

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.6
(493 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
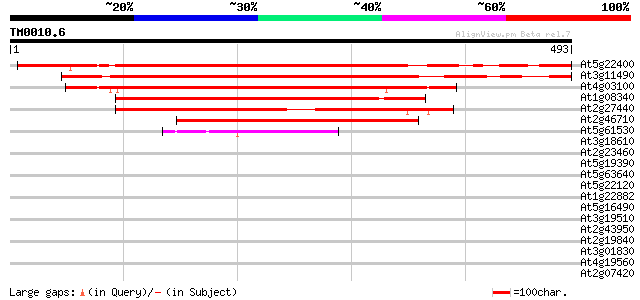
Score E
Sequences producing significant alignments: (bits) Value
At5g22400 rac GTPase activating protein 444 e-125
At3g11490 putative rac GTPase activating protein 424 e-119
At4g03100 rac GTPase activating protein like 379 e-105
At1g08340 unknown protein 358 5e-99
At2g27440 putative rac GTPase activating protein 290 1e-78
At2g46710 putative rac GTPase activating protein 272 4e-73
At5g61530 unknown protein 45 9e-05
At3g18610 unknown protein 39 0.009
At2g23460 extra-large GTP-binding protein (XLG1) 35 0.12
At5g19390 unknown protein 33 0.28
At5g63640 unknown protein (At5g63640) 32 0.80
At5g22120 unknown protein 32 1.0
At1g22882 unknown protein 32 1.0
At5g16490 putative protein 31 1.4
At3g19510 putative homeobox protein, HAT3.1 31 1.8
At2g43950 unknown protein 31 1.8
At2g19840 copia-like retroelement pol polyprotein 31 1.8
At3g01830 unknown protein 30 2.3
At4g19560 putative protein 30 3.0
At2g07420 putative retroelement pol polyprotein 30 3.0
>At5g22400 rac GTPase activating protein
Length = 466
Score = 444 bits (1143), Expect = e-125
Identities = 259/497 (52%), Positives = 316/497 (63%), Gaps = 65/497 (13%)
Query: 8 PSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEE----------KEKDR 57
PSPS S+ T LI++ + V + +D EE++ +E D
Sbjct: 24 PSPSSLSYASRSNATLLISSDHNRRNPVARFDQDVDFHASIEEQDLRRRSSTDGGEEDDG 83
Query: 58 EGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVRHVAHVTFDRFHGF 117
DQ+SLL LL+A FR+SLI SC ++ R+ SMEIGWP+NVRHVAHVTFDRF+GF
Sbjct: 84 GEDQISLLALLVAIFRRSLI-SCKSNRREL-----CSMEIGWPTNVRHVAHVTFDRFNGF 137
Query: 118 LGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQA 177
LGLPVEFEPEVPRR PSAS +VFGVSTESMQLS+D+RGN VPTILLLMQ LY++GGLQA
Sbjct: 138 LGLPVEFEPEVPRRAPSASATVFGVSTESMQLSYDSRGNCVPTILLLMQNCLYSQGGLQA 197
Query: 178 EGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQ 237
EGIFR+ AENS+EE VREQLNRG +P +DVHCLAGLIKAWFRELPT +LD LSPE+VMQ
Sbjct: 198 EGIFRLTAENSEEEAVREQLNRGFIPERIDVHCLAGLIKAWFRELPTSVLDSLSPEQVMQ 257
Query: 238 SQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAPNMTHMADP 297
Q+EEE +LVRLLPPTEAALLDWAINLMADV Q EH NKMN+RNIAMVFAPNMT M DP
Sbjct: 258 CQTEEENVELVRLLPPTEAALLDWAINLMADVVQYEHLNKMNSRNIAMVFAPNMTQMDDP 317
Query: 298 LTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSEN 357
LTALMYAVQVM+FLKTL+ KTLRER++S+V+ L D+ GHQS SQ L
Sbjct: 318 LTALMYAVQVMNFLKTLIEKTLRERQDSVVEQAHAFPLEPSDESGHQSPSQSL------- 370
Query: 358 GNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCN 417
F ++E S+ + + + E+ S SE S L +++ C
Sbjct: 371 ---------AFNTSEQSEETQSDNIENAENQSSSSEIS-----------DELTLENNAC- 409
Query: 418 VVSQLCSFAIGLQDSSIATGQAKISR-SKSLQMSTSDIDKSFKNVIEFPVVGPAEKNRGT 476
+ G+ + R S S Q ++D P P + +G
Sbjct: 410 ------------EQRETDFGKYRTGRLSDSSQQVVLNLDP--------PAQWPVGRTKGL 449
Query: 477 AIIGRINSRTELTEAWR 493
+ R+ SR E TEAWR
Sbjct: 450 TNLSRVGSRVERTEAWR 466
>At3g11490 putative rac GTPase activating protein
Length = 435
Score = 424 bits (1091), Expect = e-119
Identities = 240/449 (53%), Positives = 293/449 (64%), Gaps = 61/449 (13%)
Query: 46 EEDEEEEKEKDREGDQLS-LLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEIGWPSNVR 104
EE+EEE EK+RE +LS L +L++ R+S+IG C G SMEIG P++VR
Sbjct: 47 EEEEEERSEKERERFELSSALEILVSAIRRSVIGGCV------GEEDLCSMEIGVPTDVR 100
Query: 105 HVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTILLL 164
HVAHVTFDRFHGFLGLPVEFEPEVPRR PSAS +VFGVSTESMQLS+D RGN VPTILL+
Sbjct: 101 HVAHVTFDRFHGFLGLPVEFEPEVPRRAPSASATVFGVSTESMQLSYDTRGNIVPTILLM 160
Query: 165 MQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFRELPT 224
MQ HLY+RGGL+ EGIFRIN EN QEE +RE+LN+G++P+ +DVHCLA LIKAWFRELP+
Sbjct: 161 MQSHLYSRGGLRVEGIFRINGENGQEEYIREELNKGIIPDNIDVHCLASLIKAWFRELPS 220
Query: 225 GILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARNIA 284
G+LD LSPE+VM+S+SE+EC +LVRLLP TEA+LLDWAINLMADV +ME NKMNARNIA
Sbjct: 221 GVLDSLSPEQVMESESEDECVELVRLLPSTEASLLDWAINLMADVVEMEQLNKMNARNIA 280
Query: 285 MVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVPNLNSFDDDGHQ 344
MVFAPNMT M DPLTALMYAVQVM+FLKTL+VKTL++R+ES K P N + D +G Q
Sbjct: 281 MVFAPNMTQMLDPLTALMYAVQVMNFLKTLIVKTLKDRKESRDKLVPASNPSPRDHNGDQ 340
Query: 345 SDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLS 404
S S+ L N + T D E E + E
Sbjct: 341 SSSRQLLHLMKANKEE---------------------TLDNFEAEMKDKEESADEEEEEC 379
Query: 405 SGSRLLIDSCPCNVVSQLCSFAIGLQDSSIATGQAKISRSKSLQMSTSDIDKSFKNVIEF 464
+ S L+D ++V+ +SS GQ I + MS
Sbjct: 380 AESVELVDIKKSSLVN----------NSSGGFGQKHIGWEEQRTMS-------------- 415
Query: 465 PVVGPAEKNRGTAIIGRINSRTELTEAWR 493
+ ++I+GR+N R EL EAWR
Sbjct: 416 ---------KASSIVGRVNYRVELFEAWR 435
>At4g03100 rac GTPase activating protein like
Length = 430
Score = 379 bits (973), Expect = e-105
Identities = 202/354 (57%), Positives = 253/354 (71%), Gaps = 13/354 (3%)
Query: 50 EEEKEKDREGDQLSLLTLLIATFRKSLIGSCSTSPRDS-----GALSSS--SMEIGWPSN 102
EEE+E+++ QLSL+ L+ RKS++ SC R G +SS+ MEIGWP+N
Sbjct: 27 EEEEEEEQNQQQLSLVEFLLTALRKSVV-SCRVDNRQDDGGVGGGISSAVHHMEIGWPTN 85
Query: 103 VRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDARGNSVPTIL 162
VRH+ HVTFDRFHGFLGLP E + E+P R PSAS SVFGVS ESMQ S+D +GNSVPTIL
Sbjct: 86 VRHITHVTFDRFHGFLGLPHELQVEIPCRVPSASVSVFGVSAESMQCSYDEKGNSVPTIL 145
Query: 163 LLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAGLIKAWFREL 222
LLMQ LY++ GL+AEGIFRIN ENSQEE VR+QLNRG+VP +DVHCLAGLIKAWFREL
Sbjct: 146 LLMQERLYSQQGLKAEGIFRINPENSQEEHVRDQLNRGIVPENIDVHCLAGLIKAWFREL 205
Query: 223 PTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQMEHFNKMNARN 282
P+G+LD LSPEEV+ +E+E +L++ L PTE+ALL+WA++LMADV + E NKMNARN
Sbjct: 206 PSGVLDGLSPEEVLNCNTEDESVELIKQLKPTESALLNWAVDLMADVVEEEESNKMNARN 265
Query: 283 IAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKS---NPVPNLNS-F 338
IAMVFAPNMT M DPLTALM+AVQVM+ LKTL+ KTL EREE+ S +P + NS
Sbjct: 266 IAMVFAPNMTQMTDPLTALMHAVQVMNLLKTLITKTLAEREENATGSEGYSPSHSSNSQT 325
Query: 339 DDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGS 392
D D + + + ++C +E+ V E Q + H+ ET+ GS
Sbjct: 326 DSDSDNAQDMEVSCESQATDSECGEEEEV-EEVEQHQEHLSRHSTHEDETDIGS 378
>At1g08340 unknown protein
Length = 404
Score = 358 bits (918), Expect = 5e-99
Identities = 179/273 (65%), Positives = 219/273 (79%), Gaps = 4/273 (1%)
Query: 94 SMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDA 153
+M+IG P+N+RHVAHVTFDRF GFLGLP EFEP+VPR+ PSAS +VFGVSTESMQLS+D+
Sbjct: 73 AMDIGGPTNIRHVAHVTFDRFDGFLGLPSEFEPDVPRKAPSASATVFGVSTESMQLSYDS 132
Query: 154 RGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAG 213
RGN VP ILLL+Q LY +GGLQAEG+FRI ENS+EE VREQLN+G++P+G+DVHCLAG
Sbjct: 133 RGNCVPVILLLLQSRLYDQGGLQAEGVFRITGENSEEEFVREQLNKGIIPDGIDVHCLAG 192
Query: 214 LIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQME 273
LIKAWFRELP G+LDPL E+VMQ +S+E+ ++VRLLP TEA+LL+WAINLMADV Q E
Sbjct: 193 LIKAWFRELPRGVLDPLPSEQVMQCESDEDFVKVVRLLPQTEASLLNWAINLMADVIQFE 252
Query: 274 HFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVP 333
H NKMN+RN+A+VFAPNM+ MADPLTALMYAVQVM LK+L KT+RERE S S+ V
Sbjct: 253 HVNKMNSRNLALVFAPNMSQMADPLTALMYAVQVMKLLKSLTEKTVREREAS---SSVVD 309
Query: 334 NLNSFD-DDGHQSDSQVLPKDGSENGNDCSDED 365
S + +DG + ++ E + DED
Sbjct: 310 RRCSKEAEDGEKEKDNEEEEEDEEEEEEEEDED 342
>At2g27440 putative rac GTPase activating protein
Length = 368
Score = 290 bits (743), Expect = 1e-78
Identities = 155/307 (50%), Positives = 202/307 (65%), Gaps = 34/307 (11%)
Query: 94 SMEIGWPSNVRHVAHVTFDRFHGFLGLPVEFEPEVPRRPPSASTSVFGVSTESMQLSFDA 153
+M+I P+N+ HVAHVT+DRF GFLGLP EFEP+VP++PPSAS +VFGVSTESMQLS+D+
Sbjct: 86 AMDISRPTNISHVAHVTYDRFDGFLGLPSEFEPDVPKKPPSASATVFGVSTESMQLSYDS 145
Query: 154 RGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGVDVHCLAG 213
RGN VPTIL L+Q LY +GGLQ EGIFRI +NS+EE +RE+LN+GV+P G+D+HCLAG
Sbjct: 146 RGNCVPTILTLLQSRLYDQGGLQVEGIFRITGDNSEEEFIREELNKGVLPEGIDIHCLAG 205
Query: 214 LIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLMADVAQME 273
LIKAWFRELP G+LD L ++VMQ +S E+ V E
Sbjct: 206 LIKAWFRELPKGVLDSLPSQQVMQCESGEDF------------------------VKVFE 241
Query: 274 HFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESIVKSNPVP 333
NKM +RN+A+VFAPNM+ MADPLTALMYAVQVM+ L+ L KTLRER+ + +P
Sbjct: 242 VVNKMTSRNLALVFAPNMSQMADPLTALMYAVQVMNLLRNLTDKTLRERKIATSNVDPCD 301
Query: 334 NLNSFDDDGHQSDSQ-----VLPKDGSENGNDCSDEDT-----VFVSAEPSQPSPTHHTE 383
N + +D + +Q VL + E G D D D + + A+ +PS +
Sbjct: 302 NRSEAEDGNVEEYNQEVEIYVLEEVEEEEGEDVDDLDNEESERITLLADEHKPSSAVNAN 361
Query: 384 DGCETES 390
D + E+
Sbjct: 362 DRKKQET 368
>At2g46710 putative rac GTPase activating protein
Length = 301
Score = 272 bits (695), Expect = 4e-73
Identities = 136/214 (63%), Positives = 167/214 (77%), Gaps = 1/214 (0%)
Query: 147 MQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVREQLNRGVVPNGV 206
MQ S+D RGNSVPTILL MQ+ LY GGL+AEGIFRIN +N +EE VR QLN GVVP G+
Sbjct: 1 MQCSYDDRGNSVPTILLRMQKRLYTEGGLKAEGIFRINPDNGKEEHVRRQLNCGVVPRGI 60
Query: 207 DVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSEEECDQLVRLLPPTEAALLDWAINLM 266
DVHCLAGLIKAWFRELPTG+LD L+PE+VM+ +EE+C +LV LLPP E+A+LDWAI LM
Sbjct: 61 DVHCLAGLIKAWFRELPTGVLDVLTPEQVMRCNTEEDCSRLVILLPPVESAILDWAIGLM 120
Query: 267 ADVAQMEHFNKMNARNIAMVFAPNMTHMADPLTALMYAVQVMSFLKTLVVKTLREREESI 326
ADV + E FNKMNARN+AMVFAPNMT MADPLTAL++AVQVM+FLKTL++ L+ERE +
Sbjct: 121 ADVVEHEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILMNLKERENAD 180
Query: 327 VKSNPVPNLNSFDDDGHQSD-SQVLPKDGSENGN 359
K+ + S + +S S++L + N N
Sbjct: 181 AKARWLKKQTSDPSEEWESQHSEILSPEKPNNNN 214
>At5g61530 unknown protein
Length = 376
Score = 45.1 bits (105), Expect = 9e-05
Identities = 40/159 (25%), Positives = 72/159 (45%), Gaps = 6/159 (3%)
Query: 135 ASTSVFGVSTESMQLSFDARGNSVPTILLLMQRHLYARGGLQAEGIFRINAENSQEELVR 194
AS+ VFGV+ E + + +P IL+ +L G L + +F+ + + +
Sbjct: 132 ASSDVFGVAIE-ITVQRQESSRPIPLILVKCADYLILTG-LNSPNLFKAEGDRKLIQQLV 189
Query: 195 EQLN---RGVVPNGVDVHCLAGLIKAWFRELPTGILDPLSPEEVMQSQSE-EECDQLVRL 250
N R +P GV+ +A L+K + LPT + E+ ++S Q ++
Sbjct: 190 SAYNQDPRASIPEGVNPVDVAALLKYYLASLPTPLTTFELYNEIKDARSSIHRMRQSLQK 249
Query: 251 LPPTEAALLDWAINLMADVAQMEHFNKMNARNIAMVFAP 289
L L++ L+ V+Q NKM++ ++AM AP
Sbjct: 250 LSNVNYNTLEFITALLLRVSQKSLLNKMDSHSLAMEMAP 288
>At3g18610 unknown protein
Length = 636
Score = 38.5 bits (88), Expect = 0.009
Identities = 45/158 (28%), Positives = 66/158 (41%), Gaps = 20/158 (12%)
Query: 336 NSFDDDGHQSDSQVLP---------KDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGC 386
+S DDG SD + P KD S+ + SDE+T V +P+ E
Sbjct: 210 SSSSDDGSSSDEEPTPAKKEPIVVKKDSSDESS--SDEETPVVKKKPTTVVKDAKAESSS 267
Query: 387 -ETESGSETSPTPAE--NFLSSGSRLLIDSCPCNVVSQLCSFAIGLQDSSIATGQAKISR 443
E ES S+ PTPA+ + + DS S+ S D T +AK+S
Sbjct: 268 SEEESSSDDEPTPAKKPTVVKNAKPAAKDSSS----SEEDSDEEESDDEKPPTKKAKVSS 323
Query: 444 SKSLQMSTSD--IDKSFKNVIEFPVVGPAEKNRGTAII 479
S Q S+SD D+S K + V P +K+ ++
Sbjct: 324 KTSKQESSSDESSDESDKEESKDEKVTPKKKDSDVEMV 361
>At2g23460 extra-large GTP-binding protein (XLG1)
Length = 888
Score = 34.7 bits (78), Expect = 0.12
Identities = 27/95 (28%), Positives = 46/95 (48%), Gaps = 14/95 (14%)
Query: 9 SPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDREGDQLSLLTLL 68
SP+ +N GS+Q L+++S +V ++E EEEE+E+ +G+ L
Sbjct: 72 SPTSVIANCGSNQLELVSDSITVSP--------TSVIEHTEEEEEEEGGDGEDCEL---- 119
Query: 69 IATFRKSLIGSCSTSPR-DSGALSSSSMEIGWPSN 102
++ + L+ SCS D SS+ + W SN
Sbjct: 120 -SSSGELLLRSCSVKESLDLNESSSNPLVPDWESN 153
>At5g19390 unknown protein
Length = 870
Score = 33.5 bits (75), Expect = 0.28
Identities = 29/126 (23%), Positives = 51/126 (40%), Gaps = 10/126 (7%)
Query: 174 GLQAEGIFRINAENSQEELVREQLNRGVVPNGVDV--HCLAGLIKAWFRELPTGILDPLS 231
G + EGI R +A+ + E ++ +G D H + IK RELP+ +
Sbjct: 190 GTKIEGILRQSADVEEVERRVQEYEQGKTEFTFDEDPHVVGDCIKHVLRELPSSPVSASC 249
Query: 232 PEEVMQSQSEEECDQLVRLL--------PPTEAALLDWAINLMADVAQMEHFNKMNARNI 283
++++ E + + L P LL + +M ++ H N+MN +
Sbjct: 250 CTALLEAYRIESKEARISSLRSAIAETFPEPNRRLLQRILKMMHTISSHSHENRMNPNAV 309
Query: 284 AMVFAP 289
A AP
Sbjct: 310 AACMAP 315
>At5g63640 unknown protein (At5g63640)
Length = 447
Score = 32.0 bits (71), Expect = 0.80
Identities = 16/46 (34%), Positives = 24/46 (51%)
Query: 10 PSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEK 55
P R SN + +N+++ + + HL L EEDEEEE E+
Sbjct: 287 PVRPISNGDQKRELKASNANTESSSFISNRAHLKLEEEDEEEEPEQ 332
>At5g22120 unknown protein
Length = 383
Score = 31.6 bits (70), Expect = 1.0
Identities = 18/47 (38%), Positives = 25/47 (52%), Gaps = 3/47 (6%)
Query: 333 PNLNSFDDDGHQSDSQVLPKDGSENGNDCSDED-TVFVSAEPSQPSP 378
PN++SFD DG DS+V + +D SD+D EPS+ P
Sbjct: 126 PNVSSFDADGKTEDSKV--SSSNSAAHDSSDDDWEALADVEPSKLLP 170
>At1g22882 unknown protein
Length = 660
Score = 31.6 bits (70), Expect = 1.0
Identities = 29/119 (24%), Positives = 54/119 (45%), Gaps = 10/119 (8%)
Query: 300 ALMYAVQVMSFLKTLVVKTLREREESIVKSNPV-PNLNSFDDDGHQSDSQVLPKDGSENG 358
+L++ + V+ F TL++ K P+ ++ D D QSD +V+P DG +
Sbjct: 28 SLVFLLWVLLFFSTLLIS-----HGDGAKDEPLNDSMGMADPDDGQSDEKVVPFDGPLSL 82
Query: 359 NDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSRLLIDSCPCN 417
S V V+++ S+ + +E+ + E +E S T + N + S L+ N
Sbjct: 83 ASAS----VDVTSDLSRNDDVNLSEESEDKEQEAEISSTVSGNDIESKDTYLLKQSEIN 137
>At5g16490 putative protein
Length = 153
Score = 31.2 bits (69), Expect = 1.4
Identities = 19/75 (25%), Positives = 38/75 (50%), Gaps = 6/75 (8%)
Query: 44 LVEEDEEEEKEKDR------EGDQLSLLTLLIATFRKSLIGSCSTSPRDSGALSSSSMEI 97
++ E EEE+ K+ ++S+ I K++ + + + +S MEI
Sbjct: 40 VIREQEEEDNMKNVFKFLAVSKPEISIGINRIFKSFKTISQLFADKDEEKEEVETSGMEI 99
Query: 98 GWPSNVRHVAHVTFD 112
G P+NV+HV+H+ ++
Sbjct: 100 GVPTNVKHVSHIGWE 114
>At3g19510 putative homeobox protein, HAT3.1
Length = 723
Score = 30.8 bits (68), Expect = 1.8
Identities = 29/103 (28%), Positives = 40/103 (38%), Gaps = 9/103 (8%)
Query: 346 DSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHTEDGCETESGSETSPTPAENFLSS 405
D LP D SE+ + D T E S T TED + G ET+ + L
Sbjct: 428 DVMALPSDDSEDDDYDPDAPTCDDDKESSNSDCTSDTEDLETSFKGDETNQQAEDTPLED 487
Query: 406 GSRLLIDSCPCNVVSQLCSFAIGLQDSSIATGQAKISRSKSLQ 448
P SQL AI D + G A +SR ++++
Sbjct: 488 ---------PGRQTSQLQGDAILESDVGLDDGPAGVSRRRNVE 521
>At2g43950 unknown protein
Length = 343
Score = 30.8 bits (68), Expect = 1.8
Identities = 18/75 (24%), Positives = 28/75 (37%), Gaps = 14/75 (18%)
Query: 373 PSQPSPTHHTEDGCETESGSETSPTPAENFLSSGSR--------------LLIDSCPCNV 418
PS PSPTH + G S P +FL + +R L ++ C +
Sbjct: 17 PSSPSPTHQIQSGTSELSPPSRPPCSTLSFLKTANRPKLRVTSEFDSDSLLFLNKVSCKL 76
Query: 419 VSQLCSFAIGLQDSS 433
L + Q++S
Sbjct: 77 FDNLAKLKLSFQNNS 91
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 30.8 bits (68), Expect = 1.8
Identities = 30/136 (22%), Positives = 54/136 (39%), Gaps = 18/136 (13%)
Query: 323 EESIVKSNPVPNLNSFDDDGHQSDSQVLPKDGSENGNDCSDEDTVFVSAEPSQPSPTHHT 382
++++V+ P+P L D+ Q + ++ + D D +F++
Sbjct: 4 KDALVERAPLPPLKEEDESDPAKKKQRIEEEKARIDQDEKAMDMIFINV----------- 52
Query: 383 EDGCETESGSETSPTPAENFLSSGSRLLIDSCPCNVVSQLCSFAIGLQDS-----SIATG 437
G + E S T AE + + L+ S P V QL + +QDS ++
Sbjct: 53 --GDKVLRNIENSKTAAEAWATLDKLYLVKSLPNRVYLQLKVYNYRMQDSKTLEENVDEF 110
Query: 438 QAKISRSKSLQMSTSD 453
Q IS +LQ+ D
Sbjct: 111 QKMISDLNNLQIQVPD 126
>At3g01830 unknown protein
Length = 146
Score = 30.4 bits (67), Expect = 2.3
Identities = 14/31 (45%), Positives = 20/31 (64%)
Query: 26 NNSSSVEEGVLQLQTHLDLVEEDEEEEKEKD 56
N S ++ L+L+ + LVEE EE +KEKD
Sbjct: 56 NPEESTDDKSLELEDFVKLVEEGEEADKEKD 86
>At4g19560 putative protein
Length = 474
Score = 30.0 bits (66), Expect = 3.0
Identities = 17/53 (32%), Positives = 28/53 (52%)
Query: 6 QLPSPSRRCSNDGSHQTALINNSSSVEEGVLQLQTHLDLVEEDEEEEKEKDRE 58
++ S S N H I S + GV+QL+ L L +E+ E ++EKD++
Sbjct: 344 EVNSESEAQKNLQDHSVGNIMVEKSDDVGVVQLKKDLQLHQEEVESKQEKDKK 396
>At2g07420 putative retroelement pol polyprotein
Length = 1664
Score = 30.0 bits (66), Expect = 3.0
Identities = 11/25 (44%), Positives = 21/25 (84%)
Query: 43 DLVEEDEEEEKEKDREGDQLSLLTL 67
++ EE+EEEE+E++++G + L+TL
Sbjct: 776 NMEEEEEEEEEEEEKQGKEQELITL 800
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.130 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,202,901
Number of Sequences: 26719
Number of extensions: 505233
Number of successful extensions: 2675
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 2570
Number of HSP's gapped (non-prelim): 88
length of query: 493
length of database: 11,318,596
effective HSP length: 103
effective length of query: 390
effective length of database: 8,566,539
effective search space: 3340950210
effective search space used: 3340950210
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0010.6


