

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0020.6
(795 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
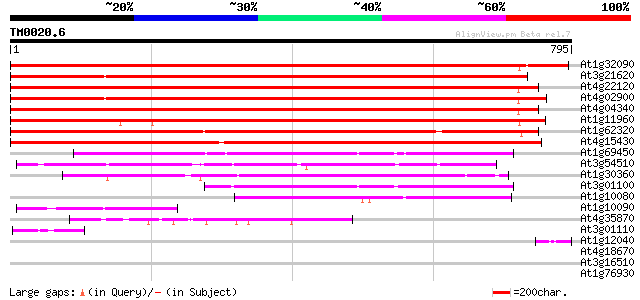
Score E
Sequences producing significant alignments: (bits) Value
At1g32090 unknown protein 1189 0.0
At3g21620 unknown protein 982 0.0
At4g22120 unknown protein 947 0.0
At4g02900 hypothetical protein 937 0.0
At4g04340 predicted protein of unknown function 937 0.0
At1g11960 unknown protein 928 0.0
At1g62320 hypothetical protein 917 0.0
At4g15430 unknown protein 913 0.0
At1g69450 hypothetical protein 381 e-106
At3g54510 putative protein 367 e-101
At1g30360 ERD4 protein (ERD4) 347 2e-95
At3g01100 unknown protein 309 3e-84
At1g10080 hypothetical protein 265 1e-70
At1g10090 hypothetical protein 104 2e-22
At4g35870 putative protein 84 3e-16
At3g01110 hypothetical protein 49 1e-05
At1g12040 leucine-rich repeat/extensin 1 (LRX1) 43 6e-04
At4g18670 extensin-like protein 42 0.001
At3g16510 unknown protein 42 0.002
At1g76930 extensin 42 0.002
>At1g32090 unknown protein
Length = 806
Score = 1189 bits (3077), Expect = 0.0
Identities = 597/807 (73%), Positives = 678/807 (83%), Gaps = 18/807 (2%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPR-STGNFVG 59
MATL DIGVSA IN+ AF FL+AFA+LRIQPINDRVYFPKWY+ GER SPR S VG
Sbjct: 1 MATLQDIGVSALINLFGAFLFLIAFAVLRIQPINDRVYFPKWYLTGERNSPRRSDRTLVG 60
Query: 60 KFVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLI 119
KFVNLN+KTY TFLNWMPQA++MSE+EII HAGLDSA+FLRIYTLGLKIF PV V+AL++
Sbjct: 61 KFVNLNYKTYFTFLNWMPQAMKMSESEIIRHAGLDSAIFLRIYTLGLKIFAPVMVLALVV 120
Query: 120 LIPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
L+PVNVSSG LFFLK++LVVS+IDKLSISNV KSS+FF HI +EY+FT W CF+LY+EY
Sbjct: 121 LVPVNVSSGTLFFLKKELVVSNIDKLSISNVQPKSSKFFFHIAVEYIFTFWACFMLYREY 180
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
+N+A+MRL +LASQRRR EQFTVVVR+VP + GHSV D+V+ FF+TNHP HY+ HQAVYN
Sbjct: 181 NNVAIMRLQYLASQRRRPEQFTVVVRNVPDMPGHSVPDTVDQFFKTNHPEHYLCHQAVYN 240
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
AN +AKLV++R +LQ W DYY LK QR+P K+PT TG LGLWG++VD+IEYY IKE
Sbjct: 241 ANTYAKLVKQRAKLQRWFDYYVLKHQRNPHKQPTCRTGFLGLWGKRVDSIEYYKQQIKEF 300
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
D M+LERQ+++KD K +LPVAF+SF SRWGA+VCAQTQQSKNPTLWLT APEPRDIYW
Sbjct: 301 DHNMSLERQKVLKDSKLMLPVAFVSFDSRWGAAVCAQTQQSKNPTLWLTSSAPEPRDIYW 360
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
+NLAIPF+SL+IRKLVI +SVFALVFFYMIPIAFVQSLANLEGL+RVAPFLRPV L FI
Sbjct: 361 QNLAIPFISLTIRKLVIGVSVFALVFFYMIPIAFVQSLANLEGLDRVAPFLRPVTRLDFI 420
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
KSFLQGFLPGLALKIFL+ILP VL+IMSKIEG+IA STLER+ AAKYYYFMLVNVFLGSI
Sbjct: 421 KSFLQGFLPGLALKIFLWILPTVLLIMSKIEGYIALSTLERRAAAKYYYFMLVNVFLGSI 480
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
+ GTAFEQLHSFLHQSP+QIPRTIGVSIPMKATFF+TYIMVDGWAGIAGEILRLKPLVI+
Sbjct: 481 IAGTAFEQLHSFLHQSPSQIPRTIGVSIPMKATFFITYIMVDGWAGIAGEILRLKPLVIF 540
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKNMFIVKTE DR +AMDPG VDF ETIPSLQLYFLLGIVY VTPILLPFIL+FFA A
Sbjct: 541 HLKNMFIVKTEEDRVRAMDPGFVDFKETIPSLQLYFLLGIVYTAVTPILLPFILIFFAFA 600
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
YLVYRHQIINVYNQQYES AFWPHVH RIIASLLISQLL++GLL++KKAA STPLL IL
Sbjct: 601 YLVYRHQIINVYNQQYESCGAFWPHVHGRIIASLLISQLLLMGLLASKKAADSTPLLIIL 660
Query: 660 PILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE 719
PILT +FHKYC+HRFEPAFR+YPLEEAMAKD LEK TEP+LN+KA LADAYLHPIF SFE
Sbjct: 661 PILTLSFHKYCKHRFEPAFRQYPLEEAMAKDKLEKETEPELNMKADLADAYLHPIFHSFE 720
Query: 720 VE---------------EELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSPPQHV 764
E EE EVRVDK ETQ +SP ++E + S H + SP H
Sbjct: 721 KEVELSSSSSSEKETHQEETPEVRVDKH-ETQSSSP-VTELGTSSHHHHVYNSTSPSSHY 778
Query: 765 NQYSHYPTSPPGYYYHPPPPPHYAYQY 791
+S Y+Y+ + Y+Y
Sbjct: 779 ASAYEQSSSQYEYHYNTHQYEEHEYRY 805
>At3g21620 unknown protein
Length = 756
Score = 982 bits (2538), Expect = 0.0
Identities = 464/737 (62%), Positives = 603/737 (80%), Gaps = 4/737 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGV+A INIL+AFAF +AFA+LR+QP+NDRVYFPKWY+ G R+SP TG F K
Sbjct: 1 MATLTDIGVAATINILTAFAFFIAFAILRLQPVNDRVYFPKWYLKGLRSSPIKTGGFASK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL+F++Y+ FLNWMPQALRM E E+I+HAGLDS V+LRIY LGLKIF P+ +A ++
Sbjct: 61 FVNLDFRSYIRFLNWMPQALRMPEPELIDHAGLDSVVYLRIYLLGLKIFFPIACIAFTVM 120
Query: 121 IPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
+PVN ++ L LK +L SDIDKLSISN+P+ SSRF+VH+ + Y+ T W CF+L +EY
Sbjct: 121 VPVNWTNSTLDQLK-NLTFSDIDKLSISNIPTGSSRFWVHLCMAYVITFWTCFVLQREYK 179
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNA 240
+IA MRL FLAS+ RR +QFTV+VR++P SVS+ VE FF+ NHP++Y+ +QAVYNA
Sbjct: 180 HIASMRLQFLASEHRRPDQFTVLVRNIPPDPDESVSELVEHFFKVNHPDYYLTYQAVYNA 239
Query: 241 NKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELD 300
NK ++LV++R +LQNWLDYYQ K R+P KRP I G LG WG +VDAI++Y I+ L
Sbjct: 240 NKLSELVQKRMKLQNWLDYYQNKHSRNPSKRPLIKIGFLGCWGEEVDAIDHYIEKIEGLT 299
Query: 301 KVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWR 360
+ ++ E++ ++ K ++P AF+SF RWGA VC+QTQQS+NPT WLT+WAPEPRDIYW
Sbjct: 300 RKISEEKETVMSSTKSLVPAAFVSFKKRWGAVVCSQTQQSRNPTEWLTEWAPEPRDIYWD 359
Query: 361 NLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIK 420
NLA+P+V L+IR+LVI+++ F L FF+MIPIAFVQ+LAN+EG+E+ PFL+P+IE+K +K
Sbjct: 360 NLALPYVQLTIRRLVIAVAFFFLTFFFMIPIAFVQTLANIEGIEKAVPFLKPLIEVKTVK 419
Query: 421 SFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIV 480
SF+QGFLPG+ALKIFL +LP++LM+MSK EG I+KS+LER+ A++YY F +NVFL SI+
Sbjct: 420 SFIQGFLPGIALKIFLIVLPSILMLMSKFEGFISKSSLERRCASRYYMFQFINVFLCSII 479
Query: 481 TGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYH 540
GTA +QL SFL+QS T+IP+TIGVSIPMKATFF+TYIMVDGWAG+AGEILRLKPL+IYH
Sbjct: 480 AGTALQQLDSFLNQSATEIPKTIGVSIPMKATFFITYIMVDGWAGVAGEILRLKPLIIYH 539
Query: 541 LKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAY 600
LKN F+VKTE+DR +AMDPG++ F P +QLYF+LG+VYA V+PILLPFILVFFALAY
Sbjct: 540 LKNFFLVKTEKDREEAMDPGTIGFNTGEPQIQLYFILGLVYAAVSPILLPFILVFFALAY 599
Query: 601 LVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILP 660
+VYRHQIINVYNQ+YESAAAFWP VH R++ +L++SQLL++GLLSTKKAA+STPLL ILP
Sbjct: 600 VVYRHQIINVYNQEYESAAAFWPDVHRRVVIALIVSQLLLMGLLSTKKAARSTPLLFILP 659
Query: 661 ILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE- 719
+LT FHK+CQ R++P F YPL++AM KD LE+ EP+LN+K +L +AY HP+F++ +
Sbjct: 660 VLTIGFHKFCQGRYQPIFVTYPLQDAMVKDTLERMREPNLNLKTFLQNAYAHPVFKAADN 719
Query: 720 VEEELV--EVRVDKQPE 734
+ E+V E DK P+
Sbjct: 720 LANEMVVEEPAPDKTPD 736
>At4g22120 unknown protein
Length = 771
Score = 947 bits (2449), Expect = 0.0
Identities = 454/763 (59%), Positives = 593/763 (77%), Gaps = 14/763 (1%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGVSA INILSAF F + FA+LR+QP NDRVYF KWY+ G R+SP G F +
Sbjct: 1 MATLQDIGVSAGINILSAFVFFIIFAVLRLQPFNDRVYFSKWYLKGLRSSPARGGAFAQR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL+F++Y+ FLNWMP+AL+M E E+I+HAGLDS V+LRIY LGLKIF P+ V+A +L
Sbjct: 61 FVNLDFRSYMKFLNWMPEALKMPEPELIDHAGLDSVVYLRIYWLGLKIFTPIAVLAWAVL 120
Query: 121 IPVNVSSGALFFLK--RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKE 178
+PVN ++ L K R++ SDIDKLS+SN+P S RF+ HI + Y FT+W C++L KE
Sbjct: 121 VPVNWTNNTLEMAKQLRNVTSSDIDKLSVSNIPEYSMRFWTHIVMAYAFTIWTCYVLMKE 180
Query: 179 YDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVY 238
Y+ IA MRL F+AS+ RR +QFTV+VR+VP + SVS+ VE FF NHP+HY+ HQ V
Sbjct: 181 YETIANMRLQFVASEARRPDQFTVLVRNVPPDADESVSELVEHFFLVNHPDHYLTHQVVC 240
Query: 239 NANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKE 298
NANK A LV+++ +LQNWLDYYQLK+ R+ +R + G LGLWG+KVDAIE+Y I +
Sbjct: 241 NANKLADLVKKKKKLQNWLDYYQLKYARNNSQRIMVKLGFLGLWGQKVDAIEHYIAEIDK 300
Query: 299 LDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIY 358
+ K ++ ER+ ++ DPK I+P AF+SF +RW A+VCAQTQQ++NPT WLT+WAPEPRD++
Sbjct: 301 ISKEISKEREEVVNDPKAIMPAAFVSFKTRWAAAVCAQTQQTRNPTQWLTEWAPEPRDVF 360
Query: 359 WRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKF 418
W NLAIP+VSL++R+L++ ++ F L FF+++PIAFVQSLA +EG+ + APFL+ +++ KF
Sbjct: 361 WSNLAIPYVSLTVRRLIMHVAFFFLTFFFIVPIAFVQSLATIEGIVKAAPFLKFIVDDKF 420
Query: 419 IKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGS 478
+KS +QGFLPG+ALK+FL LP++LMIMSK EG + S+LER+ A +YY F LVNVFL S
Sbjct: 421 MKSVIQGFLPGIALKLFLAFLPSILMIMSKFEGFTSISSLERRAAFRYYIFNLVNVFLAS 480
Query: 479 IVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVI 538
++ G AFEQL+SFL+QS QIP+TIGV+IPMKATFF+TYIMVDGWAG+AGEIL LKPL++
Sbjct: 481 VIAGAAFEQLNSFLNQSANQIPKTIGVAIPMKATFFITYIMVDGWAGVAGEILMLKPLIM 540
Query: 539 YHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFAL 598
+HLKN F+VKT++DR +AMDPGS+ F P +QLYFLLG+VYA VTP+LLPFILVFFAL
Sbjct: 541 FHLKNAFLVKTDKDREEAMDPGSIGFNTGEPRIQLYFLLGLVYAPVTPMLLPFILVFFAL 600
Query: 599 AYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAI 658
AY+VYRHQIINVYNQ+YESAAAFWP VH R+IA+L+ISQLL++GLL TK AA + P L
Sbjct: 601 AYIVYRHQIINVYNQEYESAAAFWPDVHGRVIAALVISQLLLMGLLGTKHAALAAPFLIA 660
Query: 659 LPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSF 718
LP+LT FH +C+ R+EPAF +YPL+EAM KD LE EP+LN+K YL +AY+HP+F+
Sbjct: 661 LPVLTIGFHHFCKGRYEPAFIRYPLQEAMMKDTLETAREPNLNLKGYLQNAYVHPVFKGD 720
Query: 719 E-----------VEEELVEVRVDKQPETQVASPT-LSEPSSPS 749
E E+E + V +Q +P+ +S SPS
Sbjct: 721 EDDYDIDDKLGKFEDEAIIVPTKRQSRRNTPAPSIISGDDSPS 763
>At4g02900 hypothetical protein
Length = 785
Score = 937 bits (2423), Expect = 0.0
Identities = 458/766 (59%), Positives = 590/766 (76%), Gaps = 7/766 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MA++ DIG+SAAIN+LSAFAFL AFA+LR+QP+NDRVYFPKWY+ G R SP + + +
Sbjct: 1 MASVQDIGLSAAINLLSAFAFLFAFAMLRLQPVNDRVYFPKWYLKGIRGSPTRSRGIMTR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL++ TY+ FLNWMP AL+M E E+I HAGLDSAV++RIY LGLK+FVP+T++A +L
Sbjct: 61 FVNLDWTTYVKFLNWMPAALQMPEPELIEHAGLDSAVYIRIYLLGLKMFVPITLLAFGVL 120
Query: 121 IPVNVSSGALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYD 180
+PVN + L + DL S++DKLSISNVP S RF+ HI + Y+ T W C++LY EY
Sbjct: 121 VPVNWTGETLENID-DLTFSNVDKLSISNVPPGSPRFWAHITMTYVITFWTCYILYMEYK 179
Query: 181 NIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYNA 240
+A MRL LA++ RR +Q TV+VR+VP SV++ VE FF NHP+HY+ HQ VYNA
Sbjct: 180 AVANMRLRHLAAESRRPDQLTVLVRNVPPDPDESVNEHVEHFFCVNHPDHYLCHQVVYNA 239
Query: 241 NKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELD 300
N AKLV +R +QNWL YY+ KF+R P RPT TG G WG VDAI++Y+ + L
Sbjct: 240 NDLAKLVAQRKAMQNWLTYYENKFERKPSSRPTTKTGYGGFWGTTVDAIDFYTSKMDILA 299
Query: 301 KVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWR 360
+ +ER++I+ DPK I+P AF+SF SRWG +VCAQTQQ NPT+WLT+WAPEPRD++W
Sbjct: 300 EQEAVEREKIMNDPKAIMPAAFVSFRSRWGTAVCAQTQQCHNPTIWLTEWAPEPRDVFWD 359
Query: 361 NLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIK 420
NLAIP+V LSIR+L+ ++++F L+F +MIPIAFVQSLANLEG+++V PFL+PVIE+K +K
Sbjct: 360 NLAIPYVELSIRRLLTTVALFFLIFCFMIPIAFVQSLANLEGIQKVLPFLKPVIEMKTVK 419
Query: 421 SFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIV 480
S +QGFLPG+ALKIFL ILP +LM MS+IEG+ + S L+R++A KY++F++VNVFLGSI+
Sbjct: 420 SVIQGFLPGIALKIFLIILPTILMTMSQIEGYTSLSYLDRRSAEKYFWFIIVNVFLGSII 479
Query: 481 TGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYH 540
TGTAF+QL SFL Q PT+IP+T+GVSIPMKATFF+TYIMVDGWAGIA EILR+ PLVI+H
Sbjct: 480 TGTAFQQLKSFLEQPPTEIPKTVGVSIPMKATFFITYIMVDGWAGIAAEILRVVPLVIFH 539
Query: 541 LKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAY 600
LKN F+VKTE+DR +AMDPG +DF + P +Q YFLLG+VYA V PILLPFI+VFFA AY
Sbjct: 540 LKNTFLVKTEQDRQQAMDPGHLDFATSEPRIQFYFLLGLVYAAVAPILLPFIIVFFAFAY 599
Query: 601 LVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILP 660
+V+RHQ+INVY+Q+YES A +WP VH R+I L+ISQLLM+GLLSTKK AK T LL P
Sbjct: 600 VVFRHQVINVYDQKYESGARYWPDVHRRLIICLIISQLLMMGLLSTKKFAKVTALLLPQP 659
Query: 661 ILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE- 719
ILTF F++YC RFE AF K+PL+EAM KD LEK TEP+LN+K YL DAY+HP+F+ +
Sbjct: 660 ILTFWFYRYCAGRFESAFSKFPLQEAMVKDTLEKATEPNLNLKEYLKDAYVHPVFKGNDF 719
Query: 720 -----VEEELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPSP 760
V+EE V + +Q + SE SS + + SP
Sbjct: 720 DRPRVVDEEESNPLVRTKRTSQGTTRYNSEASSSATTTPVANNDSP 765
>At4g04340 predicted protein of unknown function
Length = 772
Score = 937 bits (2421), Expect = 0.0
Identities = 453/763 (59%), Positives = 590/763 (76%), Gaps = 14/763 (1%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGVSA INIL+AF F + FA LR+QP NDRVYF KWY+ G R+SP S G F G+
Sbjct: 1 MATLKDIGVSAGINILTAFIFFIIFAFLRLQPFNDRVYFSKWYLRGLRSSPASGGGFAGR 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVNL ++YL FL+WMP+AL+M E E+I+HAGLDS V+LRIY LGLKIF P+ ++A +L
Sbjct: 61 FVNLELRSYLKFLHWMPEALKMPERELIDHAGLDSVVYLRIYWLGLKIFAPIAMLAWAVL 120
Query: 121 IPVNVSSGALFFLK--RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKE 178
+PVN ++ L K +++ SDIDKL+ISN+P S+RF+ HI + Y FT+W C++L KE
Sbjct: 121 VPVNWTNNELELAKHFKNVTSSDIDKLTISNIPEGSNRFWAHIIMAYAFTIWTCYMLMKE 180
Query: 179 YDNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVY 238
Y+ +A MRL FLAS+ RR +QFTV+VR+VP +VS+ VE FF NHP++Y+ HQ V
Sbjct: 181 YETVANMRLQFLASEGRRPDQFTVLVRNVPPDPDETVSELVEHFFLVNHPDNYLTHQVVC 240
Query: 239 NANKFAKLVRRRDRLQNWLDYYQLKFQRHPDK-RPTINTGVLGLWGRKVDAIEYYSHTIK 297
NANK A LV ++ +LQNWLDYYQLK+ R+ + RP G LGL G+KVDAIE+Y +
Sbjct: 241 NANKLADLVSKKTKLQNWLDYYQLKYTRNNSQIRPITKLGCLGLCGQKVDAIEHYIAEVD 300
Query: 298 ELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDI 357
+ K + ER+ ++ D K ++P +F+SF +RW A+VCAQT Q++NPT WLT+WA EPRDI
Sbjct: 301 KTSKEIAEERENVVNDQKSVMPASFVSFKTRWAAAVCAQTTQTRNPTEWLTEWAAEPRDI 360
Query: 358 YWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELK 417
YW NLAIP+VSL++R+LV++++ F L FF++IPIAFVQSLA +EG+E+VAPFL+ +IE
Sbjct: 361 YWPNLAIPYVSLTVRRLVMNVAFFFLTFFFIIPIAFVQSLATIEGIEKVAPFLKVIIEKD 420
Query: 418 FIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLG 477
FIKS +QG L G+ALK+FL LPA+LM MSK EG + S LER++A++YY F LVNVFLG
Sbjct: 421 FIKSLIQGLLAGIALKLFLIFLPAILMTMSKFEGFTSVSFLERRSASRYYIFNLVNVFLG 480
Query: 478 SIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLV 537
S++ G AFEQL+SFL+QSP QIP+TIG++IPMKATFF+TYIMVDGWAG+AGEIL LKPL+
Sbjct: 481 SVIAGAAFEQLNSFLNQSPNQIPKTIGMAIPMKATFFITYIMVDGWAGVAGEILMLKPLI 540
Query: 538 IYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFA 597
IYHLKN F+VKTE+DR +AM+PGS+ F P +QLYFLLG+VYA VTP+LLPFILVFFA
Sbjct: 541 IYHLKNAFLVKTEKDREEAMNPGSIGFNTGEPQIQLYFLLGLVYAPVTPMLLPFILVFFA 600
Query: 598 LAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLA 657
LAY+VYRHQIINVYNQ+YESAAAFWP VH R+I +L+ISQLL++GLL TK AA + P L
Sbjct: 601 LAYVVYRHQIINVYNQEYESAAAFWPDVHGRVITALIISQLLLMGLLGTKHAASAAPFLI 660
Query: 658 ILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRS 717
LP++T FH++C+ RFEPAF +YPL+EAM KD LE+ EP+LN+K YL DAY+HP+F+
Sbjct: 661 ALPVITIGFHRFCKGRFEPAFVRYPLQEAMMKDTLERAREPNLNLKGYLQDAYIHPVFKG 720
Query: 718 FE----------VEEELVEVRVDKQPETQVASPT-LSEPSSPS 749
+ +E E++ V +Q +P+ +S SSPS
Sbjct: 721 GDNDDDGDMIGKLENEVIIVPTKRQSRRNTPAPSRISGESSPS 763
>At1g11960 unknown protein
Length = 786
Score = 928 bits (2398), Expect = 0.0
Identities = 449/779 (57%), Positives = 596/779 (75%), Gaps = 20/779 (2%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATL DIGV+AAINIL+A FLLAFA+LRIQP NDRVYFPKWY+ G R+SP +G V K
Sbjct: 1 MATLGDIGVAAAINILTAIIFLLAFAILRIQPFNDRVYFPKWYLKGIRSSPLHSGALVSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
FVN+N +YL FLNWMP AL+M E E+I+HAGLDSAV+LRIY +GLKIFVP+ ++A IL
Sbjct: 61 FVNVNLGSYLRFLNWMPAALKMPEPELIDHAGLDSAVYLRIYLIGLKIFVPIALLAWSIL 120
Query: 121 IPVNVSSGALFFLK-RDLVVSDIDKLSISNVPSKSS-----RFFVHIGLEYMFTLWICFL 174
+PVN +S L K R++ SDIDKLSISN+ + S RF+ H+ + Y FT W C++
Sbjct: 121 VPVNWTSHGLQLAKLRNVTSSDIDKLSISNIENGSDSLYFGRFWTHLVMAYAFTFWTCYV 180
Query: 175 LYKEYDNIAMMRLHFLASQRRRVEQFT----------VVVRSVPHISGHSVSDSVESFFQ 224
L KEY+ +A MRL FL +++RR +QFT V+VR+VP S+SDSVE FF
Sbjct: 181 LMKEYEKVAAMRLAFLQNEQRRPDQFTNLGLSQLLSQVLVRNVPADPDESISDSVEHFFL 240
Query: 225 TNHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGR 284
NHP+HY+ HQ VYNAN A LV ++ QNWLDYYQLK+ R+ + +P I TG LGLWG+
Sbjct: 241 VNHPDHYLTHQVVYNANDLAALVEQKKSTQNWLDYYQLKYTRNQEHKPRIKTGFLGLWGK 300
Query: 285 KVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPT 344
KVDAI++Y I++L++ + ER+++ KD ++P AF+SF +RWGA+V AQTQQS +PT
Sbjct: 301 KVDAIDHYIAEIEKLNEQIMEERKKVKKDDTSVMPAAFVSFKTRWGAAVSAQTQQSSDPT 360
Query: 345 LWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLE 404
WLT+WAPE R+++W NLAIP+VSL++R+L++ ++ F L FF+MIPIAFVQSLA++EG+E
Sbjct: 361 EWLTEWAPEAREVFWSNLAIPYVSLTVRRLIMHIAFFFLTFFFMIPIAFVQSLASIEGIE 420
Query: 405 RVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAA 464
+ APFL+ +IE KS +QGFLPG+ LK+FL LP++LM+MSK EG ++ S+LER+ A
Sbjct: 421 KNAPFLKSIIENDLFKSVIQGFLPGIVLKLFLIFLPSILMVMSKFEGFVSLSSLERRAAF 480
Query: 465 KYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWA 524
+YY F L+NVFLGS++TG+AFEQL SFL QS +IP+T+GV+IP+KATFF+TYIMVDGWA
Sbjct: 481 RYYIFNLINVFLGSVITGSAFEQLDSFLKQSAKEIPKTVGVAIPIKATFFITYIMVDGWA 540
Query: 525 GIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVV 584
GIAGEILRLKPL+ +H+KN +VKTE+DR +AM+PG +++ T P +QLYFLLG+VYA V
Sbjct: 541 GIAGEILRLKPLIFFHIKNSLLVKTEKDREEAMNPGQINYHATEPRIQLYFLLGLVYAPV 600
Query: 585 TPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLL 644
TP+LLPFI++FFALAYLV+RHQIINVYNQ+YESAA FWP VH RII++L+I+Q+L++GLL
Sbjct: 601 TPVLLPFIIIFFALAYLVFRHQIINVYNQEYESAARFWPDVHGRIISALIIAQILLMGLL 660
Query: 645 STKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKA 704
STK AA+STP L LPI+TF FH+YC+ R+EPAF ++PL+EAM KD LE+ EP+ N+K
Sbjct: 661 STKGAAQSTPFLLFLPIITFFFHRYCKGRYEPAFLRHPLKEAMVKDTLERAREPNFNLKP 720
Query: 705 YLADAYLHPIFRSFEVE----EELVEVRVDKQPETQVASPTLSEPSSPSPLHDHVHQPS 759
YL AY+HP+F+ + E +E+ ++ E V PT + +P H + S
Sbjct: 721 YLQKAYIHPVFKDNDYEDSRFDEISGYCIEDSDEECVTVPTKRQSRINTPAVSHASRGS 779
>At1g62320 hypothetical protein
Length = 769
Score = 917 bits (2370), Expect = 0.0
Identities = 447/763 (58%), Positives = 588/763 (76%), Gaps = 23/763 (3%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MATLADIG++AAINILSA FLL FA+LRIQP NDRVYFPKWY+ G R+SP ++G FV K
Sbjct: 1 MATLADIGLAAAINILSALIFLLLFAILRIQPFNDRVYFPKWYLKGVRSSPVNSGAFVSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
+NL+F++Y+ FLNWMP AL+M E E+I+HAGLDSAV+LRIY +GLKIF P+ +++ IL
Sbjct: 61 IMNLDFRSYVRFLNWMPDALKMPEPELIDHAGLDSAVYLRIYLIGLKIFGPIALLSWSIL 120
Query: 121 IPVNVSSGALFFLK-RDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
+PVN +S L K R++ S+IDKLSISNV S RF+ H+ + Y FT W C++L KEY
Sbjct: 121 VPVNWTSDGLQLAKLRNVTSSNIDKLSISNVERGSDRFWAHLVMAYAFTFWTCYVLMKEY 180
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
+ IA MRL FL S++RR +QFTV+VR+VP S S+S++V+ FF NHP+HY+ HQ VYN
Sbjct: 181 EKIAAMRLSFLQSEKRRADQFTVLVRNVPPDSDESISENVQHFFLVNHPDHYLTHQVVYN 240
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
AN+ AKLV + ++QNWLDYYQLK+ R+ ++RP + G LGLWG+KVDA+++Y+ I++L
Sbjct: 241 ANELAKLVEDKKKMQNWLDYYQLKYTRNKEQRPRM--GFLGLWGKKVDAMDHYTAEIEKL 298
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
+ + ER+RI KD K ++ AF+SF +RWGA+VCAQTQQ+KNPT WLT+WAPE R++YW
Sbjct: 299 SEQIMEERKRIKKDDKSVMQAAFVSFKTRWGAAVCAQTQQTKNPTEWLTEWAPEAREMYW 358
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
NLA+P+VSL++R+ V+ ++ F L FF++IPIAFVQSLA++EG+E+ APFL P+++ K +
Sbjct: 359 PNLAMPYVSLTVRRFVMHIAFFFLTFFFIIPIAFVQSLASIEGIEKSAPFLSPIVKNKLM 418
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
KS +QGFLPG+ LK+FL LP +LMIMSK EG I+ S+LER+ A +YY F LVNVFLGS+
Sbjct: 419 KSLIQGFLPGIVLKLFLIFLPTILMIMSKFEGFISISSLERRAAFRYYIFNLVNVFLGSV 478
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
+TG+AFEQL SFL QS IPRT+GV+IP+KATFF+TYIMVDGWAG+AGEI RLKPLVI+
Sbjct: 479 ITGSAFEQLDSFLKQSANDIPRTVGVAIPIKATFFITYIMVDGWAGVAGEIFRLKPLVIF 538
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKN F VKTE+DR +AMDPG +DF T P +QLYFLLG+VYA VTP+LLPFI+ FF A
Sbjct: 539 HLKNFFFVKTEKDREEAMDPGQIDFYATEPRIQLYFLLGLVYAPVTPVLLPFIIFFFGFA 598
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
YLV+RH Q+YESA AFWP VH RII++L+ISQ+L+LGL+STK +STP L +L
Sbjct: 599 YLVFRH-------QKYESAGAFWPDVHGRIISALIISQILLLGLMSTKGKVQSTPFLLVL 651
Query: 660 PILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSFE 719
ILTF FH++C+ R+E AF PL+EAM KD LE+ EP+LN+K +L +AY+HP+F+ E
Sbjct: 652 AILTFGFHRFCKGRYESAFVINPLQEAMIKDTLERAREPNLNLKGFLQNAYVHPVFKDEE 711
Query: 720 VEEE-------------LVEVRVDKQPETQVASPTLSEPSSPS 749
+E +V+ + + T VAS S SS S
Sbjct: 712 DSDEEGLIEDSDDEDCVVVQTKRQRSRRTTVASSNASRGSSQS 754
>At4g15430 unknown protein
Length = 756
Score = 913 bits (2360), Expect = 0.0
Identities = 442/755 (58%), Positives = 582/755 (76%), Gaps = 7/755 (0%)
Query: 1 MATLADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGK 60
MAT+ DIGV+AAINI++AFAFLLAFA+ RIQP+NDRVYFPKWY+ G R+S TG F K
Sbjct: 1 MATINDIGVAAAINIVTAFAFLLAFAIFRIQPVNDRVYFPKWYLKGLRSSSIQTGGFGSK 60
Query: 61 FVNLNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLIL 120
F+NL+F++Y+ FLNWMP+AL+M E E+++HAGLDS V+LRIY LGLKIF P+ VA +
Sbjct: 61 FINLDFRSYIRFLNWMPEALKMPEPELVDHAGLDSVVYLRIYLLGLKIFFPIACVAFTTM 120
Query: 121 IPVNVSSGALFFLKR-DLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEY 179
+PVN ++ L L+ ++ SDIDKLS+SN+P+ S RF+VH+ + Y T W CF+L +EY
Sbjct: 121 VPVNWTNKGLDRLRHSNISFSDIDKLSLSNIPNGSPRFWVHLCMAYAITFWTCFILKREY 180
Query: 180 DNIAMMRLHFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVYN 239
NIA+MRL FLA+ +RR QFTV+VR++P S+ + VE FF+ NHP+HY+ QAV++
Sbjct: 181 QNIALMRLQFLANDQRRPNQFTVLVRNIPADPHESICELVEHFFKVNHPDHYLTFQAVHD 240
Query: 240 ANKFAKLVRRRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKEL 299
A K ++LV R ++QN LDY K R+ RP I G LG G + D I+YY+ ++
Sbjct: 241 ATKLSELVLTRKQMQNLLDYNINKHMRNLSNRPVIKMGFLGCCGEEADGIKYYTSVVE-- 298
Query: 300 DKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYW 359
++ E+QR+ K I+P AF+SF SRWGA+VCAQTQQ++NPT WLT+WA EPRDIY+
Sbjct: 299 ---ISEEKQRLRTGTKSIVPAAFVSFKSRWGAAVCAQTQQTRNPTEWLTEWAAEPRDIYY 355
Query: 360 RNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI 419
NLA+P+V L IR+L++ ++ F L FF+MIPIAFVQSLAN+EG+E+ PFL+P+IE+K +
Sbjct: 356 DNLALPYVDLKIRRLIVGVAYFFLTFFFMIPIAFVQSLANIEGIEKAFPFLKPLIEVKLL 415
Query: 420 KSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSI 479
KS +QGFLPG+ALKIFL LP +LM MSK EG ++ S+LER+ A ++Y F +NVFLGSI
Sbjct: 416 KSIIQGFLPGIALKIFLLFLPRILMQMSKFEGFVSTSSLERRAATRFYMFQFINVFLGSI 475
Query: 480 VTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIY 539
VTGTAF+QL+SFL+QS IP+TIGVSIPMKATFF+TYIMVDGWAG+AGEILRLKPL+IY
Sbjct: 476 VTGTAFQQLNSFLNQSANDIPKTIGVSIPMKATFFITYIMVDGWAGVAGEILRLKPLIIY 535
Query: 540 HLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALA 599
HLKN F+V+TE+DR +A DPG++ F P +QLYFLLG+VYA V+PILLPFILVFF LA
Sbjct: 536 HLKNSFLVRTEKDREEATDPGTIGFNTGEPQIQLYFLLGLVYAAVSPILLPFILVFFGLA 595
Query: 600 YLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAIL 659
++VYRHQ+INVYNQ+YESA FWP VH R++ +L++SQLL++GLLSTK A+KSTPLL +L
Sbjct: 596 FVVYRHQVINVYNQKYESAGKFWPDVHRRVVTALVVSQLLLMGLLSTKHASKSTPLLLVL 655
Query: 660 PILTFAFHKYCQHRFEPAFRKYPL-EEAMAKDLLEKTTEPDLNIKAYLADAYLHPIFRSF 718
P+LT FHK+C++R++PAF YPL +EAM KD L++ EP+LN+KA+L DAY HP FR
Sbjct: 656 PLLTIGFHKHCKNRYQPAFVTYPLQQEAMIKDTLDRIREPNLNLKAFLRDAYAHPEFRVG 715
Query: 719 EVEEELVEVRVDKQPETQVASPTLSEPSSPSPLHD 753
E E ++ D P VA+ S ++P P D
Sbjct: 716 EDPEPEEKLESDMSPPDLVATKRWSWRNTPLPSKD 750
>At1g69450 hypothetical protein
Length = 646
Score = 381 bits (978), Expect = e-106
Identities = 199/624 (31%), Positives = 344/624 (54%), Gaps = 9/624 (1%)
Query: 91 AGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGALFFLK-RDLVVSDIDKLSISN 149
+GLD VF+R+ T LK+F+ ++ + +L+PVN L + D + +D S++N
Sbjct: 4 SGLDGVVFMRMITFSLKVFLFAGIIGVFVLLPVNCFGDQLTVIDYADWSANSLDLFSVAN 63
Query: 150 VPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRRRVEQFTVVVRSVPH 209
+ +S +VH G Y+ T+++C LLY E+ IA+ R+ S + + EQFT++VR++P
Sbjct: 64 LKVRSQWLWVHFGAIYLVTVFVCCLLYFEFRYIALKRIEHFYSSKPKPEQFTILVRNIPS 123
Query: 210 ISGHSVSDSVESFFQTNHPNHYIGHQAVYNANKFAKLVRRRDRLQNWLDYYQLKFQRHPD 269
G SVSD+V+ FF NH + Y H ++ +K +V ++ D + ++
Sbjct: 124 SDGSSVSDTVDRFFGENHSSTYFSHVVIHRTSKLRSVVVSLEKAYLPSDKAKKLYKEVKH 183
Query: 270 KRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRW 329
K+P T + + RK + +Y ++E+++ + L Q + P + AF+SF SR+
Sbjct: 184 KKPVKKTP-MRFFSRKDNTEGHYESVLQEMEQNIRLG-QAEVSAPGKEVRAAFVSFKSRY 241
Query: 330 GASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMI 389
GA+ QS NPT WLT+ APEP D++W + F+ + K+++ + L +++
Sbjct: 242 GAATALHMPQSINPTYWLTEPAPEPHDVHWPFFSASFMQKWLAKILVVFACLLLTILFLV 301
Query: 390 PIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKI 449
P+ VQ L NL LE + PFL ++ +K + + G+LP L L+ L ++P + +S I
Sbjct: 302 PVVLVQGLTNLPALEFMFPFLSLILSMKVVSQIITGYLPSLILQTSLKVVPPTMEFLSSI 361
Query: 450 EGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPM 509
+GHI S +++ K +F + NVF ++ +G+AF +L L P QIP + V++P
Sbjct: 362 QGHICHSDIQKSACNKVIWFTIWNVFFATVFSGSAFYKLSVIL--DPKQIPLKLAVAVPA 419
Query: 510 KATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIP 569
+A+FF+ Y++ GW E+ R+ P ++ ++K F D + + P + + P
Sbjct: 420 QASFFIAYVVTTGWTDTLTELFRVVPFMVSYIKRSF---EPSDENEFVVP-PMRYHRDTP 475
Query: 570 SLQLYFLLGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRI 629
+ + LLGI Y + P++LPFIL++F LAY++YR+Q +NVY ++++ FWP +H +
Sbjct: 476 RVLFFGLLGITYFFLAPLILPFILLYFILAYIIYRNQFMNVYAPKFDTGGMFWPMIHYTM 535
Query: 630 IASLLISQLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAK 689
I SL++ Q + +GL + KK +T LL LP+ T F+++C+ RF P F YP E +
Sbjct: 536 IFSLVLMQAIAIGLFALKKMELATYLLVPLPVFTLLFNEFCRKRFMPIFTDYPAEVLTKR 595
Query: 690 DLLEKTTEPDLNIKAYLADAYLHP 713
D ++ L AY P
Sbjct: 596 DKEDRNDPTMPEFYNNLVSAYKDP 619
>At3g54510 putative protein
Length = 710
Score = 367 bits (941), Expect = e-101
Identities = 220/688 (31%), Positives = 384/688 (54%), Gaps = 42/688 (6%)
Query: 10 SAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVNLNFKTY 69
SA+INI A L F++L+ QP N VY+ + R S R + +L +
Sbjct: 9 SASINIGLAVVALWLFSVLKKQPRNAVVYYAR------RLSDRHHHRPLSLHSSLCLPRF 62
Query: 70 LTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGA 129
L + W+P+A R+ E+EI++ GLD+ V +R++ G++ F+ +++ +L+PV+ + +
Sbjct: 63 LPSVAWIPRAFRVPEDEILSRHGLDALVLIRLFKFGIRFFLMCSLLGASLLLPVDYYNES 122
Query: 130 LFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHF 189
+R+ +D +ISN+ S++ +VH + + + FLL+KEY I ++RL
Sbjct: 123 DLPTRREY---SMDAFTISNITRGSNKLWVHFSCLWCISFYALFLLHKEYKEILVIRLQQ 179
Query: 190 LASQRRRVEQFTVVVRSVPHISGHSVSD-SVESFFQTNHPNHYIGHQAVYNANKFAKLVR 248
+ R R +QFTV+VR VP H+ +V+ FF +H Y HQ +Y+ L+
Sbjct: 180 MKELRHRADQFTVLVRQVPLCPEHNTRGCAVDHFFSKHHRFSYHSHQMLYDGRDLEYLLG 239
Query: 249 RRDRLQNWLDYYQLKFQRHPDKRPTINTGVLGLWGRKVDAIEYYSHTIKELDKVMT-LER 307
++ +L+ L+ DKR +T +L ++ I ++E+ ++ L+
Sbjct: 240 KQKKLKKELE----------DKR---HTEILSNGSQEHKQISTSEEKLREITHMIYHLQS 286
Query: 308 QRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFV 367
+ ++++ + LPVAF++F SR A++ AQTQQ NP +T+ APEPRD+ WRNLAIP
Sbjct: 287 ETMLREKE--LPVAFVTFKSRRNAALAAQTQQHSNPLELITEMAPEPRDVSWRNLAIPQK 344
Query: 368 SLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFI---KSFLQ 424
L + K+ + L+ L F+ IP+ VQ +A E L++ P P + ++FI S +
Sbjct: 345 ILPLNKIGVILAAALLTIFFAIPVTAVQGIAKYEKLKKWFP---PAMAIEFIPGLSSVVT 401
Query: 425 GFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTA 484
G+LP LK F+YI+P ++ ++ + G I+ S E K +YF++ NVF S+++G+
Sbjct: 402 GYLPSAILKGFMYIIPFAMLGLAYLGGSISNSKEEIKACNMVFYFLMGNVFFLSLISGSL 461
Query: 485 FEQLHSFLHQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYHLKNM 544
+++ +L P IP + ++ +A FFMTYI+ DG +G + EIL+L L+++ +
Sbjct: 462 LDEIGEYL-THPRDIPSHLAAAVSAQAEFFMTYILTDGLSGFSLEILQL-GLILFDIIRS 519
Query: 545 FIVKTERDRGKAMDPGSVDFP--ETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAYLV 602
+ RGK P FP IP++ L ++G++YAVV P++LPF++ +F L Y+V
Sbjct: 520 YTY----GRGKERTPYLFSFPYFRVIPTVSLSIMIGMIYAVVAPLMLPFLVGYFCLGYIV 575
Query: 603 YRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILPIL 662
Y +Q +VY Y++ FWP +H I S+++ Q+ M+GL K + L ++
Sbjct: 576 YFNQ--DVYETTYDTCGRFWPFIHHYIFVSIILMQITMVGLFGLKSKPSAAIATVPLILI 633
Query: 663 TFAFHKYCQHRFEPAFRKYPLEEAMAKD 690
T A+++YC+ RF P+F+ +P++ A+ D
Sbjct: 634 TIAYNEYCKIRFLPSFKHFPIQTAVEID 661
>At1g30360 ERD4 protein (ERD4)
Length = 724
Score = 347 bits (889), Expect = 2e-95
Identities = 210/648 (32%), Positives = 341/648 (52%), Gaps = 30/648 (4%)
Query: 75 WMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGALFFLK 134
WM +AL SE +++N +G+D+AV + L IF +++ L L+P+ + + K
Sbjct: 61 WMREALTSSEQDVVNLSGVDTAVHFVFLSTVLGIFACSSLLLLPTLLPLAATDNNIKNTK 120
Query: 135 RDL------VVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLH 188
S +D LS++N+ KSSR + +G Y +L F L+K Y +++ +R
Sbjct: 121 NATDTTSKGTFSQLDNLSMANITKKSSRLWAFLGAVYWISLVTYFFLWKAYKHVSSLRAQ 180
Query: 189 FLASQRRRVEQFTVVVRSVPHI-SGHSVSDSVESFFQTNHPNHYIGHQAVYNANKFAKLV 247
L S + EQF ++VR +P G + + ++S+F+ +P + +K K+
Sbjct: 181 ALMSADVKPEQFAILVRDMPAPPDGQTQKEFIDSYFREIYPETFYRSLVATENSKVNKIW 240
Query: 248 RRRDRLQNWLDYYQLKFQRHP------DKRPTINTGVLGLWGRKVDAIEYYSHTIKELDK 301
+ L+ Y+ K R + RPT TG GL G++VD+IEYY+ I E
Sbjct: 241 EK-------LEGYKKKLARAEAILAATNNRPTNKTGFCGLVGKQVDSIEYYTELINESVA 293
Query: 302 VMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRN 361
+ E++ ++ + + V F F++R A+ AQ+ + W APEPR + W+N
Sbjct: 294 KLETEQKAVLAEKQQTAAVVF--FTTRVAAASAAQSLHCQMVDKWTVTEAPEPRQLLWQN 351
Query: 362 LAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKS 421
L I S IR+ I V + FYMIPIAFV ++ L+ L+R+ PF++PV+E+ I++
Sbjct: 352 LNIKLFSRIIRQYFIYFFVAVTILFYMIPIAFVSAITTLKNLQRIIPFIKPVVEITAIRT 411
Query: 422 FLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVT 481
L+ FLP +AL +FL +LP +L+ +SK EG ++S R + KY+YF + NVF+G +
Sbjct: 412 VLESFLPQIALIVFLAMLPKLLLFLSKAEGIPSQSHAIRAASGKYFYFSVFNVFIGVTLA 471
Query: 482 GTAFEQLHSFL-HQSPTQIPRTIGVSIPMKATFFMTYIMVDGWAGIAGEILRLKPLVIYH 540
GT F + + I + S+P ATFF+TY+ + + G E+ R+ PL+I+H
Sbjct: 472 GTLFNTVKDIAKNPKLDMIINLLATSLPKSATFFLTYVALKFFIGYGLELSRIIPLIIFH 531
Query: 541 LKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPILLPFILVFFALAY 600
LK ++ KTE + +A PG + + +P L + Y+V+ P++L F + +F L +
Sbjct: 532 LKKKYLCKTEAEVKEAWYPGDLSYATRVPGDMLILTITFCYSVIAPLILIFGITYFGLGW 591
Query: 601 LVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKKAAKSTPLLAILP 660
LV R+Q + VY YES WPH+H RI+A+L + Q++M G L K T L+ L
Sbjct: 592 LVLRNQALKVYVPSYESYGRMWPHIHQRILAALFLFQVVMFGYLGA-KTFFYTALVIPLI 650
Query: 661 ILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLN--IKAYL 706
I + F C+ +F F LE A E PDL +AY+
Sbjct: 651 ITSLIFGYVCRQKFYGGFEHTALEVACR----ELKQSPDLEEIFRAYI 694
>At3g01100 unknown protein
Length = 468
Score = 309 bits (792), Expect = 3e-84
Identities = 151/437 (34%), Positives = 259/437 (58%), Gaps = 9/437 (2%)
Query: 277 GVLGLWGRKVDAIEYYSHTIKELDKVMTLERQRIIKDPKCILPVAFLSFSSRWGASVCAQ 336
G LG++G VD +++Y + +L+ M L++ + + +P AF+SF +R GA++
Sbjct: 28 GFLGMFGNNVDVVDHYQKKLDKLEDDMRLKQSLLAGEE---VPAAFVSFRTRHGAAIATN 84
Query: 337 TQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQS 396
QQ +PT WLT+ APEP D++W FV I +V+ ++ AL+ Y++P+ VQ
Sbjct: 85 IQQGIDPTQWLTEAAPEPEDVHWPFFTASFVRRWISNVVVLVAFVALLILYIVPVVLVQG 144
Query: 397 LANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYILPAVLMIMSKIEGHIAKS 456
LANL LE PFL+ ++ +K + + G+LP L ++FL I+P +++++S ++G I+ S
Sbjct: 145 LANLHQLETWFPFLKGILNMKIVSQVITGYLPSLIFQLFLLIVPPIMLLLSSMQGFISHS 204
Query: 457 TLERKTAAKYYYFMLVNVFLGSIVTGTAFEQLHSFLHQSPTQIPRTIGVSIPMKATFFMT 516
+E+ K F + N F ++++G+A +++ FL P IPR + ++P +A+FF++
Sbjct: 205 QIEKSACIKLLIFTVWNSFFANVLSGSALYRVNVFL--EPKTIPRVLAAAVPAQASFFVS 262
Query: 517 YIMVDGWAGIAGEILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFL 576
Y++ GW G++ EILRL PL+ + +F ++ K + S F + IP + + L
Sbjct: 263 YVVTSGWTGLSSEILRLVPLLWSFITKLF----GKEDDKEFEVPSTPFCQEIPRILFFGL 318
Query: 577 LGIVYAVVTPILLPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLIS 636
LGI Y ++P++LPF+LV++ L Y++YR+Q++NVY +YE+ FWP VHS I SL++
Sbjct: 319 LGITYFFLSPLILPFLLVYYCLGYIIYRNQLLNVYAAKYETGGKFWPIVHSYTIFSLVLM 378
Query: 637 QLLMLGLLSTKKAAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTT 696
++ +GL K+ ++ L LP+LT F YCQ RF P F+ YP + + KD ++
Sbjct: 379 HIIAVGLFGLKELPVASSLTIPLPVLTVLFSIYCQRRFLPNFKSYPTQCLVNKDKADERE 438
Query: 697 EPDLNIKAYLADAYLHP 713
+ + L AY P
Sbjct: 439 QNMSEFYSELVVAYRDP 455
>At1g10080 hypothetical protein
Length = 509
Score = 265 bits (676), Expect = 1e-70
Identities = 146/422 (34%), Positives = 233/422 (54%), Gaps = 34/422 (8%)
Query: 319 PVAFLSFSSRWGASVCAQTQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISL 378
PVAF+ F SR+ A V ++ Q+ NP LW+ D APEP D++WRNL IP+ L +R++ +
Sbjct: 14 PVAFVFFKSRYDALVVSEVLQTPNPMLWVADLAPEPHDVHWRNLRIPYRQLWMRRIATLV 73
Query: 379 SVFALVFFYMIPIAFVQSLANLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYI 438
A +F ++ P+ FVQ L L L + PFL+ ++ +F++ + G+LP + L +F Y
Sbjct: 74 GAIAFMFVFLFPVTFVQGLTQLPTLSKNFPFLKDLLNRRFMEQVITGYLPSVILVLFFYT 133
Query: 439 LPAVLMIMSKIEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFEQ--LHSFLHQSP 496
+P ++M S +EG +++S ++ K YF + NVF +I++G+ Q + + + P
Sbjct: 134 VPPLMMYFSTLEGCVSRSQRKKSACLKILYFTIWNVFFVNILSGSVIRQFTVLNSVRDVP 193
Query: 497 TQ----IPRTIGVSIP------------------------MKATFFMTYIMVDGWAGIAG 528
Q +P + S+P M+A FFMTY GWAG+A
Sbjct: 194 AQLAKLVPAQVIFSVPFTRVTIFNFVIFYKANGMKLFIDYMQAGFFMTYCFTSGWAGLAC 253
Query: 529 EILRLKPLVIYHLKNMFIVKTERDRGKAMDPGSVDFPETIPSLQLYFLLGIVYAVVTPIL 588
EI++ L I++L IVK + + + + + IP L L+ LLG +V+ P++
Sbjct: 254 EIMQPVGL-IWNLIAKVIVKNKEESYETL---RFPYHTEIPRLLLFGLLGFTNSVIAPLI 309
Query: 589 LPFILVFFALAYLVYRHQIINVYNQQYESAAAFWPHVHSRIIASLLISQLLMLGLLSTKK 648
LPF+L++F AYL+Y++QIINVY +YES +WP H+ I SL++SQ++ LG K
Sbjct: 310 LPFLLIYFFFAYLIYKNQIINVYITKYESGGQYWPVFHNTTIFSLILSQVIALGFFGLKL 369
Query: 649 AAKSTPLLAILPILTFAFHKYCQHRFEPAFRKYPLEEAMAKDLLEKTTEPDLNIKAYLAD 708
+ ++ L +LT F +YC+ RF P F+KYP E +A D ++ T I L
Sbjct: 370 STVASGFTIPLILLTLLFSEYCRQRFAPIFQKYPAEILIAMDRADEMTGKMEEIHNNLKV 429
Query: 709 AY 710
AY
Sbjct: 430 AY 431
>At1g10090 hypothetical protein
Length = 246
Score = 104 bits (260), Expect = 2e-22
Identities = 67/231 (29%), Positives = 114/231 (49%), Gaps = 16/231 (6%)
Query: 10 SAAINILSAFAFLLAFALLRIQPINDRVYFPKWYING--ERTSPRSTGNFVGKFVNLNFK 67
SA INI + +++LR QP N VYF + +G +R PR ++
Sbjct: 9 SAGINIAICVVLVSLYSILRKQPANYCVYFGRLLSDGRVKRHDPRW------------YE 56
Query: 68 TYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSS 127
+ +W+ +A +E E++ AGLD+ VF+R+ ++IF V VV L ++PVN
Sbjct: 57 RFAPSPSWLVKAWETTEEEMLAAAGLDAVVFIRMVICSIRIFSIVAVVCLAFVLPVNYYG 116
Query: 128 GALFFLKRDLVVSDIDKLSISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRL 187
+ +++ + + +I N+ +S +VH Y+ + C LLY EY NIA RL
Sbjct: 117 QKMEH--KEVHLESLGVFTIENLNPRSRWLWVHCLSLYIISSAACALLYFEYKNIAKKRL 174
Query: 188 HFLASQRRRVEQFTVVVRSVPHISGHSVSDSVESFFQTNHPNHYIGHQAVY 238
++ + FTV++R++P S S++V +F + Y+ H VY
Sbjct: 175 AHISGSASKPSHFTVLIRAIPQSPDQSYSETVSKYFTNYYAPSYVSHLMVY 225
>At4g35870 putative protein
Length = 817
Score = 84.0 bits (206), Expect = 3e-16
Identities = 101/458 (22%), Positives = 188/458 (40%), Gaps = 75/458 (16%)
Query: 86 EIINHAGLDSAVFLRIYTLGLKIFVPVTVVALLILIPVNVSSGALFFLKRDLVVSDIDKL 145
EI H G D+A FL I + + V+A+ +++P+N+ +G L+ ++ K
Sbjct: 92 EIARHCGADAAQFLLIEGGSFVLLFSIAVLAVSVMLPLNLYAGTA------LLSDELSKT 145
Query: 146 SISNVPSKSSRFFVHIGLEYMFTLWICFLLYKEYDNIAMMRLHFLASQRR---------- 195
I+++ S+ ++H +++ ++ + IA + ++ R
Sbjct: 146 MITHIQKGSALLWLHF-------VFVVIVVVISHFGIAAIEARLKFTRFRDGNGNISDPN 198
Query: 196 --RVEQFTVVVRSVPHISGHSVSDSVE--SFFQTNHPNH---YIGHQAVYNANKFA-KLV 247
FT++V+ +P G SD VE F+ +P +I + + A +LV
Sbjct: 199 ANSTAVFTIMVQGLPKNLG---SDRVEFEDCFRLKYPGKVYKFIVPMDLCALDDLATELV 255
Query: 248 RRRDRLQNWLDYYQLKFQRHPDKRPTINTG-----VLGLWGRKVDAIEYYSHTIKELDKV 302
R RD + WL ++ + PD+ + V LW R + D
Sbjct: 256 RVRDEI-TWL-VAKMDSRLLPDEYENVGDNGLVFCVCSLWVRVKVLWSQITERFGFTDDE 313
Query: 303 MTLERQRIIKDPKCILP-----------VAFLSFSSRWGASVCAQ--------------- 336
+ Q + D + L VAF+ F + A+ Q
Sbjct: 314 KLRKLQELRADLESQLAAYKEGRAQGAGVAFVMFKDVYTANKAVQDFRNERSRRTGKFFS 373
Query: 337 -TQQSKNPTLWLTDWAPEPRDIYWRNLAIPFVSLSIRKLVISLSVFALVFFYMIPIAFVQ 395
T+ W D AP DIYW +L + V+L +R+++++ + ++ F+ P+A +
Sbjct: 374 VTELRLQRNQWKVDRAPLATDIYWNHLGLTKVALIVRRVIVNTILLLILVFFSSPLALIS 433
Query: 396 SLA------NLEGLERVAPFLRPVIELKFIKSFLQGFLPGLALKIFLYI-LPAVLMIMSK 448
+L N E L+ +L V +I S + FLP + + + +YI +P+ L +SK
Sbjct: 434 ALVSAGRIFNAEALDSAQYWLTWVQTSGWIGSLIFQFLPNVFIFVSMYIVIPSALSYLSK 493
Query: 449 IEGHIAKSTLERKTAAKYYYFMLVNVFLGSIVTGTAFE 486
E H+ S +R K F LVN+ + + ++ E
Sbjct: 494 FERHLTVSGEQRAALLKMVCFFLVNLIILKALVESSLE 531
>At3g01110 hypothetical protein
Length = 203
Score = 48.9 bits (115), Expect = 1e-05
Identities = 30/103 (29%), Positives = 52/103 (50%), Gaps = 9/103 (8%)
Query: 4 LADIGVSAAINILSAFAFLLAFALLRIQPINDRVYFPKWYINGERTSPRSTGNFVGKFVN 63
L+ + S IN+ F F +++LR QP N VY P+ + + S +S
Sbjct: 82 LSALLTSVGINLGLCFLFFTLYSILRKQPSNVTVYGPR-LVKKDGKSQQSN--------E 132
Query: 64 LNFKTYLTFLNWMPQALRMSENEIINHAGLDSAVFLRIYTLGL 106
N + L W+ +AL + +EI+++ GLD+ VF+R++ L
Sbjct: 133 FNLERLLPTAGWVKRALEPTNDEILSNLGLDALVFIRVFVFRL 175
>At1g12040 leucine-rich repeat/extensin 1 (LRX1)
Length = 744
Score = 43.1 bits (100), Expect = 6e-04
Identities = 23/51 (45%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P SPSP +V+ PP +V YS P PP Y Y PPPP Y Y P
Sbjct: 461 PPSPSPPPPYVYSSPPPPYV--YSSPP--PPPYVYSSPPPPPYVYSSPPPP 507
Score = 33.1 bits (74), Expect = 0.63
Identities = 24/82 (29%), Positives = 28/82 (33%), Gaps = 26/82 (31%)
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQY----SHYPTSPPGYYY---------------H 780
P P PSP++ PSPP Y + P P YYY H
Sbjct: 637 PVTPSPPPPSPVYYPPVTPSPPPPSPVYYPSETQSPPPPTEYYYSPSQSPPPTKACKEGH 696
Query: 781 PP-------PPPHYAYQYQDEP 795
PP PPP Y+Y P
Sbjct: 697 PPQATPSYEPPPEYSYSSSPPP 718
Score = 32.7 bits (73), Expect = 0.82
Identities = 21/65 (32%), Positives = 24/65 (36%), Gaps = 9/65 (13%)
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQY----SHYPTSPPGYYY-----HPPPPPHYAYQ 790
P P PSP++ PSPP Y + P P YY PPPP Y Y
Sbjct: 622 PVTPSPPPPSPVYYPPVTPSPPPPSPVYYPPVTPSPPPPSPVYYPSETQSPPPPTEYYYS 681
Query: 791 YQDEP 795
P
Sbjct: 682 PSQSP 686
Score = 32.3 bits (72), Expect = 1.1
Identities = 17/40 (42%), Positives = 21/40 (52%), Gaps = 6/40 (15%)
Query: 746 SSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPP 785
SSP P +V+ PP +V Y + PP Y Y PPPP
Sbjct: 492 SSPPP-PPYVYSSPPPPYV-----YSSPPPPYVYSSPPPP 525
Score = 32.3 bits (72), Expect = 1.1
Identities = 24/71 (33%), Positives = 27/71 (37%), Gaps = 8/71 (11%)
Query: 733 PETQVASPTLSEPSSPSPL-HDHVHQPSPPQHVNQYSHYPTSPP---GYYYHP----PPP 784
P +P P PSP+ + V Q PP Y SPP YY P PPP
Sbjct: 540 PPVVYYAPVTQSPPPPSPVYYPPVTQSPPPPSPVYYPPVTNSPPPPSPVYYPPVTYSPPP 599
Query: 785 PHYAYQYQDEP 795
P Y Q P
Sbjct: 600 PSPVYYPQVTP 610
Score = 31.2 bits (69), Expect = 2.4
Identities = 20/58 (34%), Positives = 25/58 (42%), Gaps = 8/58 (13%)
Query: 740 PTLSEPSSPSPLH--DHVHQPSPPQHV--NQYSHYPTSPPGYYYHP----PPPPHYAY 789
P + P PSP++ + P PP V Q + P P YY P PPPP Y
Sbjct: 577 PVTNSPPPPSPVYYPPVTYSPPPPSPVYYPQVTPSPPPPSPLYYPPVTPSPPPPSPVY 634
Score = 30.4 bits (67), Expect = 4.1
Identities = 23/64 (35%), Positives = 26/64 (39%), Gaps = 11/64 (17%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPG---YYYHP----PPPP 785
P +SP PS P P + SPP V Y+ SPP YY P PPPP
Sbjct: 515 PPYVYSSPPPPPPSPPPPCPES----SPPPPVVYYAPVTQSPPPPSPVYYPPVTQSPPPP 570
Query: 786 HYAY 789
Y
Sbjct: 571 SPVY 574
Score = 30.4 bits (67), Expect = 4.1
Identities = 20/58 (34%), Positives = 23/58 (39%), Gaps = 8/58 (13%)
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQY----SHYPTSPPGYYYHP----PPPPHYAY 789
P P PSP++ PSPP Y + P P YY P PPPP Y
Sbjct: 592 PVTYSPPPPSPVYYPQVTPSPPPPSPLYYPPVTPSPPPPSPVYYPPVTPSPPPPSPVY 649
Score = 30.0 bits (66), Expect = 5.3
Identities = 20/64 (31%), Positives = 24/64 (37%), Gaps = 1/64 (1%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYH-PPPPPHYAYQY 791
P + SP S SP P PP + +SPP Y + PPPP Y Y
Sbjct: 434 PSVRAYSPPPPPYSKMSPSVRAYPPPPPPSPSPPPPYVYSSPPPPYVYSSPPPPPYVYSS 493
Query: 792 QDEP 795
P
Sbjct: 494 PPPP 497
Score = 30.0 bits (66), Expect = 5.3
Identities = 19/69 (27%), Positives = 24/69 (34%), Gaps = 6/69 (8%)
Query: 733 PETQVASPTLSEPSSPSPLHDHV------HQPSPPQHVNQYSHYPTSPPGYYYHPPPPPH 786
P + SPT+ S P P + + P PP + PP P PPP
Sbjct: 410 PPSSKMSPTVRAYSPPPPPSSKMSPSVRAYSPPPPPYSKMSPSVRAYPPPPPPSPSPPPP 469
Query: 787 YAYQYQDEP 795
Y Y P
Sbjct: 470 YVYSSPPPP 478
Score = 29.6 bits (65), Expect = 6.9
Identities = 18/50 (36%), Positives = 23/50 (46%), Gaps = 5/50 (10%)
Query: 740 PTLSEPS-SPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYY-HPPPPPHY 787
P + PS P P + + P PP S++P P Y PPPPP Y
Sbjct: 697 PPQATPSYEPPPEYSYSSSPPPPSPT---SYFPPMPSVSYDASPPPPPSY 743
Score = 29.6 bits (65), Expect = 6.9
Identities = 20/58 (34%), Positives = 24/58 (40%), Gaps = 9/58 (15%)
Query: 740 PTLSEPSSPSPLH--DHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQYQDEP 795
P P PSP++ + P PP V +YP P Y PPP P Y Q P
Sbjct: 562 PVTQSPPPPSPVYYPPVTNSPPPPSPV----YYP---PVTYSPPPPSPVYYPQVTPSP 612
>At4g18670 extensin-like protein
Length = 839
Score = 42.0 bits (97), Expect = 0.001
Identities = 22/64 (34%), Positives = 29/64 (44%), Gaps = 14/64 (21%)
Query: 742 LSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPP---HYA-------YQY 791
+S P P+P +H P P H + P PP +Y+PPPPP HY+ Y Y
Sbjct: 733 ISPPPPPTP----IHSPPPQSHPPCIEYSPPPPPTVHYNPPPPPSPAHYSPPPSPPVYYY 788
Query: 792 QDEP 795
P
Sbjct: 789 NSPP 792
Score = 41.2 bits (95), Expect = 0.002
Identities = 25/54 (46%), Positives = 29/54 (53%), Gaps = 9/54 (16%)
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHY--PTSPPGYYYH-PPPPP--HYA 788
P E S P P H + P PP +HY P SPP YYY+ PPPPP HY+
Sbjct: 751 PPCIEYSPPPPPTVHYNPPPPPSP----AHYSPPPSPPVYYYNSPPPPPAVHYS 800
Score = 40.4 bits (93), Expect = 0.004
Identities = 19/44 (43%), Positives = 24/44 (54%), Gaps = 3/44 (6%)
Query: 742 LSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPP 785
L+ P P+P + QP PP H YS P +P +Y PPPPP
Sbjct: 699 LASPPPPAPYYYSSPQPPPPPH---YSLPPPTPTYHYISPPPPP 739
Score = 39.7 bits (91), Expect = 0.007
Identities = 19/41 (46%), Positives = 23/41 (55%), Gaps = 4/41 (9%)
Query: 745 PSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPP 785
P PSP H + PSPP + Y + P PP +Y PPPPP
Sbjct: 769 PPPPSPAH-YSPPPSPPVY---YYNSPPPPPAVHYSPPPPP 805
Score = 33.5 bits (75), Expect = 0.48
Identities = 18/42 (42%), Positives = 21/42 (49%), Gaps = 6/42 (14%)
Query: 748 PSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYH--PPPPPHY 787
PSP + + P PP V HY PP +H PPPPP Y
Sbjct: 781 PSPPVYYYNSPPPPPAV----HYSPPPPPVIHHSQPPPPPIY 818
Score = 30.0 bits (66), Expect = 5.3
Identities = 16/48 (33%), Positives = 19/48 (39%), Gaps = 3/48 (6%)
Query: 740 PTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P + P P+ H P PP + P P Y PPPPP Y
Sbjct: 795 PAVHYSPPPPPVIHHSQPPPPPIYEGPL---PPIPGISYASPPPPPFY 839
Score = 29.3 bits (64), Expect = 9.0
Identities = 15/43 (34%), Positives = 22/43 (50%), Gaps = 5/43 (11%)
Query: 748 PSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPP---HY 787
P+P + ++ P PP + +S P S P + PPPP HY
Sbjct: 726 PTPTYHYISPPPPPTPI--HSPPPQSHPPCIEYSPPPPPTVHY 766
>At3g16510 unknown protein
Length = 360
Score = 41.6 bits (96), Expect = 0.002
Identities = 25/69 (36%), Positives = 32/69 (46%), Gaps = 10/69 (14%)
Query: 733 PETQVASPTLSEPSSPSPLHDH-------VHQPSPPQHVNQYS--HYPTSPPGYYYHPPP 783
P +SP P+ P H H V+ P PP N Y +Y TSPP + +PPP
Sbjct: 213 PPQHYSSPPYPYPN-PYQYHSHYPEQPVAVYPPPPPSASNLYPPPYYSTSPPQHQSYPPP 271
Query: 784 PPHYAYQYQ 792
P H +Q Q
Sbjct: 272 PGHSFHQTQ 280
Score = 35.0 bits (79), Expect = 0.16
Identities = 24/69 (34%), Positives = 29/69 (41%), Gaps = 14/69 (20%)
Query: 735 TQVASPTLSEPSSPSPLHDHVHQPS-PPQHVNQ-----------YSHYPTSPPGYYYHPP 782
T P SE + PL + PS PPQH + +SHYP P Y PP
Sbjct: 187 TNYPPPQSSEANFYPPLSSIGYPPSSPPQHYSSPPYPYPNPYQYHSHYPEQPVAVY--PP 244
Query: 783 PPPHYAYQY 791
PPP + Y
Sbjct: 245 PPPSASNLY 253
Score = 32.0 bits (71), Expect = 1.4
Identities = 21/60 (35%), Positives = 26/60 (43%), Gaps = 6/60 (10%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPP---GYYYHPPPPPHYAY 789
P SP + P P H HQ P Q + ++ P+SP GY Y PP P Y Y
Sbjct: 255 PPYYSTSPPQHQSYPPPPGHSF-HQTQPSQSFHGFA--PSSPQNQHGYGYPPPTSPGYGY 311
>At1g76930 extensin
Length = 373
Score = 41.6 bits (96), Expect = 0.002
Identities = 25/76 (32%), Positives = 31/76 (39%), Gaps = 13/76 (17%)
Query: 733 PETQVASPTLSEPSSPSPLHDH-----VHQPSPPQHVNQ---YSHYPT-----SPPGYYY 779
P + P + S P P+H H P PP H + H P SPP Y
Sbjct: 260 PPVHYSPPPVVYHSPPPPVHYSPPPVVYHSPPPPVHYSPPPVVYHSPPPPVHYSPPPVVY 319
Query: 780 HPPPPPHYAYQYQDEP 795
H PPPP Y+Y+ P
Sbjct: 320 HSPPPPKKHYEYKSPP 335
Score = 37.4 bits (85), Expect = 0.033
Identities = 20/61 (32%), Positives = 26/61 (41%), Gaps = 12/61 (19%)
Query: 733 PETQVASPTLSEPSSPSPLHDH-----VHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P + P + S P P+H H P PP H + PP Y+ PPPP HY
Sbjct: 244 PPVHYSPPPVVYHSPPPPVHYSPPPVVYHSPPPPVHYSP-------PPVVYHSPPPPVHY 296
Query: 788 A 788
+
Sbjct: 297 S 297
Score = 37.0 bits (84), Expect = 0.043
Identities = 19/46 (41%), Positives = 20/46 (43%), Gaps = 9/46 (19%)
Query: 746 SSPSPLHDH----VHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
S P P+H H P PP H HY Y Y PPPPHY
Sbjct: 333 SPPPPVHYSPPTVYHSPPPPVH-----HYSPPHQPYLYKSPPPPHY 373
Score = 35.8 bits (81), Expect = 0.097
Identities = 25/70 (35%), Positives = 29/70 (40%), Gaps = 14/70 (20%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSHYPT----------SPPGYYY 779
P + SP S P P+ + P SPP V YS P SPP Y
Sbjct: 196 PPVKYYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPPVYKSPPPPVHYSPPPVVY 255
Query: 780 HPPPPP-HYA 788
H PPPP HY+
Sbjct: 256 HSPPPPVHYS 265
Score = 34.7 bits (78), Expect = 0.22
Identities = 19/64 (29%), Positives = 28/64 (43%), Gaps = 3/64 (4%)
Query: 733 PETQVASPTLSEPSSPSPL-HDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYAYQY 791
P + P + S P P H P PP H + + Y + PP +++ PPH Y Y
Sbjct: 308 PPVHYSPPPVVYHSPPPPKKHYEYKSPPPPVHYSPPTVYHSPPPPVHHY--SPPHQPYLY 365
Query: 792 QDEP 795
+ P
Sbjct: 366 KSPP 369
Score = 33.1 bits (74), Expect = 0.63
Identities = 23/63 (36%), Positives = 28/63 (43%), Gaps = 7/63 (11%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSH---YPTSPPGYYYHPPPP-P 785
P + SP S P P+ + P SPP V YS Y + PP Y PPPP
Sbjct: 44 PPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPPVYKSPPPPVYKSPPPPVK 103
Query: 786 HYA 788
HY+
Sbjct: 104 HYS 106
Score = 31.6 bits (70), Expect = 1.8
Identities = 21/58 (36%), Positives = 24/58 (41%), Gaps = 8/58 (13%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSHYPTSPPGYYYHPPPPPHY 787
P + SP S P P+ + P SPP V YS PP Y PPPP Y
Sbjct: 116 PPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKHYS-----PPPVYKSPPPPVKY 168
Score = 31.2 bits (69), Expect = 2.4
Identities = 22/60 (36%), Positives = 23/60 (37%), Gaps = 8/60 (13%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP----SPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P SP SP P H P SPP V YS PP Y PPP HY+
Sbjct: 83 PPPVYKSPPPPVYKSPPPPVKHYSPPPVYKSPPPPVKHYS----PPPVYKSPPPPVKHYS 138
Score = 31.2 bits (69), Expect = 2.4
Identities = 23/69 (33%), Positives = 26/69 (37%), Gaps = 14/69 (20%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSHYPT-----------SPPGYY 778
P + SP S P P+ + P SPP V YS P SPP Y
Sbjct: 132 PPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPPVYKSPPPPVKHYSPPPVY 191
Query: 779 YHPPPPPHY 787
PPPP Y
Sbjct: 192 KSPPPPVKY 200
Score = 30.8 bits (68), Expect = 3.1
Identities = 20/56 (35%), Positives = 24/56 (42%), Gaps = 9/56 (16%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQPSPPQHVNQYSHYPTSPPGYYYHPPPPPHYA 788
P + SP S P P++ SPP V YS PP Y PPP HY+
Sbjct: 76 PPVKYYSPPPVYKSPPPPVYK-----SPPPPVKHYS----PPPVYKSPPPPVKHYS 122
Score = 30.4 bits (67), Expect = 4.1
Identities = 20/58 (34%), Positives = 25/58 (42%), Gaps = 6/58 (10%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSH---YPTSPPGYYYHPPPP 784
P + SP S P P+ + P SPP V YS Y + PP Y+ PPP
Sbjct: 28 PPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPP 85
Score = 29.6 bits (65), Expect = 6.9
Identities = 20/58 (34%), Positives = 25/58 (42%), Gaps = 6/58 (10%)
Query: 733 PETQVASPTLSEPSSPSPLHDHVHQP---SPPQHVNQYSH---YPTSPPGYYYHPPPP 784
P + SP S P P+ + P SPP V YS Y + PP Y+ PPP
Sbjct: 148 PPVKHYSPPPVYKSPPPPVKYYSPPPVYKSPPPPVKHYSPPPVYKSPPPPVKYYSPPP 205
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,428,414
Number of Sequences: 26719
Number of extensions: 776497
Number of successful extensions: 6949
Number of sequences better than 10.0: 153
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 85
Number of HSP's that attempted gapping in prelim test: 3417
Number of HSP's gapped (non-prelim): 1362
length of query: 795
length of database: 11,318,596
effective HSP length: 107
effective length of query: 688
effective length of database: 8,459,663
effective search space: 5820248144
effective search space used: 5820248144
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0020.6


