

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0022.17
(341 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
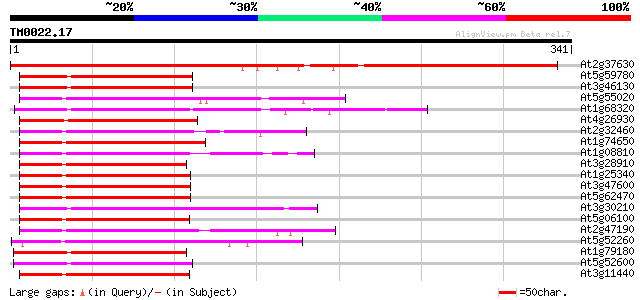
Sequences producing significant alignments: (bits) Value
At2g37630 putative MYB family transcription factor 400 e-112
At5g59780 MYB27 protein - like 103 2e-22
At3g46130 Myb DNA binding protein -like 102 2e-22
At5g55020 putative transcription factor MYB120 (MYB120) 101 5e-22
At1g68320 putative transcription factor (MYB62) 101 7e-22
At4g26930 putative myb-related protein 99 3e-21
At2g32460 putative MYB family transcription factor 99 3e-21
At1g74650 putative MYB family transcription factor 99 3e-21
At1g08810 transcription factor like protein 99 3e-21
At3g28910 MYB family transcription factor (hsr1), putative 97 1e-20
At1g25340 putative transcription factor MYB116 (MYB116) 97 1e-20
At3g47600 putative transcription factor (MYB94) 96 2e-20
At5g62470 MYB96 transcription factor-like protein 96 4e-20
At3g30210 myb-like transcription factor, putative 96 4e-20
At5g06100 transcription factor MYB33 - like protein 95 5e-20
At2g47190 MYB transcription factor (Atmyb2) 95 5e-20
At5g52260 unknown protein 94 1e-19
At1g79180 ranscription factor like protein (MYB63) 94 1e-19
At5g52600 MYB82 (gb|AAF14064.1) 94 1e-19
At3g11440 transcription factor like protein (MYB65) 94 1e-19
>At2g37630 putative MYB family transcription factor
Length = 367
Score = 400 bits (1028), Expect = e-112
Identities = 220/364 (60%), Positives = 264/364 (72%), Gaps = 37/364 (10%)
Query: 1 MKERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKG 60
MKERQRWS EEDALL AYV+Q+GPREW+LVS+RMN PLNRD KSCLERWKNYLKPGIKKG
Sbjct: 1 MKERQRWSGEEDALLRAYVRQFGPREWHLVSERMNKPLNRDAKSCLERWKNYLKPGIKKG 60
Query: 61 SLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSP 120
SLT+EEQRLVI LQ +GNKWKKIAAEVPGRTAKRLGKWWEV+KEKQQRE+ E N V P
Sbjct: 61 SLTEEEQRLVIRLQEKHGNKWKKIAAEVPGRTAKRLGKWWEVFKEKQQREEKESNKRVEP 120
Query: 121 ISDTKYEHMLEGFAEKLVKE--HTLPSFAMA-----ASSNEAFLHTNS-----SAMLPSW 168
I ++KY+ +LE FAEKLVKE + +P+ A A A+SN FLH+ + ++P W
Sbjct: 121 IDESKYDRILESFAEKLVKERSNVVPAAAAAATVVMANSNGGFLHSEQQVQPPNPVIPPW 180
Query: 169 LSNYDS----TSTPPSSISVTLSLSPSTVAT--------------PRGLENN-APFVLRN 209
L+ ++ + PP SVTL+LSPSTVA P EN VL +
Sbjct: 181 LATSNNGNNVVARPP---SVTLTLSPSTVAAAAPQPPIPWLQQQQPERAENGPGGLVLGS 237
Query: 210 VTAHNGSVPSFSDHILMSELVGFSKELEEGHRALAAHKKEAEWRLRRLELQLESEKACRR 269
+ S S+ + +SELV +ELEEGHRA A HKKEA WRLRRLELQLESEK CR+
Sbjct: 238 MMP---SCSGSSESVFLSELVECCRELEEGHRAWADHKKEAAWRLRRLELQLESEKTCRQ 294
Query: 270 RETVEEFEANIKALQEEQTAALNRIENACREQLGGLRRDAESKEQKLAEKWTSKHLRLTR 329
RE +EE EA +KAL+EEQ A+ +IE REQL GLRRDAE+K+QKLA++WTS+H+RLT+
Sbjct: 295 REKMEEIEAKMKALREEQKNAMEKIEGEYREQLVGLRRDAEAKDQKLADQWTSRHIRLTK 354
Query: 330 LLEQ 333
LEQ
Sbjct: 355 FLEQ 358
>At5g59780 MYB27 protein - like
Length = 235
Score = 103 bits (256), Expect = 2e-22
Identities = 46/105 (43%), Positives = 68/105 (63%), Gaps = 2/105 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ +ED LL +V +G R W+ V++ LNR KSC RW NYL PG+K+G +T +E
Sbjct: 13 WTEQEDILLVNFVHLFGDRRWDFVAKVSG--LNRTGKSCRLRWVNYLHPGLKRGKMTPQE 70
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREK 111
+RLV+ L A +GN+W KIA ++PGRT + +W + K+ +EK
Sbjct: 71 ERLVLELHAKWGNRWSKIARKLPGRTDNEIKNYWRTHMRKKAQEK 115
>At3g46130 Myb DNA binding protein -like
Length = 256
Score = 102 bits (255), Expect = 2e-22
Identities = 45/105 (42%), Positives = 68/105 (63%), Gaps = 2/105 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ +ED LL +V +G R W+ +++ LNR KSC RW NYL PG+K+G +T +E
Sbjct: 12 WTEQEDILLVNFVHLFGDRRWDFIAKVSG--LNRTGKSCRLRWVNYLHPGLKRGKMTPQE 69
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREK 111
+RLV+ L A +GN+W KIA ++PGRT + +W + K+ +EK
Sbjct: 70 ERLVLELHAKWGNRWSKIARKLPGRTDNEIKNYWRTHMRKKAQEK 114
>At5g55020 putative transcription factor MYB120 (MYB120)
Length = 523
Score = 101 bits (252), Expect = 5e-22
Identities = 69/211 (32%), Positives = 109/211 (50%), Gaps = 19/211 (9%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W++ ED +L AYV++ G WN V + NT L R KSC RW N+L+P +KKGS T +E
Sbjct: 31 WTAAEDEILAAYVRENGEGNWNAVQK--NTGLARCGKSCRLRWANHLRPNLKKGSFTGDE 88
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEI--NGIV------ 118
+RL+I L A GNKW ++AA++PGRT + +W ++ R+ + + I+
Sbjct: 89 ERLIIQLHAQLGNKWARMAAQLPGRTDNEIKNYWNTRLKRLLRQGLPLYPPDIIPNHQLH 148
Query: 119 -SPISDTKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPS-WLSNYDSTS 176
P + +H + ++H F +S +T SS+ LPS +N S+S
Sbjct: 149 PHPHHQQQQQHNHHHHHHQQQQQHQQMYFQPQSSQR----NTPSSSPLPSPTPANAKSSS 204
Query: 177 T---PPSSISVTLSLSPSTVATPRGLENNAP 204
+ ++ ++ LSP T TP L + P
Sbjct: 205 SFTFHTTTANLLHPLSPHTPNTPSQLSSTPP 235
>At1g68320 putative transcription factor (MYB62)
Length = 286
Score = 101 bits (251), Expect = 7e-22
Identities = 74/268 (27%), Positives = 122/268 (44%), Gaps = 27/268 (10%)
Query: 4 RQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLT 63
R W+ EED LL Y+ G WN V++ L R KSC RW NYLKP I++G+LT
Sbjct: 21 RGPWTLEEDTLLTNYILHNGEGRWNHVAKCAG--LKRTGKSCRLRWLNYLKPDIRRGNLT 78
Query: 64 KEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISD 123
+EQ L++ L + +GN+W KIA +PGRT + +W +KQ R+ + I D
Sbjct: 79 PQEQLLILELHSKWGNRWSKIAQYLPGRTDNEIKNYWRTRVQKQARQ-LNIESNSDKFFD 137
Query: 124 TKYEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLP---------------SW 168
+ EK+ + + + +N N+S +LP S
Sbjct: 138 AVRSFWVPRLIEKMEQNSSTTTTYCCPQNN-----NNNSLLLPSQSHDSLSMQKDIDYSG 192
Query: 169 LSNYDSTSTPPSSISVTLSLSPSTV--ATPRGLENNAPFVLRNVTAHNGSVPSFSDHILM 226
SN D +S+ + +S L+ P + + ++ + F NV G VP D+++
Sbjct: 193 FSNIDGSSSTSTCMS-HLTTVPHFMDQSNTNIIDGSMCFHEGNVQEFGGYVPGMEDYMVN 251
Query: 227 SELVGFSKELEEGHRALAAHKKEAEWRL 254
S+ + + +G+ A ++ W +
Sbjct: 252 SD-ISMECHVADGYSAYEDVTQDPMWNV 278
>At4g26930 putative myb-related protein
Length = 389
Score = 99.4 bits (246), Expect = 3e-21
Identities = 46/108 (42%), Positives = 69/108 (63%), Gaps = 2/108 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ ED L AYV++YG WN V ++ T L R KSC RW N+L+P ++KGS T EE
Sbjct: 24 WTVAEDETLAAYVREYGEGNWNSVQKK--TWLARCGKSCRLRWANHLRPNLRKGSFTPEE 81
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEI 114
+RL+I L + GNKW ++AA++PGRT + +W ++ QR+ + +
Sbjct: 82 ERLIIQLHSQLGNKWARMAAQLPGRTDNEIKNYWNTRLKRFQRQGLPL 129
>At2g32460 putative MYB family transcription factor
Length = 490
Score = 99.4 bits (246), Expect = 3e-21
Identities = 57/178 (32%), Positives = 98/178 (55%), Gaps = 15/178 (8%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W++ EDA+L YV+++G WN V + N+ L R KSC RW N+L+P +KKGS T +E
Sbjct: 23 WTTTEDAILTEYVRKHGEGNWNAVQK--NSGLLRCGKSCRLRWANHLRPNLKKGSFTPDE 80
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
++++I L A GNKW ++A+++PGRT + +W +++QR + P+ +
Sbjct: 81 EKIIIDLHAKLGNKWARMASQLPGRTDNEIKNYWNTRMKRRQRAGL-------PLYPHEI 133
Query: 127 EHMLEGFAEKLVKEHTLPSFAMAAS----SNEAFLHTNSSAMLPSWLSNYDSTSTPPS 180
+H +G E L SF +++ + +S+ S S++ S+S+ PS
Sbjct: 134 QH--QGIDIDDEFEFDLTSFQFQNQDLDHNHQNMIQYTNSSNTSSSSSSFSSSSSQPS 189
>At1g74650 putative MYB family transcription factor
Length = 330
Score = 99.4 bits (246), Expect = 3e-21
Identities = 48/113 (42%), Positives = 72/113 (63%), Gaps = 2/113 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L +Y+QQ+GP W V NT L R +KSC RW NYL+PGIK+G+ T+ E
Sbjct: 17 WTPEEDIILVSYIQQHGPGNWRSVPA--NTGLLRCSKSCRLRWTNYLRPGIKRGNFTQPE 74
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVS 119
++++I LQA GN+W IA+ +P RT + +W + +K+ NGI++
Sbjct: 75 EKMIIHLQALLGNRWAAIASYLPQRTDNDIKNYWNTHLKKKLVMMKFQNGIIN 127
>At1g08810 transcription factor like protein
Length = 280
Score = 99.4 bits (246), Expect = 3e-21
Identities = 63/180 (35%), Positives = 93/180 (51%), Gaps = 24/180 (13%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L +Y+Q++GP W V NT L R +KSC RW NYL+PGIK+G+ T E
Sbjct: 17 WTPEEDIILVSYIQEHGPGNWRSVPT--NTGLLRCSKSCRLRWTNYLRPGIKRGNFTPHE 74
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
+ ++I LQA GNKW IA+ +P RT + +W + +K+ + SD+
Sbjct: 75 EGMIIHLQALLGNKWASIASYLPQRTDNDIKNYWNTHLKKKLNK-----------SDSDE 123
Query: 127 EHMLEGFA-EKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPSSISVT 185
E A + +T+ + ASS E N S +L W+ ++P SS S T
Sbjct: 124 RSRSENIALQTSSTRNTINHRSTYASSTE-----NISRLLEGWM-----RASPKSSTSTT 173
>At3g28910 MYB family transcription factor (hsr1), putative
Length = 323
Score = 97.1 bits (240), Expect = 1e-20
Identities = 43/101 (42%), Positives = 66/101 (64%), Gaps = 2/101 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L Y+Q++GP W V NT L R +KSC RW NYL+PGIK+G+ T+ E
Sbjct: 17 WTPEEDIILVTYIQEHGPGNWRAVPT--NTGLLRCSKSCRLRWTNYLRPGIKRGNFTEHE 74
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
+++++ LQA GN+W IA+ +P RT + +W + +K+
Sbjct: 75 EKMIVHLQALLGNRWAAIASYLPQRTDNDIKNYWNTHLKKK 115
>At1g25340 putative transcription factor MYB116 (MYB116)
Length = 283
Score = 97.1 bits (240), Expect = 1e-20
Identities = 45/104 (43%), Positives = 66/104 (63%), Gaps = 2/104 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED LL Y+ G WNL+++ ++ L R KSC RW NYLKP IK+G+LT +E
Sbjct: 23 WTLEEDTLLTNYISHNGEGRWNLLAK--SSGLKRAGKSCRLRWLNYLKPDIKRGNLTPQE 80
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE 110
Q L++ L + +GN+W KI+ +PGRT + +W +KQ R+
Sbjct: 81 QLLILELHSKWGNRWSKISKYLPGRTDNDIKNYWRTRVQKQARQ 124
>At3g47600 putative transcription factor (MYB94)
Length = 333
Score = 96.3 bits (238), Expect = 2e-20
Identities = 42/104 (40%), Positives = 69/104 (65%), Gaps = 2/104 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L +Y+Q++GP W V +T L R +KSC RW NYL+PGIK+G+ T+ E
Sbjct: 17 WTPEEDIILVSYIQEHGPGNWRSVPT--HTGLRRCSKSCRLRWTNYLRPGIKRGNFTEHE 74
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE 110
+++++ LQA GN+W IA+ +P RT + +W + +K+ ++
Sbjct: 75 EKMILHLQALLGNRWAAIASYLPERTDNDIKNYWNTHLKKKLKK 118
>At5g62470 MYB96 transcription factor-like protein
Length = 352
Score = 95.5 bits (236), Expect = 4e-20
Identities = 42/104 (40%), Positives = 68/104 (65%), Gaps = 2/104 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED +L +Y+Q++GP W V +T L R +KSC RW NYL+PGIK+G+ T+ E
Sbjct: 17 WTPEEDIILVSYIQEHGPGNWRSVPT--HTGLRRCSKSCRLRWTNYLRPGIKRGNFTEHE 74
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE 110
++ ++ LQA GN+W IA+ +P RT + +W + +K+ ++
Sbjct: 75 EKTIVHLQALLGNRWAAIASYLPERTDNDIKNYWNTHLKKKLKK 118
>At3g30210 myb-like transcription factor, putative
Length = 276
Score = 95.5 bits (236), Expect = 4e-20
Identities = 54/181 (29%), Positives = 93/181 (50%), Gaps = 4/181 (2%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EED LL YV +G W+ V++ LNR KSC RW NYL+PG+K+G +T +E
Sbjct: 32 WTLEEDKLLAEYVTSHGEGRWSTVAKCAG--LNRSGKSCRLRWVNYLRPGLKRGQITPQE 89
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREKIEINGIVSPISDTKY 126
+ +++ L + +GNKW IA +PGRT + +W + +K Q+ + + + +S +
Sbjct: 90 EGIILELHSLWGNKWSTIARYLPGRTDNEIKNYWRTHYKKNQKSSSKQDKVKKSLSRKQQ 149
Query: 127 EHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNSSAMLPSWLSNYDSTSTPPSSISVTL 186
+ L+ + + H + + + + SS P+ S +D P S++ T
Sbjct: 150 QVDLKPQPQAQSENHQSQLVSQDHMNIDNDHNIASSLYYPT--SVFDDKLYMPQSVATTS 207
Query: 187 S 187
S
Sbjct: 208 S 208
>At5g06100 transcription factor MYB33 - like protein
Length = 520
Score = 95.1 bits (235), Expect = 5e-20
Identities = 43/103 (41%), Positives = 67/103 (64%), Gaps = 2/103 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
WSS ED +L YV ++G WN V + +T L R KSC RW N+L+P +KKG+ ++EE
Sbjct: 37 WSSAEDDILIDYVNKHGEGNWNAVQK--HTSLFRCGKSCRLRWANHLRPNLKKGAFSQEE 94
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQR 109
++L++ L A GN+W ++AA +PGRT + +W +++QR
Sbjct: 95 EQLIVELHAKMGNRWARMAAHLPGRTDNEIKNYWNTRIKRRQR 137
>At2g47190 MYB transcription factor (Atmyb2)
Length = 273
Score = 95.1 bits (235), Expect = 5e-20
Identities = 61/203 (30%), Positives = 104/203 (51%), Gaps = 19/203 (9%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+ EEDA+L +V +G WN +++ ++ L R KSC RW NYL+P +++G++T EE
Sbjct: 25 WTEEEDAILVNFVSIHGDARWNHIAR--SSGLKRTGKSCRLRWLNYLRPDVRRGNITLEE 82
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQRE-KIEINGIVSPISDTK 125
Q +++ L + +GN+W KIA +PGRT + +W +KQ + + ++N S+
Sbjct: 83 QFMILKLHSLWGNRWSKIAQYLPGRTDNEIKNYWRTRVQKQAKHLRCDVN------SNLF 136
Query: 126 YEHMLEGFAEKLVKEHTLPSFAMAASSNEAFLHTNS------SAMLPSWL---SNYDSTS 176
E M + +LV+ S E+ + S S + P ++ N+
Sbjct: 137 KETMRNVWMPRLVERINAQSLPTTCEQVESMITDPSQPVNEPSPVEPGFVQFSQNHHQQF 196
Query: 177 TPPSSISVTLSLSPS-TVATPRG 198
P + +S T S SP+ T + RG
Sbjct: 197 VPATELSATSSNSPAETFSDVRG 219
>At5g52260 unknown protein
Length = 268
Score = 94.0 bits (232), Expect = 1e-19
Identities = 57/191 (29%), Positives = 100/191 (51%), Gaps = 16/191 (8%)
Query: 2 KERQR---WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIK 58
K+RQR WS EED L +++ G W V + L R+ KSC RW NYL+PG+K
Sbjct: 9 KQRQRKGLWSPEEDQKLKSFILSRGHACWTTVP--ILAGLQRNGKSCRLRWINYLRPGLK 66
Query: 59 KGSLTKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVY-KEKQQREKIEINGI 117
+GS ++EE+ ++ L ++ GNKW +IA +PGRT + +W Y K++ + + ++
Sbjct: 67 RGSFSEEEEETILTLHSSLGNKWSRIAKYLPGRTDNEIKNYWHSYLKKRWLKSQPQLKSQ 126
Query: 118 VSPISDTKYEHMLEG--FAEKLVKEHTL--------PSFAMAASSNEAFLHTNSSAMLPS 167
+S ++++ + G E +H + P+ + + SN + NS+ +
Sbjct: 127 ISDLTESPSSLLSCGKRNLETETLDHVISFQKFSENPTSSPSKESNNNMIMNNSNNLPKL 186
Query: 168 WLSNYDSTSTP 178
+ S + S+S P
Sbjct: 187 FFSEWISSSNP 197
>At1g79180 ranscription factor like protein (MYB63)
Length = 294
Score = 94.0 bits (232), Expect = 1e-19
Identities = 43/105 (40%), Positives = 65/105 (60%), Gaps = 2/105 (1%)
Query: 3 ERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSL 62
+R WS EED L +++Q++G W + ++ L R KSC RW NYL+P +K+G+
Sbjct: 15 KRGPWSPEEDIKLISFIQKFGHENWRSLPKQSG--LLRCGKSCRLRWINYLRPDLKRGNF 72
Query: 63 TKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQ 107
T EE+ +I L NYGNKW KIA+++PGRT + W + +K+
Sbjct: 73 TSEEEETIIKLHHNYGNKWSKIASQLPGRTDNEIKNVWHTHLKKR 117
>At5g52600 MYB82 (gb|AAF14064.1)
Length = 201
Score = 93.6 bits (231), Expect = 1e-19
Identities = 43/109 (39%), Positives = 64/109 (58%), Gaps = 2/109 (1%)
Query: 3 ERQRWSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSL 62
+R W EED +L +YV+ +G W +S+R L R KSC RWKNYL+P IK+GS+
Sbjct: 13 KRGLWKPEEDMILKSYVETHGEGNWADISRRSG--LKRGGKSCRLRWKNYLRPNIKRGSM 70
Query: 63 TKEEQRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQREK 111
+ +EQ L+I + GN+W IA +PGRT + +W + K+ +
Sbjct: 71 SPQEQDLIIRMHKLLGNRWSLIAGRLPGRTDNEVKNYWNTHLNKKPNSR 119
>At3g11440 transcription factor like protein (MYB65)
Length = 553
Score = 93.6 bits (231), Expect = 1e-19
Identities = 41/103 (39%), Positives = 67/103 (64%), Gaps = 2/103 (1%)
Query: 7 WSSEEDALLHAYVQQYGPREWNLVSQRMNTPLNRDTKSCLERWKNYLKPGIKKGSLTKEE 66
W+S ED +L YV+++G WN V + +T L R KSC RW N+L+P +KKG+ ++EE
Sbjct: 46 WTSTEDGILIDYVKKHGEGNWNAVQK--HTSLARCGKSCRLRWANHLRPNLKKGAFSQEE 103
Query: 67 QRLVILLQANYGNKWKKIAAEVPGRTAKRLGKWWEVYKEKQQR 109
++L++ + A GNKW ++A +PGRT + +W +++QR
Sbjct: 104 EQLIVEMHAKMGNKWAQMAEHLPGRTDNEIKNYWNTRIKRRQR 146
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.127 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,704,755
Number of Sequences: 26719
Number of extensions: 323015
Number of successful extensions: 1605
Number of sequences better than 10.0: 247
Number of HSP's better than 10.0 without gapping: 172
Number of HSP's successfully gapped in prelim test: 75
Number of HSP's that attempted gapping in prelim test: 1187
Number of HSP's gapped (non-prelim): 338
length of query: 341
length of database: 11,318,596
effective HSP length: 100
effective length of query: 241
effective length of database: 8,646,696
effective search space: 2083853736
effective search space used: 2083853736
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0022.17


