

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0019a.4
(310 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
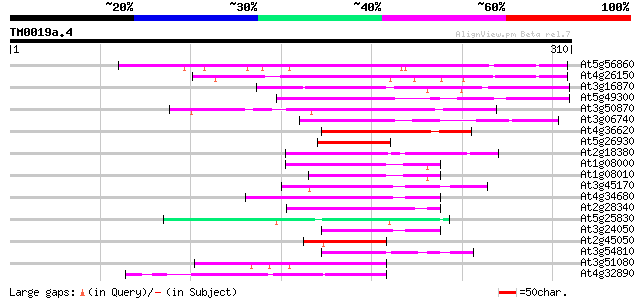
Score E
Sequences producing significant alignments: (bits) Value
At5g56860 unknown protein 138 3e-33
At4g26150 putative transcription factor 130 9e-31
At3g16870 hypothetical protein 78 5e-15
At5g49300 putative protein 77 2e-14
At3g50870 transcription factor-like protein 74 8e-14
At3g06740 unknown protein 74 1e-13
At4g36620 transcription factor like protein 72 4e-13
At5g26930 unknown protein 70 1e-12
At2g18380 putative GATA-type zinc finger transcription factor 65 5e-11
At1g08000 GATA transcription factor 3, putative 63 2e-10
At1g08010 GATA transcription factor 3, putative 63 2e-10
At3g45170 putative protein 61 7e-10
At4g34680 GATA transcription factor 3 60 2e-09
At2g28340 hypothetical protein 60 2e-09
At5g25830 GATA transcription factor - like 59 4e-09
At3g24050 GATA transcription factor 1 (AtGATA-1) 57 2e-08
At2g45050 putative GATA-type zinc finger transcription factor 57 2e-08
At3g54810 unknown protein 56 3e-08
At3g51080 transcription factor-like protein 55 4e-08
At4g32890 unknown protein 55 5e-08
>At5g56860 unknown protein
Length = 398
Score = 138 bits (348), Expect = 3e-33
Identities = 112/329 (34%), Positives = 151/329 (45%), Gaps = 84/329 (25%)
Query: 61 HDGSSSSSSSNHLYNSTFISPQPVMDNPSSSTCDP---------NLSFSKMEEED----- 106
H +++ ++HL+ S + + + N SS CD L+ K + ED
Sbjct: 72 HVAYNNTYHADHLHLSQPLKAKMFVANGGSSACDHMVPKKETRLKLTIRKKDHEDQPHPL 131
Query: 107 ----IKNVHGSAKW-MSSKMRLMKKMMST--PATDKANN-----STTIPISPRIQ-NQGH 153
K S KW MS KMRL+KK ++ D+ NN S P++ + ++ H
Sbjct: 132 HQNPTKPDSDSDKWLMSPKMRLIKKTITNNKQLIDQTNNNNHKESDHYPLNHKTNFDEDH 191
Query: 154 -------------------ENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPK 194
ENRY+ + +N++ RVCSDCNT+ TPLWRSGP GPK
Sbjct: 192 HEDLNFKNVLTRKTTAATTENRYNTINENGYSNNNGVIRVCSDCNTTKTPLWRSGPRGPK 251
Query: 195 SLCNACGIRQRKARRAMAEAA------------------------------NG-----LA 219
SLCNACGIRQRKARRA AA NG +
Sbjct: 252 SLCNACGIRQRKARRAAMAAAAAAGDQEVAVAPRVQQLPLKKKLQNKKKRSNGGEKYNHS 311
Query: 220 TPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRN 279
P+ + K ++ +EK+ A A SKST+S+ S K CF D I L
Sbjct: 312 PPMVAKAKKCKIKEEEEKEMEAETVAGDSEISKSTTSSNSSISSNK--FCFDDLTIMLSK 369
Query: 280 NSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+S +QQVFP+DE EAA+LLM LS G VH
Sbjct: 370 SSAYQQVFPQDE-KEAAVLLMALSYGMVH 397
>At4g26150 putative transcription factor
Length = 352
Score = 130 bits (327), Expect = 9e-31
Identities = 88/228 (38%), Positives = 118/228 (51%), Gaps = 31/228 (13%)
Query: 102 MEEEDIKNVHG--SAKWMSSKMRLMKKMMSTPATDKANNSTTIPISPRIQNQGHENRYSQ 159
+ + IK++ G S KW+SSK+RLMKK + T ++ T N + S
Sbjct: 134 LPQSPIKDMTGTNSLKWISSKVRLMKKKKAIITTSDSSKQHT--------NNDQSSNLSN 185
Query: 160 RSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARR-----AMAEA 214
+N N+ R+CSDCNT+ TPLWRSGP GPKSLCNACGIRQRKARR A A A
Sbjct: 186 SERQNGYNNDCVIRICSDCNTTKTPLWRSGPRGPKSLCNACGIRQRKARRAAMATATATA 245
Query: 215 ANGLATPI--NTASTKTRVHHNKEK--KPRANHFAQFKN----------KSKSTSSNAGS 260
+G++ P+ K ++ + K P K + T SN+
Sbjct: 246 VSGVSPPVMKKKMQNKNKISNGVYKILSPLPLKVNTCKRMITLEETALAEDLETQSNSTM 305
Query: 261 SQETKKLECFKDFAISLRNNSTFQQVFPRDEVAEAALLLMDLSCGYVH 308
+ + F D A+ L +S +QQVFP+DE EAA+LLM LS G VH
Sbjct: 306 LSSSDNI-YFDDLALLLSKSSAYQQVFPQDE-KEAAILLMALSHGMVH 351
>At3g16870 hypothetical protein
Length = 190
Score = 78.2 bits (191), Expect = 5e-15
Identities = 62/192 (32%), Positives = 92/192 (47%), Gaps = 27/192 (14%)
Query: 137 NNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSL 196
+ T + + + + +EN S S ++S +T R C DC T TPLWR GP GPKSL
Sbjct: 7 DTKTKLDSAGELSDVDNENCSSSGSG-GGSSSGDTKRTCVDCGTIRTPLWRGGPAGPKSL 65
Query: 197 CNACGIRQRKARRAMAEAANGLATPINTASTKT------RVHHNKEKKPRANHFAQFK-- 248
CNACGI+ RK R +AA G+ + + K+ + H KK + N K
Sbjct: 66 CNACGIKSRKKR----QAALGMRSEEKKKNRKSNCNNDLNLDHRNAKKYKINIVDDGKID 121
Query: 249 ----------NKSKSTSSNAGSSQETKKLECFKDFAISLRNNSTFQQVFPR-DEVAEAAL 297
+S S+SSN G S K L+ + R+ ++++ + E AA+
Sbjct: 122 IDDDPKICNNKRSSSSSSNKGVS---KFLDLGFKVPVMKRSAVEKKRLWRKLGEEERAAV 178
Query: 298 LLMDLSCGYVHS 309
LLM LSC V++
Sbjct: 179 LLMALSCSSVYA 190
>At5g49300 putative protein
Length = 139
Score = 76.6 bits (187), Expect = 2e-14
Identities = 52/163 (31%), Positives = 77/163 (46%), Gaps = 35/163 (21%)
Query: 148 IQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKA 207
+ ++ + R +NN + ++ + C+DC TS TPLWR GP GPKSLCNACGIR RK
Sbjct: 11 VDSETMKTRAEDMIEQNNTSVNDKKKTCADCGTSKTPLWRGGPVGPKSLCNACGIRNRKK 70
Query: 208 RRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKL 267
RR E NK+ K K+ S + G S + +
Sbjct: 71 RRGGTE-------------------DNKKLK---------KSSSGGGNRKFGESLKQSLM 102
Query: 268 ECFKDFAISLRNNSTFQQVFPR-DEVAEAALLLMDLSCGYVHS 309
+ + +R ST ++ + E +AA+LLM LS G V++
Sbjct: 103 D------LGIRKRSTVEKQRQKLGEEEQAAVLLMALSYGSVYA 139
>At3g50870 transcription factor-like protein
Length = 295
Score = 74.3 bits (181), Expect = 8e-14
Identities = 57/197 (28%), Positives = 82/197 (40%), Gaps = 31/197 (15%)
Query: 89 SSSTCDPNLSF----SKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNSTTIPI 144
SSS D LS +++ EED K ++ SS + ++ T K NNS T P
Sbjct: 63 SSSLVDCTLSLGTPSTRLCEEDEKRRRSTSSGASSCISNFWDLIHT----KNNNSKTAPY 118
Query: 145 SPRIQNQGHENRYSQRSPRNN------------NNSSNTTRVCSDCNTSSTPLWRSGPNG 192
+ + +S P S R C++C+T+STPLWR+GP G
Sbjct: 119 N-------NVPSFSANKPSRGCSGGGGGGGGGGGGDSLLARRCANCDTTSTPLWRNGPRG 171
Query: 193 PKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSK 252
PKSLCNACGIR +K R A T HHN N++ N +
Sbjct: 172 PKSLCNACGIRFKKEERRTTAATGNTVVGAAPVQTDQYGHHNS----GYNNYHAATNNNN 227
Query: 253 STSSNAGSSQETKKLEC 269
+ + T+++ C
Sbjct: 228 NNGTPWAHHHSTQRVPC 244
>At3g06740 unknown protein
Length = 149
Score = 73.6 bits (179), Expect = 1e-13
Identities = 49/143 (34%), Positives = 67/143 (46%), Gaps = 29/143 (20%)
Query: 161 SPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLAT 220
S +N SN + C+ C TS TPLWR GP GPKSLCNACGIR RK RR +
Sbjct: 29 SSSSNEAISNEKKSCAICGTSKTPLWRGGPAGPKSLCNACGIRNRKKRRTLIS------- 81
Query: 221 PINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECFKDFAISLRNN 280
N + K + HN+ K G S + + +E ++ + R+
Sbjct: 82 --NRSEDKKKKSHNRNPK-------------------FGDSLKQRLMELGREVMMQ-RST 119
Query: 281 STFQQVFPRDEVAEAALLLMDLS 303
+ Q+ E +AA+LLM LS
Sbjct: 120 AENQRRNKLGEEEQAAVLLMALS 142
>At4g36620 transcription factor like protein
Length = 211
Score = 72.0 bits (175), Expect = 4e-13
Identities = 36/83 (43%), Positives = 52/83 (62%), Gaps = 4/83 (4%)
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVH 232
R C++C+T+STPLWR+GP GPKSLCNACGIR +K R + A N + +TA+ +
Sbjct: 75 RRCANCDTTSTPLWRNGPRGPKSLCNACGIRFKKEERRASTARNSTSGGGSTAAGVPTLD 134
Query: 233 HNKEKKPRANHFAQFKNKSKSTS 255
H + AN++ N+ S+S
Sbjct: 135 H----QASANYYYNNNNQYASSS 153
>At5g26930 unknown protein
Length = 120
Score = 70.1 bits (170), Expect = 1e-12
Identities = 28/40 (70%), Positives = 32/40 (80%)
Query: 171 TTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRA 210
T R CS+C T+ TP+WR GP GPKSLCNACGIR RK RR+
Sbjct: 24 TIRCCSECKTTKTPMWRGGPTGPKSLCNACGIRHRKQRRS 63
>At2g18380 putative GATA-type zinc finger transcription factor
Length = 207
Score = 65.1 bits (157), Expect = 5e-11
Identities = 41/118 (34%), Positives = 59/118 (49%), Gaps = 4/118 (3%)
Query: 153 HENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMA 212
H S + + + R C+ C+T+STPLWR+GP GPKSLCNACGIR +K R A
Sbjct: 71 HGGNAKTSSYKKGGVAHSLPRRCASCDTTSTPLWRNGPKGPKSLCNACGIRFKKEER-RA 129
Query: 213 EAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQETKKLECF 270
A N T S+ V N+++ + S SS + + Q T+++ F
Sbjct: 130 TARN--LTISGGGSSAAEVPVENSYNGGGNYYSHHHHHYAS-SSPSWAHQNTQRVPYF 184
>At1g08000 GATA transcription factor 3, putative
Length = 308
Score = 63.2 bits (152), Expect = 2e-10
Identities = 32/89 (35%), Positives = 46/89 (50%), Gaps = 12/89 (13%)
Query: 153 HENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMA 212
H +++ S ++ S R+C+ C T +TP WR GP+GPK+LCNACG+R + R
Sbjct: 198 HLITHTESSTLESSKSDGIVRICTHCETITTPQWRQGPSGPKTLCNACGVRFKSGR---- 253
Query: 213 EAANGLATPINTASTKT---RVHHNKEKK 238
L AS+ T VH N +K
Sbjct: 254 -----LVPEYRPASSPTFIPSVHSNSHRK 277
>At1g08010 GATA transcription factor 3, putative
Length = 303
Score = 62.8 bits (151), Expect = 2e-10
Identities = 32/76 (42%), Positives = 40/76 (52%), Gaps = 12/76 (15%)
Query: 166 NNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTA 225
+NS R C+ C T+ TP WR GP+GPK+LCNACG+R R R L A
Sbjct: 213 SNSDGIVRKCTHCETTKTPQWREGPSGPKTLCNACGVRFRSGR---------LVPEYRPA 263
Query: 226 STKT---RVHHNKEKK 238
S+ T VH N +K
Sbjct: 264 SSPTFIPAVHSNSHRK 279
>At3g45170 putative protein
Length = 204
Score = 61.2 bits (147), Expect = 7e-10
Identities = 37/116 (31%), Positives = 55/116 (46%), Gaps = 13/116 (11%)
Query: 151 QGHENRYSQRSPRN--NNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
+G + R S +SP + ++ T + CS C T TPLWR GP G +LCNACG+R R R
Sbjct: 91 RGRQKRLSFKSPSDLFDSKFGITDKSCSHCGTRKTPLWREGPRGAGTLCNACGMRYRTGR 150
Query: 209 RAMAEAANGLATPINTASTKTRVHHNKEKKPRANHFAQFKNKSKSTSSNAGSSQET 264
P ++ K VH N +K + + + KS+ N+ E+
Sbjct: 151 LLPE------YRPASSPDFKPNVHSNFHRK-----VMEIRRERKSSPPNSFGFSES 195
>At4g34680 GATA transcription factor 3
Length = 269
Score = 59.7 bits (143), Expect = 2e-09
Identities = 33/109 (30%), Positives = 49/109 (44%), Gaps = 7/109 (6%)
Query: 131 PATDKANNSTT-IPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSG 189
P T ++ NS T + P + Q + R CS C T++TP WR+G
Sbjct: 137 PRTKRSRNSLTGSRVWPLVSTNHQHAATEQLRKKKQETVLVFQRRCSHCGTNNTPQWRTG 196
Query: 190 PNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVHHNKEKK 238
P GPK+LCNACG+R + R P ++ + +H N +K
Sbjct: 197 PVGPKTLCNACGVRFKSGRLCPE------YRPADSPTFSNEIHSNLHRK 239
>At2g28340 hypothetical protein
Length = 315
Score = 59.7 bits (143), Expect = 2e-09
Identities = 34/86 (39%), Positives = 44/86 (50%), Gaps = 7/86 (8%)
Query: 154 ENRYSQRSPRNNNNSSNTTRV-CSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMA 212
E+ S+ S + + R+ C+ C T++TP WR GPNG K+LCNACGIR R R +
Sbjct: 195 ESESSEISALKKRKKNKSRRLKCTHCETTTTPQWREGPNGRKTLCNACGIRFRSGRLVLE 254
Query: 213 EAANGLATPINTASTKTRVHHNKEKK 238
T I T VH N KK
Sbjct: 255 YRPAASPTFIPT------VHSNLHKK 274
>At5g25830 GATA transcription factor - like
Length = 331
Score = 58.5 bits (140), Expect = 4e-09
Identities = 49/177 (27%), Positives = 72/177 (39%), Gaps = 23/177 (12%)
Query: 86 DNPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRLMKKMMSTPATDKANNSTTIPIS 145
+NP+SS+ S + K +A +S+ L + +P T + S+ +S
Sbjct: 121 ENPNSSSPIFTTDVSVPAKARSKRSRAAACNWASRGLLKETFYDSPFTGETILSSQQHLS 180
Query: 146 P--------------RIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPN 191
P + + GH + SP + R C C T TP WR+GP
Sbjct: 181 PPTSPPLLMAPLGKKQAVDGGHRRKKDVSSPESGGAEE---RRCLHCATDKTPQWRTGPM 237
Query: 192 GPKSLCNACGIRQRKAR-----RAMAEAANGLATPINTASTKTRVHHNKEKKPRANH 243
GPK+LCNACG+R + R R A LA N+ + KE RA+H
Sbjct: 238 GPKTLCNACGVRYKSGRLVPEYRPAASPTFVLAKHSNSHRKVMELRRQKEMS-RAHH 293
>At3g24050 GATA transcription factor 1 (AtGATA-1)
Length = 274
Score = 56.6 bits (135), Expect = 2e-08
Identities = 25/66 (37%), Positives = 33/66 (49%), Gaps = 6/66 (9%)
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVH 232
R C C TP WR+GP GPK+LCNACG+R + R P N+ + +H
Sbjct: 194 RKCQHCGAEKTPQWRAGPAGPKTLCNACGVRYKSGRLVPE------YRPANSPTFTAELH 247
Query: 233 HNKEKK 238
N +K
Sbjct: 248 SNSHRK 253
>At2g45050 putative GATA-type zinc finger transcription factor
Length = 264
Score = 56.6 bits (135), Expect = 2e-08
Identities = 25/51 (49%), Positives = 34/51 (66%), Gaps = 5/51 (9%)
Query: 163 RNNNNSSNTT-----RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKAR 208
R+ ++SS TT R C+ C + TP WR+GP GPK+LCNACG+R + R
Sbjct: 164 RHQSSSSETTEGGGMRRCTHCASEKTPQWRTGPLGPKTLCNACGVRFKSGR 214
>At3g54810 unknown protein
Length = 322
Score = 55.8 bits (133), Expect = 3e-08
Identities = 32/84 (38%), Positives = 42/84 (49%), Gaps = 11/84 (13%)
Query: 173 RVCSDCNTSSTPLWRSGPNGPKSLCNACGIRQRKARRAMAEAANGLATPINTASTKTRVH 232
R C C + TP WR GP GPK+LCNACG+R + R + A+P T + +H
Sbjct: 229 RKCMHCEVTKTPQWRLGPMGPKTLCNACGVRYKSGR--LFPEYRPAASPTFTPA----LH 282
Query: 233 HNKEKKPRANHFAQFKNKSKSTSS 256
N KK A+ +NK S S
Sbjct: 283 SNSHKK-----VAEMRNKRCSDGS 301
>At3g51080 transcription factor-like protein
Length = 312
Score = 55.5 bits (132), Expect = 4e-08
Identities = 36/126 (28%), Positives = 55/126 (43%), Gaps = 20/126 (15%)
Query: 103 EEEDIKNVHGSAKWMSSKMRLMKKMMSTPA---TDKANNSTTI------PISPRIQNQGH 153
EE K+ H + K + R ++ S + TD +++STT P SP G
Sbjct: 131 EETCFKSQHPAVKTRPKRARTGVRVWSHGSQSLTDSSSSSTTSSSSSPRPSSPLWLASGQ 190
Query: 154 -----------ENRYSQRSPRNNNNSSNTTRVCSDCNTSSTPLWRSGPNGPKSLCNACGI 202
+ + + + + + TR C C TP WR+GP G K+LCNACG+
Sbjct: 191 FLDEPMTKTQKKKKVWKNAGQTQTQTQTQTRQCGHCGVQKTPQWRAGPLGAKTLCNACGV 250
Query: 203 RQRKAR 208
R + R
Sbjct: 251 RYKSGR 256
>At4g32890 unknown protein
Length = 308
Score = 55.1 bits (131), Expect = 5e-08
Identities = 41/144 (28%), Positives = 63/144 (43%), Gaps = 21/144 (14%)
Query: 65 SSSSSSNHLYNSTFISPQPVMDNPSSSTCDPNLSFSKMEEEDIKNVHGSAKWMSSKMRLM 124
++ S+ HL I P+P +D+ + + D NV AK S + R
Sbjct: 110 TTGSTLTHL-----IKPEPELDH-------------QFIDIDESNVAVPAKARSKRSRSA 151
Query: 125 KKMMSTPATDKANNSTTIPISPRIQNQGHENRYSQRSPRNNNNSSNTTRVCSDCNTSSTP 184
++ A++ T P + Q + E ++ + S R C C T TP
Sbjct: 152 ASTWASRLLSLADSDETNP--KKKQRRVKEQDFAGDMDVDCGESGGGRR-CLHCATEKTP 208
Query: 185 LWRSGPNGPKSLCNACGIRQRKAR 208
WR+GP GPK+LCNACG+R + R
Sbjct: 209 QWRTGPMGPKTLCNACGVRYKSGR 232
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.125 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,538,874
Number of Sequences: 26719
Number of extensions: 327505
Number of successful extensions: 1177
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 1117
Number of HSP's gapped (non-prelim): 95
length of query: 310
length of database: 11,318,596
effective HSP length: 99
effective length of query: 211
effective length of database: 8,673,415
effective search space: 1830090565
effective search space used: 1830090565
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0019a.4


