I++VS TL GAE+RY +EK A A++ +AR+LRP
Sbjct: 1059 LYLYIAVSTSAVSGVLVREDRGEQHPIFYVSKTLDGAELRYPTLEKLAFAVVISARKLRP 1118
Query: 72 YFQSFQIKVKTDILLRQVWQKPDLSG 97
YF+S+ ++V T+ LR + P SG
Sbjct: 1119 YFKSYTVEVLTNQPLRTILHSPSQSG 1144
Score = 66.2 bits (160), Expect = 2e-11
Identities = 31/77 (40%), Positives = 45/77 (58%), Gaps = 1/77 (1%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P E L T P+PF W DI+ F ++ Q K+I+V DYFTKW EAE+ I +++
Sbjct: 1490 PSEVLQTAMPPYPFMRWAMDIIRPFPASR-QKKYILVMTDYFTKWFEAESYARIQSKEVQ 1548
Query: 159 NFLWRRITSTGETPYKL 175
NF+W+ I PY++
Sbjct: 1549 NFVWKNIICHHGLPYEI 1565
Score = 46.2 bits (108), Expect = 2e-05
Identities = 33/89 (37%), Positives = 48/89 (53%), Gaps = 2/89 (2%)
Query: 167 STGETPYKLTYGSDAMLPVEVENQS-WRVITMNEEENNDN-LIANLIMLPELQREAHLRN 224
ST TP+ LTYG +A+ P E + R + +N+ ND L+ NL L E + +A LR
Sbjct: 1643 STDRTPFSLTYGMEALAPCEAGLPTVRRSMLINDPTLNDQMLLDNLDTLEEQRDQALLRI 1702
Query: 225 EAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ + A+ Y KV +R DLVLR+
Sbjct: 1703 QNYQQAAAKFYNKKVKNRLFAEGDLVLRK 1731
>At3g31530 hypothetical protein
Length = 831
Score = 80.5 bits (197), Expect = 9e-16
Identities = 42/93 (45%), Positives = 60/93 (64%)
Query: 5 KPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILK 64
KP P LY+AV+ TAVS L++E+ + K+I++VS T AE RY ++EK ALA++
Sbjct: 274 KPVIGEPQYLYVAVSDTAVSGELVREDRGEQKLIFYVSQTFTSAESRYPQMEKLALAVVM 333
Query: 65 TARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
+A++LRPYFQS I V + LR + P SG
Sbjct: 334 SAQKLRPYFQSHSIIVMGSMPLRVILHSPSQSG 366
Score = 64.3 bits (155), Expect = 6e-11
Identities = 30/77 (38%), Positives = 48/77 (61%), Gaps = 1/77 (1%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P E L+++ +P+PF W DI+G +K Q K ++V DYF+KWIEAE+ +I A++
Sbjct: 648 PAELLSSIASPYPFMRWSMDIIGPMHPSK-QKKLVLVLTDYFSKWIEAESYASIKDAQVE 706
Query: 159 NFLWRRITSTGETPYKL 175
NF+ + I PY++
Sbjct: 707 NFVLKHILCRHGIPYEI 723
>At1g20390 hypothetical protein
Length = 1791
Score = 79.7 bits (195), Expect = 1e-15
Identities = 44/93 (47%), Positives = 60/93 (64%)
Query: 5 KPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILK 64
KP L LY+AV+ AVS++L++E+ + + I++ S +L AE RY IEKAALA++
Sbjct: 1067 KPEVGETLYLYIAVSDHAVSSVLVREDRGEQRPIFYTSKSLVEAETRYPVIEKAALAVVT 1126
Query: 65 TARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
+AR+LRPYFQS I V TD LR P SG
Sbjct: 1127 SARKLRPYFQSHTIAVLTDQPLRVALHSPSQSG 1159
Score = 71.2 bits (173), Expect = 5e-13
Identities = 37/99 (37%), Positives = 55/99 (55%), Gaps = 4/99 (4%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P E L AP+PF W DI+G ++ Q +FI+V DYFTKW+EAE+ TI ++
Sbjct: 1494 PTELLRAGVAPYPFMRWAMDIVGPMPASR-QKRFILVMTDYFTKWVEAESYATIRANDVQ 1552
Query: 159 NFLWRRITSTGETPYK-LTYGSDAMLPVEVEN--QSWRV 194
NF+W+ I PY+ +T + + EN SW++
Sbjct: 1553 NFVWKFIICRHGLPYEIITDNGSQFISLSFENFCASWKI 1591
Score = 45.8 bits (107), Expect = 2e-05
Identities = 30/90 (33%), Positives = 47/90 (51%), Gaps = 2/90 (2%)
Query: 166 TSTGETPYKLTYGSDAMLPVEVENQSWR--VITMNEEENNDNLIANLIMLPELQREAHLR 223
++T +TP+ YG +AM P EV S R ++ N E N+ ++ L L E++ A R
Sbjct: 1646 SATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVKNPELNDRMMLDRLDDLEEIRNAALCR 1705
Query: 224 NEAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ + A+ Y KV +R DLVLR+
Sbjct: 1706 IQNYQLAAAKHYNQKVHNRHFDVGDLVLRK 1735
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 75.9 bits (185), Expect = 2e-14
Identities = 58/201 (28%), Positives = 93/201 (45%), Gaps = 46/201 (22%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAV--------- 149
P E ++++AP+PF W DI+G + +++++V DYF+KWIEAEA+
Sbjct: 1030 PSELFSSISAPYPFMRWSMDIIGPLHRSTRGVQYLLVLTDYFSKWIEAEAILLNWGIKVS 1089
Query: 150 -----------------ETIT--------------VAKIRNFLWRRIT----STGETPYK 174
+TI +++ LW T STGETP+
Sbjct: 1090 YSTLRYPQGNGQAEAANKTILSNLKKRLNHLKGGWYDELQPVLWAYRTTPRRSTGETPFS 1149
Query: 175 LTYGSDAMLPVEVENQSWR--VITMNEEENNDNLIANLIMLPELQREAHLRNEAGKGRVA 232
L YG A++P E+ R +NE+EN+ L +L + E + +A ++ + + A
Sbjct: 1150 LVYGMKAVVPAELNVPGLRRTEAPLNEKENSAMLDDSLDTINERRDQALIQIQNYQHAAA 1209
Query: 233 RKYASKVDSRKMKARDLVLRQ 253
R Y SKV SR D VL++
Sbjct: 1210 RYYNSKVKSRPFFVGDYVLKR 1230
Score = 33.9 bits (76), Expect = 0.092
Identities = 20/74 (27%), Positives = 39/74 (52%), Gaps = 2/74 (2%)
Query: 5 KPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILK 64
KP + L LY++V++ AVS +L++E+ + K IY++ + Y+ + A L
Sbjct: 754 KPELDEKLYLYVSVSNHAVSGVLIREDRGEQKPIYYIIVNQFNGD--YEAKDSRMEAYLH 811
Query: 65 TARRLRPYFQSFQI 78
+ L F+ F++
Sbjct: 812 VVKDLAKNFRKFEL 825
>At2g11940 putative retroelement gag/pol polyprotein
Length = 1212
Score = 73.6 bits (179), Expect = 1e-13
Identities = 38/93 (40%), Positives = 56/93 (59%)
Query: 5 KPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILK 64
KP L LY++V+ A+S +L++ E + I++VS + E RY +EK ALA++
Sbjct: 611 KPEFGERLFLYISVSENAISGVLVRVERSDERPIFYVSKSFSDVETRYPMMEKLALAVVT 670
Query: 65 TARRLRPYFQSFQIKVKTDILLRQVWQKPDLSG 97
AR+LRPYFQS I V T + LR + P+ SG
Sbjct: 671 AARKLRPYFQSHPIVVLTSLPLRTILHSPNQSG 703
Score = 55.5 bits (132), Expect = 3e-08
Identities = 43/180 (23%), Positives = 75/180 (40%), Gaps = 40/180 (22%)
Query: 108 APWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNFLWRRITS 167
+P PF W DI+G V+ ++F+++ +W+EA A IT ++R+F+W+ I
Sbjct: 984 SPLPFMKWSMDIVGPLHVSTRGVRFLLL------EWVEAAAYSYITQVQVRHFIWKEIIC 1037
Query: 168 TGETPYKLTYGS--------------------------------DAMLPVEVENQSWRVI 195
PY++ + +A++P E++ R I
Sbjct: 1038 RHGLPYEIVTDNGPQFISEQFKAFYAEWQIRLNRSTPRYPQGNVEAVIPAEIKVPRTRRI 1097
Query: 196 --TMNEEENNDNLIANLIMLPELQREAHLRNEAGKGRVARKYASKVDSRKMKARDLVLRQ 253
NE NN ++ + + E + A +R + A+ Y S V SR LVLR+
Sbjct: 1098 RNPKNEAANNKMMVDVIDTIDERRNRALVRMQNYHNAAAQYYNSNVRSRSFDVGTLVLRR 1157
>At1g37200 hypothetical protein
Length = 1564
Score = 63.5 bits (153), Expect = 1e-10
Identities = 38/157 (24%), Positives = 79/157 (50%), Gaps = 3/157 (1%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVA--KAQMKFIVVAVDYFTKWIEAEAVETITVAK 156
PPE+++++++P+PF W D++ + K ++K ++V DY TKWIE +A + +T +
Sbjct: 1352 PPEEMSSISSPYPFMKWSMDVVRPMEASGGKKRLKNLLVLTDYSTKWIEGKAFQQVTEKQ 1411
Query: 157 IRNFLWRRITSTGETPYKLTYGSDAMLPVEVENQSWRVITMNEEENNDNLIANLIMLPEL 216
+ FL I PY++ + P E +++ + + + L+ + E
Sbjct: 1412 VEVFLLENIVYRHGIPYEIVTDNGLTSPRE-KSKHFATSGRSALPHPPPLLPTRDFIDER 1470
Query: 217 QREAHLRNEAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ A ++ + + R Y S + R+ + +LVLR+
Sbjct: 1471 RDHAMIQVQNYQQPAVRYYNSNIKIRRFEVGELVLRK 1507
Score = 52.8 bits (125), Expect = 2e-07
Identities = 27/59 (45%), Positives = 38/59 (63%)
Query: 31 EHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRPYFQSFQIKVKTDILLRQV 89
+H+ K IY+VS TL AE Y+ +EK ALA++ +AR+LR Y QS I V T +R +
Sbjct: 927 DHEGQKPIYYVSKTLIDAETHYRAMEKLALAVVMSARKLRQYLQSHPIVVMTSQPIRTI 985
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 61.2 bits (147), Expect = 5e-10
Identities = 34/86 (39%), Positives = 51/86 (58%), Gaps = 7/86 (8%)
Query: 12 LILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRP 71
L LY+AV+ +AV+ + ++ E + I++V + RY +EK ALA++ AR+LRP
Sbjct: 661 LFLYIAVSESAVTGVQVRVERSDKRPIFYV-------KTRYPMMEKLALAVVTAARKLRP 713
Query: 72 YFQSFQIKVKTDILLRQVWQKPDLSG 97
YFQS I V T + LR + P SG
Sbjct: 714 YFQSHPIVVLTSLPLRTILHSPTQSG 739
Score = 53.5 bits (127), Expect = 1e-07
Identities = 36/104 (34%), Positives = 55/104 (52%), Gaps = 6/104 (5%)
Query: 156 KIRNFLWRRITS----TGETPYKLTYGSDAMLPVEVENQSWRVI--TMNEEENNDNLIAN 209
+++N LW T+ T ETP+ L YG +A++P E++ S R I NE ENN+ +I
Sbjct: 1170 ELQNVLWAVRTTPRRATDETPFSLIYGMEAVIPAEIKVPSARRIRNPQNETENNEMIIDV 1229
Query: 210 LIMLPELQREAHLRNEAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ + E + A R + AR Y S V +R + LVLR+
Sbjct: 1230 IDTIDERRNRALARMQNYHNAGARYYNSNVRNRSFEVGTLVLRR 1273
Score = 47.8 bits (112), Expect = 6e-06
Identities = 29/99 (29%), Positives = 47/99 (47%), Gaps = 17/99 (17%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P E LTT++AP+PF W DI+G V+ T+ +EA A IT ++
Sbjct: 1045 PAELLTTVSAPYPFMKWLMDIVGPLHVS--------------TRGVEAAAYSNITHVQVW 1090
Query: 159 NFLWRRITSTGETPYKLTYGSDAML---PVEVENQSWRV 194
NF+W+ I PY++ + + EV + W++
Sbjct: 1091 NFIWKDIICRHGLPYEIVTDNGSQFISEQFEVFCEEWQI 1129
>At2g13230 pseudogene
Length = 353
Score = 59.3 bits (142), Expect = 2e-09
Identities = 27/77 (35%), Positives = 46/77 (59%), Gaps = 1/77 (1%)
Query: 99 PPEQLTTMTAPWPFAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIR 158
P E L+++ +P+P W DI+G +K Q K ++V DYF+K EAE+ +I A++
Sbjct: 199 PAELLSSIASPYPLMRWSMDIIGPMHASK-QKKLVLVLTDYFSKLREAESYASIKEAQVE 257
Query: 159 NFLWRRITSTGETPYKL 175
+F+W+ I PY++
Sbjct: 258 SFMWKHILCRHGVPYEI 274
>At2g12920 pseudogene
Length = 863
Score = 54.3 bits (129), Expect = 7e-08
Identities = 25/53 (47%), Positives = 39/53 (73%)
Query: 11 PLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
PL LY++V+ T VS +L++E+ + K I++VS T GAE RY ++EK ALA++
Sbjct: 441 PLYLYVSVSDTTVSKVLVREDRGEQKPIFYVSQTFTGAESRYPQMEKLALAVI 493
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 50.8 bits (120), Expect = 7e-07
Identities = 30/92 (32%), Positives = 47/92 (50%), Gaps = 1/92 (1%)
Query: 4 QKPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
Q P + P + AV A+L Q+ K +IY+ S T+ A+ RY IEK LA++
Sbjct: 342 QAPNWDHPFEIMCDAFDYAVGAVLDQKIDDKLHVIYYASRTMDEAQTRYATIEKELLAVV 401
Query: 64 KTARRLRPYFQSFQIKVKTD-ILLRQVWQKPD 94
++R Y F++KV D LR ++ K +
Sbjct: 402 FAFEKIRSYLVGFKVKVYIDHAALRHIYAKKE 433
Score = 34.7 bits (78), Expect = 0.054
Identities = 30/96 (31%), Positives = 43/96 (44%), Gaps = 18/96 (18%)
Query: 115 WGADIL-----GLFSVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNFLWRRITS-- 167
W AD++ G V+ QMK D TK + + T K+ LW T+
Sbjct: 515 WYADLVNYLTSGQVEVSNRQMK------DILTKVVGVSKRDWST--KLDETLWAYRTAYK 566
Query: 168 --TGETPYKLTYGSDAMLPVEVENQS-WRVITMNEE 200
G TP+++ YG LPVEVE ++ W +N E
Sbjct: 567 TPIGRTPFQMLYGKSCHLPVEVEYKAIWATKLLNLE 602
>At2g11680 putative retroelement pol polyprotein
Length = 301
Score = 48.9 bits (115), Expect = 3e-06
Identities = 29/92 (31%), Positives = 46/92 (49%), Gaps = 1/92 (1%)
Query: 4 QKPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
Q P + P + + AV A+L Q+ +K +IY+ S T+ +VRY EK LA++
Sbjct: 97 QAPNRDYPFEIMCDDSDYAVGAVLGQKIDRKLHVIYYASGTMDDTQVRYATTEKELLAVV 156
Query: 64 KTARRLRPYFQSFQIKVKTD-ILLRQVWQKPD 94
+ R Y ++ V TD LR ++ K D
Sbjct: 157 FAFEKFRSYLVGSKVTVYTDHAALRHIYAKKD 188
>At2g12040 T10J7.2
Length = 976
Score = 48.5 bits (114), Expect = 4e-06
Identities = 50/201 (24%), Positives = 87/201 (42%), Gaps = 12/201 (5%)
Query: 4 QKPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
Q P ++P + + AV A+L Q + KK IY+ S TL A+ Y EK LA++
Sbjct: 233 QPPDWDLPFEVMCDASDFAVGAVLGQRKDKKLHAIYYASRTLDDAKRNYATTEKEFLAVV 292
Query: 64 KTARRLRPYFQSFQIKVKTD-ILLRQVWQKPDLSGAPPEQLTTMTAPWPFAIWGADILGL 122
+ R Y ++ V TD L+ + Q D A P L + F I D G+
Sbjct: 293 FAFEKFRSYLVGLKVIVHTDHAALKYLMQNKD---AKPRLLRWILLLQEFDIEVRDKKGV 349
Query: 123 FSVAKAQMKFIVVAVDY-FTKWIEAEAVETITVAKIRNF----LWRRITSTGETPYKLTY 177
+ I + D ++ E + I A+ + L R++ + +TP +
Sbjct: 350 EKCVADHLSRIRIDDDVPINDFLPEENIYMIDTAEEYDCKCDKLQNRVSLSIDTP---SM 406
Query: 178 GSDAMLPVEVENQSWRVITMN 198
D + EV+ +SW +++++
Sbjct: 407 SIDTHISEEVDIRSWGMVSID 427
Score = 37.7 bits (86), Expect = 0.006
Identities = 25/86 (29%), Positives = 43/86 (49%), Gaps = 3/86 (3%)
Query: 166 TSTGETPYKLTYGSDAMLPVEVENQ-SWRVITMNEE--ENNDNLIANLIMLPELQREAHL 222
T G TP+ L YG LPVE+E++ +W V +N + + + L L E+Q A+
Sbjct: 688 TPLGTTPFHLLYGKACHLPVELEHKAAWTVKMLNFDIKSAGERRLIQLNELDEIQIHAYD 747
Query: 223 RNEAGKGRVARKYASKVDSRKMKARD 248
++ K R + K+ +R + +D
Sbjct: 748 NSKLYKERTKAYHDKKILTRTFEPKD 773
Score = 31.6 bits (70), Expect = 0.46
Identities = 14/37 (37%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Query: 112 FAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEA 148
F WG + +G + + +I+V VDY +KW+EA A
Sbjct: 569 FDCWGINFMGPIPPSNKNL-YILVVVDYVSKWVEAIA 604
>At1g36830 hypothetical protein
Length = 243
Score = 47.4 bits (111), Expect = 8e-06
Identities = 20/52 (38%), Positives = 32/52 (61%)
Query: 124 SVAKAQMKFIVVAVDYFTKWIEAEAVETITVAKIRNFLWRRITSTGETPYKL 175
S K ++K ++V DY TKWIEA+A + +T ++ +FLW I PY++
Sbjct: 10 SGGKKKLKNLLVLTDYSTKWIEAKAFQQVTEKQVEDFLWENIVCQHGIPYEI 61
Score = 43.9 bits (102), Expect = 9e-05
Identities = 28/89 (31%), Positives = 46/89 (51%), Gaps = 2/89 (2%)
Query: 167 STGETPYKLTYGSDAMLPVEVENQSWR--VITMNEEENNDNLIANLIMLPELQREAHLRN 224
ST ETP+ L YG +A++P+ S R N + N L N+ + E + +A +R
Sbjct: 139 STQETPFSLAYGLEAVIPIVTIIPSVRRTASHANSDMNTQMLRDNMDFIDERRDQAMIRV 198
Query: 225 EAGKGRVARKYASKVDSRKMKARDLVLRQ 253
+ + V R Y S + R+ + +LVLR+
Sbjct: 199 QNYQQAVTRYYNSNIKIRRFEVGELVLRK 227
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 42.0 bits (97), Expect = 3e-04
Identities = 32/104 (30%), Positives = 51/104 (48%), Gaps = 7/104 (6%)
Query: 155 AKIRNFLWRRITS----TGETPYKLTYGSDAMLPVEVENQS-WRVITMNEE--ENNDNLI 207
AK+ + LW T+ G TP+ L YG LPVE+E ++ W V +N + + +
Sbjct: 1560 AKLNDTLWAYRTAFKTPIGTTPFNLLYGKSCHLPVELEYKAMWAVKLLNFDIKTAEEKRL 1619
Query: 208 ANLIMLPELQREAHLRNEAGKGRVARKYASKVDSRKMKARDLVL 251
L L +++ EA+ ++ K R + K+ SR K D VL
Sbjct: 1620 IQLNDLNKIRLEAYESSKIYKERTKSFHDKKIVSRDFKVGDQVL 1663
Score = 40.4 bits (93), Expect = 0.001
Identities = 18/40 (45%), Positives = 26/40 (65%), Gaps = 1/40 (2%)
Query: 112 FAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVET 151
F +WG D +G F + K+I+VA+DY +KW+EA A T
Sbjct: 1444 FDVWGIDFMGPFPSSYGN-KYILVAIDYVSKWVEAIASHT 1482
Score = 32.7 bits (73), Expect = 0.21
Identities = 17/52 (32%), Positives = 27/52 (51%), Gaps = 1/52 (1%)
Query: 44 TLQGAEVRYQKIEKAALAILKTARRLRPYFQSFQIKVKTD-ILLRQVWQKPD 94
T+ A+VRY EK LA++ + R Y ++ + TD LR ++ K D
Sbjct: 1218 TMDDAQVRYATTEKELLAVVFAFEKFRSYLVGSKVTIYTDHAALRHIYAKKD 1269
>At4g07610
Length = 276
Score = 41.6 bits (96), Expect = 4e-04
Identities = 20/40 (50%), Positives = 26/40 (65%), Gaps = 1/40 (2%)
Query: 112 FAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVET 151
F +WG D +G F + K+I+VAVDY +KWIEA A T
Sbjct: 153 FDVWGIDFIGPFPSSYGN-KYILVAVDYVSKWIEATASPT 191
>At4g07660 putative athila transposon protein
Length = 724
Score = 40.8 bits (94), Expect = 8e-04
Identities = 24/80 (30%), Positives = 38/80 (47%), Gaps = 1/80 (1%)
Query: 16 LAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAILKTARRLRPYFQS 75
+ V + M+ + K K IY+ S TL A+ RY EK L ++ ++ R Y
Sbjct: 576 IEVDKEKIKVMMQLQPPKTVKDIYYASRTLDEAQGRYATTEKELLVVVFAFKKFRSYLVE 635
Query: 76 FQIKVKTD-ILLRQVWQKPD 94
++ V TD LR ++ K D
Sbjct: 636 SKVTVYTDHAALRHMYAKKD 655
>At2g13060 pseudogene
Length = 930
Score = 39.7 bits (91), Expect = 0.002
Identities = 30/100 (30%), Positives = 49/100 (49%), Gaps = 5/100 (5%)
Query: 166 TSTGETPYKLTYGSDAMLPVEVENQ-SWRVITMNEEENNDNLIANLIMLPELQREAHLRN 224
T G TP+ L YG LPVE+E++ +W V MN + + A L E++ A+ +
Sbjct: 406 TPLGTTPFHLLYGKACHLPVELEHKAAWAVKMMNFDIKS----AGESELDEIRIHAYDNS 461
Query: 225 EAGKGRVARKYASKVDSRKMKARDLVLRQRTMTSRCNKLT 264
+ K R + K+ +R + D +LR T++ LT
Sbjct: 462 KLYKERTKAYHDKKILTRTFEPNDQILRFFTISRDFRLLT 501
Score = 34.3 bits (77), Expect = 0.071
Identities = 16/37 (43%), Positives = 23/37 (61%), Gaps = 1/37 (2%)
Query: 112 FAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEA 148
F WG D +G F + + +I+V VDY +KW+EA A
Sbjct: 267 FDRWGIDFMGPFPPSNKNL-YILVDVDYVSKWVEAIA 302
>At4g07590
Length = 860
Score = 38.9 bits (89), Expect = 0.003
Identities = 30/104 (28%), Positives = 52/104 (49%), Gaps = 7/104 (6%)
Query: 155 AKIRNFLWRRITS----TGETPYKLTYGSDAMLPVEVENQS-WRVITMNEE--ENNDNLI 207
AK+ + LW +T+ G T +KL YG LPVE+E ++ W V +N + + +
Sbjct: 44 AKLDDALWAYMTAFKTPIGTTSFKLLYGKSCHLPVELEYKAMWTVKLLNFDIKTAEEKRL 103
Query: 208 ANLIMLPELQREAHLRNEAGKGRVARKYASKVDSRKMKARDLVL 251
L L E++ EA+ R++ K + K+ ++ + D VL
Sbjct: 104 IQLRDLDEIRLEAYERSKIYKEGTKLFHDKKIITKDFQVGDQVL 147
>At2g06320 putative retroelement pol polyprotein
Length = 466
Score = 38.9 bits (89), Expect = 0.003
Identities = 18/40 (45%), Positives = 25/40 (62%), Gaps = 1/40 (2%)
Query: 112 FAIWGADILGLFSVAKAQMKFIVVAVDYFTKWIEAEAVET 151
F +WG D +G F + K+I+V VDY +KW+EA A T
Sbjct: 168 FDVWGIDFMGPFPSSYGN-KYILVVVDYVSKWVEAIASPT 206
>At3g30800 hypothetical protein
Length = 361
Score = 37.7 bits (86), Expect = 0.006
Identities = 21/67 (31%), Positives = 33/67 (48%)
Query: 4 QKPTPEVPLILYLAVTSTAVSAMLLQEEHKKYKMIYFVSHTLQGAEVRYQKIEKAALAIL 63
Q P ++P + AV A+L Q + KK IY+ S TL A++ Y +EK L ++
Sbjct: 124 QPPYWDLPFEEMCDASDFAVGAVLRQRKDKKVHDIYYASRTLDYAQMNYATVEKELLVVV 183
Query: 64 KTARRLR 70
+ R
Sbjct: 184 FAFEKFR 190
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,450,764
Number of Sequences: 26719
Number of extensions: 203770
Number of successful extensions: 581
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 519
Number of HSP's gapped (non-prelim): 58
length of query: 266
length of database: 11,318,596
effective HSP length: 98
effective length of query: 168
effective length of database: 8,700,134
effective search space: 1461622512
effective search space used: 1461622512
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0023.5


