

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0011a.2
(391 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
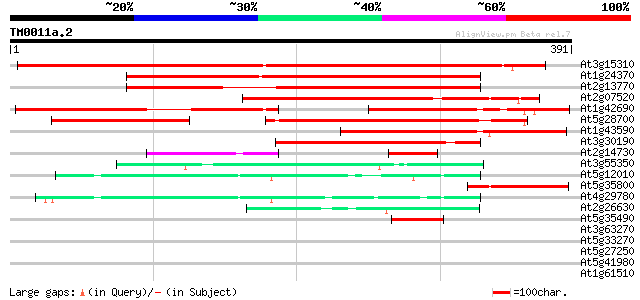
Score E
Sequences producing significant alignments: (bits) Value
At3g15310 unknown protein 419 e-117
At1g24370 hypothetical protein 312 2e-85
At2g13770 hypothetical protein 244 5e-65
At2g07520 pseudogene 197 6e-51
At1g42690 unknown protein 189 2e-48
At5g28700 putative protein 178 4e-45
At1g43590 hypothetical protein 159 2e-39
At3g30190 hypothetical protein 159 2e-39
At2g14730 hypothetical protein 69 6e-12
At3g55350 unknown protein 61 1e-09
At5g12010 putative protein 56 3e-08
At5g35800 putative protein 49 4e-06
At4g29780 unknown protein 48 8e-06
At2g26630 En/Spm-like transposon protein 47 2e-05
At5g35490 putative protein 42 8e-04
At3g63270 unknown protein 41 0.001
At5g33270 putative protein 40 0.002
At5g27250 putative protein 40 0.002
At5g41980 unknown protein 39 0.004
At1g61510 hypothetical protein 38 0.008
>At3g15310 unknown protein
Length = 415
Score = 419 bits (1077), Expect = e-117
Identities = 204/373 (54%), Positives = 273/373 (72%), Gaps = 7/373 (1%)
Query: 6 RQRSHIERNREEGHVRLWNDYFSTNPVYTEEQFQRRYRMRKHMFLRIVEAISNTDDYFQM 65
R+R++ +R REEGHV LWNDYFS N ++ + F+RR+RM+K +FLRIV+ +S+ +FQ
Sbjct: 43 RKRAYFDRKREEGHVLLWNDYFSDNAIFPLQTFRRRFRMKKPLFLRIVDRLSSELMFFQH 102
Query: 66 RNDATGRMGLSPLQKQTAAIRMLAYGSSTDSVDEYVCIGESTARKCMARFVKGINEVFGP 125
R DATGR G SP+QK TAAIR+LAYG ++D+VDEY+ +GE+TA C+ F KGI FG
Sbjct: 103 RRDATGRFGHSPIQKCTAAIRLLAYGYASDAVDEYLRMGETTAMSCLENFTKGIISFFGD 162
Query: 126 EYLRKTNNKDIARLMELGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIIL 185
EYLR ++ RL+ +GK RGFPGM+GS+DCMHW W+N PTAWKGQY RG GKPTI+L
Sbjct: 163 EYLRAPTATNLRRLLNIGKIRGFPGMIGSLDCMHWEWKNCPTAWKGQYTRGS-GKPTIVL 221
Query: 186 EAIASQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLA 245
EAIASQDLWIWH FFG G+ NDIN+L++S +F+D+LQG+AP V++ +NG Y++ YYL
Sbjct: 222 EAIASQDLWIWHVFFGPPGTLNDINILDRSPIFDDILQGRAPNVKYKVNGREYHLAYYLT 281
Query: 246 DEIYPEWATFVKTISMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLH 305
D IYP+WATF+++I +PQ K LFA HQE RKDVERAFGVL +RF I++ PA W
Sbjct: 282 DGIYPKWATFIQSIRLPQNRKATLFATHQEADRKDVERAFGVLQARFHIIKNPALVWDKE 341
Query: 306 ILKDIIYACIILHNMIVEDEREMCDGNIDFS-YDQLENDASTPEV----FSGPHPDFATY 360
+ +I+ ACIILHNMIVEDER+ + D S + Q+E + TP+V +G +
Sbjct: 342 KIGNIMKACIILHNMIVEDERDGYNIQFDVSEFLQVEGN-QTPQVDLSYSTGMPLNIENM 400
Query: 361 LQKRAQVREKTVH 373
+ R Q+R++ +H
Sbjct: 401 MGMRNQLRDQNMH 413
>At1g24370 hypothetical protein
Length = 413
Score = 312 bits (800), Expect = 2e-85
Identities = 149/247 (60%), Positives = 184/247 (74%), Gaps = 1/247 (0%)
Query: 82 TAAIRMLAYGSSTDSVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKTNNKDIARLME 141
TAA RMLAYG DS DEY+ IGESTA + + RF + I EVF YLR + D+ARL+
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACRYLRSPDANDVARLLH 61
Query: 142 LGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIILEAIASQDLWIWHAFFG 201
+G++RGFP MLGS+DCMHW W+N PTAW GQY G PTIILEA+A DLWIWHA+FG
Sbjct: 62 IGESRGFPRMLGSLDCMHWKWKNCPTAWGGQYA-GRSRSPTIILEAVADYDLWIWHAYFG 120
Query: 202 VAGSNNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISM 261
+ GSNNDINVL S +F ++ +G AP + +N YNM YYLAD IYP+W+T V+TI
Sbjct: 121 LPGSNNDINVLEASHLFANLAEGTAPPASYVINEKPYNMSYYLADGIYPKWSTLVQTIHD 180
Query: 262 PQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMI 321
P+G K+KLFA QE RKDVERAFGVL SRFAIV GP+R W+ +L DI+ +CII+HNMI
Sbjct: 181 PRGPKKKLFAMKQEACRKDVERAFGVLQSRFAIVAGPSRLWNKTVLHDIMTSCIIMHNMI 240
Query: 322 VEDEREM 328
+EDER++
Sbjct: 241 IEDERDI 247
>At2g13770 hypothetical protein
Length = 244
Score = 244 bits (624), Expect = 5e-65
Identities = 125/247 (50%), Positives = 158/247 (63%), Gaps = 36/247 (14%)
Query: 82 TAAIRMLAYGSSTDSVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKTNNKDIARLME 141
TAA RMLAYG DS DEY+ IGESTA + + RF + I EVF YLR + D+ARL+
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACCYLRSPDANDVARLLH 61
Query: 142 LGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIILEAIASQDLWIWHAFFG 201
+G++RGFP EA+A DLWIWHA+FG
Sbjct: 62 IGESRGFP------------------------------------EAVADYDLWIWHAYFG 85
Query: 202 VAGSNNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISM 261
+ GSNN INVL S +F ++ +G AP + +NG YNMGYYLAD IYP+W+T V+TI
Sbjct: 86 LPGSNNGINVLEASHLFANLAEGTAPPASYVINGKPYNMGYYLADGIYPKWSTLVQTIHD 145
Query: 262 PQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMI 321
P+G K+KLFA QE RKDVERAFGVL RFAIV GP+R W+ +L DI+ +CII+HNMI
Sbjct: 146 PRGPKKKLFAMKQEACRKDVERAFGVLQLRFAIVAGPSRLWNKTVLHDIMTSCIIMHNMI 205
Query: 322 VEDEREM 328
+EDER++
Sbjct: 206 IEDERDI 212
>At2g07520 pseudogene
Length = 222
Score = 197 bits (502), Expect = 6e-51
Identities = 108/210 (51%), Positives = 140/210 (66%), Gaps = 11/210 (5%)
Query: 163 ENGPTAWKGQYCRGDHGKPTIILEAIASQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVL 222
+N P AW+GQ+ RGD G TI EA+AS DLWIWH FFG A + NDIN L++S VF+D+
Sbjct: 16 KNCPIAWEGQFTRGDKGTSTISFEAVASHDLWIWHTFFGCASTLNDINFLDRSPVFDDIE 75
Query: 223 QGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGEKRKLFAQHQEGARKDVE 282
QG P V F + YNM YYLAD IYP + TFVK+I +PQ E KLF Q QEG RKD+E
Sbjct: 76 QGNTPRVNFFVYQRQYNMAYYLADGIYPSYPTFVKSIRLPQSEPDKLFVQLQEGCRKDIE 135
Query: 283 RAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDEREMCDGNIDFSYDQ-LE 341
RAFGVL +RF I+ W++ L I+ +CIILHNMIVE+ER+ + YDQ E
Sbjct: 136 RAFGVLHARFKII------WNIADLAIIMRSCIILHNMIVENERDTYAQHWT-DYDQSKE 188
Query: 342 NDASTPEVFSGP--HPDFATYLQKRAQVRE 369
+ +STP+ FS H FA ++ R+++R+
Sbjct: 189 SGSSTPQPFSTEVLHA-FANHVHARSKLRD 217
>At1g42690 unknown protein
Length = 333
Score = 189 bits (480), Expect = 2e-48
Identities = 95/183 (51%), Positives = 118/183 (63%), Gaps = 31/183 (16%)
Query: 5 RRQRSHIERNREEGHVRLWNDYFSTNPVYTEEQFQRRYRMRKHMFLRIVEAISNTDDYFQ 64
R+++ +IERNREEGH++L NDYF+ NP Y F+RR+RM K +F+RIVE SN YF+
Sbjct: 38 RKKKLYIERNREEGHIQLVNDYFTENPTYPPHIFRRRFRMNKSLFMRIVERFSNEVPYFK 97
Query: 65 MRNDATGRMGLSPLQKQTAAIRMLAYGSSTDSVDEYVCIGESTARKCMARFVKGINEVFG 124
R DAT R+G S LQK TAAIRMLAYG + D+
Sbjct: 98 QRRDATRRLGFSALQKSTAAIRMLAYGIAADA---------------------------- 129
Query: 125 PEYLRKTNNKDIARLMELGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTII 184
YLR+ D+ RL+ +G+ RGFPGM+GSIDCMHW W+N PTAWKGQY RG GKPTI+
Sbjct: 130 --YLRRPTRDDLIRLLHIGEQRGFPGMIGSIDCMHWEWKNCPTAWKGQYTRGS-GKPTIV 186
Query: 185 LEA 187
LEA
Sbjct: 187 LEA 189
Score = 110 bits (275), Expect = 1e-24
Identities = 69/150 (46%), Positives = 90/150 (60%), Gaps = 17/150 (11%)
Query: 251 EWATFVKTISMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDI 310
E ATF+++IS+ QG K LFA QE RKDVERAFGVL +RFAI++ PA + +I
Sbjct: 188 EAATFIQSISILQGNKASLFATTQEACRKDVERAFGVLQARFAIIKHPALFHDKVKIGNI 247
Query: 311 IYACIILHNMIVEDEREMCDGNIDFSYDQLENDASTPEVFSGPHPDF--ATYLQKR---- 364
+ ACIILHNMIVED+R DG F + + PE S DF AT +
Sbjct: 248 MRACIILHNMIVEDKR---DGYTQFDVSEFVH----PESASSSQVDFTYATVMPSNLGNM 300
Query: 365 ----AQVREKTVHLHLQADLVEHVWEHFRR 390
A+VR++ H L+ADLVEHVW+H+ +
Sbjct: 301 MATGARVRDRIKHEELKADLVEHVWQHYNQ 330
>At5g28700 putative protein
Length = 292
Score = 178 bits (452), Expect = 4e-45
Identities = 95/186 (51%), Positives = 128/186 (68%), Gaps = 13/186 (6%)
Query: 179 GKPTIILEAIASQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTY 238
G+ T+++ +ASQDLWIWHA FG G+ NDINVL++S VF+D+LQG+A +V++ +NG Y
Sbjct: 105 GESTLLV--VASQDLWIWHAIFGPPGTLNDINVLDRSPVFDDILQGRASKVKYVVNGKDY 162
Query: 239 NMGYYLADEIYPEWATFVKTISMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGP 298
N+ YYL D IYP+WATF+++IS PQG+K LF Q+ RKDVERAFGV +RF+IV+ P
Sbjct: 163 NLAYYLTDGIYPKWATFIQSISNPQGDKASLFVTTQKACRKDVERAFGVFQARFSIVKHP 222
Query: 299 ARAWHLHILKDIIYACIILHNMIVEDEREMCDGNIDFSYDQLE-NDASTPEVFSGPHPDF 357
A + +I+ ACIILHNMIVEDER+ Y Q + ++ PE S DF
Sbjct: 223 ALFHDKVKIGNIMRACIILHNMIVEDERD--------GYTQCDVSEFVHPEFASSSQVDF 274
Query: 358 --ATYL 361
ATY+
Sbjct: 275 TYATYM 280
Score = 94.4 bits (233), Expect = 1e-19
Identities = 49/96 (51%), Positives = 63/96 (65%)
Query: 30 NPVYTEEQFQRRYRMRKHMFLRIVEAISNTDDYFQMRNDATGRMGLSPLQKQTAAIRMLA 89
N Y F+ R+RM K +F+RIVE SN YF+ R DATGR+ S LQK TAAIRMLA
Sbjct: 31 NATYPPHIFRHRFRMNKPLFMRIVERFSNEVPYFKQRRDATGRLDFSALQKSTAAIRMLA 90
Query: 90 YGSSTDSVDEYVCIGESTARKCMARFVKGINEVFGP 125
YG + D+VDEY+ IGEST ++ + + +FGP
Sbjct: 91 YGIAADAVDEYLRIGESTLLVVASQDLWIWHAIFGP 126
>At1g43590 hypothetical protein
Length = 168
Score = 159 bits (403), Expect = 2e-39
Identities = 82/164 (50%), Positives = 110/164 (67%), Gaps = 9/164 (5%)
Query: 231 FTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGEKRKLFAQHQEGARKDVERAFGVL*S 290
F +NG YN+ YYL DEIYP+WATF+++IS+PQ EK LFA +QE RKDVERAFGVL +
Sbjct: 3 FVVNGNEYNLAYYLTDEIYPKWATFIQSISLPQDEKASLFATNQEACRKDVERAFGVLQA 62
Query: 291 RFAIVRGPARAWHLHILKDIIYACIILHNMIVEDEREMCDGNI-----DFSYDQLENDAS 345
RFAIV+ PA W + +I+ ACIILHNMIVEDER DG DF++ + + +
Sbjct: 63 RFAIVKHPALIWDKIKIGNIMRACIILHNMIVEDER---DGYTQYDVSDFAHPESASSSQ 119
Query: 346 TPEVFSGPHP-DFATYLQKRAQVREKTVHLHLQADLVEHVWEHF 388
+S P + + R ++R++T H HL+ADLVEH+W+ F
Sbjct: 120 VDFTYSTDMPSNLGNMMATRTRLRDRTKHEHLKADLVEHIWQKF 163
>At3g30190 hypothetical protein
Length = 263
Score = 159 bits (402), Expect = 2e-39
Identities = 77/143 (53%), Positives = 100/143 (69%), Gaps = 6/143 (4%)
Query: 186 EAIASQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLA 245
+A+A D+WIWHA+FG+ GSNNDINVL +F ++ + AP + NG YNMGYYLA
Sbjct: 104 QAVADYDMWIWHAYFGLPGSNNDINVLEAYHLFANLAEDTAPPASYVSNGKPYNMGYYLA 163
Query: 246 DEIYPEWATFVKTISMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLH 305
D IY +W+T V+TI P+G K+KLFA E RKDVE AF VL SRFAIV P+R W+
Sbjct: 164 DGIYSKWSTLVQTIHDPRGPKKKLFAMKLETCRKDVEPAFEVLHSRFAIVAEPSRLWNK- 222
Query: 306 ILKDIIYACIILHNMIVEDEREM 328
+ +CII+HNMI+EDER++
Sbjct: 223 -----MTSCIIMHNMIIEDERDI 240
>At2g14730 hypothetical protein
Length = 117
Score = 68.6 bits (166), Expect = 6e-12
Identities = 37/92 (40%), Positives = 55/92 (59%), Gaps = 4/92 (4%)
Query: 96 SVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKTNNKDIARLMELGKARGFPGMLGSI 155
+VD+Y+ IGE+T CM V+ I +FG EYLR+ +D+ RL+ +G+ RGF GM GSI
Sbjct: 19 AVDKYLRIGENTLMSCMIHSVEAIIYLFGKEYLRRPTRQDLKRLLRIGELRGFLGMTGSI 78
Query: 156 DCMHWIWENGPTAWKGQYCRGDHGKPTIILEA 187
DC ++ A + CR D + +L+A
Sbjct: 79 DCD----KDSLFATNQEVCRKDVERAFGVLQA 106
Score = 45.4 bits (106), Expect = 5e-05
Identities = 23/34 (67%), Positives = 26/34 (75%)
Query: 265 EKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGP 298
+K LFA +QE RKDVERAFGVL +RFAIV P
Sbjct: 81 DKDSLFATNQEVCRKDVERAFGVLQARFAIVTNP 114
>At3g55350 unknown protein
Length = 406
Score = 60.8 bits (146), Expect = 1e-09
Identities = 68/267 (25%), Positives = 106/267 (39%), Gaps = 22/267 (8%)
Query: 75 LSPLQKQTAAIRMLAYGSSTDSVDEYVCIGESTARKCMARFVKGINE-----VFGPEYLR 129
LS + A+R L G S + E + +ST + RFV+ + E + P L
Sbjct: 109 LSLNDRVAVALRRLGSGESLSVIGETFGMNQSTVSQITWRFVESMEERAIHHLSWPSKLD 168
Query: 130 KTNNKDIARLMELGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIILEAIA 189
+ +K K G P G+ID H + + ++ L+A+
Sbjct: 169 EIKSK-------FEKISGLPNCCGAIDITHIVMNLPAVEPSNKVWLDGEKNFSMTLQAVV 221
Query: 190 SQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVLQGKAPEVQ-FTLNGTTYNMGYYLADEI 248
D+ G GS ND VL S + V +GK + L+ T Y + D
Sbjct: 222 DPDMRFLDVIAGWPGSLNDDVVLKNSGFYKLVEKGKRLNGEKLPLSERTELREYIVGDSG 281
Query: 249 YPEWATFV-----KTISMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWH 303
+P + K S+PQ E K +H E A K + A L R+ I+ G
Sbjct: 282 FPLLPWLLTPYQGKPTSLPQTEFNK---RHSE-ATKAAQMALSKLKDRWRIINGVMWMPD 337
Query: 304 LHILKDIIYACIILHNMIVEDEREMCD 330
+ L II+ C +LHN+I++ E + D
Sbjct: 338 RNRLPRIIFVCCLLHNIIIDMEDQTLD 364
>At5g12010 putative protein
Length = 502
Score = 56.2 bits (134), Expect = 3e-08
Identities = 70/308 (22%), Positives = 121/308 (38%), Gaps = 33/308 (10%)
Query: 33 YTEEQFQRRYRMRKHMFLRIVEAISNTDDYFQMRNDATGRMGLSPLQKQTAAIRMLAYGS 92
Y EE F++ +RM K F I + +++ + D R + Q+ I LA G
Sbjct: 170 YPEEDFKKAFRMSKSTFELICDELNSA----VAKEDTALRNAIPVRQRVAVCIWRLATGE 225
Query: 93 STDSVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKTNNKDIARLME-LGKARGFPGM 151
V + +G ST K + K I +V P+YL+ +++ + + E G P +
Sbjct: 226 PLRLVSKKFGLGISTCHKLVLEVCKAIKDVLMPKYLQWPDDESLRNIRERFESVSGIPNV 285
Query: 152 LGSIDCMHWIWENGPTAWKGQYCRGDHGKP------TIILEAIASQDLWIWHAFFGVAGS 205
+GS+ H I P Y H + +I ++A+ + G GS
Sbjct: 286 VGSMYTTH-IPIIAPKISVASYFNKRHTERNQKTSYSITIQAVVNPKGVFTDLCIGWPGS 344
Query: 206 NNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGE 265
D VL +S ++ G + + G G+ L D W + +P +
Sbjct: 345 MPDDKVLEKSLLYQRANNGGLLKGMWVAGGP----GHPLLD-----W------VLVPYTQ 389
Query: 266 KRKLFAQHQEGARKD-----VERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNM 320
+ + QH + + AFG L R+A ++ L L ++ AC +LHN+
Sbjct: 390 QNLTWTQHAFNEKMSEVQGVAKEAFGRLKGRWACLQKRTEV-KLQDLPTVLGACCVLHNI 448
Query: 321 IVEDEREM 328
E +M
Sbjct: 449 CEMREEKM 456
>At5g35800 putative protein
Length = 73
Score = 49.3 bits (116), Expect = 4e-06
Identities = 29/72 (40%), Positives = 47/72 (65%), Gaps = 3/72 (4%)
Query: 320 MIVEDEREMCDGNIDFSYDQLE-NDASTPEVFS-GPHPDFATYLQKRAQVREKTVHLHLQ 377
MIV DERE + Y+Q E + +STP+ FS P+FA++++ R+++R+ +H LQ
Sbjct: 1 MIVNDERETYAQHWS-DYNQSEASGSSTPQPFSIEVLPEFASHVRARSELRDSNLHHELQ 59
Query: 378 ADLVEHVWEHFR 389
A LV+++W FR
Sbjct: 60 AGLVKYIWTKFR 71
>At4g29780 unknown protein
Length = 540
Score = 48.1 bits (113), Expect = 8e-06
Identities = 71/324 (21%), Positives = 126/324 (37%), Gaps = 30/324 (9%)
Query: 19 HVRLW-----NDYFS--TNPVYTEEQFQRRYRMRKHMFLRIVEAISNTDDYFQMRNDATG 71
H RLW D++ + P + E++F+R +RM K F I E + T + +
Sbjct: 187 HRRLWVKERTTDWWDRVSRPDFPEDEFRREFRMSKSTFNLICEELDTT----VTKKNTML 242
Query: 72 RMGLSPLQKQTAAIRMLAYGSSTDSVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKT 131
R + ++ + LA G+ V E +G ST K + + I +V P+YL
Sbjct: 243 RDAIPAPKRVGVCVWRLATGAPLRHVSERFGLGISTCHKLVIEVCRAIYDVLMPKYLLWP 302
Query: 132 NNKDI-ARLMELGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKP------TII 184
++ +I + + P ++GSI H I P Y H + +I
Sbjct: 303 SDSEINSTKAKFESVHKIPNVVGSIYTTH-IPIIAPKVHVAAYFNKRHTERNQKTSYSIT 361
Query: 185 LEAIASQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYL 244
++ + + D G GS D +L +S + +A + N G+ L
Sbjct: 362 VQGVVNADGIFTDVCIGNPGSLTDDQILEKSSLSRQ----RAARGMLRDSWIVGNSGFPL 417
Query: 245 ADEIYPEWATFVKTISMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHL 304
D + + + ++ Q + + Q A ER G R+A ++ L
Sbjct: 418 TDYLLVPYTR--QNLTWTQHAFNESIGEIQGIATAAFERLKG----RWACLQKRTEV-KL 470
Query: 305 HILKDIIYACIILHNMIVEDEREM 328
L ++ AC +LHN+ + EM
Sbjct: 471 QDLPYVLGACCVLHNICEMRKEEM 494
>At2g26630 En/Spm-like transposon protein
Length = 292
Score = 46.6 bits (109), Expect = 2e-05
Identities = 46/174 (26%), Positives = 69/174 (39%), Gaps = 24/174 (13%)
Query: 166 PTAWKGQYCRGDHGKPTIILEAIASQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVLQGK 225
P + + +G +PT+ + AI + D+ +A+ GV G +D VLN
Sbjct: 57 PPSQTAKKYKGRKLEPTMNVLAICNFDMKFIYAYVGVPGRAHDTKVLNYCAT-------N 109
Query: 226 APEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISM------------PQGEKRKLFAQH 273
P NG YYL D YP ++ P R+LF +
Sbjct: 110 EPYFSHPPNGK-----YYLVDSGYPTRTGYLGPHRRMRYHLGQFGRGGPPVTARELFNRK 164
Query: 274 QEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDERE 327
G R +ER FGV +++ IV + L I+ A + LHN I + RE
Sbjct: 165 HSGLRSVIERTFGVWKAKWRIVDRKHPKYGLAKWIKIVTATMALHNFIRDSHRE 218
>At5g35490 putative protein
Length = 106
Score = 41.6 bits (96), Expect = 8e-04
Identities = 19/36 (52%), Positives = 24/36 (65%)
Query: 267 RKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAW 302
+K+FA QE RKDVER F VL S+FAI+ + W
Sbjct: 69 KKMFAAKQEACRKDVERVFRVLQSKFAIIARSSNCW 104
>At3g63270 unknown protein
Length = 396
Score = 41.2 bits (95), Expect = 0.001
Identities = 59/258 (22%), Positives = 96/258 (36%), Gaps = 16/258 (6%)
Query: 75 LSPLQKQTA-AIRMLAYGSSTDSVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKTNN 133
L ++KQ A A+R LA G S SV +G+ST + RF++ + E +LR ++
Sbjct: 102 LLSVEKQVAIALRRLASGDSQVSVGAAFGVGQSTVSQVTWRFIEALEE-RAKHHLRWPDS 160
Query: 134 KDIARL-MELGKARGFPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIILEAIASQD 192
I + + + G P G+ID H I +C + ++ L+ + +
Sbjct: 161 DRIEEIKSKFEEMYGLPNCCGAIDTTHIIMTLPAVQASDDWCDQEKNY-SMFLQGVFDHE 219
Query: 193 LWIWHAFFGVAGSNNDINVLNQSDVFN-----DVLQGKAPEVQFTLNGTTYNMGYYLADE 247
+ + G G +L S F +L G TL+ Y +
Sbjct: 220 MRFLNMVTGWPGGMTVSKLLKFSGFFKLCENAQILDGNPK----TLSQGAQIREYVVGGI 275
Query: 248 IYP--EWATFVKTISMPQGEKRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLH 305
YP W P + F + E R AF L + I+
Sbjct: 276 SYPLLPWLITPHDSDHP-SDSMVAFNERHEKVRSVAATAFQQLKGSWRILSKVMWRPDRR 334
Query: 306 ILKDIIYACIILHNMIVE 323
L II C +LHN+I++
Sbjct: 335 KLPSIILVCCLLHNIIID 352
>At5g33270 putative protein
Length = 343
Score = 40.0 bits (92), Expect = 0.002
Identities = 46/206 (22%), Positives = 83/206 (39%), Gaps = 7/206 (3%)
Query: 152 LGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIILEAIASQDLWIWHAFFGVAGSNNDINV 211
+G++D H + P+ +Y RG + T+ + A+ + + +A+ GV G +D V
Sbjct: 117 IGALDGTH-VSVRPPSGDVERY-RGRKSEATMNILALCNFSMKFTYAYVGVPGRAHDTKV 174
Query: 212 LNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGEKRKLFA 271
L + ++ YL + + P R+LF
Sbjct: 175 LTYCATHEASFPHPPAGKYYLVDSGYPTRSGYLGPHRRTRYHLELFNRGGPPTNSRELFN 234
Query: 272 QHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDEREMCD- 330
+ R +ER FGV +++ I+ + + I+ + + LHN I + ++E D
Sbjct: 235 RRHSSLRSVIERTFGVWKAKWRILDRKHLKYEVKKWIKIVTSTMALHNYIRDSQQEDSDF 294
Query: 331 --GNIDFSYDQL--ENDASTPEVFSG 352
I SY+Q END P V +G
Sbjct: 295 RHWEIVESYEQHGDENDGHVPYVPTG 320
>At5g27250 putative protein
Length = 348
Score = 40.0 bits (92), Expect = 0.002
Identities = 53/216 (24%), Positives = 85/216 (38%), Gaps = 41/216 (18%)
Query: 187 AIASQDLWIWHAFFGVAGSNNDINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLAD 246
A + DL + G GS +D S V D L + +Q YYLAD
Sbjct: 147 AACNFDLEFIYVLSGWEGSAHD------SKVLQDALTRRTNRLQVPEGK------YYLAD 194
Query: 247 EIYPEWATFVKTISMPQ-------GEKR------KLFAQHQEGARKDVERAFGVL*SRFA 293
+P F+ + + GE R +LF R +ER FG+ SRF
Sbjct: 195 CGFPNRRNFLAPLRSTRYHLQDFRGEGRDPTNQNELFNLRHASLRNVIERIFGIFKSRFL 254
Query: 294 IVRGPARAWHLHILKDIIYACIILHNMIVEDEREMCDGNIDFSYDQLE----NDASTPEV 349
I + A + +I+ +C LHN + R+ C + +FS D+ + ++A+
Sbjct: 255 IFKS-APPFSFKTQAEIVLSCAALHNFL----RQKCRSD-EFSSDEEDETDVDNANQNSE 308
Query: 350 FSGPHPDFATYLQKRAQVREKTVHLHLQADLVEHVW 385
+G + T Q+R R H +A + +W
Sbjct: 309 ENGGEENVETQEQEREHAR------HWRATIAADMW 338
>At5g41980 unknown protein
Length = 374
Score = 39.3 bits (90), Expect = 0.004
Identities = 63/296 (21%), Positives = 115/296 (38%), Gaps = 44/296 (14%)
Query: 36 EQFQRRYRMRKHMFLRIVEAISNTDDYFQMRNDATGRMGLSPLQKQTAA-IRMLAYGSST 94
EQ +RM K +F ++ + + +R+ T R+ ++ Q A + ++ + T
Sbjct: 40 EQCFENFRMDKPVFYKLCDLLQTRG---LLRH--TNRI---KIEAQLAIFLFIIGHNLRT 91
Query: 95 DSVDEYVCIGESTARKCMARFVKGINEVFGPEYLRKTNNKDIARLMELGKARGFPGMLGS 154
+V E C T + + + + ++ + +N D F +G
Sbjct: 92 RAVQELFCYSGETISRHFNNVLNAVIAI-SKDFFQPNSNSDTLE----NDDPYFKDCVGV 146
Query: 155 IDCMH-----WIWENGPTAWKGQYCRGDHGKPTIILEAIASQDLWIWHAFFGVAGSNNDI 209
+D H + E GP R +G T + A +S DL + G GS +D
Sbjct: 147 VDSFHIPVMVGVDEQGPF-------RNGNGLLTQNVLAASSFDLRFNYVLAGWEGSASDQ 199
Query: 210 NVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTI----SMPQGE 265
VLN + + LQ P+ + YY+ D YP F+ + + E
Sbjct: 200 QVLNAALTRRNKLQ--VPQGK-----------YYIVDNKYPNLPGFIAPYHGVSTNSREE 246
Query: 266 KRKLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMI 321
+++F + + + + R FG L RF I+ A + L ++ A LHN +
Sbjct: 247 AKEMFNERHKLLHRAIHRTFGALKERFPILLS-APPYPLQTQVKLVIAACALHNYV 301
>At1g61510 hypothetical protein
Length = 608
Score = 38.1 bits (87), Expect = 0.008
Identities = 47/210 (22%), Positives = 83/210 (39%), Gaps = 7/210 (3%)
Query: 148 FPGMLGSIDCMHWIWENGPTAWKGQYCRGDHGKPTIILEAIASQDLWIWHAFFGVAGSNN 207
F +G++D H + P+ +Y RG + T+ + A+ + + A+ GV G +
Sbjct: 378 FIDCIGALDGTH-VSVRPPSGDVERY-RGRKSEATMNILAVCNFSMKFNIAYVGVPGRAH 435
Query: 208 DINVLNQSDVFNDVLQGKAPEVQFTLNGTTYNMGYYLADEIYPEWATFVKTISMPQGEKR 267
D VL + ++ YL + + P R
Sbjct: 436 DTKVLTYCATHEASFPHPPAGKYYLVDSGYPTRSGYLGPHRRTRYHLELFNRGGPPTNSR 495
Query: 268 KLFAQHQEGARKDVERAFGVL*SRFAIVRGPARAWHLHILKDIIYACIILHNMIVEDERE 327
+LF + R +ER FGV +++ I+ + + I+ + + LHN I + ++E
Sbjct: 496 ELFNRRHSSLRSVIERTFGVWKAKWRILDRKHPKYEVKKWIKIVTSTMALHNYIRDSQQE 555
Query: 328 MCD---GNIDFSYDQL--ENDASTPEVFSG 352
D I SY+Q END P V +G
Sbjct: 556 DSDFRHWEIVESYEQHGDENDGHVPYVPTG 585
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.138 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,285,254
Number of Sequences: 26719
Number of extensions: 403480
Number of successful extensions: 797
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 766
Number of HSP's gapped (non-prelim): 33
length of query: 391
length of database: 11,318,596
effective HSP length: 101
effective length of query: 290
effective length of database: 8,619,977
effective search space: 2499793330
effective search space used: 2499793330
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0011a.2


