

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0016.15
(1469 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
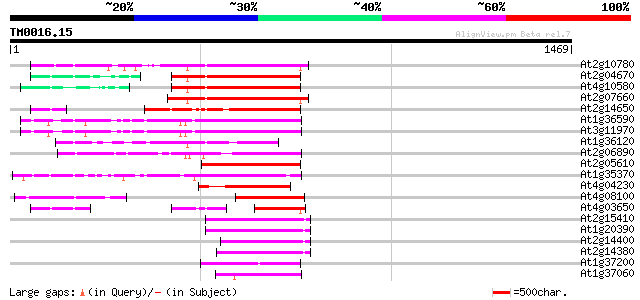
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 446 e-125
At2g04670 putative retroelement pol polyprotein 396 e-110
At4g10580 putative reverse-transcriptase -like protein 392 e-109
At2g07660 putative retroelement pol polyprotein 381 e-105
At2g14650 putative retroelement pol polyprotein 353 3e-97
At1g36590 hypothetical protein 300 3e-81
At3g11970 hypothetical protein 299 7e-81
At1g36120 putative reverse transcriptase gb|AAD22339.1 299 9e-81
At2g06890 putative retroelement integrase 270 5e-72
At2g05610 putative retroelement pol polyprotein 268 2e-71
At1g35370 hypothetical protein 268 2e-71
At4g04230 putative transposon protein 196 1e-49
At4g08100 putative polyprotein 190 5e-48
At4g03650 putative reverse transcriptase 171 2e-42
At2g15410 putative retroelement pol polyprotein 162 1e-39
At1g20390 hypothetical protein 159 9e-39
At2g14400 putative retroelement pol polyprotein 159 1e-38
At2g14380 putative retroelement pol polyprotein 145 2e-34
At1g37200 hypothetical protein 144 4e-34
At1g37060 Athila retroelment ORF 1, putative 139 2e-32
>At2g10780 pseudogene
Length = 1611
Score = 446 bits (1147), Expect = e-125
Identities = 289/776 (37%), Positives = 394/776 (50%), Gaps = 96/776 (12%)
Query: 55 FSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTRGIMEDNHEE 114
F G TDP AD W +++ F + E + +A + L GDA WW D
Sbjct: 182 FMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVEKRRGDEVRS 241
Query: 115 IN*NSFRMAFLEKYFPTSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNE 174
F F +KYFP A D E +L L QG +V E+ + L ++ + E
Sbjct: 242 FA--DFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVG--RELEEE 297
Query: 175 RYMCKRFVNGLRPDIEDSVRPLGIMRFQSLVEKATEVE--LMKNRRLNRAGTGGPMRSGS 232
+ +RF+ GLR +I + LVE+A +E + + R LNR +
Sbjct: 298 QAQLRRFIRGLRIEIRNHCLVRTFNSVSELVERAAMIEEGIEEERYLNREKAPIRNNQST 357
Query: 233 QNFQNRGKFQNRRPYQRPAGRGFA-------SGSYRPMVGAAGGSGDQTQNRELTCFKCG 285
+ + KF + A G SG+ +GA G +CG
Sbjct: 358 KPADKKRKFDKVDNTKSDAKTGECVTCGKNHSGTCWKAIGACG--------------RCG 403
Query: 286 KPGHFARACPDTRP-----------QCYNCNKLGHTAAQCRVPKAEPTVNTA--RG---- 328
H ++CP P C+ C K GH +C AE RG
Sbjct: 404 SKDHAIQSCPKMEPGQSKVLGEETRTCFYCGKTGHLKRECPKLTAEKQAGQRDNRGGNGL 463
Query: 329 ----KRPAAKARVYTMDGDDTEGVDGLIRGDCEIDGNLLSVLFDSGATHSFISRECAIRL 384
KR A +RVY + + + GN ++ T F
Sbjct: 464 PPPPKRQAVASRVYELSEEANDA------------GNFRAI------TGGFRKE------ 499
Query: 385 RLPITALSFDLIVTTPAKTLLANTACMYCSVTYNNRVYHANLVCLTLKNLDVILGMDWLS 444
P T + ++ + + SV N A+L+ + LK DVILGMDWL
Sbjct: 500 --PNT--DYGMVRAAGGQAMYPTGLVRGISVVVNGVNMPADLIIVPLKKHDVILGMDWLG 555
Query: 445 HYHCLLDCN*KRVVFP--DSVL----------SEYLFANHISVSLNGGSQEYSVLLSLES 492
Y +DC+ R+ F + +L S + A L G + Y ++ +
Sbjct: 556 KYKAHIDCHRGRIQFERDEGMLKFQGIRTTSGSLVISAIQAERMLGKGCEAYLATITTKE 615
Query: 493 KG-NPEVDSIPVVKEFTDVFPGDVPGLPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVE 551
G + E+ IP+V EF+DVF V G+PP R F I++ PGT PIS APYRMAPAE+ +
Sbjct: 616 VGASAELKDIPIVNEFSDVFAA-VSGVPPDRSDPFTIELEPGTTPISKAPYRMAPAEMAK 674
Query: 552 LKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDD 611
LK QLE+LL KGFIRPS SPWGAPVL VKKKDG RLC+DYR LNKVTVKN+YPLPRID+
Sbjct: 675 LKKQLEELLDKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDE 734
Query: 612 LMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMD 671
LMDQL GA FSKIDL SGYHQI ++ D++KTAFRTRY H+E++VMPFG+TNAPA FM
Sbjct: 735 LMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYDHFEFVVMPFGLTNAPAAFMK 794
Query: 672 YMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEE 731
MN F FLD FV++FI+DIL+YSK+ H+ HLR VL+ LRE EL+A SKC FW
Sbjct: 795 MMNGVFRDFLDEFVIIFINDILVYSKSWEAHQEHLRAVLERLREHELFAKLSKCSFWQRS 854
Query: 732 VKFLGHVISKEGIAVDPAKVETVLSWERP------KTVLGVSLVWRDIIVALSRIS 781
V FLGHVIS +G++VDP K+ ++ W RP ++ LG++ +R +++ + ++
Sbjct: 855 VGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMA 910
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 396 bits (1018), Expect = e-110
Identities = 201/350 (57%), Positives = 247/350 (70%), Gaps = 14/350 (4%)
Query: 424 ANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVF--PDSVL----------SEYLFAN 471
A+L+ ++ DVILGMDWL HY LDC+ RV F P+ L S + A
Sbjct: 393 ADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGRLVYQGVRPTSGSLVISAV 452
Query: 472 HISVSLNGGSQEYSVLLSL-ESKGNPEVDSIPVVKEFTDVFPGDVPGLPPVRDIEFAIDI 530
+ G + Y V +S+ ES G V I V++EF DVF + GLPP R F I++
Sbjct: 453 QAKKMIEKGCEAYLVTISMPESLGQVAVSDIRVIQEFEDVFQS-LQGLPPSRSDPFTIEL 511
Query: 531 VPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDGKSRLCV 590
PGT P+S APYRMAPAE+ ELK QLEDLL KGFIRPS SPWGAPVL VKKKDG RLC+
Sbjct: 512 EPGTAPLSKAPYRMAPAEMTELKKQLEDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCI 571
Query: 591 DYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRY 650
DYR LN VTVKN+YPLPRID+L+DQLRGA FSKIDL SGYHQI + D++KTAFRTRY
Sbjct: 572 DYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRY 631
Query: 651 GHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVL 710
GH+E++VMPF +TNAPA FM MN F FLD FV++FIDDIL+YSK+ EHE HLR+V+
Sbjct: 632 GHFEFVVMPFALTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEHEVHLRRVM 691
Query: 711 QVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP 760
+ LRE++L+A SKC FW E+ FLGH++S EG++VDP K+E + W RP
Sbjct: 692 EKLREQKLFAKLSKCSFWQREIGFLGHIVSAEGVSVDPEKIEAIRDWPRP 741
Score = 86.7 bits (213), Expect = 9e-17
Identities = 77/287 (26%), Positives = 114/287 (38%), Gaps = 41/287 (14%)
Query: 55 FSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTRGIMEDNHEE 114
FS GT P++AD W +++ FG + ++ +A + L GDA WW+ +
Sbjct: 138 FSSGTSPEEADSWRSRVQRNFGSSRCPAEYRVDLAVHFLEGDAHLWWRSVTA--RRRQAD 195
Query: 115 IN*NSFRMAFLEKYFPTSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNE 174
++ F F KYFP A D E++FL L QG SV E+ + L + ++
Sbjct: 196 MSWADFVAEFNAKYFPQEALDRMEARFLELTQGEWSVREYDREFNRLLAYAG--RGMEDD 253
Query: 175 RYMCKRFVNGLRPDIEDSVRPLGIMRFQSLVEKATEVELMKNRRLNRAGTGGPMRSGSQN 234
+ +RF+ GLRPD+ R +LVE A EVE L R G ++N
Sbjct: 254 QAQMRRFLRGLRPDLRVQCRVSQYATKAALVETAAEVE----EDLQRQVVGVSPAVQTKN 309
Query: 235 FQNRGKFQNRRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARAC 294
Q + + + GG Q Q R K H +RA
Sbjct: 310 TQQQ------------------------VTPSKGGKPAQGQKR--------KWDHPSRAG 337
Query: 295 PDTRPQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYTMD 341
R ++C L HT A C + V A G+ A R +D
Sbjct: 338 QGGRAGYFSCGSLDHTVADC-TERGHLHVKVAGGQFLAVLGRAKGVD 383
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 392 bits (1007), Expect = e-109
Identities = 199/350 (56%), Positives = 249/350 (70%), Gaps = 14/350 (4%)
Query: 424 ANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVF--PDSVL----------SEYLFAN 471
A+L+ ++ DVILGMDWL +Y LD + RV F P+ L S + A
Sbjct: 367 ADLIISPVELYDVILGMDWLDYYRVHLDWHRGRVFFERPEGRLVYQGVRPISGSLVISAV 426
Query: 472 HISVSLNGGSQEYSVLLSL-ESKGNPEVDSIPVVKEFTDVFPGDVPGLPPVRDIEFAIDI 530
+ G + Y V +S+ ES G V I VV+EF DVF + GLPP + F I++
Sbjct: 427 QAEKMIEKGCEAYLVTISMPESVGQVAVSDIRVVQEFQDVFQS-LQGLPPSQSDPFTIEL 485
Query: 531 VPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDGKSRLCV 590
PGT P+S APYRMAPAE+ ELK QL+DLL KGFIRPS SPWGAPVL VKKKDG RLC+
Sbjct: 486 EPGTAPLSKAPYRMAPAEMAELKKQLKDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCI 545
Query: 591 DYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRY 650
DYR+LN+VTVKNRYPLPRID+L+DQLRGA FSKIDL SGYHQI + D++KTAFRTRY
Sbjct: 546 DYRELNRVTVKNRYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRY 605
Query: 651 GHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVL 710
GH+E++VMPFG+TNAPA+FM MN F FLD FV++FIDDIL+YSK+ E E HLR+V+
Sbjct: 606 GHFEFVVMPFGLTNAPAVFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEQEVHLRRVM 665
Query: 711 QVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP 760
+ LRE++L+A SKC FW E+ FLGH++S EG++VDP K+E + W RP
Sbjct: 666 EKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPRP 715
Score = 66.2 bits (160), Expect = 1e-10
Identities = 68/285 (23%), Positives = 104/285 (35%), Gaps = 67/285 (23%)
Query: 28 EQHQRQREVTLDQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLG 87
+Q E L + + +R D FSGGT P++AD W + + FG + ++
Sbjct: 125 QQRAAVAEEVLSYLRMMEQLQRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVD 184
Query: 88 MATYLLLGDAEYWWKGTRGIMEDNHEEIN*NSFRMAFLEKYFPTSARDERESQFLTLRQG 147
+A + L GDA WW+ +++ F F KYFP A D Q
Sbjct: 185 LAVHFLEGDAHLWWRSVTA--RRRQTDMSWADFVAEFKAKYFPQEALDPYAGQ------- 235
Query: 148 GMSVPEFASKLESLAKHFQFFHDHVNERYMCKRFVNGLRPDIEDSVRPLGIMRFQSLVEK 207
GM +++ +RF+ GLRPD+ R +LVE
Sbjct: 236 GME----------------------DDQAQMRRFLRGLRPDLRVRCRVSQYATKAALVET 273
Query: 208 ATEVELMKNRRLNRAGTGGPMRSGSQNFQNRGKFQNRRPYQRPAGRGFASGSYRPMVGAA 267
A EVE ++FQ + P +P + + + +
Sbjct: 274 AAEVE--------------------EDFQR--QVVGVSPVVQP------KKTQQQVTPSK 305
Query: 268 GGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYNCNKLGHTAA 312
GG Q Q R K H +RA R +C++C L H A
Sbjct: 306 GGKPAQGQKR--------KWDHPSRAGQGGRARCFSCGSLDHKDA 342
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 381 bits (978), Expect = e-105
Identities = 202/387 (52%), Positives = 262/387 (67%), Gaps = 20/387 (5%)
Query: 414 SVTYNNRVYHANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVFP--DSVL------- 464
SV N A+L+ + LK DVILGMDWL Y +DC+ +R+ F + +L
Sbjct: 20 SVVVNGVNMPADLIIVPLKKHDVILGMDWLGKYKGHIDCHRERIQFERDEGMLKFQGIRT 79
Query: 465 ---SEYLFANHISVSLNGGSQEYSVLLSLESKG-NPEVDSIPVVKEFTDVFPGDVPGLPP 520
S + A L G + Y ++ + G + E+ I +V EF+DVF V G+PP
Sbjct: 80 TSGSLVISAIQAERMLGKGCEAYLATITTKEVGASAELKDILIVNEFSDVFAA-VSGVPP 138
Query: 521 VRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVK 580
R F I++ PGT PIS APYRMAPAE+ ELK QLE+LL+KGFIRPS SPWGAPVL VK
Sbjct: 139 DRSDPFTIELEPGTTPISKAPYRMAPAEMAELKKQLEELLAKGFIRPSSSPWGAPVLFVK 198
Query: 581 KKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTED 640
KKDG RLC+DYR LNKVTVKN+YPLPRID+LMDQL GA FSKIDL SGYHQI ++ D
Sbjct: 199 KKDGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTD 258
Query: 641 IQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVV 700
++KTAFRTRYGH+E++VMPFG+TNAPA FM MN F FLD FV++FIDDIL++SK+
Sbjct: 259 VRKTAFRTRYGHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFIDDILVHSKSWE 318
Query: 701 EHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP 760
H+ HLR VL+ LRE EL+A SK FW V FLGHVIS +G++VDP K+ ++ W RP
Sbjct: 319 AHQEHLRAVLERLREHELFAKLSKFSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRP 378
Query: 761 ------KTVLGVSLVWRDIIVALSRIS 781
++ LG++ +R +++ + ++
Sbjct: 379 RNATEIRSFLGLAGYYRRFVMSFASMA 405
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 353 bits (907), Expect = 3e-97
Identities = 202/412 (49%), Positives = 257/412 (62%), Gaps = 30/412 (7%)
Query: 354 GDCEIDGNLLSVLFDSGATHSFISRECAIRLRL---PITALSFDLIVTTPAKTLLANTAC 410
G + G VLFDSGA+H FI+ E A R + P L + +L T
Sbjct: 304 GTLLVGGVEAHVLFDSGASHCFITPESASRGNIRGDPGEQLGAVKVAGGQFVAVLGRTKG 363
Query: 411 MYCSVTYNNRVYHANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVF--PDSVLSEYL 468
+ + + A+L+ ++ DVILGMDWL HY LDC+ RV F P+ L Y
Sbjct: 364 VDIQIAGESMP--ADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGSLV-YQ 420
Query: 469 FANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVVKEFTDVFPGDVPGLPPVRDIEFAI 528
S SL V+ +++++ E EF DVF + GLPP R F I
Sbjct: 421 GVRPTSGSL--------VISAVQAEKMIE-KGCEAYLEFEDVFQS-LQGLPPSRSDPFTI 470
Query: 529 DIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDGKSRL 588
++ PGT P+S APYRMAPAE+ ELK QLEDLL GAPVL VKKKDG RL
Sbjct: 471 ELEPGTAPLSKAPYRMAPAEMAELKKQLEDLL------------GAPVLFVKKKDGSFRL 518
Query: 589 CVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRT 648
C+DYR LN VTVKN+YPLPRID+L+DQLRGA FSKIDL SGYH I + D++KTAFRT
Sbjct: 519 CIDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEADVRKTAFRT 578
Query: 649 RYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQ 708
RYGH+E++VMPFG+TNAPA FM MN F LD FV++FIDDIL+YSK++ EHE HLR+
Sbjct: 579 RYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLEEHEVHLRR 638
Query: 709 VLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERP 760
V++ LRE++L+A SKC FW E+ FLGH++S EG++VDP K+E + W P
Sbjct: 639 VMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTP 690
Score = 65.1 bits (157), Expect = 3e-10
Identities = 34/93 (36%), Positives = 46/93 (48%), Gaps = 2/93 (2%)
Query: 55 FSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTRGIMEDNHEE 114
FSGGT P+ AD W +E+ FG + ++ +A + L GDA WW+ +
Sbjct: 156 FSGGTSPEVADSWRSRVERNFGSSRCPAEYRIDLAVHFLEGDAHLWWRSVTA--RRRQAD 213
Query: 115 IN*NSFRMAFLEKYFPTSARDERESQFLTLRQG 147
I+ F F KYFP A D E+ FL L QG
Sbjct: 214 ISWADFVAEFNAKYFPQEALDRMEAHFLELTQG 246
>At1g36590 hypothetical protein
Length = 1499
Score = 300 bits (769), Expect = 3e-81
Identities = 231/798 (28%), Positives = 362/798 (44%), Gaps = 123/798 (15%)
Query: 29 QHQRQREVTLDQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGM 88
+HQ+ R + L + D P+F G + W+ ++E+ FGV T E K+ M
Sbjct: 91 RHQQVRSDHFNAYNNLTRLGKIDFPRFDG----TRLKEWLFKVEEFFGVDSTPEDMKVKM 146
Query: 89 ATYLLLGDAEYW----------------WKGTRGIMEDNHEEIN*NSFRMAFLEKYFPTS 132
A A W WKG ++++ E+
Sbjct: 147 AAIHFDSHASTWHQSFIQSGVGLEVLYDWKGYVKLLKERFED------------------ 188
Query: 133 ARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNERYMCKRFVNGLRPDIEDS 192
D+ ++ L++ + ++ K E + +++E Y+ ++ GLR D +
Sbjct: 189 DCDDPMAELKHLQETD-GIIDYHQKFELIKTRV-----NLSEEYLVSVYLAGLRTDTQMH 242
Query: 193 VRP-----------LGIMRFQSLVEKATEVELMKNRRLNRAGTGGPMRSGSQNFQNRGKF 241
VR LG ++ ++K NR G + G + G
Sbjct: 243 VRMFQPQTVRHCLFLGKTYEKAHLKKPANTTWSTNRSAPTGGYNKYQKEGESKTDHYGNK 302
Query: 242 QNRRPY-QRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGK---PGHFARACPDT 297
N +P Q+P S R G C+ C + P H+
Sbjct: 303 GNFKPVSQQPKKMSQQEMSDRRSKGL--------------CYFCDEKYTPEHYL---VHK 345
Query: 298 RPQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYT-MDGDDTEGVDGLIRGDC 356
+ Q + + + R E VN P + + G T V G
Sbjct: 346 KTQLFRMD-VDEEFEDAR----EELVNDDDEHMPQISVNAVSGIAGYKTMRVKGTY---- 396
Query: 357 EIDGNLLSVLFDSGATHSFISRECAIRLRLPITALSFDLIVTTPAKTLLANTACMYCSVT 416
D ++ +L DSG+TH+F+ A +L + + + L S
Sbjct: 397 --DKKIIFILIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFSWK 454
Query: 417 YNNRVYHANLVCLTLKNLDVILGMDWLSHY-----------------------HCLLDCN 453
+ ++++ + L+ +D++LG+ WL H L +
Sbjct: 455 LQTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGS 514
Query: 454 *KRV-------VFPDSVLSEYLFANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVVKE 506
+ V + D V L +S S G E + +L S+ E V+ E
Sbjct: 515 VREVKAQKLQKLQEDQVQLAMLCVQEVSESTEG---ELCTINALTSELGEESVVEEVLNE 571
Query: 507 FTDVFPGDVPGLPPVRDIE-FAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFI 565
+ D+F + LPP R+ I ++ G+ P++ PYR + + E+ +EDLL+ G +
Sbjct: 572 YPDIFI-EPTALPPFREKHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTV 630
Query: 566 RPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKI 625
+ S SP+ +PV+LVKKKDG RLCVDYR+LN +TVK+ +P+P I+DLMD+L GA IFSKI
Sbjct: 631 QASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKI 690
Query: 626 DLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFV 685
DL++GYHQ+R+ +DIQKTAF+T GH+EYLVMPFG+TNAPA F MN F FL +FV
Sbjct: 691 DLRAGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFV 750
Query: 686 VVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIA 745
+VF DDIL+YS ++ EH HL+QV +V+R +L+A SKC F + +V++LGH IS +GI
Sbjct: 751 LVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIE 810
Query: 746 VDPAKVETVLSWERPKTV 763
DPAK++ V W +P T+
Sbjct: 811 TDPAKIKAVKEWPQPTTL 828
>At3g11970 hypothetical protein
Length = 1499
Score = 299 bits (766), Expect = 7e-81
Identities = 232/800 (29%), Positives = 359/800 (44%), Gaps = 127/800 (15%)
Query: 29 QHQRQREVTLDQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGM 88
+HQ+ R + L + D P+F G + W+ ++E+ FGV T E K+ M
Sbjct: 91 RHQQVRSDHFNAYNNLTRLGKIDFPRFDG----TRLKEWLFKVEEFFGVDSTPEDMKVKM 146
Query: 89 ATYLLLGDAEYW----------------WKGTRGIMEDNHEEIN*NSFRMAFLEKYFPTS 132
A A W WKG ++++ E
Sbjct: 147 AAIHFDSHASTWHQSFIQSGVGLEVLYDWKGYVKLLKERFE------------------- 187
Query: 133 ARDERESQFLTLR--QGGMSVPEFASKLESLAKHFQFFHDHVNERYMCKRFVNGLRPDIE 190
D+ + L+ Q + ++ K E + +++E Y+ ++ GLR D +
Sbjct: 188 --DDCDDPMAELKHLQETDGIIDYHQKFELIKTRV-----NLSEEYLVSVYLAGLRTDTQ 240
Query: 191 DSVRP-----------LGIMRFQSLVEKATEVELMKNRRLNRAGTGGPMRSGSQNFQNRG 239
VR LG ++ +K NR G + G + G
Sbjct: 241 MHVRMFQPQTVRHCLFLGKTYEKAHPKKPANTTWSTNRSAPTGGYNKYQKEGESKTDHYG 300
Query: 240 KFQNRRPY-QRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGK---PGHFARACP 295
N +P Q+P S R G C+ C + P H+
Sbjct: 301 NKGNFKPVSQQPKKMSQQEMSDRRSKGL--------------CYFCDEKYTPEHYL---V 343
Query: 296 DTRPQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYT-MDGDDTEGVDGLIRG 354
+ Q + + + R E VN P + + G T V G
Sbjct: 344 HKKTQLFRMD-VDEEFEDAR----EELVNDDDEHMPQISVNAVSGIAGYKTMRVKGTY-- 396
Query: 355 DCEIDGNLLSVLFDSGATHSFISRECAIRLRLPITALSFDLIVTTPAKTLLANTACMYCS 414
D ++ +L DSG+TH+F+ A +L + + + L S
Sbjct: 397 ----DKKIIFILIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVSVADGRKLRVEGKVTDFS 452
Query: 415 VTYNNRVYHANLVCLTLKNLDVILGMDWLSHY-----------------------HCLLD 451
+ ++++ + L+ +D++LG+ WL H L
Sbjct: 453 WKLQTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTS 512
Query: 452 CN*KRV-------VFPDSVLSEYLFANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVV 504
+ + V + D V L +S S G E + +L S+ E V+
Sbjct: 513 GSVREVKAQKLQKLQEDQVQLAMLCVQEVSESTEG---ELCTINALTSELGEESVVEEVL 569
Query: 505 KEFTDVFPGDVPGLPPVRDIE-FAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKG 563
E+ D+F + LPP R+ I ++ G+ P++ PYR + + E+ +EDLL+ G
Sbjct: 570 NEYPDIFI-EPTALPPFREKHNHKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNG 628
Query: 564 FIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFS 623
++ S SP+ +PV+LVKKKDG RLCVDYR+LN +TVK+ +P+P I+DLMD+L GA IFS
Sbjct: 629 TVQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFS 688
Query: 624 KIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDR 683
KIDL++GYHQ+R+ +DIQKTAF+T GH+EYLVMPFG+TNAPA F MN F FL +
Sbjct: 689 KIDLRAGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRK 748
Query: 684 FVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEG 743
FV+VF DDIL+YS ++ EH HL+QV +V+R +L+A SKC F + +V++LGH IS +G
Sbjct: 749 FVLVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQG 808
Query: 744 IAVDPAKVETVLSWERPKTV 763
I DPAK++ V W +P T+
Sbjct: 809 IETDPAKIKAVKEWPQPTTL 828
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 299 bits (765), Expect = 9e-81
Identities = 220/600 (36%), Positives = 288/600 (47%), Gaps = 86/600 (14%)
Query: 120 FRMAFLEKYFPTSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNERYMCK 179
F F KYFP A D E++FL L +G +SV E+ + + L + +++ +
Sbjct: 157 FVAEFNAKYFPQEALDRMEARFLELTKGELSVREYGREFKRLLVYAG--RGMEDDQAQMR 214
Query: 180 RFVNGLRPDIEDSVRPLGIMRFQSLVEKATEVELMKNRRLNRAGTGGPMRSGSQNFQNRG 239
RF+ GLRPD+ R +L E +L + + + S + Q +
Sbjct: 215 RFLRGLRPDLRVRCRVSQYATKAALTAAEVEEDLQRQ-----------VVAVSPSVQPKK 263
Query: 240 KFQNRRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTC-FKCGKPGHFARACPDTR 298
Q P + GG Q Q R+ F+ G+ G A D+
Sbjct: 264 IQQQVAP-------------------SKGGKPAQVQKRKWDHPFRAGQGGRAGSAADDSS 304
Query: 299 PQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYTMDGDDTEGVDGLIRGDCEI 358
TA A + AR A Y G LI G C
Sbjct: 305 TS---------TAGG-----AAQSAGAARSAAECAAVSTYC-SGTTWLYYADLISGRCR- 348
Query: 359 DGNLLSVLFDSGATHSFISRECAIRLRLP-ITALSFDLIVTTPAKTLLANTACMYCSVTY 417
G+ VLFDSGA+H FI+ E A R + F + + L ++
Sbjct: 349 -GH---VLFDSGASHCFITPESASRDNIRGEPGEQFGAVKIAGGQFLTVLGRAKGVNIQI 404
Query: 418 NNRVYHANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVF--PDSVL--------SEY 467
A+L+ + D ILGMDWL HY LDC+ RV F P+ L S
Sbjct: 405 EEESMPADLIINPVVLYDAILGMDWLDHYRVHLDCHRGRVSFERPEGRLVYQGVRPTSRS 464
Query: 468 LFANHISVS--LNGGSQEYSVLLSL-ESKGNPEVDSIPVVKEFTDVFPGDVPGLPPVRDI 524
L + + V + G + Y V +S+ ES G V I VV+EF DVF + GLPP R
Sbjct: 465 LVISEVQVEKMIEKGCEAYLVTISMSESVGQVAVSDIWVVQEFEDVFQS-LQGLPPSRSD 523
Query: 525 EFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDG 584
F I++ GT P+S PYRM PAE+ ELK QLEDLL KGFIRPS S WGAP
Sbjct: 524 PFTIELELGTAPLSKTPYRMVPAEIAELKKQLEDLLGKGFIRPSTSRWGAP--------- 574
Query: 585 KSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKT 644
LN+VT+KN+YPLPRID+L+DQLRGA FSKIDL GYHQ + D++KT
Sbjct: 575 ---------GLNRVTLKNKYPLPRIDELLDQLRGATCFSKIDLTPGYHQFPIAEADVRKT 625
Query: 645 AFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEG 704
AFRTRYGH+E++VMPFG+TNAP M MN F FLD FV++FIDDIL+Y K+ EHEG
Sbjct: 626 AFRTRYGHFEFVVMPFGLTNAPTALMRLMNSVFQEFLDEFVIIFIDDILVYFKSPEEHEG 685
>At2g06890 putative retroelement integrase
Length = 1215
Score = 270 bits (690), Expect = 5e-72
Identities = 202/685 (29%), Positives = 312/685 (45%), Gaps = 92/685 (13%)
Query: 126 EKYFPTSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNERYMCKRFVNGL 185
+++ P+ E + L QG SV E+ ++E+L D RF+ L
Sbjct: 7 KRFVPSHYHRELHLKLRNLTQGNRSVEEYYKEMETLMLRADISEDR---EATLSRFLGDL 63
Query: 186 RPDIEDSVRPLGIMRFQSLVEKATEVEL-MKNRRLNRAGTG-GPMRSGSQNFQNRGKFQN 243
DI+D + ++ + ++ KA E +K + +R+ G G + + + R +
Sbjct: 64 NRDIQDRLETQYYVQIEEMLHKAILFEQQVKRKSSSRSSYGSGTIAKPTYQREERTSSYH 123
Query: 244 RRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGKPGHFARACPDTRPQCYN 303
+P P + + G A S + R++ C+KC GH+A CP+ R
Sbjct: 124 NKPIVSPRAESKPYAAVQDHKGKAEISTSRV--RDVRCYKCQGKGHYANECPNKRVMILL 181
Query: 304 CNKLGHTAAQCRVPKAEPTVNT-----ARGKRPAAKARVYTMDG-DDTEGVDGLIRGDCE 357
N G + +P + ++ A+G+ A+ + D+ E L C
Sbjct: 182 DN--GEIEPEEEIPDSPSSLKENEELPAQGELLVARRTLSVQTKTDEQEQRKNLFHTRCH 239
Query: 358 IDGNLLSVLFDSGATHSFISRECAIRLRLPITALSFDLIVTTPAKTLLANTACMYCSVTY 417
+ G + S++ D G+ + S +L L + K + N + +
Sbjct: 240 VHGKVCSLIIDGGSCTNVASETMVKKLGLKW--------LNDSGKMRVKNQVVVPIVIGK 291
Query: 418 NNRVYHANLVC--LTLKNLDVILGMDWLSHYHCLLDCN*KRVVF-------------PDS 462
Y ++C L ++ ++LG W S + D R F P
Sbjct: 292 ----YEDEILCDVLPMEAGHILLGRPWQSDRKVMHDGFTNRHSFEFKGGKTILVSMTPHE 347
Query: 463 VLSEYL---------------FANHISVSLNGGSQEYSVLLSLESKGNPEVDSIPVV--- 504
V + + FA V S++ +L + + PV+
Sbjct: 348 VYQDQIHLKQKKEQVVKQPNFFAKSGEVKSAYSSKQPMLLFVFKEALTSLTNFAPVLPSE 407
Query: 505 -----KEFTDVFPGDVP-GLPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLED 558
+++ DVFP D P GLPP+R IE ID VPG + YR P E EL+ Q
Sbjct: 408 MTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVETKELQRQ--- 464
Query: 559 LLSKGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRG 618
KDG R+C D R +N VTVK +P+PR+DD++D+L G
Sbjct: 465 -----------------------KDGSWRMCFDCRAINNVTVKYCHPIPRLDDMLDELHG 501
Query: 619 AAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFH 678
++IFSKIDLKSGYHQIR+ D KTAF+T++G YE+LVMPFG+T+AP+ FM MN
Sbjct: 502 SSIFSKIDLKSGYHQIRMNEGDEWKTAFKTKHGLYEWLVMPFGLTHAPSTFMRLMNHVLR 561
Query: 679 TFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHV 738
F+ FV+V+ DDIL+YS+++ EH HL VL VLR++ELYAN KC F + + FLG V
Sbjct: 562 AFIGIFVIVYFDDILVYSESLREHIEHLDSVLNVLRKEELYANLKKCTFCTDNLVFLGFV 621
Query: 739 ISKEGIAVDPAKVETVLSWERPKTV 763
+S +G+ VD KV+ + W PKTV
Sbjct: 622 VSADGVKVDEEKVKAIRDWPSPKTV 646
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 268 bits (685), Expect = 2e-71
Identities = 134/259 (51%), Positives = 182/259 (69%), Gaps = 2/259 (0%)
Query: 503 VVKEFTDVFPGDVPGLPPVR-DIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLS 561
VV EF D+F + LPP R + I+++ + P++ PYR + E+ ++D+L+
Sbjct: 10 VVTEFPDIFV-EPTDLPPFRAPHDHKIELLEDSNPVNQRPYRYVVHQKNEIDKIVDDMLA 68
Query: 562 KGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAI 621
G I+ S SP+ +PV+LVKKKDG RLCVDYR+LN +TVK+R+P+P I+DLMD+L G+ +
Sbjct: 69 SGTIQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELGGSNV 128
Query: 622 FSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFL 681
+SKIDL++GYHQ+R+ DI KTAF+T GHYEYLVMPFG+TNAPA F MN F FL
Sbjct: 129 YSKIDLRAGYHQVRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQSLMNSFFKPFL 188
Query: 682 DRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISK 741
+FV+VF DDILIYS ++ EH+ HL V +V+R L+A SKC F + V++LGH IS
Sbjct: 189 RKFVLVFFDDILIYSTSMEEHKKHLEAVFEVMRVHHLFAKMSKCAFAVPRVEYLGHFISG 248
Query: 742 EGIAVDPAKVETVLSWERP 760
EGIA DPAK++ V W P
Sbjct: 249 EGIATDPAKIKAVQDWPVP 267
>At1g35370 hypothetical protein
Length = 1447
Score = 268 bits (684), Expect = 2e-71
Identities = 229/808 (28%), Positives = 363/808 (44%), Gaps = 129/808 (15%)
Query: 8 QAVTVQANDSAMRRAAEEAREQHQRQ------REVTLDQNKGLNDFRRHDPPKFSGGTDP 61
++V + S R E + QR R+ +Q L + D P+F G
Sbjct: 69 RSVPRSSTSSMARSGFVEHQRDFQRDFRPEMVRQEVNNQYGNLTRLGKIDFPRFDGS--- 125
Query: 62 DKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTRGIMEDNHEEIN*NSFR 121
+ + W+ ++E+ FGV T E K+ M A W I ++ N
Sbjct: 126 -RINEWLFKVEEFFGVDFTPEEMKVKMVAIHFDSHAATWHHSF--IQSGIGLDVFFNWPE 182
Query: 122 MAFLEKYFPTSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNERYMCKRF 181
L K A D+ ++ L++ + E+ + E + +++E Y+ +
Sbjct: 183 YVKLLKDRFEDACDDPMAELKKLQETD-GIVEYHQQFELIKVRL-----NLSEEYLVSVY 236
Query: 182 VNGLRPDIEDSVRPLGIMRFQSLVEKATEVELMK-NRRLNRAGTGGPMRSGSQNFQNRGK 240
+ GLR D + VR + E T + ++ + RA P ++ S + +G
Sbjct: 237 LAGLRTDTQMHVR---------MFEPKTVRDCLRLGKYYERAH---PKKTVSSTWSQKGT 284
Query: 241 FQNRRPYQRPAGRGFASGSYRPMVGAAGGSGDQTQNRELTCFKCGK---PGHFARACP-- 295
+ GSYRP+ +Q + C+ C + P H+
Sbjct: 285 R--------------SGGSYRPVKEV-----EQKSDHLGLCYFCDEKFTPEHYLVHKKTQ 325
Query: 296 ----DTRPQCYNCNKLGHTAAQCRVPKAEPTVNTARGKRPAAKARVYTMDGDDTEGVDGL 351
D + + ++ + P + +VN G + G T GV G
Sbjct: 326 LFRMDVDEEFEDAVEVLSDDDHEQKPMPQISVNAVSG-----------ISGYKTMGVKGT 374
Query: 352 IRGDCEIDGNLLSVLFDSGATHSFISRECAIRLRLPITALSFDLIVTTPAKTLLANTACM 411
+ D L +L DSG+TH+FI A +L + + + + L +
Sbjct: 375 V------DKRDLFILIDSGSTHNFIDSTVAAKLGCHVESAGLTKVAVADGRKLNVDGQIK 428
Query: 412 YCSVTYNNRVYHANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVFPDSVLSEYLFAN 471
+ + + ++++ + L+ +D++LG+ WL R+ + L F
Sbjct: 429 GFTWKLQSTTFQSDILLIPLQGVDMVLGVQWLETLG--------RISWEFKKLEMQFFYK 480
Query: 472 HISVSLNG---GS--------------------------------QEYSVLLSLESKGNP 496
+ V L+G GS QE + +L S
Sbjct: 481 NQRVWLHGIITGSVRDIKAHKLQKTQADQIQLAMVCVREVVSDEEQEIGSISALTSDVVE 540
Query: 497 EVDSIPVVKEFTDVFPGDVPGLPPVRDI-EFAIDIVPGTGPISIAPYRMAPAELVELKSQ 555
E +V+EF DVF + LPP R+ + I ++ G P++ PYR + E+
Sbjct: 541 ESVVQNIVEEFPDVF-AEPTDLPPFREKHDHKIKLLEGANPVNQRPYRYVVHQKDEIDKI 599
Query: 556 LEDLLSKGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQ 615
++D++ G I+ S SP+ +PV+LVKKKDG RLCVDY +LN +TVK+R+ +P I+DLMD+
Sbjct: 600 VQDMIKSGTIQVSSSPFASPVVLVKKKDGTWRLCVDYTELNGMTVKDRFLIPLIEDLMDE 659
Query: 616 LRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNR 675
L G+ +FSKIDL++GYHQ+R+ +DIQKTAF+T GH+EYLVM FG+TNAPA F MN
Sbjct: 660 LGGSVVFSKIDLRAGYHQVRMDPDDIQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNS 719
Query: 676 TFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFL 735
F FL +FV+VF DDILIYS ++ EH+ HLR V +V+R +L+A SK + L
Sbjct: 720 VFRDFLRKFVLVFFDDILIYSSSIEEHKEHLRLVFEVMRLHKLFAKGSK--------EHL 771
Query: 736 GHVISKEGIAVDPAKVETVLSWERPKTV 763
GH IS I DPAK++ V W P TV
Sbjct: 772 GHFISAREIETDPAKIQAVKEWPTPTTV 799
>At4g04230 putative transposon protein
Length = 315
Score = 196 bits (497), Expect = 1e-49
Identities = 105/244 (43%), Positives = 150/244 (61%), Gaps = 42/244 (17%)
Query: 494 GNPEVDS-IPVVKE-FTDVFPGDVP-GLPPVRDIEFAIDIVPGTGPISIAPYRMAPAELV 550
G+PE+ + + VV + D+FP ++P GLPP+
Sbjct: 95 GDPELSAEVQVVLHWYKDLFPEEIPPGLPPIH---------------------------- 126
Query: 551 ELKSQLEDLLSKGFIRPSVSPWGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRID 610
KG+IR S+SP PVLLV KKDG R+C+D R +N +T+K R+P+PR+
Sbjct: 127 -----------KGYIRESLSPCAVPVLLVPKKDGTWRMCLDCRAINNITIKYRHPIPRLY 175
Query: 611 DLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFM 670
D++D+L GA IFSK+DL+SGYHQ+R++ D KTAF+T+ G YE LVMPFG+TNAP+ FM
Sbjct: 176 DMLDELSGAIIFSKVDLRSGYHQVRMREGDEWKTAFKTKQGLYECLVMPFGLTNAPSTFM 235
Query: 671 DYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLE 730
MN+ +F+ +FVVV+ DDILIY+K+ +H L +L+ LR++ LYAN KC F +
Sbjct: 236 RLMNQVLRSFIGKFVVVYFDDILIYNKSYSDHIQPLELILKTLRKEGLYANLKKCTFCSD 295
Query: 731 EVKF 734
+ F
Sbjct: 296 KFFF 299
>At4g08100 putative polyprotein
Length = 1054
Score = 190 bits (483), Expect = 5e-48
Identities = 93/184 (50%), Positives = 124/184 (66%), Gaps = 2/184 (1%)
Query: 591 DYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRY 650
D R +N +TVK R+P+PR+DD +D+L G++IFSKIDLKSGYHQ R+K D KTA +T+
Sbjct: 527 DCRAINNITVKYRHPIPRLDDTLDKLHGSSIFSKIDLKSGYHQTRMKEGDEWKTAIKTKQ 586
Query: 651 GHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVL 710
YE+LVMPFG+TNAP FM MN + FV+V+ DDIL+YSKN+ H HL+ VL
Sbjct: 587 RLYEWLVMPFGLTNAPNTFMRLMNHVLRKHIGVFVIVYFDDILVYSKNLEYHVMHLKLVL 646
Query: 711 QVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERPKTV--LGVSL 768
+LR+++LYAN KC F + + FLG V+S +GI VD KV+ + W PK V +
Sbjct: 647 DLLRKEKLYANLKKCTFCTDNLVFLGFVVSADGIKVDEEKVKAIREWPNPKNVSEKDIGF 706
Query: 769 VWRD 772
W D
Sbjct: 707 KWED 710
Score = 67.8 bits (164), Expect = 4e-11
Identities = 67/301 (22%), Positives = 123/301 (40%), Gaps = 25/301 (8%)
Query: 13 QANDSAMRRAAEEAREQHQRQREVTLDQNKGLNDFRRHDPPKFSGGTDPDKADLWIQEIE 72
+A + ++ ++ +H+R+R D GL + P F G +PD W Q+IE
Sbjct: 61 EARNYYSHASSHSSQRRHRRERPPPRDPLGGL----KLKIPAFHGTDNPDTYLEWEQKIE 116
Query: 73 KIFGVLQTAEGAKLGMATYLLLGDAEYWWKG--TRGIMEDNHEEIN*NSFRMAFLEKYFP 130
+F + + K+ +A A WW T ++ N + +++ P
Sbjct: 117 LVFLCQECLQSNKVKIAATKFYNYALSWWDQLVTSRRRTRDYPIKTWNQLKFVMRKRFVP 176
Query: 131 TSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNERYMCKRFVNGLRPDIE 190
+ E + L QG +V E+ ++E+L D E M RF+ GL +I+
Sbjct: 177 SYYHRELHQRLRNLVQGSKTVEEYFLEMETLMLRADLQED--GEAVM-SRFMGGLNREIQ 233
Query: 191 DSVRPLGIMRFQSLVEKATEVE-LMKNRRLNRAGTGGPMRSGSQNFQNRGKF---QNRRP 246
D + + + ++ KA E +K + + T SG ++Q KF + +P
Sbjct: 234 DRLETQHYVELEEMLHKAVMFEQQIKRKNARSSHTKTNYSSGKPSYQKEEKFGYQNDSKP 293
Query: 247 YQRPAGRGFASGSYRPMVGAAGGSGDQ--TQNRELTCFKCGKPGHFARACPDTRPQCYNC 304
+ +P +P+ G G + T+ R++ CFK H+A C + R +
Sbjct: 294 FVKP----------KPVDLDPKGKGKEVITRARDIRCFKSQGLRHYASKCSNKRIMVHRD 343
Query: 305 N 305
N
Sbjct: 344 N 344
>At4g03650 putative reverse transcriptase
Length = 839
Score = 171 bits (434), Expect = 2e-42
Identities = 78/141 (55%), Positives = 106/141 (74%), Gaps = 6/141 (4%)
Query: 640 DIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNV 699
D++KTAFRTRYGH+E++VMPFG+TNAPA FM MN F FLD FV++FIDDIL+YSK+
Sbjct: 520 DVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSKSP 579
Query: 700 VEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWER 759
EHE HLR+V++ LRE++L+A SKC FW E+ FLGH++S EG++VDP K+E + W R
Sbjct: 580 EEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPR 639
Query: 760 P------KTVLGVSLVWRDII 774
P ++ LG++ +R I
Sbjct: 640 PTNATEIRSFLGLAGYYRRFI 660
Score = 92.8 bits (229), Expect = 1e-18
Identities = 60/143 (41%), Positives = 79/143 (54%), Gaps = 6/143 (4%)
Query: 424 ANLVCLTLKNLDVILGMDWLSHYHCLLDCN*KRVVFPDSVLSEYLFANHISVSLNGGSQE 483
A+L+ ++ DVILGMDWL HY LDC+ RV F S + + G
Sbjct: 386 ADLIISHVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGSAGRENDREGLRGLPGDDI 445
Query: 484 YSVLLSLESKGNPEVDSIPVVKEFTDVFPGDVPGLPPVRDIEFAIDIVPGTGPISIAPYR 543
Y+ + G V I VV+EF VF + GLPP R F I++ P T P+S APYR
Sbjct: 446 YAGVC-----GQVAVSDIRVVQEFQYVFQS-LQGLPPSRSDPFTIELEPKTAPLSKAPYR 499
Query: 544 MAPAELVELKSQLEDLLSKGFIR 566
MAPA++ ELK QLEDLL++ +R
Sbjct: 500 MAPAKMAELKKQLEDLLAEADVR 522
Score = 84.3 bits (207), Expect = 5e-16
Identities = 51/158 (32%), Positives = 77/158 (48%), Gaps = 4/158 (2%)
Query: 55 FSGGTDPDKADLWIQEIEKIFGVLQTAEGAKLGMATYLLLGDAEYWWKGTRGIMEDNHEE 114
FSGGT P++AD W ++E+ FG + ++ + + L GDA WW+ +
Sbjct: 156 FSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLEGDAHLWWRSVTA--RRRQAD 213
Query: 115 IN*NSFRMAFLEKYFPTSARDERESQFLTLRQGGMSVPEFASKLESLAKHFQFFHDHVNE 174
++ F F KYFP A D E++FL L QG SV E+ K L + ++
Sbjct: 214 MSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRLLVYAG--RGMEDD 271
Query: 175 RYMCKRFVNGLRPDIEDSVRPLGIMRFQSLVEKATEVE 212
+ +RF+ GLRP++ R +LVE A EVE
Sbjct: 272 QAQMRRFLRGLRPNLRVRCRVSQYATKAALVETAAEVE 309
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 162 bits (411), Expect = 1e-39
Identities = 97/275 (35%), Positives = 149/275 (53%), Gaps = 5/275 (1%)
Query: 514 DVPGLPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKGFIRPSVSP-W 572
D+PG+ +++ P P+ ++ P ++ +++ LL G I P W
Sbjct: 761 DMPGIDSAVTCH-ELNVDPTYKPLKQKRRKLGPDRTKDVNEEVKKLLDAGSIVEVRYPDW 819
Query: 573 GAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYH 632
++VKKK+GK R+C+D+ LNK K+ +PLP ID L++ G + S +D SGY+
Sbjct: 820 LRNPVVVKKKNGKWRVCIDFTDLNKACPKDSFPLPHIDRLVEATAGNELLSFMDAFSGYN 879
Query: 633 QIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDI 692
QI + D +KT F T G Y Y VMPFG+ NA A + +N+ F LD + V+IDD+
Sbjct: 880 QILMHQNDREKTVFITDQGTYCYKVMPFGLKNAGATYPRLVNQMFTDQLDHSMEVYIDDM 939
Query: 693 LIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVE 752
L+ S EH HLRQ QVL + NPSKC F + +FLG+++++ GI +P ++
Sbjct: 940 LVKSLRAEEHITHLRQCFQVLNRYNMKLNPSKCTFGVTSGEFLGYLVTRRGIEANPKQIS 999
Query: 753 TVLSWERPKTVLGVS-LVWRDIIVALSRISPRSLD 786
++ P+ V L+ R I AL+R RS D
Sbjct: 1000 AIIDLPSPRNTREVQRLIGR--IAALNRFISRSTD 1032
>At1g20390 hypothetical protein
Length = 1791
Score = 159 bits (403), Expect = 9e-39
Identities = 95/274 (34%), Positives = 145/274 (52%), Gaps = 3/274 (1%)
Query: 514 DVPGLPPVRDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQLEDLLSKG-FIRPSVSPW 572
D+ G+ P +++ P P+ ++ P + ++E LL G I W
Sbjct: 756 DMKGIDPAITAH-ELNVDPTFKPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEW 814
Query: 573 GAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYH 632
A ++VKKK+GK R+CVDY LNK K+ YPLP ID L++ G + S +D SGY+
Sbjct: 815 LANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYN 874
Query: 633 QIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDI 692
QI + +D +KT+F T G Y Y VM FG+ NA A + ++N+ + R V V+IDD+
Sbjct: 875 QILMHKDDQEKTSFVTDRGTYCYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDM 934
Query: 693 LIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVE 752
L+ S +H HL + VL + NP+KC F + +FLG+V++K GI +P ++
Sbjct: 935 LVKSLKPEDHVEHLSKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIR 994
Query: 753 TVLSWERPKTVLGVSLVWRDIIVALSRISPRSLD 786
+L P+ V + I AL+R RS D
Sbjct: 995 AILELPSPRNAREVQRL-TGRIAALNRFISRSTD 1027
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 159 bits (401), Expect = 1e-38
Identities = 88/236 (37%), Positives = 135/236 (56%), Gaps = 2/236 (0%)
Query: 552 LKSQLEDLLSKGFIRPSVSP-WGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRID 610
+ +++ LL G IR P W A ++VKKK+GK R+C+D+ LNK K+ +PLP ID
Sbjct: 480 VNDEVDKLLKIGSIREVQYPDWLANTVVVKKKNGKDRVCIDFTDLNKACPKDSFPLPHID 539
Query: 611 DLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFM 670
L++ G + S +D SGY+QI + ED +KT F T G Y Y VMPFG+ NA A +
Sbjct: 540 RLVESTAGNELLSFMDAFSGYNQIMMNPEDQEKTLFITDRGIYCYKVMPFGLRNAGATYP 599
Query: 671 DYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLE 730
+N+ F + + + V+IDD+LI S +H HL + +L + ++ NP+KC F +
Sbjct: 600 RLVNKMFSEHVGKTMEVYIDDMLIKSLKKEDHVKHLEECFAILNQYQMKLNPAKCTFGVP 659
Query: 731 EVKFLGHVISKEGIAVDPAKVETVLSWERPKTVLGVSLVWRDIIVALSRISPRSLD 786
+FLG++++K GI +P ++ L+ PK V + I AL+R RS D
Sbjct: 660 SGEFLGYIVTKRGIEANPNQINAFLNMPSPKNFKEVQRL-TGRIAALNRFISRSTD 714
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 145 bits (366), Expect = 2e-34
Identities = 81/245 (33%), Positives = 134/245 (54%), Gaps = 2/245 (0%)
Query: 543 RMAPAELVELKSQLEDLLSKGFIRPSVSP-WGAPVLLVKKKDGKSRLCVDYRQLNKVTVK 601
++ P + +++ LL G I P W A ++VKKK+ K R+C+D+ LNK K
Sbjct: 504 KLGPERSKAVNDEVDKLLDAGSIVEVKYPEWLANPVVVKKKNDKWRVCIDFTDLNKACPK 563
Query: 602 NRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFG 661
+ +PLP ID +++ G + S +D SGY+QI + +D +KT+F G Y Y VMPFG
Sbjct: 564 DSFPLPHIDRMVEATTGNELLSFMDAFSGYNQIPMHKDDQEKTSFIIDRGTYCYKVMPFG 623
Query: 662 VTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYAN 721
+ N A + +N+ F L + + V+IDD+L+ S +H HL+ + L + + N
Sbjct: 624 LKNVGARYQRLVNQMFAPQLGKTMEVYIDDMLVKSTRSADHIDHLKACFETLNKYNMKLN 683
Query: 722 PSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSWERPKTVLGVSLVWRDIIVALSRIS 781
P+KC F + +FLG++++K GI +P ++ +L + P+ V + I L+R
Sbjct: 684 PAKCLFGVTSGEFLGYIVTKRGIEANPKQIRAILDLQSPRNKKEVQRL-TGRIAGLNRFI 742
Query: 782 PRSLD 786
RS D
Sbjct: 743 ARSTD 747
>At1g37200 hypothetical protein
Length = 1564
Score = 144 bits (363), Expect = 4e-34
Identities = 86/266 (32%), Positives = 141/266 (52%), Gaps = 7/266 (2%)
Query: 499 DSIPVVKEFTDVFPGDVPGLPPV--RDIEFAIDIVPGTGPISIAPYRMAPAELVELKSQL 556
D I +KE D F L + +++ P PI ++ P ++ ++
Sbjct: 646 DFITFLKENKDSFAWSSANLQGISLEVTSHELNVDPTYRPIKQKRRKLGPERAKAVQDEV 705
Query: 557 EDLLSKGFIRPSVSP-WGAPVLLVKKKDGKSRLCVDYRQLNKVTVKNRYPLPRIDDLMDQ 615
+ LL G IR P W A ++VK K+GK R+C+D+ LNK K+ +PLP ID L+
Sbjct: 706 DRLLKIGSIREVKYPDWLANPVVVKNKNGKWRVCIDFMDLNKACPKDSFPLPHIDRLVKA 765
Query: 616 LRGAAIFSKIDLKSGYHQIRVKTEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNR 675
G + S +D SGY+QI ++ +D +KTAF T Y VMPFG+ N A + +NR
Sbjct: 766 TAGHELLSFMDAFSGYNQILMRPDDQEKTAFITDC----YKVMPFGLKNTGATYQRLVNR 821
Query: 676 TFHTFLDRFVVVFIDDILIYSKNVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFL 735
F L + + ++IDD+L+ S + +H LR+ ++L + E+ NP KC F + +FL
Sbjct: 822 MFADQLGKTMELYIDDMLVKSAHEKDHLPQLRECFKILNKFEMKLNPEKCSFGVPSGEFL 881
Query: 736 GHVISKEGIAVDPAKVETVLSWERPK 761
G+++++ GI +P ++ + PK
Sbjct: 882 GYLVTQRGIEANPKQIAAFIDMPSPK 907
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 139 bits (349), Expect = 2e-32
Identities = 88/246 (35%), Positives = 124/246 (49%), Gaps = 20/246 (8%)
Query: 538 SIAPYRMAPAELVEL-KSQLEDLLSKGFIRP-SVSPWGAPVLLVKKKDGKS--------- 586
SI P R L E+ K ++ LL G I P S S W +PV V KK G +
Sbjct: 900 SIEPQRRLNPNLKEVVKKEILKLLDAGVIYPISDSTWVSPVHCVPKKGGMTVVKNSKDEL 959
Query: 587 ---------RLCVDYRQLNKVTVKNRYPLPRIDDLMDQLRGAAIFSKIDLKSGYHQIRVK 637
R+C++YR+LN + K +PLP ID ++++L + +D SG+ QI +
Sbjct: 960 IPTRTITGHRMCIEYRKLNVASRKEHFPLPFIDHMLERLANHPYYCFLDSYSGFFQIPIH 1019
Query: 638 TEDIQKTAFRTRYGHYEYLVMPFGVTNAPAIFMDYMNRTFHTFLDRFVVVFIDDILIYSK 697
D KT F YG + Y MPFG+ NAPA F M F ++ V VF+DD +Y
Sbjct: 1020 PNDQGKTTFTCPYGTFAYKRMPFGLCNAPATFQRCMTSIFSDLIEEMVEVFMDDFSVYGS 1079
Query: 698 NVVEHEGHLRQVLQVLREKELYANPSKCEFWLEEVKFLGHVISKEGIAVDPAKVETVLSW 757
+ +L +VL+ E L N KC F + E LG IS+EGI VD AK++ ++
Sbjct: 1080 SFSSCLLNLCRVLKRCEETNLVLNWEKCHFMVREGIVLGRKISEEGIEVDKAKIDVMMQL 1139
Query: 758 ERPKTV 763
+ PKTV
Sbjct: 1140 QPPKTV 1145
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.345 0.152 0.510
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,749,899
Number of Sequences: 26719
Number of extensions: 1204751
Number of successful extensions: 5595
Number of sequences better than 10.0: 117
Number of HSP's better than 10.0 without gapping: 87
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 5200
Number of HSP's gapped (non-prelim): 289
length of query: 1469
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1357
effective length of database: 8,326,068
effective search space: 11298474276
effective search space used: 11298474276
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0016.15


