

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.23
(1569 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
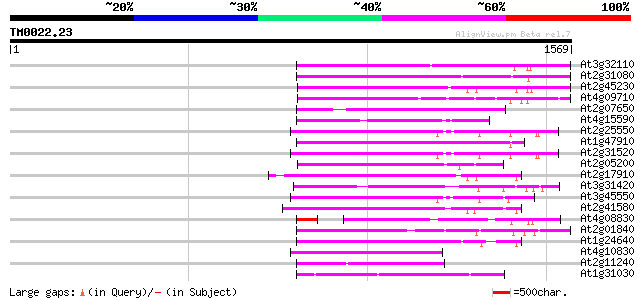
Score E
Sequences producing significant alignments: (bits) Value
At3g32110 non-LTR reverse transcriptase, putative 402 e-112
At2g31080 putative non-LTR retroelement reverse transcriptase 401 e-111
At2g45230 putative non-LTR retroelement reverse transcriptase 293 5e-79
At4g09710 RNA-directed DNA polymerase -like protein 290 3e-78
At2g07650 putative non-LTR retrolelement reverse transcriptase 282 1e-75
At4g15590 reverse transcriptase like protein 280 4e-75
At2g25550 putative non-LTR retroelement reverse transcriptase 274 3e-73
At1g47910 reverse transcriptase, putative 274 3e-73
At2g31520 putative non-LTR retroelement reverse transcriptase 273 6e-73
At2g05200 putative non-LTR retroelement reverse transcriptase 272 1e-72
At2g17910 putative non-LTR retroelement reverse transcriptase 266 5e-71
At3g31420 hypothetical protein 259 9e-69
At3g45550 putative protein 258 1e-68
At2g41580 putative non-LTR retroelement reverse transcriptase 248 3e-65
At4g08830 putative protein 245 1e-64
At2g01840 putative non-LTR retroelement reverse transcriptase 243 5e-64
At1g24640 hypothetical protein 228 2e-59
At4g10830 putative protein 222 1e-57
At2g11240 pseudogene 216 6e-56
At1g31030 F17F8.5 215 2e-55
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 402 bits (1034), Expect = e-112
Identities = 262/815 (32%), Positives = 406/815 (49%), Gaps = 53/815 (6%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLE 862
TKVMV RL+ + +LIGP Q+SFIPGR + DN +++Q++VH M ++G ++ KLDLE
Sbjct: 1058 TKVMVGRLKGVINKLIGPAQTSFIPGRLSTDNIVVVQEVVHSMRRKKGVKGWMLLKLDLE 1117
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDP 922
KA + W L DTL G P T V IM CV S+ LLWNG K P RGLR GDP
Sbjct: 1118 KAYDRIRWDLLEDTLKAAGLPGTWVQWIMKCVEGPSMRLLWNGEKTDAFKPLRGLRQGDP 1177
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
LSPYLFVLC+ERL I+ ++ + + ++IS+ G +SH+ FADD++L A+A + Q+R
Sbjct: 1178 LSPYLFVLCIERLCHLIESSIAAKKWKPIKISQSGPRLSHICFADDLILFAEASIDQIRV 1237
Query: 983 VQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQG 1042
++ +L FC ASG V++ KS+ + S +V R I + + T +L +YLG PI Q
Sbjct: 1238 LRGVLEKFCGASGQKVSLEKSKIYFSKNVLRDLGKRISEESGMKATKDLGKYLGVPILQK 1297
Query: 1043 RQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHL 1102
R +E F V+ R + + A WKGR+L+ AG+ L K VL +I V+ M + D L
Sbjct: 1298 RINKETFGEVIKRFSSRLAGWKGRMLSFAGRLTLTKAVLTSILVHTMSTIKLPQSTLDGL 1357
Query: 1103 DSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHD 1161
D +++ F+W S E++ ++VAW V LP+ GGLGIR A +N +++ K+ W ++N
Sbjct: 1358 DKVSRAFLWGSTLEKKKQHLVAWTRVCLPRREGGLGIRSATAMNKALIAKVGWRVLNDGS 1417
Query: 1162 KLWV*VLLDLYCTNSDFLH-----APRRRGSCVWNAV-MKTRDIIQDGFTWRIGRG-DLS 1214
LW V+ Y +H + S W +V + RD+I W IG G +
Sbjct: 1418 SLWAQVVRSKYKVGD--VHDRNWTVAKSNWSSTWRSVGVGLRDVIWREQHWVIGDGRQIR 1475
Query: 1215 FWFSKCHGSKALC-EKVPYVHISDTALRVRDVFQDGV-CDIRGLSTPITPELWALLQNLF 1272
FW + + + + + ++ RD+++DG D+ ++ +T L +
Sbjct: 1476 FWTDRWLSETPIADDSIVPLSLAQMLCTARDLWRDGTGWDMSQIAPFVTDNKRLDLLAVI 1535
Query: 1273 VYVSLDDEDLVVWAPSLEGKYSVQSCYRWLL-DMEGGTAVSVDWA*LWKFQVPEKMKLFA 1331
V D + W + +G+++V+S + L D +S + +WK Q PE++++F
Sbjct: 1536 VDSVTGAHDRLAWGMTSDGRFTVKSAFAMLTNDDSPRQDMSSLYGRVWKVQAPERVRVFL 1595
Query: 1332 WQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTPGF 1391
W ++++ T R + C +C +E+ +H LRDC A +W +
Sbjct: 1596 WLVVNQAIMTNSERKRRHLCDSDVCQVCRGGIESILHVLRDCPAMSGIWDRIVPRRLQQS 1655
Query: 1392 FTASSFLDWIASVPRHDV--------AGLLVHAWFIWCRRRRFVFAGE----LEPWRLIL 1439
F S L+W+ S R + + W+ W R +F GE + R I
Sbjct: 1656 FFTMSLLEWLYSNLRQGLMTEGSDWSTMFAMAVWWGWKWRCSNIF-GENKTCRDRVRFIK 1714
Query: 1440 HRAADV-----------LRCLRATSP----------------SMPAADPGRAEVGGVVRD 1472
A +V L LR P +PG A GGV+R+
Sbjct: 1715 DLAVEVSIAYSREVELRLSGLRVNKPIRWTPPMEGWYKINTDGASRGNPGLASAGGVLRN 1774
Query: 1473 AVGCWLFGYSGFIGMAEILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLLRHGVTR 1532
+ G W G++ IG AEL V +GL W + L + DS VV+ L+ G+
Sbjct: 1775 SAGAWCGGFAVNIGRCSAPLAELWGVYYGLYMAWAKQLTHLELEVDSEVVVGFLKTGIGE 1834
Query: 1533 FHVYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
H + +V L +DW V + H+ RE N +AD
Sbjct: 1835 THPLSFLVRLCHNFLSKDWTVRISHVYREANSLAD 1869
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 401 bits (1031), Expect = e-111
Identities = 270/813 (33%), Positives = 422/813 (51%), Gaps = 53/813 (6%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLE 862
TK+MV RL+ + +LIGP Q+SFIPGR +IDN +++Q+ VH M ++G+ ++ KLDLE
Sbjct: 382 TKMMVTRLKNVISKLIGPAQASFIPGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLDLE 441
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDP 922
KA V W FL++TL G S IM V+ S+S+LWNG + P+RGLR GDP
Sbjct: 442 KAYDRVRWDFLQETLEAAGLSEGWTSRIMAGVTDPSMSVLWNGERTDSFVPARGLRQGDP 501
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
LSPYLFVLC+ERL I+ +V + + + + +S GS +SH+ FADD++L A+A V+Q+R
Sbjct: 502 LSPYLFVLCLERLCHLIEASVGKREWKPIAVSCGGSKLSHVCFADDLILFAEASVAQIRI 561
Query: 983 VQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQG 1042
++ +L FC ASG V++ KS+ F S +V R + I + I T L +YLG PI Q
Sbjct: 562 IRRVLERFCEASGQKVSLEKSKIFFSHNVSREMEQLISEESGIGCTKELGKYLGMPILQK 621
Query: 1043 RQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHL 1102
R +E F VL+RV+ + A WKGR L+ AG+ L K VL++IPV+ M + D L
Sbjct: 622 RMNKETFGEVLERVSARLAGWKGRSLSLAGRITLTKAVLSSIPVHVMSAILLPVSTLDTL 681
Query: 1103 DSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHD 1161
D ++ F+W S E++ ++++WR + PK GG+G+R AR +N +++ K+ W L+ +
Sbjct: 682 DRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGLRSARDMNKALVAKVGWRLLQDKE 741
Query: 1162 KLWV*VLLDLY----CTNSDFLHAPRRRGSCVWNAV-MKTRDIIQDGFTWRIGRG-DLSF 1215
LW V+ Y ++ +L P+ R S W +V + R+++ G W G G + F
Sbjct: 742 SLWARVVRKKYKVGGVQDTSWL-KPQPRWSSTWRSVAVGLREVVVKGVGWVPGDGCTIRF 800
Query: 1216 WFSKCHGSKALCEKVPYVHISDTALRV-RDVFQDGV---CDIRGLSTPITPELWALLQNL 1271
W + + L E + ++V D + G +I GL P T + L ++
Sbjct: 801 WLDRWLLQEPLVELGTDMIPEGERIKVAADYWLPGSGWNLEILGLYLPETVK--RRLLSV 858
Query: 1272 FVYVSLDDEDLVVWAPSLEGKYSVQSCYRWLL-DMEGGTAVSVDWA*LWKFQVPEKMKLF 1330
V V L + D + W + +G ++V+S Y L D+ + + +WK PE++++F
Sbjct: 859 VVQVFLGNGDEISWKGTQDGAFTVRSAYSLLQGDVGDRPNMGSFFNRIWKLITPERVRVF 918
Query: 1331 AWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTPG 1390
W + + T R ++ CS+C+ ET +H LRDC A +W+ L
Sbjct: 919 IWLVSQNVIMTNVERVRRHLSENAICSVCNGAEETILHVLRDCPAMEPIWRRLLPLRRHH 978
Query: 1391 FFTASSFLDWIASVPRHDVAGLL-----VHAWFIWCRRRRFVFAGELEPWR----LILHR 1441
F + S L+W+ + V G+ + W+ W R VF GE + R I
Sbjct: 979 EFFSQSLLEWLFT-NMDPVKGIWPTLFGMGIWWAWKWRCCDVF-GERKICRDRLKFIKDM 1036
Query: 1442 AADVLRC---------------------------LRATSPSMPAADPGRAEVGGVVRDAV 1474
A +V R ++ T+ + G A GG +R+
Sbjct: 1037 AEEVRRVHVGAVGNRPNGVRVERMIRWQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQ 1096
Query: 1475 GCWLFGYSGFIGMAEILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLLRHGVTRFH 1534
G WL G++ IG AEL +GL WD+GFR++ + D +V+ L GV+ H
Sbjct: 1097 GEWLGGFALNIGSCAAPLAELWGAYYGLLIAWDKGFRRVELDLDCKLVVGFLSTGVSNAH 1156
Query: 1535 VYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
+ +V + RDW V + H+ RE N +AD
Sbjct: 1157 PLSFLVRLCQGFFTRDWLVRVSHVYREANRLAD 1189
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 293 bits (750), Expect = 5e-79
Identities = 241/835 (28%), Positives = 369/835 (43%), Gaps = 83/835 (9%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQN--VVFKLDL 861
K+M NRL++ L LI Q++F+ GR DN +I +++H ++S KC + K D+
Sbjct: 519 KLMANRLKKILPSLISETQAAFVKGRLISDNILIAHELLHALSSNN-KCSEEFIAIKTDI 577
Query: 862 EKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GD 921
KA V W FL + GF + LIM CV S +L NG I PSRGLR GD
Sbjct: 578 SKAYDRVEWPFLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGD 637
Query: 922 PLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVR 981
PLSPYLFV+C E L +Q A +NQ ++++R PISHL FADD + K +
Sbjct: 638 PLSPYLFVICTEMLVKMLQSAEQKNQITGLKVARGAPPISHLLFADDSMFYCKVNDEALG 697
Query: 982 KVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFP-IF 1040
++ I+ ++ +ASG VN KS + + ++ + I YLG P F
Sbjct: 698 QIIRIIEEYSLASGQRVNYLKSSIYFGKHISEERRCLVKRKLGIEREGGEGVYLGLPESF 757
Query: 1041 QGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCD 1100
QG + ++ DR+ +K W+ L+ GK IL K V A+P Y M F +C
Sbjct: 758 QGSKVAT-LSYLKDRLGKKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQ 816
Query: 1101 HLDSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNS 1159
++S+ F W ++ E RG++ AW ++ PK GGLG ++ N ++LGK +W ++
Sbjct: 817 QIESVMAEFWWKNKKEGRGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITE 876
Query: 1160 HDKLWV*VLLDLYCTNSDFLHAP-RRRGSCVWNAVMKTRDIIQDGFTWRIGRGD-LSFWF 1217
D L V Y + SD L+AP R S W ++ + + +I+ G IG G+ ++ W
Sbjct: 877 KDSLMAKVFKSRYFSKSDPLNAPLGSRPSFAWKSIYEAQVLIKQGIRAVIGNGETINVWT 936
Query: 1218 SKCHGSKALCEKVPYVHISDTALRVRDVFQDGVCDIRGLSTPITPE----LWALLQNLFV 1273
G+K + R V Q I + + P+ W L+ LF
Sbjct: 937 DPWIGAKP-------AKAAQAVKRSHLVSQYAANSIHVVKDLLLPDGRDWNWNLVSLLFP 989
Query: 1274 YVSLDD-----------EDLVVWAPSLEGKYSVQSCYRWLL--------DMEGGTAVSVD 1314
+ ++ D W S G YSV+S Y W++ + + S+D
Sbjct: 990 DNTQENILALRPGGKETRDRFTWEYSRSGHYSVKSGY-WVMTEIINQRNNPQEVLQPSLD 1048
Query: 1315 --WA*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRD 1372
+ +WK VP K+ F W+ + L R +A + C C H ET H L
Sbjct: 1049 PIFQQIWKLDVPPKIHHFLWRCVNNCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFK 1108
Query: 1373 CDASLQVWQNLCVSLTPGFFTASSF---LDWIASVPRHDVAGLLVHA------WFIWCRR 1423
C + W + PG A S + + SV + HA W +W R
Sbjct: 1109 CPFARLTWAISPLPAPPGGEWAESLFRNMHHVLSVHKSQPEESDHHALIPWILWRLWKNR 1168
Query: 1424 RRFVFAG-ELEPWRLILHRAADV------------------LRCLRATSPSMP------- 1457
VF G E ++IL D+ RC++ PS
Sbjct: 1169 NDLVFKGREFTAPQVILKATEDMDAWNNRKEPQPQVTSSTRDRCVKWQPPSHGWVKCNTD 1228
Query: 1458 ---AADPGRAEVGGVVRDAVG--CWLFGYSGFIGMAEILKAELLAVLFGLQSCWDRGFRQ 1512
+ D G VG V+R+ G WL G +L+ E+ A+ + + S +R+
Sbjct: 1229 GAWSKDLGNCGVGWVLRNHTGRLLWL-GLRALPSQQSVLETEVEALRWAVLSLSRFNYRR 1287
Query: 1513 LVVSSDSMVVLSLLRHGVTRFHVYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
++ SDS ++SL+++ + A + IR LLR V+ + REGN VAD
Sbjct: 1288 VIFESDSQYLVSLIQNEMD-IPSLAPRIQDIRNLLRHFEEVKFQFTRREGNNVAD 1341
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 290 bits (743), Expect = 3e-78
Identities = 236/815 (28%), Positives = 374/815 (44%), Gaps = 69/815 (8%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMN-SRRGKCQNVVFKLDLE 862
K++ RL+ +L ELI QS+F+PGR DN +I +I+H + S K ++ K D+
Sbjct: 460 KILTRRLQPWLSELISLHQSAFVPGRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDMS 519
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDP 922
KA + W FL++ L GF + +M CV + S S L NG + PSRGLR GDP
Sbjct: 520 KAYDRIKWNFLQEVLMRLGFHDKWIRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDP 579
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
LSPYLF+LC E L+ + A ++ +R++R ++HL FADD + K +
Sbjct: 580 LSPYLFILCTEVLSGLCRKAQEKGVMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCGA 639
Query: 983 VQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQG 1042
+ NIL + +ASG ++N+AKS S P+ K + + I + +YLG P G
Sbjct: 640 LSNILKKYELASGQSINLAKSAITFSSKTPQDIKRRVKLSLRIDNEGGIGKYLGLPEHFG 699
Query: 1043 RQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHL 1102
R+ ++ F ++DR+ ++ SW R L+ AGK IL K VL+++P Y M F ++C +
Sbjct: 700 RRKRDIFSSIVDRIRQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPASLCKQI 759
Query: 1103 DSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHD 1161
S+ F W S+ ++R M V+W +TLP GGLG R+ KL W ++
Sbjct: 760 QSVLTRFWWDSKPDKRKMAWVSWDKLTLPINEGGLGFREIE-------AKLSWRILKEPH 812
Query: 1162 KLWV*VLLDLYCTNSDFL--HAPRRRGSCVWNAVMKTRDIIQDGFTWRIGRGD-LSFWFS 1218
L VLL YC S F+ A S W ++ RD+++ G W IG+GD ++ W
Sbjct: 813 SLLSRVLLGKYCNTSSFMDCSASPSFASHGWRGILAGRDLLRKGLGWSIGQGDSINVWTE 872
Query: 1219 KCHGSKALCEKVPYVHISDTALRVRDV-FQDGVC-DIRGLSTPI----TPELWALLQNLF 1272
L P I +D+ D +C D++ + P+ ++ +
Sbjct: 873 AW-----LSPSSPQTPIGPPTETNKDLSVHDLICHDVKSWNVEAIRKHLPQYEDQIRKIT 927
Query: 1273 VYVSLDDEDLVVWAPSLEGKYSVQSCYRWLLDMEGGTAVSVDW---A*LWKFQVPEKMKL 1329
+ +L +D +VW P G+Y+ ++ Y L + A +D+ +WK K+K
Sbjct: 928 IN-ALPLQDSLVWLPVKSGEYTTKTGYA-LAKLNSFPASQLDFNWQKNIWKIHTSPKVKH 985
Query: 1330 FAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTP 1389
F W+A +LP + R + C C E+ +H + C + +VW+ V P
Sbjct: 986 FLWKAMKGALPVGEALSRRNIEAEVTCKRC-GQTESSLHLMLLCPYAKKVWELAPVLFNP 1044
Query: 1390 GFFTASSF------LDWIASVPRHDVAGLLVHAWF---IWCRRRRFVFAG---------- 1430
T SS + ++P + ++ W +W R R +F
Sbjct: 1045 SEATHSSVALLLVDAKRMVALPPTGLGSAPLYPWLLWHLWKARNRLIFDNHSCSEEGLVL 1104
Query: 1431 ----ELEPW---RLILHRAADV------LRCLRATSPSMPAA--DPGRAEVGGVVRDAVG 1475
+ W +L++H + + L+ TS + AA G +G ++D
Sbjct: 1105 KAILDARAWMEAQLLIHHPSPISDYPSPTPNLKVTSCFVDAAWTTSGYCGMGWFLQDPYK 1164
Query: 1476 CWL---FGYSGFIGMAEILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLLRHGVTR 1532
+ S F+G A L AE LAV L G RQL V SD ++SLL G +
Sbjct: 1165 VKIKENQSSSSFVGSA--LMAETLAVHLALVDALSTGVRQLNVFSDCKELISLLNSGKSI 1222
Query: 1533 FHVYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
+ ++ IREL + + R NVVAD
Sbjct: 1223 VEL-RGLLHDIRELSVSFTHLCFFFIPRLSNVVAD 1256
>At2g07650 putative non-LTR retrolelement reverse transcriptase
Length = 732
Score = 282 bits (721), Expect = 1e-75
Identities = 181/594 (30%), Positives = 300/594 (50%), Gaps = 44/594 (7%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLE 862
TK +V RL++ + +LI P Q+SFI GR DN +I+Q+ VH M ++G+ ++ KLDLE
Sbjct: 79 TKTLVMRLKRVISKLIRPAQASFISGRLAADNIVIMQEAVHSMRRKKGRKGWMLLKLDLE 138
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDP 922
KA + W FL DTL P + IM CV+ S+SLLWN
Sbjct: 139 KAYDRIRWEFLEDTLRAVRLPEKWIVWIMQCVTEPSMSLLWN------------------ 180
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
+ ++ + + + +S+ G +SH+ FADD++L A+A V+Q+R
Sbjct: 181 -----------------EHSIARKDWKPISLSQGGPKLSHICFADDLILFAEASVAQIRV 223
Query: 983 VQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQG 1042
++ +L FC+ASG V++ KS+ F S +V R I + I T L +YLG P+ Q
Sbjct: 224 IRRVLERFCVASGQKVSLEKSKIFFSENVSRDLGKLISDESGISSTRELGKYLGMPVLQR 283
Query: 1043 RQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHL 1102
R ++ F +L+++ + A WKGR L+ AG+ L K VL++IPV+ M + D L
Sbjct: 284 RINKDTFGDILEKLTTRLAGWKGRFLSLAGRVTLTKAVLSSIPVHTMSTIALPKSTLDGL 343
Query: 1103 DSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHD 1161
D ++++F+W S +R ++++W+ V P+ GGLGIRKA+ +N ++L K+ W L+ +
Sbjct: 344 DKVSRSFLWGSSVTQRKQHLISWKRVCKPRSEGGLGIRKAQDMNKALLSKVGWRLIQDYH 403
Query: 1162 KLWV*VLLDLY---CTNSDFLHAPRRRGSCVWNAV-MKTRDIIQDGFTWRIGRG-DLSFW 1216
LW ++ Y R S W +V + R+++ G +W IG G ++ FW
Sbjct: 404 SLWARIMRCNYRVQDVRDGAWTKVRSVCSSTWRSVALGMREVVIPGLSWVIGDGREILFW 463
Query: 1217 FSKCHGSKALCE-KVPYVHISDTALRVRDVFQDGV-CDIRGLSTPITPELWALLQNLFVY 1274
K + L E V +R RD+ ++G+ D+ ++ I+ LQ+ +
Sbjct: 464 MDKWLTNIPLAELLVQEPPAGWKGMRARDLRRNGIGWDMATIAPYISSYNRLQLQSFVLD 523
Query: 1275 VSLDDEDLVVWAPSLEGKYSVQSCYRWLL-DMEGGTAVSVDWA*LWKFQVPEKMKLFAWQ 1333
D + W S +G++ V + Y +L D + + +W PE+++LF W
Sbjct: 524 DITGARDRISWGESQDGRFKVSTTYSFLTRDEAPRQDMKRFFDRVWSVTAPERVRLFLWL 583
Query: 1334 AAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSL 1387
A +++ T H R ++ C +C E+ +H LRDC A +W L L
Sbjct: 584 VAQQAIMTNQERHRRHLSTTTICQVCKGAKESIIHVLRDCPAMEGIWLRLISDL 637
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 280 bits (717), Expect = 4e-75
Identities = 182/547 (33%), Positives = 289/547 (52%), Gaps = 41/547 (7%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLDLE 862
TK+MV RL+ + +LIGP Q+SFIPGR + DN +++Q+ VH M ++G+ ++ KLDLE
Sbjct: 334 TKMMVIRLKNVISKLIGPAQASFIPGRLSFDNIVVVQEAVHSMRRKKGRKGWMLLKLDLE 393
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDP 922
KA + W FL +TL G + IM CV+ +SLLWNG K P RGLR GDP
Sbjct: 394 KAYDRIRWDFLAETLEAAGLSEGWIKRIMECVAGPEMSLLWNGEKTDSFTPERGLRQGDP 453
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
+SPYLFVLC+ERL I+ AV + + + IS+ G +SH+ FADD++L A+A V+Q
Sbjct: 454 ISPYLFVLCIERLCHQIETAVGRGDWKSISISQGGPKVSHVCFADDLILFAEASVAQ--- 510
Query: 983 VQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQG 1042
V++ KS+ F S +V R + I T I T L +YLG P+ Q
Sbjct: 511 --------------KVSLEKSKIFFSNNVSRDLEGLITAETGIGSTRELGKYLGMPVLQK 556
Query: 1043 RQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHL 1102
R ++ F VL+RV+ + + WK R L+ AG+ L K VL +IP++ M ++ + L
Sbjct: 557 RINKDTFGEVLERVSSRLSGWKSRSLSLAGRITLTKAVLMSIPIHTMSSILLPASLLEQL 616
Query: 1103 DSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHD 1161
D +++NF+W S E+R ++++W+ V PK GGLG+R ++ +N ++L K+ W L+N
Sbjct: 617 DKVSRNFLWGSTVEKRKQHLLSWKKVCRPKAAGGLGLRASKDMNRALLAKVGWRLLNDKV 676
Query: 1162 KLWV*VLLDLY----CTNSDFLHAPRRRGSCVWNAV-MKTRDIIQDGFTWRIGRGDLSFW 1216
LW VL Y +S +L P+ S W ++ + R+ + G W +
Sbjct: 677 SLWARVLRRKYKVTDVHDSSWL-VPKATWSSTWRSIGVGLREGVAKG--WILHE------ 727
Query: 1217 FSKCHGSKALCEKVPYVHISDTALRVRDVFQDGV-CDIRGLSTPITPELWALLQNLFVYV 1275
C ++A C P + RV + + +GV D+ L + + L + +
Sbjct: 728 -PLC--TRATCLLSP----EELNARVEEFWTEGVGWDMVKLGQCLPRSVTDRLHAVVIKG 780
Query: 1276 SLDDEDLVVWAPSLEGKYSVQSCYRWLL-DMEGGTAVSVDWA*LWKFQVPEKMKLFAWQA 1334
L D + W + +G ++V S Y L + E + + +W PE++++F W
Sbjct: 781 VLGLRDRISWQGTSDGDFTVGSAYVLLTQEEESKPCMESFFKRIWGVIAPERVRVFLWLV 840
Query: 1335 AHRSLPT 1341
+ + T
Sbjct: 841 GQQVIMT 847
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 274 bits (700), Expect = 3e-73
Identities = 224/818 (27%), Positives = 362/818 (43%), Gaps = 81/818 (9%)
Query: 785 PSGSSGQSAYAMCYI--S*FTKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIV 842
P+ S A+C + +K +VNRL+ L ++ Q++FIPGR DN +I +++
Sbjct: 880 PTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVM 939
Query: 843 HHMNSRRGKCQN-VVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSL 901
H + R+ + + K D+ KA V W FL T+ FGF + IM V S S+
Sbjct: 940 HSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHYSV 999
Query: 902 LWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPIS 961
L NG I P+RG+R GDPLSPYLF+LC + L+ I VRI I+
Sbjct: 1000 LINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPAIT 1059
Query: 962 HLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFV 1021
HL FADD L +A V + ++++ + SG +N+ KS V + ++ +
Sbjct: 1060 HLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSRLKQ 1119
Query: 1022 VTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVL 1081
+ IP +YLG P GR+ +E F++++DRV ++ ++W R L+ AGK I+ K V
Sbjct: 1120 ILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVA 1179
Query: 1082 NAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSR-GERRGMNMVAWRTVTLPKCNGGLGIRK 1140
A+PVY M F + ++SL NF W + +RG+ VAW+ + K GGLG R
Sbjct: 1180 LAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGGLGFRD 1239
Query: 1141 ARLINTSMLGKLVWDLVNSHDKLWV*VLLDLYCTNSDFLHAP-RRRGSCVWNAVMK---- 1195
N ++L K W L+ + L+ V+ Y + L A R++ S W +++
Sbjct: 1240 LAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLLDGIAL 1299
Query: 1196 ----TRDIIQDGFTWRIGRGDLSFWFSKCHGSKALCEKVPYVHISDTALRVRDVFQDG-- 1249
TR +I DG RIG ++ H + L + Y ++ + ++F+
Sbjct: 1300 LKKGTRHLIGDGQNIRIGLDNI----VDSHPPRPLNTEETYKEMT-----INNLFERKGS 1350
Query: 1250 --VCDIRGLSTPITPELWALLQNLFVYVSLDDEDLVVWAPSLEGKYSVQSCYRWLLDMEG 1307
D +S + + +++ S D ++W + G+Y+V+S Y WLL +
Sbjct: 1351 YYFWDDSKISQFVDQSDHGFIHRIYLAKS-KKPDKIIWNYNTTGEYTVRSGY-WLLTHDP 1408
Query: 1308 GTAV--------SVDW-A*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSL 1358
T + S+D +W + K+K F W+A ++L T + RG+ I C
Sbjct: 1409 STNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCPR 1468
Query: 1359 CHHHVET*MHCLRDCDASLQVWQNLCVSLTPGFFTASSF-------LDWIASVPRHDVAG 1411
CH E+ H L C + W+ SL ++ F L+++ D
Sbjct: 1469 CHRENESINHALFTCPFATMAWRLSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHK 1528
Query: 1412 LLV--HAWFIWCRRRRFVFAGELE-PWRLILHRAADVLRCLRATSPSMPAADPGRAEVGG 1468
LL W IW R VF E P + +L A+ L AT P R
Sbjct: 1529 LLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAEN 1588
Query: 1469 VVR----------------------DAVGCWL----------FGYSGFIGMAEILKAELL 1496
+ +A G W+ +G + L+AE
Sbjct: 1589 KIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLEAETK 1648
Query: 1497 AVLFGLQSCWDRGFRQLVVSSDSMVVLSLLRHGVTRFH 1534
A+L LQ W RG+ Q+ + D +++L+ +G++ FH
Sbjct: 1649 ALLAALQQTWIRGYTQVFMEGDCQTLINLI-NGIS-FH 1684
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 274 bits (700), Expect = 3e-73
Identities = 201/657 (30%), Positives = 306/657 (45%), Gaps = 23/657 (3%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRG-KCQNVVFKLDL 861
+K++ RL+ L LI QS+F+ GR DN +I Q++ H + + K + + K D+
Sbjct: 302 SKILCQRLKTVLPNLISETQSAFVDGRLISDNILIAQEMFHGLRTNSSCKDKFMAIKTDM 361
Query: 862 EKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GD 921
KA V W F+ L GF +S IM+C+++ +L NG I P RGLR GD
Sbjct: 362 SKAYDQVEWNFIEALLRKMGFCEKWISWIMWCITTVQYKVLINGQPKGLIIPERGLRQGD 421
Query: 922 PLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVR 981
PLSPYLF+LC E L +I+ A QN ++++ +SHL FADD L KA Q
Sbjct: 422 PLSPYLFILCTEVLIANIRKAERQNLITGIKVATPSPAVSHLLFADDSLFFCKANKEQCG 481
Query: 982 KVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQ 1041
+ IL + SG +N +KS V + KADI ++ I + YLG P
Sbjct: 482 IILEILKQYESVSGQQINFSKSSIQFGHKVEDSIKADIKLILGIHNLGGMGSYLGLPESL 541
Query: 1042 GRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDH 1101
G + F FV DR+ + W + L+K GK ++ K V +P Y M F A+
Sbjct: 542 GGSKTKVFSFVRDRLQSRINGWSAKFLSKGGKEVMIKSVAATLPRYVMSCFRLPKAITSK 601
Query: 1102 LDSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSH 1160
L S F W S G+ RGM+ +AW + K +GGLG R N+++L K +W L+ +
Sbjct: 602 LTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDGGLGFRNVDDFNSALLAKQLWRLITAP 661
Query: 1161 DKLWV*VLLDLYCTNSDFLHAPRRRG-SCVWNAVMKTRDIIQDGFTWRIGRG-DLSFWFS 1218
D L+ V Y S+ L + + S W +++ R ++ G R+G G +S W
Sbjct: 662 DSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSMISARSLVYKGLIKRVGSGASISVWND 721
Query: 1219 KCHGSKALCEKVPYVHISDTALRVRDVF--QDGVCDIRGLSTPITPELWALLQNLFVYVS 1276
++ I D +L+V+ + + +I L PE L+ L + +
Sbjct: 722 PWIPAQFPRPAKYGGSIVDPSLKVKSLIDSRSNFWNIDLLKELFDPEDVPLISALPI-GN 780
Query: 1277 LDDEDLVVWAPSLEGKYSVQSCYRWL-LDM-EGGTAVSVDW----A*LWKFQVPEKMKLF 1330
+ ED + W + G Y+V+S Y LD+ EG T + D A +WK Q P K++ F
Sbjct: 781 PNMEDTLGWHFTKAGNYTVKSGYHTARLDLNEGTTLIGPDLTTLKAYIWKVQCPPKLRHF 840
Query: 1331 AWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTPG 1390
WQ +P + RG+ + C C E+ H L C + Q+W + PG
Sbjct: 841 LWQILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPARQIWALSQIPTAPG 900
Query: 1391 FFTASSF------LDWIASVPRH-DVAGLLVHAWFIWCRRRRFVFAG-ELEPWRLIL 1439
F ++S L W +P D A W+IW R VF + +P ++L
Sbjct: 901 IFPSNSIFTNLDHLFW--RIPSGVDSAPYPWIIWYIWKARNEKVFENVDKDPMEILL 955
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 273 bits (698), Expect = 6e-73
Identities = 224/818 (27%), Positives = 361/818 (43%), Gaps = 81/818 (9%)
Query: 785 PSGSSGQSAYAMCYI--S*FTKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIV 842
P+ S A+C + +K +VNRL+ L ++ Q++FIPGR DN +I +++
Sbjct: 654 PTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVM 713
Query: 843 HHMNSRRGKCQN-VVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSL 901
H + R+ + + K D+ KA V W FL T+ FGF + IM V S S+
Sbjct: 714 HSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHYSV 773
Query: 902 LWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPIS 961
L NG I P+RG+R GDPLSPYLF+LC + L+ I VRI I+
Sbjct: 774 LINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPAIT 833
Query: 962 HLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFV 1021
HL FADD L +A V + ++++ + SG +N+ KS V + ++ +
Sbjct: 834 HLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSKLKQ 893
Query: 1022 VTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVL 1081
+ IP +YLG P GR+ +E F++++DRV ++ ++W R L+ AGK I+ K V
Sbjct: 894 ILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVA 953
Query: 1082 NAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSR-GERRGMNMVAWRTVTLPKCNGGLGIRK 1140
A+PVY M F + ++SL NF W + +RG+ VAW+ + K GGLG R
Sbjct: 954 LAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGGLGFRD 1013
Query: 1141 ARLINTSMLGKLVWDLVNSHDKLWV*VLLDLYCTNSDFLHAP-RRRGSCVWNAVMK---- 1195
N ++L K W L+ + L+ V+ Y + L A R++ S W +++
Sbjct: 1014 LAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLLDGIAL 1073
Query: 1196 ----TRDIIQDGFTWRIGRGDLSFWFSKCHGSKALCEKVPYVHISDTALRVRDVFQDG-- 1249
TR +I DG RIG ++ H + L + Y ++ + ++F+
Sbjct: 1074 LKKGTRHLIGDGQNIRIGLDNI----VDSHPPRPLNTEETYKEMT-----INNLFERKGS 1124
Query: 1250 --VCDIRGLSTPITPELWALLQNLFVYVSLDDEDLVVWAPSLEGKYSVQSCYRWLLDMEG 1307
D +S + + +++ S D ++W + G+Y+V+S Y WLL +
Sbjct: 1125 YYFWDDSKISQFVDQSDHGFIHRIYLAKS-KKPDKIIWNYNTTGEYTVRSGY-WLLTHDP 1182
Query: 1308 GTAV--------SVDW-A*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSL 1358
T + S+D +W + K+K F W+A ++L T + RG+ I C
Sbjct: 1183 STNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPICPR 1242
Query: 1359 CHHHVET*MHCLRDCDASLQVWQNLCVSLTPGFFTASSF-------LDWIASVPRHDVAG 1411
CH E+ H L C + W SL ++ F L+++ D
Sbjct: 1243 CHRENESINHALFTCPFATMAWWLSDSSLIRNQLMSNDFEENISNILNFVQDTTMSDFHK 1302
Query: 1412 LLV--HAWFIWCRRRRFVFAGELE-PWRLILHRAADVLRCLRATSPSMPAADPGRAEVGG 1468
LL W IW R VF E P + +L A+ L AT P R
Sbjct: 1303 LLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAEN 1362
Query: 1469 VVR----------------------DAVGCWL----------FGYSGFIGMAEILKAELL 1496
+ +A G W+ +G + L+AE
Sbjct: 1363 KIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLEAETK 1422
Query: 1497 AVLFGLQSCWDRGFRQLVVSSDSMVVLSLLRHGVTRFH 1534
A+L LQ W RG+ Q+ + D +++L+ +G++ FH
Sbjct: 1423 ALLAALQQTWIRGYTQVFMEGDCQTLINLI-NGIS-FH 1458
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 272 bits (695), Expect = 1e-72
Identities = 187/594 (31%), Positives = 289/594 (48%), Gaps = 27/594 (4%)
Query: 804 KVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQ-NVVFKLDLE 862
K+M R++ L +LI QS+F+PGR DN +I +++H + + K ++ K D+
Sbjct: 414 KIMTKRMQLILPKLISENQSAFVPGRVISDNVLITHEVLHFLRTSSAKKHCSMAVKTDMS 473
Query: 863 KACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDP 922
KA V W FL+ L FGF + ++ CV+S S S L NG + P+RGLR GDP
Sbjct: 474 KAYDRVEWDFLKKVLQRFGFHSIWIDWVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDP 533
Query: 923 LSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRK 982
LSP LF+LC E L+ A Q VR+S +G ++HL FADD + +K+ K
Sbjct: 534 LSPCLFILCTEVLSGLCTRAQRLRQLPGVRVSINGPRVNHLLFADDTMFFSKSDPESCNK 593
Query: 983 VQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQG 1042
+ IL+ + ASG ++N KS S PR+ K + + I +YLG P G
Sbjct: 594 LSEILSRYGKASGQSINFHKSSVTFSSKTPRSVKGQVKRILKIRKEGGTGKYLGLPEHFG 653
Query: 1043 RQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHL 1102
R+ ++ F ++D++ +K SW R L++AGK ++ K VL ++P+Y M F A+C +
Sbjct: 654 RRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQVMLKAVLASMPLYSMSCFKLPSALCRKI 713
Query: 1103 DSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHD 1161
SL F W ++ + R + VAW +T PK GGLG R N S+L KL W L+NS +
Sbjct: 714 QSLLTRFWWDTKPDVRKTSWVAWSKLTNPKNAGGLGFRDIERCNDSLLAKLGWRLLNSPE 773
Query: 1162 KLWV*VLLDLYCTNSDFLHAP-RRRGSCVWNAVMKTRDIIQDGFTWRIGRGD-LSFWFSK 1219
L +LL YC +S F+ + S W +++ R+I+++G W I G+ +S W
Sbjct: 774 SLLSRILLGKYCHSSSFMECKLPSQPSHGWRSIIAGREILKEGLGWLITNGEKVSIW--- 830
Query: 1220 CHGSKALCEKVPYVHISDTALRVRDVFQDGVCDIRGLS------TPITPELWALLQNLFV 1273
L P V I +D+ + + L I P L++ L
Sbjct: 831 --NDPWLSISKPLVPIGPALREHQDLRVSALINQNTLQWDWNKIAVILPNYENLIKQL-P 887
Query: 1274 YVSLDDEDLVVWAPSLEGKYSVQSCYRWLLDMEGGTAVSV-----DW-A*LWKFQVPEKM 1327
S D + W P G+Y+ +S Y + ++ + +W + LWK Q K+
Sbjct: 888 APSSRGVDKLAWLPVKSGQYTSRSGY----GIASVASIPIPQTQFNWQSNLWKLQTLPKI 943
Query: 1328 KLFAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQ 1381
K W+AA +LP R ++ C C T H C+ + QVW+
Sbjct: 944 KHLMWKAAMEALPVGIQLVRRHISPSAACHRCGAPEST-THLFFHCEFAAQVWE 996
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 266 bits (681), Expect = 5e-71
Identities = 197/741 (26%), Positives = 322/741 (42%), Gaps = 67/741 (9%)
Query: 725 WALIRHRVLMAFNRFSSSSIGILWGMMFGVWFVMLLSRVLLTLSWPRL*FVLSLRLTSPL 784
W L++H++L F + + L W L ++TSP
Sbjct: 438 WDLVKHQILTEIFGFFETGV--------------------LPQDWNHTHICLIPKITSPQ 477
Query: 785 PSGSSGQSAYAMCYIS*FTKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHH 844
+ +K++ RL++ L ++ QS+F+P R DN ++ +++H
Sbjct: 478 RMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLISDNILVAHEMIHS 537
Query: 845 M--NSRRGKCQNVVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLL 902
+ N R K +++ FK D+ KA V W FL + GF +S IM CV+S S S+L
Sbjct: 538 LRTNDRISK-EHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIMNCVTSVSYSVL 596
Query: 903 WNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISH 962
NG I P+RG+R GDPLSP LFVLC E L + A + ++ ++H
Sbjct: 597 INGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKITGIQFQDKKVSVNH 656
Query: 963 LFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVV 1022
L FADD LL+ KA + ++ L+ + SG +N+ KS +V K I
Sbjct: 657 LLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNVDIQIKDWIKSR 716
Query: 1023 TSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLN 1082
+ I +YLG P ++ F F+ +++ + W + L++ GK +L K +
Sbjct: 717 SGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQGGKEVLLKSIAL 776
Query: 1083 AIPVYPMQLFWFH*AVCDHLDSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKA 1141
A+PVY M F +C L ++ +F W S ++R ++ ++W+ +TLPK GG G +
Sbjct: 777 ALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGGFGFKDL 836
Query: 1142 RLINTSMLGKLVWDLVNSHDKLWV*VLLDLYCTNSDFLHAPR-RRGSCVWNAVMKTRDII 1200
+ N ++L K W ++ L+ V Y +NSDFL A R R S W +++ R+++
Sbjct: 837 QCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSILFGRELL 896
Query: 1201 QDGFTWRIGRGDLSF-WFSKCHGSKALCEKVPYVHISDTALRVRDVFQDGVCDIRGLSTP 1259
G IG G +F W K + + + L+V +
Sbjct: 897 MQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLNRRRFINVDLKVSQLIDP----------- 945
Query: 1260 ITPELWAL--LQNLFVYVSLD----------DEDLVVWAPSLEGKYSVQSCYRWL----- 1302
T W L L++LF + ++ ED W S G YSV++ Y +L
Sbjct: 946 -TSRNWNLNMLRDLFPWKDVEIILKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQVH 1004
Query: 1303 --LDMEGGTAVSVD--WA*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSL 1358
L E SV+ + +W K+++F W+A H ++P D RG+ C +
Sbjct: 1005 HRLYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLM 1064
Query: 1359 CHHHVET*MHCLRDCDASLQVWQNLCVSLTPGFFTASSFLDWIASVPRHDVAGLLVH--- 1415
C ET H L +C + QVW +S F+ S + + + L H
Sbjct: 1065 CDTENETINHILFECPLARQVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRF 1124
Query: 1416 -----AWFIWCRRRRFVFAGE 1431
WF+W R +F G+
Sbjct: 1125 VSPWILWFLWKNRNALLFEGK 1145
>At3g31420 hypothetical protein
Length = 1491
Score = 259 bits (662), Expect = 9e-69
Identities = 212/805 (26%), Positives = 348/805 (42%), Gaps = 133/805 (16%)
Query: 795 AMCYIS*--FTKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKC 852
A+C +S +K++VNRL++ L I Q++F+PGR +NAII ++ + + +R+ +
Sbjct: 693 ALCNVSYKIISKILVNRLKKHLGGAITENQAAFVPGRLITNNAIIAHEVYYALKARKRQA 752
Query: 853 QN-VVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPI 911
+ + K D+ KA + W FL +T+ GF + IM CV+ S+L NG I
Sbjct: 753 NSYMALKTDITKAYDRLEWDFLEETMRQMGFNTKWIERIMICVTMVRFSVLINGSPHGTI 812
Query: 912 APSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLL 971
P RG+R GDPLSPYLF+LC E L+ I+ A + + +R+S G ISHL FADD +
Sbjct: 813 KPERGIRHGDPLSPYLFILCAEVLSHMIKQAEINKKLKGIRLSTQGPFISHLLFADDSIF 872
Query: 972 LAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANL 1031
+A R +++ P K+ ++ I
Sbjct: 873 FT--------------------------LANQRSCTAIKEPDTKRRMRHLL-GIHNEGGE 905
Query: 1032 ERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQL 1091
+YLG P ++ +E F++++++V K W + L++ GK +L K V A+PVY M +
Sbjct: 906 GKYLGLPEQFNKKKKELFNYIIEKVKDKTQGWSKKFLSQGGKEVLLKAVALAMPVYSMNI 965
Query: 1092 FWFH*AVCDHLDSLAKNFIWSRG-ERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLG 1150
F VC+ +DSL F WS G E +GM+ W+ +++PK GGLG ++ N ++LG
Sbjct: 966 FKLTKEVCEEIDSLLARFWWSSGNETKGMHWFTWKRMSIPKKEGGLGFKELENFNLALLG 1025
Query: 1151 KLVWDLVNSHDKLWV*VLLDLYCTNSDFLHAPR-RRGSCVWNAVMKTRDIIQDGFTWRIG 1209
K W L+ + L VL Y ++ ++A + RR S VW +++ R++++ G +G
Sbjct: 1026 KQTWHLLQHPNCLMARVLRGRYFPETNVMNAVQGRRASFVWKSILHGRNLLKKGLRCCVG 1085
Query: 1210 RGDL-SFWFSKCHGSKALCEKVPYVHISDTALRVRDVFQDGVCDIRGLSTPITPELWALL 1268
G L + W P++ L +P P
Sbjct: 1086 DGSLINAWLD------------PWL---------------------PLHSPRAPYKQEDA 1112
Query: 1269 QNLFVYVSLDDEDLVVWAPSLEGKYSVQSCYRWLLDMEGGTA--------VSVDWA*LWK 1320
+ S +DL+ W + +G Y+V+S Y WL GT + + LWK
Sbjct: 1113 PEQLLVCSTARDDLIGWHYTKDGMYTVKSAY-WLATHLPGTTGTHPPPGDIKLKQL-LWK 1170
Query: 1321 FQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVW 1380
+ K+K F W+ ++ T + R + C C ET H +CD + W
Sbjct: 1171 TKTAPKIKHFCWKILSGAIATGEMLRYRHINKQSICKRCCRDEETSQHLFFECDYAKATW 1230
Query: 1381 ----------QNLCVSLTPGFFTASSFLDWIASVPRHDVAGLLVHAWFIWCRRRRFVFAG 1430
Q+ V+L F +F + R +L W +W R F
Sbjct: 1231 RGAGLPNLIFQDSIVTLEEKFRAMFTFNPSTTNYWRQLPFWIL---WRLWRSRNILTFQQ 1287
Query: 1431 ELEPWRLILHRA-------------ADVLRCLRATSPSMPAADPG--------------- 1462
+ PW + + A V+ L + AA+
Sbjct: 1288 KHIPWEVTVQLAKQDALEWQDIEDRVQVINPLSRRHSNRYAANRWTRPVCGWKKCNYDGS 1347
Query: 1463 -----RAEVGGVVRDAVGCWLFGYSGFIGMAE------ILKAELLAVLFGLQSCWDRGFR 1511
++ G V+RD G ++ G G A+ L++ L A++ +QSCW G
Sbjct: 1348 YSTIINSKAGWVIRDEHGQFIGG-----GQAKGKHTNNALESALQALIIAMQSCWSHGHT 1402
Query: 1512 QLVVSSDSMVVLSLLRHGVTRFHVY 1536
++ D++ V +L G RF VY
Sbjct: 1403 KVCFEGDNIEVYQILNEGKARFDVY 1427
>At3g45550 putative protein
Length = 851
Score = 258 bits (660), Expect = 1e-68
Identities = 207/723 (28%), Positives = 327/723 (44%), Gaps = 53/723 (7%)
Query: 785 PSGSSGQSAYAMCYI--S*FTKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIV 842
P S A+C + +K MVNRL+ L ++ Q++FIPGR DN +I +I+
Sbjct: 119 PQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGRIINDNVMIAHEIM 178
Query: 843 HHMNSRRGKCQN-VVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSL 901
H + R+ + + K D+ KA V W FL T+ FGF + IM V S S+
Sbjct: 179 HSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGWIMAAVKSVHYSV 238
Query: 902 LWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPIS 961
L NG I+P+RG+R GDPLSPYLF+LC + L+ I+ VRI I+
Sbjct: 239 LINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIRGVRIGNGAPAIT 298
Query: 962 HLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFV 1021
HL FADD L +A V + ++++ + SG +N+ KS V + + +
Sbjct: 299 HLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGSTQTRLKT 358
Query: 1022 VTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVL 1081
+ +IP +YLG P GR+ +E F++++DRV + ASW + L+ AGK IL K V
Sbjct: 359 LLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEILLKSVA 418
Query: 1082 NAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSR-GERRGMNMVAWRTVTLPKCNGGLGIRK 1140
A+PVY M F + ++SL NF W + +RG+ VAW+ + K GGLG R
Sbjct: 419 LAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRLQYSKKEGGLGFRD 478
Query: 1141 ARLINTSMLGKLVWDLVNSHDKLWV*VLLDLYCTNSDFLHA-PRRRGSCVWNAVMK---- 1195
N ++L K W ++ + L+ V+ Y ++ + A R + S W++++
Sbjct: 479 LAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKDNSIIDAKTRSQQSYGWSSLLSGIAL 538
Query: 1196 ----TRDIIQDGFTWRIGRGDLSFWFSKCHGSKALCEKVPYVHISDTALRVRDVFQ---- 1247
TR +I DG T R+G ++ H + L + L + ++FQ
Sbjct: 539 LRKGTRYVIGDGKTIRLGIDNV----VDSHPPRPLLTDEQH-----NGLSLDNLFQHRGH 589
Query: 1248 DGVCDIRGLSTPITPELWALLQNLFVYVSLDDEDLVVWAPSLEGKYSVQSCYRWLLDMEG 1307
D L T + ++ +++ + D ++W+ + G Y+V+S Y WL +
Sbjct: 590 SRCWDNAKLQTFVDQSDHDYIKRIYL-STRSKTDRLIWSYNSTGDYTVRSGY-WLSTHDP 647
Query: 1308 GTAV--------SVDW-A*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSL 1358
+ SVD +W + K+K F W+ ++LPT D RG+ I C
Sbjct: 648 SNTIPTMAKPHGSVDLKTKIWNLPIMPKLKHFLWRILSKALPTTDRLTTRGMRIDPGCPR 707
Query: 1359 CHHHVET*MHCLRDCDASLQVWQNLCVSLTPGFFTA----------SSFLDWIASVPRHD 1408
C E+ H L C + W+ +S TP + ++ S+ L + + D
Sbjct: 708 CRRENESINHALFTCPFATMAWR---LSDTPLYRSSILSNNIEDNISNILLLLQNTTITD 764
Query: 1409 VAGLLVH--AWFIWCRRRRFVFAGELE-PWRLILHRAADVLRCLRATSPSMPAADPGRAE 1465
L+ W IW R VF E P ++ A+ L AT P P R
Sbjct: 765 SQKLIPFWLLWRIWKARNNVVFNNLRESPSITVVRAKAETNEWLNATQTQGPRRLPKRTT 824
Query: 1466 VGG 1468
G
Sbjct: 825 AAG 827
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 248 bits (632), Expect = 3e-65
Identities = 199/708 (28%), Positives = 312/708 (43%), Gaps = 61/708 (8%)
Query: 764 LLTLSWPRL*FVLSLRLTSPLPSGSSGQSAYAMCYIS*FTKVMVNRLRQFLQELIGPLQS 823
+L W L ++T P + +K++ RL+++L ++ P QS
Sbjct: 208 ILPEDWNHTHLCLIPKITKPARMADIRPISLCSVMYKIISKILSARLKKYLPVIVSPTQS 267
Query: 824 SFIPGRGTIDNAIILQKIVHHMNSRRGKCQN-VVFKLDLEKACGSVSWAFLRDTLCFFGF 882
+F+ R DN I+ +IVH++ + ++ +VFK D+ KA V W FL+ L GF
Sbjct: 268 AFVAERLVSDNIILAHEIVHNLRTNEKISKDFMVFKTDMSKAYDRVEWPFLKGILLALGF 327
Query: 883 PPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDA 942
T ++ +M CVSS S S+L NG I P RGLR GDPLSP+LFVLC E L + A
Sbjct: 328 NSTWINWMMACVSSVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEALIHILNQA 387
Query: 943 VDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAK 1002
+ ++ + G ++HL FADD LL+ KA + ++ + L+ + SG +N K
Sbjct: 388 EKIGKISGIQFNGTGPSVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISGQMINSEK 447
Query: 1003 SRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFP-IFQGRQTQEHFDFVLDRVNRKFA 1061
S V K I + I +YLG P FQG + Q F F+ +++ + +
Sbjct: 448 SAITFGAKVNEETKQWIMNRSGIQTEGGTGKYLGLPECFQGSK-QVLFGFIKEKLQSRLS 506
Query: 1062 SWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHLDSLAKNFIW-SRGERRGMN 1120
W + L++ GK IL K + A PVY M F +C L S+ +F W S +++ ++
Sbjct: 507 GWYAKTLSQGGKDILLKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFWWNSVQDKKKIH 566
Query: 1121 MVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHDKLWV*VLLDLYCTNSDFLH 1180
+ + + LPK GG G + + N ++L K L D L +L Y NSDFL
Sbjct: 567 WIGAQKLMLPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRYYMNSDFLS 626
Query: 1181 APR-RRGSCVWNAVMKTRDIIQDGFTWRIGRGDLSF-WFSKCHGSKALCEKVPYVHISDT 1238
A + R S W +++ R+++ G IG G+ ++ W I D
Sbjct: 627 ATKGTRPSYAWQSILYGRELLVSGLKKIIGNGENTYVWMDN--------------WIFDD 672
Query: 1239 ALRVRDVFQDGVCDIRGLSTPITP--ELWAL--LQNLFVYVSL----------DDEDLVV 1284
R + Q V +S I P W L L++LF + + +D
Sbjct: 673 KPRRPESLQIMVDIQLKVSQLIDPFSRNWNLNMLRDLFPWKEIQIICQQRPMASRQDSFC 732
Query: 1285 WAPSLEGKYSVQSCY---------RWLLDMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAA 1335
W + G Y+V+S Y + + E +++ + +W K+K+F W+
Sbjct: 733 WFGTNHGLYTVKSEYDLCSRQVHKQMFKEAEEQPSLNPLFGKIWNLNSAPKIKVFLWKVL 792
Query: 1336 HRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVSLTPGFFTAS 1395
++ D RGV I CS+C ET H L C + QVW +LTP
Sbjct: 793 KGAVAVEDRLRTRGVLIEDGCSMCPEKNETLNHILFQCPLARQVW-----ALTPMQSPNH 847
Query: 1396 SFLDWIASVPRHDVAG-----LLVH--------AWFIWCRRRRFVFAG 1430
F D I + H + L H W +W R + +F G
Sbjct: 848 GFGDSIFTNVNHVIGNCHNTELSPHLRYVSPWIIWILWKNRNKRLFEG 895
>At4g08830 putative protein
Length = 947
Score = 245 bits (626), Expect = 1e-64
Identities = 193/653 (29%), Positives = 302/653 (45%), Gaps = 83/653 (12%)
Query: 933 ERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVRKVQNILADFCM 992
ERL I AV + + + +S+ G ISH+ FADD++L A+A VSQ+R ++ IL FC+
Sbjct: 326 ERLCHMIDRAVAVKEWKSIGLSQGGPKISHICFADDLILFAEASVSQIRVIRRILETFCI 385
Query: 993 ASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQGRQTQEHFDFV 1052
ASG V++ KS+ F S +V R + I + I T L +YLG PI Q R ++ F V
Sbjct: 386 ASGQKVSLDKSKIFFSKNVSRDLEKLISKESGIKSTRELGKYLGMPILQRRINKDTFGEV 445
Query: 1053 LDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDHLDSLAKNF-IW 1111
L+RV+ + A WKGR L+ AG+ L K VL+ IP++ M + + LD LA+ F +
Sbjct: 446 LERVSSRLAGWKGRSLSFAGRLTLTKSVLSLIPIHTMSTISLPQSTLEGLDKLARVFLLG 505
Query: 1112 SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSHDKLWV*VLLDL 1171
S E++ +++VAW V LPK GGLGIR ++ +N +++ K+ W L+N LW +L
Sbjct: 506 SSAEKKKLHLVAWDRVCLPKSEGGLGIRTSKCMNKALVSKVGWRLINDRYSLWARILRSK 565
Query: 1172 YCTNSDFLHAPRRRGSCVWNAVMKTRDIIQDGFTWRIGRG-DLSFWFSKCHGSKALCEK- 1229
Y R+++ G W +G G D+ FW +AL +
Sbjct: 566 YRVG--------------------LREVVSRGSRWVVGNGRDILFWSDNWLSHEALINRA 605
Query: 1230 VPYVHISDTALRVRDVFQDGV-CDIRGLSTPITPELWALLQNLFVYVSLDDEDLVVWAPS 1288
V + S+ LRV+D++ +G+ + + I+ L + V D + W S
Sbjct: 606 VIEIPNSEKELRVKDLWANGLGWKLDKIEPYISYHTRLELAAVVVDSVTGARDRLSWGYS 665
Query: 1289 LEGKYSVQSCYRWLL-DMEGGTAVSVDWA*LWKFQVPEKMKLFAWQAAHRSLPTVDTFHD 1347
+G ++V+S YR L D + ++ + LW+ E++K F W S+
Sbjct: 666 ADGVFTVKSAYRLLTEDHDPRPNMAAFFDRLWRVVALERVKTFLWHIGDTSV-------- 717
Query: 1348 RGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLC-VSLTPGFFTASSFLDWI----- 1401
C +C ET +H L+DC + +W+ L V + FF S F W+
Sbjct: 718 --------CQVCKGGDETILHVLKDCPSIAGIWRRLVQVQRSYDFFNGSLF-GWLYVNLG 768
Query: 1402 ---ASVPRHDVAGLLVHAWFIWCRRRRFVFAGELEPWR----LILHRAADVLRC------ 1448
A + W+ W R +VF GE+ R AA+V
Sbjct: 769 MKNAETGYAWATLFAIVVWWSWKWRCGYVF-GEVGKCRDRVKFFRDLAAEVSHAHAIHSQ 827
Query: 1449 ---LRATSPSMPAADP------------------GRAEVGGVVRDAVGCWLFGYSGFIGM 1487
LR + A P G A GGV+RD +G W G++ IG+
Sbjct: 828 NGGLRTRVERLVAWKPPDGEWVKLNTDGASRGNLGLATTGGVLRDGIGHWCGGFALDIGV 887
Query: 1488 AEILKAELLAVLFGLQSCWDRGFRQLVVSSDSMVVLSLLRHGVTRFHVYAAIV 1540
AEL V +GL W+R F ++ + DS +V+ L G++ H + +V
Sbjct: 888 CSAPLAELWGVYYGLYMAWERRFTRVELEVDSELVVGFLTTGISDTHSLSFLV 940
Score = 60.8 bits (146), Expect = 6e-09
Identities = 26/58 (44%), Positives = 44/58 (75%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQNVVFKLD 860
TK+MV RL++ + +LIGP QSSFIPGR ++DN +++Q+ VH +N ++G+ + + +D
Sbjct: 276 TKMMVLRLKKVIGKLIGPSQSSFIPGRLSLDNIVVVQEAVHSLNRKKGRKERLCHMID 333
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 243 bits (621), Expect = 5e-64
Identities = 212/831 (25%), Positives = 367/831 (43%), Gaps = 94/831 (11%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSRRGKCQN--VVFKLD 860
+K+++ RL+Q L ++I Q++F+PG+ DN ++ +++H + SRR +CQ+ V K D
Sbjct: 880 SKILIKRLKQCLGDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRR-ECQSGYVAVKTD 938
Query: 861 LEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*G 920
+ KA V W FL + GF P V IM CV+S S +L NG I PSRG+R G
Sbjct: 939 ISKAYDRVEWNFLEKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQG 998
Query: 921 DPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQV 980
DPLSPYLF+ C E L+ ++ A Q ++I++D ISHL FADD L +A +
Sbjct: 999 DPLSPYLFLFCAEVLSNMLRKAEVNKQIHGMKITKDCLAISHLLFADDSLFFCRASNQNI 1058
Query: 981 RKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIF 1040
++ I + ASG +N AKS +P ++ + + I +YLG P
Sbjct: 1059 EQLALIFKKYEEASGQKINYAKSSIIFGQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQ 1118
Query: 1041 QGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCD 1100
GR+ E F++++ +V + W L+ AGK I+ K + A+PVY M F +C+
Sbjct: 1119 LGRRKVELFEYIVTKVKERTEGWAYNYLSPAGKEIVIKAIAMALPVYSMNCFLLPTLICN 1178
Query: 1101 HLDSLAKNFIWSRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSH 1160
++SL F W + + G LG + N ++L K W ++ +
Sbjct: 1179 EINSLITAFWWGK-----------------ENEGDLGFKDLHQFNRALLAKQAWRILTNP 1221
Query: 1161 DKLWV*VLLDLYCTNSDFLHAPR-RRGSCVWNAVMKTRDIIQDGFTWRIGRGDLS-FWFS 1218
L + LY N+ +L A + S WN++ + + ++Q G R+G G + W
Sbjct: 1222 QSLLARLYKGLYYPNTTYLRANKGGHASYGWNSIQEGKLLLQQGLRVRLGDGQTTKIW-- 1279
Query: 1219 KCHGSKALCEKVPYVHISDTALRVRDVFQDG--VCDIRGLSTPITPELWALLQNLFVYVS 1276
+ L + I D ++V D++++ D + PE L ++L++ +
Sbjct: 1280 EDPWLPTLPPRPARGPILDEDMKVADLWRENKREWDPVIFEGVLNPEDQQLAKSLYL-SN 1338
Query: 1277 LDDEDLVVWAPSLEGKYSVQSCYRWLL------------DMEGGTAVSVDWA*LWKFQVP 1324
D WA + +Y+V+S Y W+ +EG + + +W+ ++
Sbjct: 1339 YAARDSYKWAYTRNTQYTVRSGY-WVATHVNLTEEEIINPLEGDVPLKQE---IWRLKIT 1394
Query: 1325 EKMKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLC 1384
K+K F W+ +L T +R + C C + ET H + C + VW++
Sbjct: 1395 PKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQVVWRSAN 1454
Query: 1385 VSLTPGFFTASSFLDWIASVPRHD-------VAGLLVH--AWFIWCRRRRFVFAG-ELEP 1434
S + + + I + + + GL+ W +W R ++F + P
Sbjct: 1455 FSGSNRLCFTDNLEENIRLILQGKKNQNLPILNGLMPFWIMWRLWKSRNEYLFQQLDRFP 1514
Query: 1435 WRL--------------------ILHRAAD-----VLRCLRATSPSMPAA----DPGRAE 1465
W++ I H A + R + +SP D G +
Sbjct: 1515 WKVAQKAEQEATEWVETMVNDTAISHNTAQSNDRPLSRSKQWSSPPEGFLKCNFDSGYVQ 1574
Query: 1466 ------VGGVVRDAVGCWLFGYSGFIGMAE---ILKAELLAVLFGLQSCWDRGFRQLVVS 1516
G ++RD G L +SG + + L+AE L L LQ W RG+ +
Sbjct: 1575 GRDYTSTGWILRDCNGRVL--HSGCAKLQQSYSALQAEALGFLHALQMVWIRGYCYVWFE 1632
Query: 1517 SDSMVVLSLLRHGVTRFHVYAAIVGAIRELLRRDWRVELKHMLREGNVVAD 1567
D++ + +L+ + H+ ++ IR + + + ++ RE N+ AD
Sbjct: 1633 GDNLELTNLI-NKTEDHHLLETLLYDIRFWMTKLPFSSIGYVNRERNLAAD 1682
>At1g24640 hypothetical protein
Length = 1270
Score = 228 bits (582), Expect = 2e-59
Identities = 190/653 (29%), Positives = 282/653 (43%), Gaps = 52/653 (7%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNSR-RGKCQNVVFKLDL 861
+K+M RL+ +L E++ QS+F+ R DN ++ ++VH + R + + K D+
Sbjct: 481 SKIMAKRLQPWLPEIVSDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEFMAVKSDM 540
Query: 862 EKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*GD 921
KA V W++LR L GF V+ IM CVSS + S+L N I RGLR GD
Sbjct: 541 SKAYDRVEWSYLRSLLLSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGD 600
Query: 922 PLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPISHLFFADDVLLLAKAKVSQVR 981
PLSP+LFVLC E LT + A + E ++ S +G + HL FADD L L KA Q
Sbjct: 601 PLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCKASREQSL 660
Query: 982 KVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFPIFQ 1041
+Q IL + A+G +N+ KS V K I I YLG P
Sbjct: 661 VLQKILKVYGNATGQTINLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPECF 720
Query: 1042 GRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AVCDH 1101
+ ++ DR+ K W R L++ GK +L K V A+PV+ M F C++
Sbjct: 721 SGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCEN 780
Query: 1102 LDSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLVNSH 1160
L+S +F W S R ++ +W + LPK +GGLG R + N ++L K W L++
Sbjct: 781 LESAMASFWWDSCDHSRKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFP 840
Query: 1161 DKLWV*VLLDLYCTNSDFLHAP-RRRGSCVWNAVMKTRDIIQDGFTWRIGRG-DLSFWFS 1218
D L +L Y +DFL A +R S W +++ R+++ G R+G G L W
Sbjct: 841 DCLLSRLLKSRYFDATDFLDAALSQRPSFGWRSILFGRELLSKGLQKRVGDGASLFVWID 900
Query: 1219 KCHGSKALCEKVPYVHISDTALRVRDVF--QDGVCDIRGLSTPITPELWALLQNLFVYVS 1276
I D L+V+ + + G D L PE +L+ +
Sbjct: 901 PWIDDNGFRAPWRKNLIYDVTLKVKALLNPRTGFWDEEVLHDLFLPE--DILRIKAIKPV 958
Query: 1277 LDDEDLVVWAPSLEGKYSVQSCYRWLL----DMEGGTAVSVDWA*L------WKFQVPEK 1326
+ D VW + G +SV+S Y WL + VS+ + L W Q K
Sbjct: 959 ISQADFFVWKLNKSGDFSVKSAY-WLAYQTKSQNLRSEVSMQPSTLGLKTQVWNLQTDPK 1017
Query: 1327 MKLFAWQAAHRSLPTVDTFHDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNLCVS 1386
+K+F W+ +C E+ H L C S Q+W
Sbjct: 1018 IKIFLWK------------------------VCGELGESTNHTLFLCPLSRQIWALSDYP 1053
Query: 1387 LTP-GFFTASSFLDWIASVPRHDVAGLLVH--------AWFIWCRRRRFVFAG 1430
P GF S + + + D ++ W IW R F+F G
Sbjct: 1054 FPPDGFSNGSIYSNINHLLENKDNKEWPINLRKIFPWILWRIWKNRNSFIFEG 1106
>At4g10830 putative protein
Length = 1294
Score = 222 bits (566), Expect = 1e-57
Identities = 133/432 (30%), Positives = 225/432 (51%), Gaps = 5/432 (1%)
Query: 785 PSGSSGQSAYAMCYI--S*FTKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIV 842
P S A+C + +K +V RL+ L ++ Q++FIPGR DN +I +++
Sbjct: 860 PETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMM 919
Query: 843 HHMNSRRGKCQN-VVFKLDLEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSL 901
H + +R+ Q+ + K D+ KA V W FL T+ FGF T + IM V S + S+
Sbjct: 920 HSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIMGAVKSVNYSV 979
Query: 902 LWNGVKLPPIAPSRGLR*GDPLSPYLFVLCVERLTISIQDAVDQNQGELVRISRDGSPIS 961
L NG+ I P RG+R GDPLSPYLF+LC + L I++ V + +RI ++
Sbjct: 980 LVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDIRGIRIGNGVPGVT 1039
Query: 962 HLFFADDVLLLAKAKVSQVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFV 1021
HL FADD L ++ V + ++++ + SG +N++KS V + +
Sbjct: 1040 HLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGSRVHGTTQNRLKN 1099
Query: 1022 VTSIPFTANLERYLGFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVL 1081
+ I +YLG P GR+ ++ F+++++RV ++ +SW + L+ AGK I+ K V
Sbjct: 1100 ILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVA 1159
Query: 1082 NAIPVYPMQLFWFH*AVCDHLDSLAKNFIWSR-GERRGMNMVAWRTVTLPKCNGGLGIRK 1140
++PVY M F + +++L NF W + ++R + +AW+ + K GGLG R
Sbjct: 1160 MSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQYSKKEGGLGFRD 1219
Query: 1141 ARLINTSMLGKLVWDLVNSHDKLWV*VLLDLYCTNSDFLHAPRRR-GSCVWNAVMKTRDI 1199
N ++L K VW ++N+ + L+ ++ Y L A R+R S W +++ D+
Sbjct: 1220 LAKFNDALLAKQVWRMINNPNSLFARIMKARYFREDSILDAKRQRYQSYGWTSMLAGLDV 1279
Query: 1200 IQDGFTWRIGRG 1211
I+ G + +G G
Sbjct: 1280 IKKGSRFIVGDG 1291
>At2g11240 pseudogene
Length = 1044
Score = 216 bits (551), Expect = 6e-56
Identities = 136/421 (32%), Positives = 216/421 (51%), Gaps = 10/421 (2%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIVHHMNS--RRGKCQNVVFKLD 860
+K++ RL+ LQE+I QS+F+P R + DN +I + +H++ S +C V K +
Sbjct: 344 SKLLSRRLQPILQEIISENQSAFVPKRASNDNVLITHEALHYLKSLGAEKRCFMAV-KTN 402
Query: 861 LEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*G 920
+ KA + W F++ + GF T +S I+ C+++ S S L NG + P RGLR G
Sbjct: 403 MSKAYDRIEWDFIKLVMQEMGFHQTWISWILQCITTVSYSFLLNGSAQGAVTPERGLRQG 462
Query: 921 DPLSPYLFVLCVERLTISIQDAVDQNQGELV--RISRDGSPISHLFFADDVLLLAKAKVS 978
DPLSP+LF++C E L+ + A Q G L+ R+S+ ++HL FADD + ++ +
Sbjct: 463 DPLSPFLFIICSEVLSGLCRKA--QLDGSLLGLRVSKGNPRVNHLLFADDTIFFCRSDLK 520
Query: 979 QVRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKKADIFVVTSIPFTANLERYLGFP 1038
+ IL + ASG +N +KS S P K + + I L +YLG P
Sbjct: 521 SCKTFLCILKKYEEASGQMINKSKSAITFSRKTPDHIKTEAQQILGIQLVGGLGKYLGLP 580
Query: 1039 IFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH*AV 1098
GR+ ++ F+ ++DR+ ++ SW R L+ AGK + K VL ++P Y M F ++
Sbjct: 581 KMFGRKKRDLFNQIVDRIRQRSLSWSSRFLSTAGKTTMLKSVLASMPTYTMSCFKLLVSL 640
Query: 1099 CDHLDSLAKNFIW-SRGERRGMNMVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVWDLV 1157
C + S +F W S +++ M +AW + K GGLG + N ++L KL W +V
Sbjct: 641 CKRIQSALTHFWWDSSADKKKMCWIAWSKMAKNKKEGGLGFKDITNFNDALLAKLSWRIV 700
Query: 1158 NSHDKLWV*VLLDLYCTNSDFLH-APRRRGSCVWNAVMKTRDIIQDGFTWRIGRG-DLSF 1215
S + V +LL YC S FL + S W + +D+I+ IG G D
Sbjct: 701 QSPSCVLVRILLGKYCRTSSFLDCSVTAASSHGWRGICTGKDLIKSQLGKVIGSGLDTLV 760
Query: 1216 W 1216
W
Sbjct: 761 W 761
>At1g31030 F17F8.5
Length = 872
Score = 215 bits (547), Expect = 2e-55
Identities = 167/600 (27%), Positives = 274/600 (44%), Gaps = 34/600 (5%)
Query: 803 TKVMVNRLRQFLQELIGPLQSSFIPGRGTIDNAIILQKIV--HHMNSRRGKCQNVVFKLD 860
+K++ NRL+ L I QS+F+ R I+N ++ ++V +H +S +C K+D
Sbjct: 190 SKIIANRLKLLLPRFIAENQSAFVKDRLLIENLLLATELVKDYHKDSISARC---AIKID 246
Query: 861 LEKACGSVSWAFLRDTLCFFGFPPTSVSLIMFCVSSSSLSLLWNGVKLPPIAPSRGLR*G 920
+ KA SV W+FL +TL F PT + I C++++S S+ NG + RGLR G
Sbjct: 247 ISKAFDSVQWSFLTNTLVAMNFSPTFIHWINLCITTASFSVQVNGDLVGYFQSKRGLRQG 306
Query: 921 DPLSPYLFVLCVERLTISIQDAVDQNQ-GELVRISRDGSPISHLFFADDVLLLAKAKVSQ 979
LSPYLFV+C++ L+ + A + G + R G ++HL FADD+++L+ K
Sbjct: 307 CSLSPYLFVICMDVLSKMLDKAAGVRKFGFHPKCQRLG--LTHLSFADDLMVLSDGKTRS 364
Query: 980 VRKVQNILADFCMASGLNVNIAKSRDFSSVSVPRAKK----ADIFVVTSIPFTANLERYL 1035
+ + + +FC SGL +++ KS + + P K+ +F V +P RYL
Sbjct: 365 IEGILEVFDEFCKRSGLRISLEKSTLYMAGVSPIIKQEIAAKFLFDVGQLPV-----RYL 419
Query: 1036 GFPIFQGRQTQEHFDFVLDRVNRKFASWKGRLLNKAGKFILFKLVLNAIPVYPMQLFWFH 1095
G P+ R T + +L+++ ++ A+W R + AG+F L K VL +I + + F
Sbjct: 420 GLPLVTKRLTSADYSPLLEQIKKRIATWTFRFFSFAGRFNLIKSVLWSICNFWLAAFRLP 479
Query: 1096 *AVCDHLDSLAKNFIWSRGERRGMN-MVAWRTVTLPKCNGGLGIRKARLINTSMLGKLVW 1154
+D L +F+WS E ++W V PK GGLG+R + N KLVW
Sbjct: 480 RQCIREIDKLCSSFLWSGSEMSSHKAKISWDIVCKPKAEGGLGLRNLKEANDVSCLKLVW 539
Query: 1155 DLVNSHDKLWV*VLLDLYCTNSDF--LHAPRRRGSCVWNAVMKTRDIIQDGFTWRIGRGD 1212
++++ + LW + + L GS +W ++K RD+ + +G G+
Sbjct: 540 RIISNSNSLWTKWVAEYLIRKKSIWSLKQSTSMGSWIWRKILKIRDVAKSFSRVEVGNGE 599
Query: 1213 -LSFWFSKCHGSKALCEKVPYVHISDTALRVRDVFQDGVC--DIRGLSTPITPELWALLQ 1269
SFW+ L + V D + D R T + E+ ++
Sbjct: 600 SASFWYDHWSAHGRLIDTVGDKGTIDLGIPREASVADAWTRRSRRRHRTSLLNEIEEMMA 659
Query: 1270 NLFVYVSLDDEDLVVWAPSLEGKYSVQSCY---RWLLDMEGGTAVSVDW-A*LWKFQVPE 1325
++ S D ED V+W GK V + R + T+ +V W +W
Sbjct: 660 YQRIHHS-DAEDTVLW----RGKNDVFKPHFSTRDTWHLIKATSSTVSWHKGVWFRHATP 714
Query: 1326 KMKLFAWQAAHRSLPTVDTF--HDRGVAIPRWCSLCHHHVET*MHCLRDCDASLQVWQNL 1383
K L W A H LPT D + ++ C LC ++ +T H C + VW L
Sbjct: 715 KYALCTWLAIHNRLPTGDRMLKWNSSGSVSGNCVLCTNNSKTLEHLFFSCSYASTVWAAL 774
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.338 0.147 0.507
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,344,970
Number of Sequences: 26719
Number of extensions: 1429805
Number of successful extensions: 4611
Number of sequences better than 10.0: 119
Number of HSP's better than 10.0 without gapping: 106
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 4162
Number of HSP's gapped (non-prelim): 242
length of query: 1569
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1457
effective length of database: 8,326,068
effective search space: 12131081076
effective search space used: 12131081076
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0022.23


