

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0014.18
(181 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
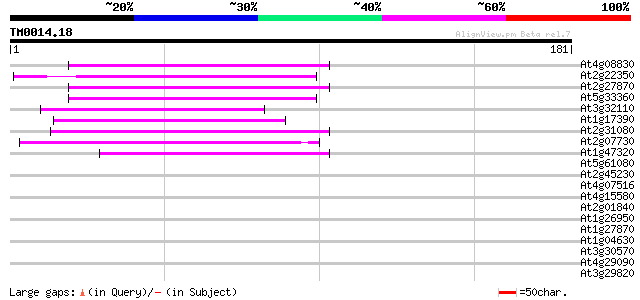
Score E
Sequences producing significant alignments: (bits) Value
At4g08830 putative protein 57 5e-09
At2g22350 putative non-LTR retroelement reverse transcriptase 52 1e-07
At2g27870 putative non-LTR retroelement reverse transcriptase 51 4e-07
At5g33360 putative protein 49 2e-06
At3g32110 non-LTR reverse transcriptase, putative 49 2e-06
At1g17390 hypothetical protein 48 3e-06
At2g31080 putative non-LTR retroelement reverse transcriptase 47 6e-06
At2g07730 putative non-LTR retroelement reverse transcriptase 44 6e-05
At1g47320 hypothetical protein 40 9e-04
At5g61080 putative protein 37 0.006
At2g45230 putative non-LTR retroelement reverse transcriptase 36 0.013
At4g07516 putative protein 34 0.049
At4g15580 splicing factor like protein 32 0.19
At2g01840 putative non-LTR retroelement reverse transcriptase 32 0.19
At1g26950 hypothetical protein 32 0.19
At1g27870 hypothetical protein 31 0.42
At1g04630 unknown protein 30 0.54
At3g30570 putative reverse transcriptase 30 0.71
At4g29090 putative protein 29 1.2
At3g29820 hypothetical protein 29 1.2
>At4g08830 putative protein
Length = 947
Score = 57.0 bits (136), Expect = 5e-09
Identities = 29/84 (34%), Positives = 45/84 (53%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
+ W P D WVKLN+DGA + +A G++RD + +A +G S AELWG+
Sbjct: 839 VAWKPPDGEWVKLNTDGASRGNLGLATTGGVLRDGIGHWCGGFALDIGVCSAPLAELWGV 898
Query: 80 LCGARLLQERDY*RVLIETNLSIL 103
G + ER + RV +E + ++
Sbjct: 899 YYGLYMAWERRFTRVELEVDSELV 922
>At2g22350 putative non-LTR retroelement reverse transcriptase
Length = 321
Score = 52.4 bits (124), Expect = 1e-07
Identities = 33/98 (33%), Positives = 50/98 (50%), Gaps = 9/98 (9%)
Query: 2 LRVAQVTRVVSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIA 61
+RV++V R+VS W + WVKLN+DGA + A G++RD A++
Sbjct: 142 VRVSRVERLVS---------WVSPEDGWVKLNTDGASRGNPGFATAGGVLRDHNGAWIGG 192
Query: 62 YARRLGSYSTLQAELWGILCGARLLQERDY*RVLIETN 99
+A +G S AELWG+ G + R RV +E +
Sbjct: 193 FAVNIGVCSAPLAELWGVYYGLFIAWGRGARRVELEVD 230
>At2g27870 putative non-LTR retroelement reverse transcriptase
Length = 314
Score = 50.8 bits (120), Expect = 4e-07
Identities = 29/84 (34%), Positives = 44/84 (51%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
I W + W KLN+DGA + +A+ G++RD A+ +A +G S AELWG+
Sbjct: 144 IAWSKPEEGWWKLNTDGASRGNPGLASAGGVLRDEEGAWRGGFALNIGVCSAPLAELWGV 203
Query: 80 LCGARLLQERDY*RVLIETNLSIL 103
G + ER R+ IE + I+
Sbjct: 204 YYGLYIAWERRVTRLEIEVDSEIV 227
>At5g33360 putative protein
Length = 306
Score = 48.5 bits (114), Expect = 2e-06
Identities = 24/80 (30%), Positives = 40/80 (50%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGI 79
+RW W KLN+DGA + +A G +R+ + + +A +G S AELWG+
Sbjct: 136 VRWSKPSLGWCKLNTDGASHGNPGLAIAGGALRNEYGEWCFGFALNIGRCSAPLAELWGV 195
Query: 80 LCGARLLQERDY*RVLIETN 99
G + +R R+ +E +
Sbjct: 196 YYGLFMAWDRGITRLELEVD 215
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 48.5 bits (114), Expect = 2e-06
Identities = 25/72 (34%), Positives = 39/72 (53%)
Query: 11 VSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYS 70
+S +R IRW P W K+N+DGA + +A+ G++R+ A+ +A +G S
Sbjct: 1732 LSGLRVNKPIRWTPPMEGWYKINTDGASRGNPGLASAGGVLRNSAGAWCGGFAVNIGRCS 1791
Query: 71 TLQAELWGILCG 82
AELWG+ G
Sbjct: 1792 APLAELWGVYYG 1803
>At1g17390 hypothetical protein
Length = 322
Score = 47.8 bits (112), Expect = 3e-06
Identities = 25/75 (33%), Positives = 36/75 (47%)
Query: 15 RDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQA 74
R+ I W P W KLN+DGA + +A G++RD + ++ +G S A
Sbjct: 97 REERLIAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICSAPLA 156
Query: 75 ELWGILCGARLLQER 89
ELWG G + ER
Sbjct: 157 ELWGAYYGLNIAWER 171
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 47.0 bits (110), Expect = 6e-06
Identities = 27/90 (30%), Positives = 46/90 (51%)
Query: 14 VRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQ 73
VR IRW WVK+ +DGA + +AA G IR+ ++ +A +GS +
Sbjct: 1055 VRVERMIRWQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALNIGSCAAPL 1114
Query: 74 AELWGILCGARLLQERDY*RVLIETNLSIL 103
AELWG G + ++ + RV ++ + ++
Sbjct: 1115 AELWGAYYGLLIAWDKGFRRVELDLDCKLV 1144
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 43.5 bits (101), Expect = 6e-05
Identities = 31/97 (31%), Positives = 43/97 (43%), Gaps = 2/97 (2%)
Query: 4 VAQVTRVVSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYA 63
V + V R IRW WVKL +DGA +AA G I + ++ +A
Sbjct: 784 VGTLNNHVKRARVERMIRWKAPSDRWVKLTTDGASRGHQGLAAASGAILNLQGEWLGGFA 843
Query: 64 RRLGSYSTLQAELWGILCGARLLQERDY*RVLIETNL 100
+GS AELWG G + ++ + RV E NL
Sbjct: 844 LNIGSCDAPLAELWGAYYGLLIAWDKGFRRV--ELNL 878
>At1g47320 hypothetical protein
Length = 259
Score = 39.7 bits (91), Expect = 9e-04
Identities = 21/74 (28%), Positives = 37/74 (49%)
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQER 89
+K+N+DGA + +A G+++D + ++ +G S AELWG G L ER
Sbjct: 101 LKINTDGASRGNPGLATAGGVLQDNEGRWCGGFSLNIGRSSAPMAELWGAYYGLYLAWER 160
Query: 90 DY*RVLIETNLSIL 103
+ +E + I+
Sbjct: 161 KSSHIELEVDSEIV 174
>At5g61080 putative protein
Length = 348
Score = 37.0 bits (84), Expect = 0.006
Identities = 30/103 (29%), Positives = 44/103 (42%), Gaps = 9/103 (8%)
Query: 30 VKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQER 89
VKLN G +D+ GI+RD+ +V Y R S + A L I G + L +
Sbjct: 195 VKLNIQGTSNPLSDLTRSAGIVRDQSGKWVFGYIRCHKSIPEVVAGLLAIYQGLKYLWDS 254
Query: 90 DY*RVLIETNLSILGVVLVLIPVLLLLTK*SRLFSRVSPRLGA 132
+ R+ +ET ++ LT S LF + LGA
Sbjct: 255 GFRRIHLET---------TSFEIINALTTKSSLFCKSKTLLGA 288
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 35.8 bits (81), Expect = 0.013
Identities = 26/96 (27%), Positives = 42/96 (43%), Gaps = 2/96 (2%)
Query: 9 RVVSPVRDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGS 68
+V S RD ++W P WVK N+DGA+ ++R+ + R L S
Sbjct: 1203 QVTSSTRDR-CVKWQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLLWLGLRALPS 1261
Query: 69 -YSTLQAELWGILCGARLLQERDY*RVLIETNLSIL 103
S L+ E+ + L +Y RV+ E++ L
Sbjct: 1262 QQSVLETEVEALRWAVLSLSRFNYRRVIFESDSQYL 1297
>At4g07516 putative protein
Length = 740
Score = 33.9 bits (76), Expect = 0.049
Identities = 22/83 (26%), Positives = 37/83 (44%)
Query: 35 DGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQERDY*RV 94
DGA + +A G++RD + +A +G S AELWG+ + E R+
Sbjct: 606 DGASPGNPGLATASGVLRDEHGNWRGDFALNIGICSAPLAELWGVYYKLYIAWEMRITRL 665
Query: 95 LIETNLSILGVVLVLIPVLLLLT 117
+E + I+ + I L + T
Sbjct: 666 ELEVDSEIVSAFFMCIERLTVTT 688
>At4g15580 splicing factor like protein
Length = 559
Score = 32.0 bits (71), Expect = 0.19
Identities = 17/47 (36%), Positives = 25/47 (53%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRL 66
I W WVK+++DGA + AA G+IRD +V +A +L
Sbjct: 26 IAWTKPPEGWVKVSTDGASRGNPGPAAAGGVIRDEDGLWVGGFALQL 72
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 32.0 bits (71), Expect = 0.19
Identities = 24/80 (30%), Positives = 38/80 (47%), Gaps = 2/80 (2%)
Query: 29 WVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLG-SYSTLQAELWGILCGARLLQ 87
++K N D Y+ D + I+RD + + +L SYS LQAE G L +++
Sbjct: 1563 FLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSYSALQAEALGFLHALQMVW 1622
Query: 88 ERDY*RVLIE-TNLSILGVV 106
R Y V E NL + ++
Sbjct: 1623 IRGYCYVWFEGDNLELTNLI 1642
>At1g26950 hypothetical protein
Length = 158
Score = 32.0 bits (71), Expect = 0.19
Identities = 16/56 (28%), Positives = 30/56 (53%)
Query: 48 RGIIRDRFSAFVIAYARRLGSYSTLQAELWGILCGARLLQERDY*RVLIETNLSIL 103
RG +RD + + ++A LG + AELWG+ G + E+ R+ +E + ++
Sbjct: 16 RGALRDEYGDWRGSFALNLGRCTAPLAELWGVYYGLVIAWEKGITRLELEVDSKLV 71
>At1g27870 hypothetical protein
Length = 213
Score = 30.8 bits (68), Expect = 0.42
Identities = 18/85 (21%), Positives = 42/85 (49%), Gaps = 3/85 (3%)
Query: 21 RWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSY--STLQAELWG 78
RW + W+K N DG++++ + ++RD ++++A + +G + L++E+
Sbjct: 51 RWRRPERGWIKCNFDGSFVNGDVKSKAGWVVRDSNGSYLLA-GQAIGRKVDNALESEIQA 109
Query: 79 ILCGARLLQERDY*RVLIETNLSIL 103
++ + Y RV E + +L
Sbjct: 110 LIISMQHCWSHGYKRVCFEGDNKML 134
>At1g04630 unknown protein
Length = 356
Score = 30.4 bits (67), Expect = 0.54
Identities = 24/83 (28%), Positives = 39/83 (46%), Gaps = 5/83 (6%)
Query: 18 PAIRWWPLDAV-WVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYST--LQA 74
P W + W K N DG Y S+A A ++RD F+ A A +GS +T +++
Sbjct: 190 PTANLWTTPPIGWTKCNYDGTYHSNAPSKA-GWLLRDDRGTFLGA-AHAIGSITTNPMES 247
Query: 75 ELWGILCGARLLQERDY*RVLIE 97
EL ++ + R Y ++ E
Sbjct: 248 ELQALVMAMQHCWSRGYRKIYFE 270
>At3g30570 putative reverse transcriptase
Length = 1099
Score = 30.0 bits (66), Expect = 0.71
Identities = 15/34 (44%), Positives = 18/34 (52%)
Query: 20 IRWWPLDAVWVKLNSDGAYISSADMAACRGIIRD 53
I W A W KLN DGA + +AA G +RD
Sbjct: 1019 IGWRVPSAGWYKLNMDGASRGNPGLAAAGGALRD 1052
>At4g29090 putative protein
Length = 575
Score = 29.3 bits (64), Expect = 1.2
Identities = 23/88 (26%), Positives = 36/88 (40%), Gaps = 1/88 (1%)
Query: 21 RWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSY-STLQAELWGI 79
RW P WVK N+D + + ++R+ AR L S L+AEL +
Sbjct: 419 RWRPPPHQWVKCNTDATWNRDNERCGIGWVLRNEKGEVKWMGARALPKLKSVLEAELEAM 478
Query: 80 LCGARLLQERDY*RVLIETNLSILGVVL 107
L Y V+ E++ +L +L
Sbjct: 479 RWAVLSLSRFQYNYVIFESDSQVLIEIL 506
>At3g29820 hypothetical protein
Length = 332
Score = 29.3 bits (64), Expect = 1.2
Identities = 20/89 (22%), Positives = 37/89 (41%), Gaps = 17/89 (19%)
Query: 15 RDAPAIRWWPLDAVWVKLNSDGAYISSADMAACRGIIRDRFSAFVIAYARRLGSYSTLQA 74
R+ I W W+K+N++GA + +A G+ ++
Sbjct: 200 REERMIGWSAPQVGWIKVNTNGASRGNLGLATSAGVCE-----------------IVMEL 242
Query: 75 ELWGILCGARLLQERDY*RVLIETNLSIL 103
ELWG+ G L ER +V +E + +++
Sbjct: 243 ELWGVYYGLYLAWERMATQVELEIDSNMV 271
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.336 0.145 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,361,096
Number of Sequences: 26719
Number of extensions: 116472
Number of successful extensions: 359
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 340
Number of HSP's gapped (non-prelim): 25
length of query: 181
length of database: 11,318,596
effective HSP length: 93
effective length of query: 88
effective length of database: 8,833,729
effective search space: 777368152
effective search space used: 777368152
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0014.18


