

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
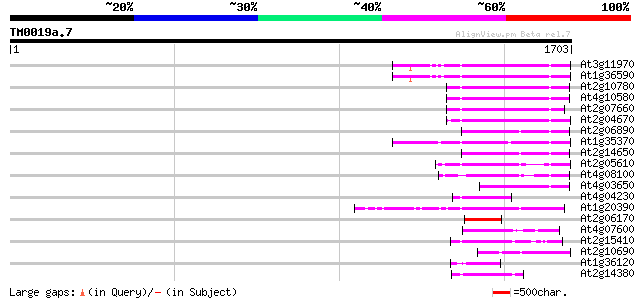
Query= TM0019a.7
(1703 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g11970 hypothetical protein 192 2e-48
At1g36590 hypothetical protein 192 2e-48
At2g10780 pseudogene 181 3e-45
At4g10580 putative reverse-transcriptase -like protein 180 6e-45
At2g07660 putative retroelement pol polyprotein 177 4e-44
At2g04670 putative retroelement pol polyprotein 177 4e-44
At2g06890 putative retroelement integrase 170 7e-42
At1g35370 hypothetical protein 159 1e-38
At2g14650 putative retroelement pol polyprotein 156 1e-37
At2g05610 putative retroelement pol polyprotein 151 3e-36
At4g08100 putative polyprotein 138 3e-32
At4g03650 putative reverse transcriptase 126 9e-29
At4g04230 putative transposon protein 118 3e-26
At1g20390 hypothetical protein 111 4e-24
At2g06170 putative Ty3-gypsy-like retroelement pol polyprotein 110 5e-24
At4g07600 102 2e-21
At2g15410 putative retroelement pol polyprotein 98 5e-20
At2g10690 putative retroelement pol polyprotein 97 1e-19
At1g36120 putative reverse transcriptase gb|AAD22339.1 94 7e-19
At2g14380 putative retroelement pol polyprotein 90 1e-17
>At3g11970 hypothetical protein
Length = 1499
Score = 192 bits (487), Expect = 2e-48
Identities = 149/562 (26%), Positives = 265/562 (46%), Gaps = 43/562 (7%)
Query: 1162 KLSAAEGSRLKIKYKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIKKL-------- 1213
++S A+G +L+++ K+ K + ++ LL+ +++G +++ L
Sbjct: 433 RVSVADGRKLRVEGKVTDFSWKLQTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFK 492
Query: 1214 -----FPYNTDE---NGIT---VQHLGQPILFKFSEPPIDKTLNVISYKEKQINFLKEEI 1262
F +N + +G+T V+ + L K E + L ++ +E + E
Sbjct: 493 KLEMRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQEDQVQ--LAMLCVQEVSESTEGELC 550
Query: 1263 SYRTIEEQLQQPSVKSRIEDILENIQSSICSD--LPNAFWERKSHMVELPYEKDFSDKQI 1320
+ + +L + SV +E++L LP F E+ +H ++L +
Sbjct: 551 TINALTSELGEESV---VEEVLNEYPDIFIEPTALP-PFREKHNHKIKL------LEGSN 600
Query: 1321 STKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVIN 1380
RP + + + K + DLL ++ S SP++ V K+ GT RL ++
Sbjct: 601 PVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKKD----GTWRLCVD 656
Query: 1381 YKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFG 1440
Y+ LN +PIP +DL+ A +FSK D+++G+ Q+++ D KTAF G
Sbjct: 657 YRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHSG 716
Query: 1441 QYEWNVMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFIS 1499
+E+ VMPFGL NAP+ FQ +MN IF P+ K +V+ DD+L++S +L++H +HL
Sbjct: 717 HFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQHLKQVFE 776
Query: 1500 VIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFL 1559
V++ N L +K + K+ +LGH I I I+ ++P K QL+ FL
Sbjct: 777 VMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLK-QLRGFL 835
Query: 1560 GCLNYVADFCPQLSTIIKPLHDRLKKDPPP*SDIHTNVVKQIKLRVKNLPCLYLPNPQAF 1619
G Y F I PLH K D + + + +K + P L LP
Sbjct: 836 GLAGYYRRFVRSFGVIAGPLHALTKTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQ 895
Query: 1620 KIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQS 1679
++ETDA G G +L Q + +A+ S+ Q + S +KE+LA++ ++ K++
Sbjct: 896 FVVETDACGQGIGAVLMQ----EGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRH 951
Query: 1680 DLINQKFLVCVDCKSAKEILQK 1701
L+ F++ D +S K +L++
Sbjct: 952 YLLQSHFIIKTDQRSLKYLLEQ 973
>At1g36590 hypothetical protein
Length = 1499
Score = 192 bits (487), Expect = 2e-48
Identities = 149/562 (26%), Positives = 265/562 (46%), Gaps = 43/562 (7%)
Query: 1162 KLSAAEGSRLKIKYKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIKKL-------- 1213
++S A+G +L+++ K+ K + ++ LL+ +++G +++ L
Sbjct: 433 RVSVADGRKLRVEGKVTDFSWKLQTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFK 492
Query: 1214 -----FPYNTDE---NGIT---VQHLGQPILFKFSEPPIDKTLNVISYKEKQINFLKEEI 1262
F +N + +G+T V+ + L K E + L ++ +E + E
Sbjct: 493 KLEMRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQEDQVQ--LAMLCVQEVSESTEGELC 550
Query: 1263 SYRTIEEQLQQPSVKSRIEDILENIQSSICSD--LPNAFWERKSHMVELPYEKDFSDKQI 1320
+ + +L + SV +E++L LP F E+ +H ++L +
Sbjct: 551 TINALTSELGEESV---VEEVLNEYPDIFIEPTALP-PFREKHNHKIKL------LEGSN 600
Query: 1321 STKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVIN 1380
RP + + + K + DLL ++ S SP++ V K+ GT RL ++
Sbjct: 601 PVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKKD----GTWRLCVD 656
Query: 1381 YKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFG 1440
Y+ LN +PIP +DL+ A +FSK D+++G+ Q+++ D KTAF G
Sbjct: 657 YRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHSG 716
Query: 1441 QYEWNVMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFIS 1499
+E+ VMPFGL NAP+ FQ +MN IF P+ K +V+ DD+L++S +L++H +HL
Sbjct: 717 HFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQHLKQVFE 776
Query: 1500 VIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFL 1559
V++ N L +K + K+ +LGH I I I+ ++P K QL+ FL
Sbjct: 777 VMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLK-QLRGFL 835
Query: 1560 GCLNYVADFCPQLSTIIKPLHDRLKKDPPP*SDIHTNVVKQIKLRVKNLPCLYLPNPQAF 1619
G Y F I PLH K D + + + +K + P L LP
Sbjct: 836 GLAGYYRRFVRSFGVIAGPLHALTKTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQ 895
Query: 1620 KIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQS 1679
++ETDA G G +L Q + +A+ S+ Q + S +KE+LA++ ++ K++
Sbjct: 896 FVVETDACGQGIGAVLMQ----EGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRH 951
Query: 1680 DLINQKFLVCVDCKSAKEILQK 1701
L+ F++ D +S K +L++
Sbjct: 952 YLLQSHFIIKTDQRSLKYLLEQ 973
>At2g10780 pseudogene
Length = 1611
Score = 181 bits (459), Expect = 3e-45
Identities = 116/375 (30%), Positives = 194/375 (50%), Gaps = 11/375 (2%)
Query: 1326 PIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLN 1385
P +M + +K++ +LL+K IR S SPW +V K+ G+ RL I+Y+ LN
Sbjct: 664 PYRMAPAEMAKLKKQLEELLDKGFIRPSSSPWGAPVLFVKKKD----GSFRLCIDYRGLN 719
Query: 1386 QALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWN 1445
+ +YP+P +L+ + A+ FSK D+ SG+ QI ++ D KTAF + +E+
Sbjct: 720 KVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYDHFEFV 779
Query: 1446 VMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRN 1504
VMPFGL NAP+ F ++MN +F + + I++I+D+L++S++ + H +HL + ++ +
Sbjct: 780 VMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFINDILVYSKSWEAHQEHLRAVLERLREH 839
Query: 1505 GLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNY 1564
L +K S +Q + FLGH I + I ++P + + T+++ FLG Y
Sbjct: 840 ELFAKLSKCSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWP-RPRNATEIRSFLGLAGY 898
Query: 1565 VADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIE 1623
F +++ +PL KD SD ++K + N P L LP +
Sbjct: 899 YRRFVMSFASMAQPLTRLTGKDTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGEPYTVY 958
Query: 1624 TDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLIN 1683
TDAS +G G +L Q K +IA+ S+ ++NY T E+ A+V + ++S L
Sbjct: 959 TDASIVGLGCVLMQ----KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLYG 1014
Query: 1684 QKFLVCVDCKSAKEI 1698
K + D KS K I
Sbjct: 1015 AKVQIYTDHKSLKYI 1029
Score = 30.8 bits (68), Expect = 7.0
Identities = 13/31 (41%), Positives = 17/31 (53%), Gaps = 5/31 (16%)
Query: 896 PETRPSQGKDV-----TCYNCGKPGHISRYC 921
P+ P Q K + TC+ CGK GH+ R C
Sbjct: 413 PKMEPGQSKVLGEETRTCFYCGKTGHLKREC 443
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 180 bits (457), Expect = 6e-45
Identities = 119/375 (31%), Positives = 192/375 (50%), Gaps = 11/375 (2%)
Query: 1326 PIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLN 1385
P +M + +K++ DLL K IR S SPW +V K+ G+ RL I+Y+ LN
Sbjct: 496 PYRMAPAEMAELKKQLKDLLGKGFIRPSTSPWGAPVLFVKKKD----GSFRLCIDYRELN 551
Query: 1386 QALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWN 1445
+ RYP+P +LL + A FSK D+ SG+ QI + E D KTAF +G +E+
Sbjct: 552 RVTVKNRYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFV 611
Query: 1446 VMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRN 1504
VMPFGL NAP+ F R+MN +F + + I++IDD+L++S++ ++ HL + ++
Sbjct: 612 VMPFGLTNAPAVFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEQEVHLRRVMEKLREQ 671
Query: 1505 GLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNY 1564
L +K S +Q ++ FLGH + + IE +P + + T+++ FLG Y
Sbjct: 672 KLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWP-RPTNATEIRSFLGWAGY 730
Query: 1565 VADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIE 1623
F +++ +P+ KD P S +K + + P L LP ++
Sbjct: 731 YRRFVKGFASMAQPMTKLTGKDVPFVWSQECEEGFVSLKEMLTSTPVLALPEHGQPYMVY 790
Query: 1624 TDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLIN 1683
TDAS +G G +L Q ++IA+ S+ + NY T E+ A++ ++ ++S L
Sbjct: 791 TDASRVGLGCVLMQ----HGKVIAYASRQLMKHEGNYPTHDLEMAAVIFALKIWRSYLYG 846
Query: 1684 QKFLVCVDCKSAKEI 1698
K V D KS K I
Sbjct: 847 GKVQVFTDHKSLKYI 861
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 177 bits (450), Expect = 4e-44
Identities = 113/359 (31%), Positives = 188/359 (51%), Gaps = 11/359 (3%)
Query: 1326 PIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLN 1385
P +M + +K++ +LL K IR S SPW +V K+ G+ RL I+Y+ LN
Sbjct: 159 PYRMAPAEMAELKKQLEELLAKGFIRPSSSPWGAPVLFVKKKD----GSFRLCIDYRGLN 214
Query: 1386 QALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWN 1445
+ +YP+P +L+ + A+ FSK D+ SG+ QI ++ D KTAF +G +E+
Sbjct: 215 KVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYGHFEFV 274
Query: 1446 VMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRN 1504
VMPFGL NAP+ F ++MN +F + + I++IDD+L+ S++ + H +HL + ++ +
Sbjct: 275 VMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFIDDILVHSKSWEAHQEHLRAVLERLREH 334
Query: 1505 GLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNY 1564
L +K S +Q + FLGH I + I ++P + + T+++ FLG Y
Sbjct: 335 ELFAKLSKFSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWP-RPRNATEIRSFLGLAGY 393
Query: 1565 VADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIE 1623
F +++ +PL KD SD ++K + N P L LP +
Sbjct: 394 YRRFVMSFASMAQPLTRLTGKDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVY 453
Query: 1624 TDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLI 1682
TDAS +G G +L Q K +IA+ S+ ++NY T E+ A+V ++ ++S LI
Sbjct: 454 TDASIVGLGCVLMQ----KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYLI 508
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 177 bits (450), Expect = 4e-44
Identities = 118/378 (31%), Positives = 191/378 (50%), Gaps = 16/378 (4%)
Query: 1326 PIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLN 1385
P +M E +K++ DLL K IR S SPW +V K+ G+ RL I+Y+ LN
Sbjct: 527 PAEMTE-----LKKQLEDLLGKGFIRPSTSPWGAPVLFVKKKD----GSFRLCIDYRGLN 577
Query: 1386 QALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWN 1445
+YP+P +LL + A FSK D+ SG+ QI + E D KTAF +G +E+
Sbjct: 578 WVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFV 637
Query: 1446 VMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRN 1504
VMPF L NAP+ F R+MN +F + + I++IDD+L++S++ ++H HL + ++
Sbjct: 638 VMPFALTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEHEVHLRRVMEKLREQ 697
Query: 1505 GLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNY 1564
L +K S +Q +I FLGH + + IE +P + + T+++ FL Y
Sbjct: 698 KLFAKLSKCSFWQREIGFLGHIVSAEGVSVDPEKIEAIRDWP-RPTNATEIRSFLRLTGY 756
Query: 1565 VADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIE 1623
F +++ +P+ KD P S +K + + P L LP ++
Sbjct: 757 YRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGQPYMVY 816
Query: 1624 TDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLIN 1683
TDAS +G G +L Q + ++IA+ S+ + NY T E+ ++ ++ ++S L
Sbjct: 817 TDASRVGLGCVLMQ----RGKVIAYASRQLRKHEGNYPTHDLEMAVVIFALKIWRSYLYG 872
Query: 1684 QKFLVCVDCKSAKEILQK 1701
K V D KS K I +
Sbjct: 873 GKVQVFTDHKSLKYIFNQ 890
>At2g06890 putative retroelement integrase
Length = 1215
Score = 170 bits (430), Expect = 7e-42
Identities = 105/330 (31%), Positives = 174/330 (51%), Gaps = 11/330 (3%)
Query: 1373 GTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYK 1432
G+ R+ + + +N +PIP D+L H + +FSK D+KSG+ QI++ E D++K
Sbjct: 467 GSWRMCFDCRAINNVTVKYCHPIPRLDDMLDELHGSSIFSKIDLKSGYHQIRMNEGDEWK 526
Query: 1433 TAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPYSKL-TIVYIDDVLIFSQTLDQHF 1491
TAF G YEW VMPFGL +APS F R+MN + + + IVY DD+L++S++L +H
Sbjct: 527 TAFKTKHGLYEWLVMPFGLTHAPSTFMRLMNHVLRAFIGIFVIVYFDDILVYSESLREHI 586
Query: 1492 KHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIID 1551
+HL++ ++V+++ L + K + + FLG + + ++ +P
Sbjct: 587 EHLDSVLNVLRKEELYANLKKCTFCTDNLVFLGFVVSADGVKVDEEKVKAIRDWPS---P 643
Query: 1552 KT--QLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNL 1608
KT +++ F G + F STI+ PL + +KKD + +K ++ N
Sbjct: 644 KTVGEVRSFHGLAGFYRRFFKDFSTIVAPLTEVMKKDVGFKWEKAQEEAFQSLKDKLTNA 703
Query: 1609 PCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVL 1668
P L L IE DAS IG G +L Q +++IAF S+ A NY T KE+
Sbjct: 704 PVLILSEFLKTFEIECDASGIGIGAVLMQ----DQKLIAFFSEKLGGATLNYPTYDKELY 759
Query: 1669 AIVLSISKFQSDLINQKFLVCVDCKSAKEI 1698
A+V ++ ++Q L + F++ D +S K +
Sbjct: 760 ALVRALQRWQHYLWPKVFVIHTDHESLKHL 789
Score = 42.7 bits (99), Expect = 0.002
Identities = 36/132 (27%), Positives = 60/132 (45%), Gaps = 18/132 (13%)
Query: 864 PKQQWRKRSSRNDHR---KPKPRSKPQSS-QIPKNPPETRPSQGKDVTCYNCGKPGHISR 919
P Q +R+S ++ P+ SKP ++ Q K E S+ +DV CY C GH +
Sbjct: 111 PTYQREERTSSYHNKPIVSPRAESKPYAAVQDHKGKAEISTSRVRDVRCYKCQGKGHYAN 170
Query: 920 YCRLKRRISELHLEPEIEDKINNLLIQTSDEEESNPSDSEVSEDLNQIQNDDSQSSSSVN 979
C KR + ++N I+ +E +PS + +E+L + +
Sbjct: 171 ECPNKR----------VMILLDNGEIEPEEEIPDSPSSLKENEELPA----QGELLVARR 216
Query: 980 TLSINTLTNEQD 991
TLS+ T T+EQ+
Sbjct: 217 TLSVQTKTDEQE 228
>At1g35370 hypothetical protein
Length = 1447
Score = 159 bits (403), Expect = 1e-38
Identities = 140/554 (25%), Positives = 253/554 (45%), Gaps = 35/554 (6%)
Query: 1162 KLSAAEGSRLKIKYKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIKKLFPYNTDEN 1221
K++ A+G +L + +I K S ++ LL+ +++G +++ L + +
Sbjct: 412 KVAVADGRKLNVDGQIKGFTWKLQSTTFQSDILLIPLQGVDMVLGVQWLETLGRISWEFK 471
Query: 1222 GITVQHLGQPILFKFSEPPIDKTLNVISYKEK-----QINFLKEEISYRTIEEQLQQPSV 1276
+ +Q + ++ ++K + QI + +E+ + S+
Sbjct: 472 KLEMQFFYKNQRVWLHGIITGSVRDIKAHKLQKTQADQIQLAMVCVREVVSDEEQEIGSI 531
Query: 1277 KSRIEDILE-NIQSSICSDLPNAFWERKSHMVELPYEKDFSDKQISTK--ARPIQMNEEL 1333
+ D++E ++ +I + P+ F E +LP ++ D +I A P+
Sbjct: 532 SALTSDVVEESVVQNIVEEFPDVFAEP----TDLPPFREKHDHKIKLLEGANPVNQRPYR 587
Query: 1334 LHFCQKE-----INDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQAL 1388
QK+ + D+++ I+ S SP++ V K+ GT RL ++Y LN
Sbjct: 588 YVVHQKDEIDKIVQDMIKSGTIQVSSSPFASPVVLVKKKD----GTWRLCVDYTELNGMT 643
Query: 1389 CWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMP 1448
R+ IP +DL+ + VFSK D+++G+ Q+++ D KTAF G +E+ VM
Sbjct: 644 VKDRFLIPLIEDLMDELGGSVVFSKIDLRAGYHQVRMDPDDIQKTAFKTHNGHFEYLVML 703
Query: 1449 FGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLA 1507
FGL NAP+ FQ +MN +F + K +V+ DD+LI+S ++++H +HL V++ + L
Sbjct: 704 FGLTNAPATFQSLMNSVFRDFLRKFVLVFFDDILIYSSSIEEHKEHLRLVFEVMRLHKLF 763
Query: 1508 VSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVAD 1567
+K LGH I I I+ ++P K Q++ FLG Y
Sbjct: 764 AKGSK--------EHLGHFISAREIETDPAKIQAVKEWPTPTTVK-QVRGFLGFAGYYRR 814
Query: 1568 FCPQLSTIIKPLHDRLKKDPPP*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDAS 1627
F I PLH K D S + +K + N P L LP ++ETDA
Sbjct: 815 FVRNFGVIAGPLHALTKTDGFCWSLEAQSAFDTLKAVLCNAPVLALPVFDKQFMVETDAC 874
Query: 1628 DIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFL 1687
G +L Q K +A+ S+ Q + S +KE+LA + ++ K++ L+ F+
Sbjct: 875 GQGIRAVLMQ----KGHPLAYISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYLLPSHFI 930
Query: 1688 VCVDCKSAKEILQK 1701
+ D +S K +L++
Sbjct: 931 IKTDQRSLKYLLEQ 944
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 156 bits (394), Expect = 1e-37
Identities = 102/329 (31%), Positives = 168/329 (51%), Gaps = 9/329 (2%)
Query: 1373 GTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYK 1432
G+ RL I+Y+ LN +YP+P +LL + A FSK D+ SG+ I + E D K
Sbjct: 514 GSFRLCIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEADVRK 573
Query: 1433 TAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNP-YSKLTIVYIDDVLIFSQTLDQHF 1491
TAF +G +E+ VMPFGL NAP+ F R+MN +F + I++IDD+L++S++L++H
Sbjct: 574 TAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLEEHE 633
Query: 1492 KHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIH-QGTIIPINRAIEFTDKFPDQII 1550
HL + ++ L +K S +Q ++ FLGH + +G + + D
Sbjct: 634 VHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTP--T 691
Query: 1551 DKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIKLRVKNLP 1609
+ T+++ FLG Y F +++ +P+ KD P S +K + + P
Sbjct: 692 NATEIRSFLGLAGYYRRFVKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEMLTSTP 751
Query: 1610 CLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLA 1669
L LP ++ TDAS +G G +L Q + ++IA+ S+ + NY T E+ A
Sbjct: 752 VLALPEHGEPYMVYTDASGVGLGCVLMQ----RGKVIAYASRQLRKHEGNYPTHDLEMAA 807
Query: 1670 IVLSISKFQSDLINQKFLVCVDCKSAKEI 1698
++ ++ ++S L V D KS K I
Sbjct: 808 VIFALKIWRSYLYGGNVQVFTDHKSLKYI 836
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 151 bits (382), Expect = 3e-36
Identities = 112/410 (27%), Positives = 186/410 (45%), Gaps = 68/410 (16%)
Query: 1293 SDLPNAFWERKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQKEINDLLEKKLIRR 1352
+DLP F H +EL + + RP + + K ++D+L I+
Sbjct: 22 TDLP-PFRAPHDHKIELLEDSN------PVNQRPYRYVVHQKNEIDKIVDDMLASGTIQA 74
Query: 1353 SKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFS 1412
S SP++ V K+ GT RL ++Y+ LN R+PIP +DL+ + V+S
Sbjct: 75 SSSPYASPVVLVKKKD----GTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELGGSNVYS 130
Query: 1413 KFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPY-SK 1471
K D+++G+ Q+++ D +KTAF G YE+ VMPFGL NAP+ FQ +MN F P+ K
Sbjct: 131 KIDLRAGYHQVRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQSLMNSFFKPFLRK 190
Query: 1472 LTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGT 1531
+V+ DD+LI+S ++++H KHL V++ + L +K + ++ +LGH I
Sbjct: 191 FVLVFFDDILIYSTSMEEHKKHLEAVFEVMRVHHLFAKMSKCAFAVPRVEYLGHFISGEG 250
Query: 1532 IIPINRAIEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPPP*S 1591
I I+ +P ++ QL FLG Y F
Sbjct: 251 IATDPAKIKAVQDWPVP-VNLKQLCGFLGLTGYYRRFF---------------------- 287
Query: 1592 DIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSK 1651
++E DA G G +L Q + +AF S+
Sbjct: 288 -----------------------------VVEMDACGHGIGAVLMQ----EGHPLAFISR 314
Query: 1652 HWNPAQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVDCKSAKEILQK 1701
Q + S +KE+LA++ + K++ LI+ F++ D +S K +L++
Sbjct: 315 QLKGKQLHLSIYEKELLAVIFVVRKWRHYLISSHFIIKTDQRSLKYLLEQ 364
>At4g08100 putative polyprotein
Length = 1054
Score = 138 bits (347), Expect = 3e-32
Identities = 105/398 (26%), Positives = 177/398 (44%), Gaps = 59/398 (14%)
Query: 1302 RKSHMVELPYEKDFSDKQISTKARPIQMNEELLHFCQKEINDLLEKKLIRRSKSPWSCAA 1361
RK+ ++ + + + D Q+S K R I+ E + K ++ + + + +K W C A
Sbjct: 473 RKTTLIPMTLHEVYLD-QLSMKQRAIKPTEPI-DTKGKNKHESVCRSGLTSTKERWDCRA 530
Query: 1362 FYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFW 1421
+N R+PIP D L + H + +FSK D+KSG+
Sbjct: 531 ----------------------INNITVKYRHPIPRLDDTLDKLHGSSIFSKIDLKSGYH 568
Query: 1422 QIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDV 1480
Q +++E D++KTA YEW VMPFGL NAP+ F R+MN + + IVY DD+
Sbjct: 569 QTRMKEGDEWKTAIKTKQRLYEWLVMPFGLTNAPNTFMRLMNHVLRKHIGVFVIVYFDDI 628
Query: 1481 LIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIE 1540
L++S+ L+ H HL + ++++ L + K + + FLG + I ++
Sbjct: 629 LVYSKNLEYHVMHLKLVLDLLRKEKLYANLKKCTFCTDNLVFLGFVVSADGIKVDEEKVK 688
Query: 1541 FTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPPP*SDIHTNVVKQ 1600
++P+ K ++ +G D N +
Sbjct: 689 AIREWPN---PKNVSEKDIGF---------------------------KWEDAQENAFQA 718
Query: 1601 IKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNY 1660
+K ++ N LPN IE DAS +G G +L Q + IA+ S+ A NY
Sbjct: 719 LKEKLTNSSVPILPNFMKSFEIECDASGLGIGAVLMQ----DHKPIAYFSEKLGGATLNY 774
Query: 1661 STVKKEVLAIVLSISKFQSDLINQKFLVCVDCKSAKEI 1698
T KE+ A+V ++ +Q L +KF++ D +S K +
Sbjct: 775 PTYDKELYALVRALQTWQHYLWPKKFVIHTDHESLKHL 812
>At4g03650 putative reverse transcriptase
Length = 839
Score = 126 bits (317), Expect = 9e-29
Identities = 84/276 (30%), Positives = 140/276 (50%), Gaps = 7/276 (2%)
Query: 1425 LQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDDVLIF 1483
L E D KTAF +G +E+ VMPFGL NAP+ F R+MN +F + + I++IDD+L++
Sbjct: 516 LAEADVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVY 575
Query: 1484 SQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTD 1543
S++ ++H HL + ++ L +K S +Q ++ FLGH + + IE
Sbjct: 576 SKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIR 635
Query: 1544 KFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPP-P*SDIHTNVVKQIK 1602
+P + + T+++ FLG Y F +++ +P+ KD P S +K
Sbjct: 636 DWP-RPTNATEIRSFLGLAGYYRRFIKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLK 694
Query: 1603 LRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYST 1662
+ + P L LP ++ TDAS +G G L Q + ++IA+ S+ + NY T
Sbjct: 695 EMLTSTPVLALPEHGEPYMVYTDASGVGLGCALMQ----RGKVIAYASRQLRKHEGNYPT 750
Query: 1663 VKKEVLAIVLSISKFQSDLINQKFLVCVDCKSAKEI 1698
E+ A++ ++ ++S L K V D KS K I
Sbjct: 751 HDLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYI 786
>At4g04230 putative transposon protein
Length = 315
Score = 118 bits (296), Expect = 3e-26
Identities = 63/179 (35%), Positives = 100/179 (55%), Gaps = 5/179 (2%)
Query: 1345 LEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNKKDLLAR 1404
+ K IR S SP + V K+ GT R+ ++ + +N R+PIP D+L
Sbjct: 125 IHKGYIRESLSPCAVPVLLVPKKD----GTWRMCLDCRAINNITIKYRHPIPRLYDMLDE 180
Query: 1405 QHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNE 1464
A +FSK D++SG+ Q++++E D++KTAF G YE VMPFGL NAPS F R+MN+
Sbjct: 181 LSGAIIFSKVDLRSGYHQVRMREGDEWKTAFKTKQGLYECLVMPFGLTNAPSTFMRLMNQ 240
Query: 1465 IFNPY-SKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRF 1522
+ + K +VY DD+LI++++ H + L + +++ GL + K + K F
Sbjct: 241 VLRSFIGKFVVVYFDDILIYNKSYSDHIQPLELILKTLRKEGLYANLKKCTFCSDKFFF 299
>At1g20390 hypothetical protein
Length = 1791
Score = 111 bits (277), Expect = 4e-24
Identities = 155/664 (23%), Positives = 284/664 (42%), Gaps = 50/664 (7%)
Query: 1046 KPVTIQDLHFEINSLKT---EVKSLKQIQKSQQLILEKLTKNYEEDDSSIPDSNP-APND 1101
+PV ++ +H + L+ V+S+KQ +K ++++ + S+ + N AP
Sbjct: 500 EPVVVRQVHVIMGGLQNCSDSVRSIKQYRKKAEMVVA-----WPSSTSTTRNPNQSAPIS 554
Query: 1102 NCEDFLENINQVTIQKFFIHVKILIGDFVLEIPALFDTGADSSCISEGL-----IPTRYF 1156
+ LE ++ T + V+++I D + L DTG+ I + + I R
Sbjct: 555 FTDVDLEGLD--TPHNDPLVVELIISDSRVT-RVLIDTGSSVDLIFKDVLTAMNITDRQI 611
Query: 1157 EKTTEKLSAAEGSRLKIKYKIPSAIIKNGSLEIETPFLLVRNLSQKIIIGTPFIKKL--F 1214
+ ++ L+ +G + I I G + ++ + +I+GTP+I ++
Sbjct: 612 KPVSKPLAGFDGDFVMTIGTIKLPIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAI 671
Query: 1215 PYNTDENGITVQHLGQPILFKFSEPPIDKTLNVISYKEKQINF-----LKEEISYRTIEE 1269
P + H G +F P KT + SY+E ++ + E R +
Sbjct: 672 PSTYHQCVKFPTHNG---IFTLRAPKEAKTPSR-SYEESELCRTEMVNIDESDPTRCVGV 727
Query: 1270 QLQ-QPSVKSRIEDILENIQSSIC---SDLPNAFWERKSHMVELPYEKDFSDKQISTKAR 1325
+ PS++ + +L+ + D+ +H EL + F K + K R
Sbjct: 728 GAEISPSIRLELIALLKRNSKTFAWSIEDMKGIDPAITAH--ELNVDPTF--KPVKQKRR 783
Query: 1326 PIQMNEELLHFCQKEINDLLEKKLIRRSKSP-WSCAAFYVNKQAEIERGTPRLVINYKPL 1384
++ E +E+ LL+ I K P W V K+ G R+ ++Y L
Sbjct: 784 --KLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKK----NGKWRVCVDYTDL 837
Query: 1385 NQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEW 1444
N+A YP+P+ L+ + S D SG+ QI + + D+ KT+F G Y +
Sbjct: 838 NKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTDRGTYCY 897
Query: 1445 NVMPFGLKNAPSEFQRIMNEIFNPYSKLTI-VYIDDVLIFSQTLDQHFKHLNTFISVIKR 1503
VM FGLKNA + +QR +N++ T+ VYIDD+L+ S + H +HL+ V+
Sbjct: 898 KVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHLSKCFDVLNT 957
Query: 1504 NGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQLQRFLGCLN 1563
G+ ++ TK + T FLG+ + + I + I + P + ++QR G +
Sbjct: 958 YGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIRAILELPSP-RNAREVQRLTGRIA 1016
Query: 1564 YVADFCPQLSTIIKPLHDRLKKDPPP*SDIHT-NVVKQIKLRVKNLPCLYLPNPQAFKII 1622
+ F + + P ++ LK+ D + +++K + P L P +
Sbjct: 1017 ALNRFISRSTDKCLPFYNLLKRRAQFDWDKDSEEAFEKLKDYLSTPPILVKPEVGETLYL 1076
Query: 1623 ETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISK----FQ 1678
SD +L ++ +++ I +TSK A+ Y ++K LA+V S K FQ
Sbjct: 1077 YIAVSDHAVSSVLVREDRGEQRPIFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQ 1136
Query: 1679 SDLI 1682
S I
Sbjct: 1137 SHTI 1140
>At2g06170 putative Ty3-gypsy-like retroelement pol polyprotein
Length = 587
Score = 110 bits (276), Expect = 5e-24
Identities = 50/113 (44%), Positives = 76/113 (67%), Gaps = 1/113 (0%)
Query: 1382 KPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQ 1441
K +N+ R+PIP D+L +KVFSK D++SG+ QI+++ D++KT F G
Sbjct: 401 KAINKITIKYRFPIPQLDDMLDELSGSKVFSKIDLRSGYHQIRIRPGDEWKTDFKSKDGL 460
Query: 1442 YEWNVMPFGLKNAPSEFQRIMNEIFNPYS-KLTIVYIDDVLIFSQTLDQHFKH 1493
YEW VMPFG+ NAPS F R+MN+I P++ +VY DD+LI+S+T +++ +H
Sbjct: 461 YEWQVMPFGMSNAPSTFMRLMNQILRPFTGSFVVVYFDDILIYSKTKEEYLEH 513
>At4g07600
Length = 630
Score = 102 bits (253), Expect = 2e-21
Identities = 85/299 (28%), Positives = 131/299 (43%), Gaps = 31/299 (10%)
Query: 1376 RLVINYKPLNQALCWIRYPIPNKKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAF 1435
R+ I+Y+ LN A +P+P +L R + + D SGF+QI + D KT F
Sbjct: 356 RMCIDYRNLNAASRNDHFPLPFTDQMLERLANHPYYCFLDGYSGFFQIPIHPNDHEKTTF 415
Query: 1436 TVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNP-YSKLTIVYIDDVLIFSQTLDQHFKHL 1494
T P+G + + MPFGL NAP+ FQR M IF+ ++ V++DD ++ + +L
Sbjct: 416 TCPYGTFAYERMPFGLCNAPATFQRCMTSIFSDIIEEMVEVFMDDFSVYGPSFSSCLLNL 475
Query: 1495 NTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIIDKTQ 1554
++ + L ++ K + LGH I + + IE +DK
Sbjct: 476 GRVLTRCEETNLVLNWEKCHFMVKEGIMLGHKISE-------KGIE---------VDK-- 517
Query: 1555 LQRFLGCLNYVADFCPQLSTIIKPLHDRLKK------DPPP*SDIHTNVVKQIKLRVKNL 1608
FLG + F S I +PL L K D HT IK + +
Sbjct: 518 -GCFLGHAGFYRRFIKDFSKIARPLTRLLCKETEFEFDEDCPKSFHT-----IKEALVSA 571
Query: 1609 PCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEV 1667
P + N I DA D G +L QK+ K +I + S+ + AQ Y+T +KE+
Sbjct: 572 PVVRATNWDYPFEIMCDALDYAVGAVLGQKIDKKLHVIYYASRTLDEAQGRYATTEKEL 630
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 97.8 bits (242), Expect = 5e-20
Identities = 90/348 (25%), Positives = 157/348 (44%), Gaps = 38/348 (10%)
Query: 1339 KEINDLLEKKLIRRSKSP-WSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPN 1397
+E+ LL+ I + P W V K+ G R+ I++ LN+A +P+P+
Sbjct: 800 EEVKKLLDAGSIVEVRYPDWLRNPVVVKKK----NGKWRVCIDFTDLNKACPKDSFPLPH 855
Query: 1398 KKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSE 1457
L+ ++ S D SG+ QI + + D+ KT F G Y + VMPFGLKNA +
Sbjct: 856 IDRLVEATAGNELLSFMDAFSGYNQILMHQNDREKTVFITDQGTYCYKVMPFGLKNAGAT 915
Query: 1458 FQRIMNEIFNPYSKLTI-VYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLF 1516
+ R++N++F ++ VYIDD+L+ S ++H HL V+ R + ++ +K +
Sbjct: 916 YPRLVNQMFTDQLDHSMEVYIDDMLVKSLRAEEHITHLRQCFQVLNRYNMKLNPSKCTFG 975
Query: 1517 QTKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIID------KTQLQRFLGCLNYVADFCP 1570
T FLG+ + R IE K IID ++QR +G + + F
Sbjct: 976 VTSGEFLGY-------LVTRRGIEANPKQISAIIDLPSPRNTREVQRLIGRIAALNRFIS 1028
Query: 1571 QLSTIIKPLHDRLKKDPPP*SDIHTNVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIG 1630
+ + P + L+ N + + + LYL + + T A
Sbjct: 1029 RSTDKCLPFYQLLR----------ANKRFEWDEKCEEGETLYL-----YIAVSTSA---- 1069
Query: 1631 FGGILKQKVFDKEQIIAFTSKHWNPAQQNYSTVKKEVLAIVLSISKFQ 1678
G+L ++ ++ I + SK + A+ Y T++K A+V+S K +
Sbjct: 1070 VSGVLVREDRGEQHPIFYVSKTLDGAELRYPTLEKLAFAVVISARKLR 1117
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 96.7 bits (239), Expect = 1e-19
Identities = 84/286 (29%), Positives = 131/286 (45%), Gaps = 13/286 (4%)
Query: 1421 WQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEFQRIMNEIFNPY-SKLTIVYIDD 1479
+++Q K KT FT P G + + M FGL NAP FQR M IF+ + ++ V++DD
Sbjct: 154 FKLQFTLMIKKKTTFTCPNGTFAYKRMSFGLCNAPGTFQRSMTSIFSDFIEEIMEVFMDD 213
Query: 1480 VLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQTKIRFLGHNIHQGTIIPINRA- 1538
FS L +L + + L ++ K + LGH I G I +++A
Sbjct: 214 FSGFSSCL----LNLCRVLERCEETNLVLNWEKCHFMVHEGIVLGHKI-SGKGIEVDKAK 268
Query: 1539 --IEFTDKFPDQIIDKTQLQRFLGCLNYVADFCPQLSTIIKPLHDRLKKDPPP*SDIHT- 1595
+ + P + D ++ FLG + F S I +PL L K+ D
Sbjct: 269 IDVMIQLQPPKTVKD---IRSFLGHAEFYRRFIKDFSKIARPLTRLLCKETEFNFDEDCL 325
Query: 1596 NVVKQIKLRVKNLPCLYLPNPQAFKIIETDASDIGFGGILKQKVFDKEQIIAFTSKHWNP 1655
IK + + P PN I DA D G +L QK+ DK +I + S+ +
Sbjct: 326 KAFHLIKETLVSAPIFQAPNWDHPFEIMCDAFDYAVGAVLDQKIDDKLHVIYYASRTMDE 385
Query: 1656 AQQNYSTVKKEVLAIVLSISKFQSDLINQKFLVCVDCKSAKEILQK 1701
AQ Y+T++KE+LA+V + K +S L+ K V +D + + I K
Sbjct: 386 AQTRYATIEKELLAVVFAFEKIRSYLVGFKVKVYIDHAALRHIYAK 431
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 94.0 bits (232), Expect = 7e-19
Identities = 53/154 (34%), Positives = 81/154 (52%), Gaps = 23/154 (14%)
Query: 1338 QKEINDLLEKKLIRRSKSPWSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPN 1397
+K++ DLL K IR S S W G P L N+ +YP+P
Sbjct: 552 KKQLEDLLGKGFIRPSTSRW---------------GAPGL-------NRVTLKNKYPLPR 589
Query: 1398 KKDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSE 1457
+LL + A FSK D+ G+ Q + E D KTAF +G +E+ VMPFGL NAP+
Sbjct: 590 IDELLDQLRGATCFSKIDLTPGYHQFPIAEADVRKTAFRTRYGHFEFVVMPFGLTNAPTA 649
Query: 1458 FQRIMNEIFNPY-SKLTIVYIDDVLIFSQTLDQH 1490
R+MN +F + + I++IDD+L++ ++ ++H
Sbjct: 650 LMRLMNSVFQEFLDEFVIIFIDDILVYFKSPEEH 683
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 89.7 bits (221), Expect = 1e-17
Identities = 68/229 (29%), Positives = 108/229 (46%), Gaps = 19/229 (8%)
Query: 1340 EINDLLEKKLIRRSKSP-WSCAAFYVNKQAEIERGTPRLVINYKPLNQALCWIRYPIPNK 1398
E++ LL+ I K P W V K+ + R+ I++ LN+A +P+P+
Sbjct: 516 EVDKLLDAGSIVEVKYPEWLANPVVVKKKND----KWRVCIDFTDLNKACPKDSFPLPHI 571
Query: 1399 KDLLARQHDAKVFSKFDMKSGFWQIQLQEKDKYKTAFTVPFGQYEWNVMPFGLKNAPSEF 1458
++ ++ S D SG+ QI + + D+ KT+F + G Y + VMPFGLKN + +
Sbjct: 572 DRMVEATTGNELLSFMDAFSGYNQIPMHKDDQEKTSFIIDRGTYCYKVMPFGLKNVGARY 631
Query: 1459 QRIMNEIFNP-YSKLTIVYIDDVLIFSQTLDQHFKHLNTFISVIKRNGLAVSKTKVSLFQ 1517
QR++N++F P K VYIDD+L+ S H HL + + + ++ K
Sbjct: 632 QRLVNQMFAPQLGKTMEVYIDDMLVKSTRSADHIDHLKACFETLNKYNMKLNPAKCLFGV 691
Query: 1518 TKIRFLGHNIHQGTIIPINRAIEFTDKFPDQIID------KTQLQRFLG 1560
T FLG+ I R IE K I+D K ++QR G
Sbjct: 692 TSGEFLGY-------IVTKRGIEANPKQIRAILDLQSPRNKKEVQRLTG 733
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,431,077
Number of Sequences: 26719
Number of extensions: 1911335
Number of successful extensions: 10598
Number of sequences better than 10.0: 220
Number of HSP's better than 10.0 without gapping: 91
Number of HSP's successfully gapped in prelim test: 137
Number of HSP's that attempted gapping in prelim test: 8876
Number of HSP's gapped (non-prelim): 1314
length of query: 1703
length of database: 11,318,596
effective HSP length: 113
effective length of query: 1590
effective length of database: 8,299,349
effective search space: 13195964910
effective search space used: 13195964910
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0019a.7


