

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0010.16
(405 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
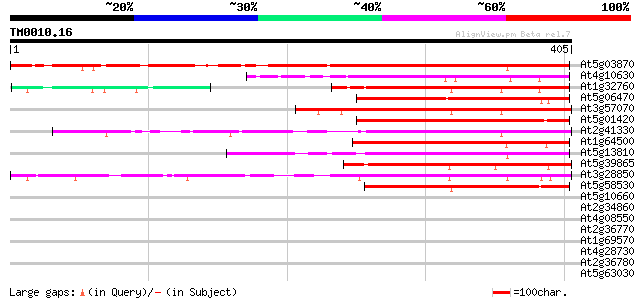
Score E
Sequences producing significant alignments: (bits) Value
At5g03870 putative protein 340 8e-94
At4g10630 putative protein 180 1e-45
At1g32760 unknown protein 172 3e-43
At5g06470 putative protein 162 3e-40
At3g57070 unknown protein 162 4e-40
At5g01420 unknown protein 158 6e-39
At2g41330 unknown protein 158 6e-39
At1g64500 unknown protein 151 5e-37
At5g13810 unknown protein 149 3e-36
At5g39865 unknown protein 136 2e-32
At3g28850 hypothetical protein 135 5e-32
At5g58530 unknown protein 128 6e-30
At5g10660 vacuolar calcium binding protein - like 37 0.019
At2g34860 unknown protein 36 0.033
At4g08550 hypothetical protein 36 0.043
At2g36770 putative glucosyl transferase 35 0.096
At1g69570 H-protein promoter binding factor-2b, putative 34 0.16
At4g28730 unknown protein 33 0.21
At2g36780 putative glucosyl transferase 33 0.21
At5g63030 glutaredoxin-like protein 33 0.28
>At5g03870 putative protein
Length = 384
Score = 340 bits (872), Expect = 8e-94
Identities = 201/424 (47%), Positives = 270/424 (63%), Gaps = 61/424 (14%)
Query: 1 MGCVSSKLIRKDIKQEHVIIDNCGGGGRYLSHVVSLTSSTYGALKLDKDND--------- 51
MGCVSSKL +K + +E I N GG H+VSLTS+TYG L LD+ +
Sbjct: 1 MGCVSSKLGKKKLIRE--IRVNNGG-----DHIVSLTSTTYGHLDLDERAETSPKSLEVT 53
Query: 52 ------QPVIAAKP--REEPEAITTINAWELMEGLEDGVPISNQPKKSPKSSSPFLRGFM 103
+I + R++PE I N WELME LED + +SN K SPKS RG
Sbjct: 54 KGEVFESEIIPRRSIKRDDPEII---NTWELMEDLEDSMHVSNPQKISPKS-----RGIF 105
Query: 104 NSDTRSPLKFLNQLGSPKTVKKSTGKENKVQVNGVRTGGVRRLDYSNSPKGILKPSNLSP 163
++P+K + + + K+ GKEN + GV SP ILKP N+
Sbjct: 106 GKSWKTPVKSVVESPKRGSSKRFGGKENNSR-------GV-------SPNQILKPKNILE 151
Query: 164 NASKNSGIPAKGSPICARRKSFGGNEKDTLQCSSRRKSQSPLFDPELVASYEKELSEEEE 223
+ G+ P+ +E+ ++ + RRKS SP+FDP+LVASYE+ELS+E+E
Sbjct: 152 TPKR--GVMRLSFPL--------KSEESSVTVTQRRKSYSPMFDPDLVASYERELSQEKE 201
Query: 224 QVKRMVWATPKTRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKV 283
Q+K ++ +P +RK+ + +++ F +KCPPGGE+SVVIY TTLRGIRKTFEDCN V
Sbjct: 202 QIKMVI--SPVVHESRKTEKTERILEKFPEKCPPGGEHSVVIYITTLRGIRKTFEDCNVV 259
Query: 284 RAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVDEVVKLEDEE 343
R+I++S+ V+ ERDVSM S FKEE+R +MG + VK+P VFVKGR+VG V+EV++LE+E
Sbjct: 260 RSILDSHEVRFSERDVSMHSVFKEEIRGIMGTKHVKIPAVFVKGRMVGSVEEVMRLEEEG 319
Query: 344 KLGVLLEGIPKA-VGG--CEGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQ 400
KLG+LLEGIP A +GG C GCGG+RF+MCV CNGSCKV +E+KK VKC KCNENG++
Sbjct: 320 KLGILLEGIPAARLGGSCCRGCGGMRFMMCVVCNGSCKVREEEKKSMVKCLKCNENGLVL 379
Query: 401 CPIC 404
CPIC
Sbjct: 380 CPIC 383
>At4g10630 putative protein
Length = 334
Score = 180 bits (456), Expect = 1e-45
Identities = 112/247 (45%), Positives = 147/247 (59%), Gaps = 26/247 (10%)
Query: 172 PAKGSPICARRKSFGGNEKDTLQCSSRRKSQSPLFDPELVASYEKELSEEEEQVKRMVWA 231
P +P+ + K FGG K+ SRR + L D L +L+ E+
Sbjct: 99 PLPKTPV--KYKVFGGENKENSD-PSRRNPRKNLNDEVLKPL---DLNREDSDSN----- 147
Query: 232 TPKTRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYC 291
+ R++ K LD L L + FE+ CPPGGEN VV+YTT+LRG+R+TFE CN VRA +ES+
Sbjct: 148 SRSPRKSFKPLD-LKLDEKFERICPPGGENRVVMYTTSLRGVRQTFEACNAVRAAVESFG 206
Query: 292 VQMRERDVSMDSGFKEELRKLM----GAEQVKV--PVVFVKGRLVGGVDEVVKLEDEEKL 345
V + ERDVSMD F+EEL LM G E V P VFVKGR +GG +EV++L +E
Sbjct: 207 VVVCERDVSMDRRFREELVSLMAKRVGDEGVAALPPRVFVKGRYIGGGEEVLRLVEEGSF 266
Query: 346 GVLLEGIP-KAVGGCE-----GCGGVRFVMCVECNGSCKVLD--EDKKKTVKCGKCNENG 397
G L+ GIP K GGCE GCGG+ F+ C CNGSCK++ V+C +CNENG
Sbjct: 267 GELISGIPRKKAGGCESGACDGCGGLFFLPCFRCNGSCKMVKGWGSASVVVRCNECNENG 326
Query: 398 IMQCPIC 404
++ CPIC
Sbjct: 327 LVPCPIC 333
>At1g32760 unknown protein
Length = 314
Score = 172 bits (436), Expect = 3e-43
Identities = 95/181 (52%), Positives = 120/181 (65%), Gaps = 11/181 (6%)
Query: 233 PKTRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCV 292
P K LD L + E+ C GGEN VVIYTT+LRG+R+TFE CN VRA IES+ V
Sbjct: 135 PNDYEVLKPLDP-KLAEESERLCD-GGENRVVIYTTSLRGVRRTFEACNAVRAAIESFGV 192
Query: 293 QMRERDVSMDSGFKEELRKLMGAEQ---VKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL 349
+ ERDVSMD GF+EEL LM E V P VFVKG+ +GG +EV++L +E LG LL
Sbjct: 193 VVCERDVSMDRGFREELSNLMAVESTAAVLPPRVFVKGKYIGGAEEVMRLVEEGLLGELL 252
Query: 350 EGIP----KAVGGCEGCGGVRFVMCVECNGSCKVLD--EDKKKTVKCGKCNENGIMQCPI 403
+ IP + GGC GCGG+ F+ C CNGSCKV++ + VKC +CNENG+++CPI
Sbjct: 253 KEIPRKKDRCGGGCGGCGGLAFLPCSGCNGSCKVVEGWGNDAVVVKCKECNENGLVRCPI 312
Query: 404 C 404
C
Sbjct: 313 C 313
Score = 42.7 bits (99), Expect = 4e-04
Identities = 51/182 (28%), Positives = 69/182 (37%), Gaps = 47/182 (25%)
Query: 2 GCVSSKLIRK--DIKQEHVIIDNCGGGGRYLSHVVSLTSSTYGALKLDKDNDQPVIAAK- 58
GCVSS L+ D H+ + H V LTS+TYG L LD + P +
Sbjct: 3 GCVSSNLLTPTDDSSFSHLTSSSLS------QHFVKLTSTTYGLLNLDSSSPSPPSPSSS 56
Query: 59 --------PREEPEAIT------------------TINAWELMEGLE-DGVPISNQPKK- 90
P PE T IN WELM GL+ D S P K
Sbjct: 57 SSAKSVITPMTPPERFTINGKEAAMMMVKSEPTTEVINFWELMSGLDGDTCRFSPIPVKC 116
Query: 91 -------SPKSSSPFLRGFMNSDTRSPLKFLNQLGSPKTVKKSTGKENKVQVNGVRTGGV 143
++S P L+ N + LK L+ + ++ + G EN+V + GV
Sbjct: 117 NGFSGGLKKENSDPNLK---NPNDYEVLKPLDPKLAEESERLCDGGENRVVIYTTSLRGV 173
Query: 144 RR 145
RR
Sbjct: 174 RR 175
>At5g06470 putative protein
Length = 239
Score = 162 bits (410), Expect = 3e-40
Identities = 78/161 (48%), Positives = 113/161 (69%), Gaps = 8/161 (4%)
Query: 251 FEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELR 310
F++ CPPGGE+SVV YTT LR +RKTFE C +VR ++E++ V RERDVSMDS F+EE+
Sbjct: 79 FKENCPPGGEDSVVFYTTGLRSVRKTFEACRRVRFLLENHQVMYRERDVSMDSEFREEMW 138
Query: 311 KLMGAEQVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKAVGGCEGCGGVRFVMC 370
+L+G +V P +F++GR +GG +EVV L + KL LL+GI + CE C RF++C
Sbjct: 139 RLLGG-KVTSPRLFIRGRYIGGAEEVVALNENGKLKKLLQGISQVDSPCESCENERFLIC 197
Query: 371 VECNGSCKVLDE--DKKKT-----VKCGKCNENGIMQCPIC 404
CNGS ++L E D++ + +C +CNENG+++CP+C
Sbjct: 198 SSCNGSTRLLAEHHDEESSNDNMWTRCRECNENGLVKCPLC 238
>At3g57070 unknown protein
Length = 417
Score = 162 bits (409), Expect = 4e-40
Identities = 86/219 (39%), Positives = 133/219 (60%), Gaps = 20/219 (9%)
Query: 207 DPELVASYEKELSEE----------EEQVKRMVWATPKTRRA------RKSLDSLSLIQM 250
DP +V+SY+K LS + + + + +P + + KS+ S LI
Sbjct: 198 DPSIVSSYKKALSSKLLSNHSNTRNPHRPTKSLSCSPSSNPSILISEEPKSVSSSQLISS 257
Query: 251 FEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELR 310
K PG E+ +V+Y TTLRGIRKT+EDC VRAI+ V + ERD+SMDS +++EL+
Sbjct: 258 PAKPRLPGTEDKIVLYFTTLRGIRKTYEDCCCVRAILRGVQVSVDERDISMDSKYRKELQ 317
Query: 311 KLMGAEQ--VKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIP--KAVGGCEGCGGVR 366
++GA + V +P VF++G +GGV+E+++L D +L +L+ P + +G C CG R
Sbjct: 318 SVLGAAEKPVCLPQVFIRGTHIGGVEEIMQLNDGGELAEMLKDFPACERLGTCRSCGDAR 377
Query: 367 FVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPICC 405
FV C C+GS KV +E ++ +C KCNENG+++C +CC
Sbjct: 378 FVPCTNCDGSTKVFEEQDERFKRCPKCNENGLVRCRVCC 416
Score = 40.8 bits (94), Expect = 0.001
Identities = 28/80 (35%), Positives = 43/80 (53%), Gaps = 14/80 (17%)
Query: 32 HVVSLTSSTYGALKL-------DKDNDQ---PVIAAKPREEPEAIT---TINAWELMEGL 78
H+VSLTS++YG+L L + +DQ P I+ + P+ ++ IN WELM+GL
Sbjct: 60 HLVSLTSTSYGSLLLLDLDGSKNSSSDQQTLPRISISGKNTPDPVSPDSVINTWELMDGL 119
Query: 79 EDGVPISNQPKKSPKSSSPF 98
+D PK + +S F
Sbjct: 120 DDEFEF-EIPKPGKRLNSDF 138
>At5g01420 unknown protein
Length = 401
Score = 158 bits (399), Expect = 6e-39
Identities = 78/156 (50%), Positives = 107/156 (68%), Gaps = 3/156 (1%)
Query: 251 FEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELR 310
FE++CPPGGE SVV YTTTLRGIRKTF+DCN +R +++S+ V+ ERDVSM ++EELR
Sbjct: 246 FEERCPPGGEESVVFYTTTLRGIRKTFDDCNMIRFLLDSFKVKYYERDVSMHREYREELR 305
Query: 311 KLMGAE-QVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIP-KAVGGCEGCGGVRFV 368
++ AE +V PV+FVKGR +GG V+ L ++ K +L EGIP C C G RF+
Sbjct: 306 RISAAETEVLPPVLFVKGRCIGGAQRVLGLHEQGKFKILFEGIPITGDERCRRCDGFRFL 365
Query: 369 MCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
MC C GS +++ D + ++C CNENG++ C C
Sbjct: 366 MCDGCRGSRRIISGDGSR-IQCLICNENGLIVCVGC 400
>At2g41330 unknown protein
Length = 402
Score = 158 bits (399), Expect = 6e-39
Identities = 125/401 (31%), Positives = 187/401 (46%), Gaps = 85/401 (21%)
Query: 32 HVVSLTSSTYGALKLDKDNDQPVIAAKPREEPEAITT------------INAWELMEGLE 79
H VSLTS+TYG+L LD P I+ + + T IN WELM L+
Sbjct: 59 HFVSLTSTTYGSLVLDDRQTLPHISVSGKSNKKMPETEEARDSFSPDSVINTWELMNDLD 118
Query: 80 DGVPISNQPKKSPKSSSPFLRGFMNSDTRSPLKFLNQLGSPKTVKKSTGKENKVQVNGVR 139
D +N + KS+S +N D+ S K T ++ V +NG
Sbjct: 119 DEFDSANSD--TSKSNS-----VVNLDSFS--------------KPITNRD--VVINGSA 155
Query: 140 TGGVR-RLDYSNSPKGILKP-----------SNLSPNASKNSGIPAKGSPICARRKSFGG 187
G D+ P +P S+L PN +S A S + S G
Sbjct: 156 YGSYEDEEDWRLLPFKPKQPLWKHMSEESFLSDLDPNII-SSYKRALSSKQLGKNSSNGH 214
Query: 188 NEKDTLQCSSRRKSQSPLFDPELVASYEKELSEEEEQVKRMVWATPKTRRARKSLDSLSL 247
+ L S SPL + SE++++ R++
Sbjct: 215 SNHSVLVSLPEYVSSSPL---------SSQTSEKDQEKPRLL------------------ 247
Query: 248 IQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKE 307
EK+ EN +V+Y T+LRGIRKT+EDC VRAI+ + V + ERD+SMDS +++
Sbjct: 248 ----EKE---DNENKIVVYFTSLRGIRKTYEDCCCVRAILRGFQVAVEERDISMDSKYRK 300
Query: 308 ELRKLMGAEQ-VKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIP--KAVGGCEGCGG 364
EL+ +G E+ V +P VF++G +GG++E+ L D +L +L+ P +++G C+ CG
Sbjct: 301 ELQNALGEEKPVCLPQVFIRGVRIGGIEEIKILNDGGELAEMLKDFPACESIGACDSCGD 360
Query: 365 VRFVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPICC 405
RFV C C GS KV +E + +C CNENG+++C CC
Sbjct: 361 ARFVPCTNCGGSTKVFEEQEDGFKRCNGCNENGLVRCNKCC 401
>At1g64500 unknown protein
Length = 368
Score = 151 bits (382), Expect = 5e-37
Identities = 79/167 (47%), Positives = 109/167 (64%), Gaps = 10/167 (5%)
Query: 248 IQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKE 307
++ F +KCPPGG +++YTT+L+G+R+T+EDC +VRAI+E V + ERDVS+D+G
Sbjct: 199 LREFPEKCPPGGGEGLIVYTTSLQGVRRTYEDCMRVRAIMEQQGVVVDERDVSLDAGVLS 258
Query: 308 ELRKLMGAE-QVKVPVVFVKGRLVGGVDEVVKLEDEEKLG-VLLEGIPKAVG-----GCE 360
EL++L+ E V P VFVKGR +GG EV + + KLG VL + VG CE
Sbjct: 259 ELKELLQDEASVAPPRVFVKGRYLGGAAEVTAMNENGKLGRVLRWARVERVGEEGRLTCE 318
Query: 361 GCGGVRFVMCVECNGSCKVLDEDKKK---TVKCGKCNENGIMQCPIC 404
GCGG R++ C EC GSCKV K +C KCNENG+++CP+C
Sbjct: 319 GCGGARWLPCFECGGSCKVAAVGAAKGERWERCVKCNENGLIRCPVC 365
>At5g13810 unknown protein
Length = 274
Score = 149 bits (376), Expect = 3e-36
Identities = 85/250 (34%), Positives = 140/250 (56%), Gaps = 20/250 (8%)
Query: 157 KPSNLSPNASKNSGIPAKGSPICARRKSFGGNEKDTLQCSSRRKSQSPLFDPELVASYEK 216
K NL+P+ ++ + I +P+ + S G K+ + K P
Sbjct: 41 KSHNLNPSLNRTTSITKFYTPVESMGTSLKGKVKNLCRLFETSKPVKPAL---------A 91
Query: 217 ELSEEEEQVKRMVWATPKTRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKT 276
E+ ++++ K ++ P++R + S + S+I++ PG E+ +V+Y T+LRGIR+T
Sbjct: 92 EIPQKQKSGKSLL---PESRISPFSSLNNSVIRL------PGTEDRIVVYFTSLRGIRRT 142
Query: 277 FEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQVKVPVVFVKGRLVGGVDEV 336
+EDC VR I + V + ERDVSMD +++EL+ MG + V +P VF+ G+ VGG D +
Sbjct: 143 YEDCYAVRMIFRGFRVWIDERDVSMDIAYRKELQIAMGEKSVSLPQVFIMGKYVGGADVI 202
Query: 337 VKLEDEEKLGVLLEGIPKAVGG--CEGCGGVRFVMCVECNGSCKVLDEDKKKTVKCGKCN 394
L + +L +L+ P G C CG +RFV C C+GS K+ DED+ + +C +CN
Sbjct: 203 KSLFEIGELAKILKEFPMRQPGFVCHCCGDIRFVPCSNCSGSKKLFDEDEDRVKRCPECN 262
Query: 395 ENGIMQCPIC 404
ENG+++CP C
Sbjct: 263 ENGLIRCPDC 272
>At5g39865 unknown protein
Length = 390
Score = 136 bits (343), Expect = 2e-32
Identities = 73/184 (39%), Positives = 112/184 (60%), Gaps = 22/184 (11%)
Query: 242 LDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSM 301
LD ++ F++K G+ VV+Y T+LRGIRKT+EDC +R I++S +++ ERDVSM
Sbjct: 208 LDPPDIVSRFKRKTL--GKERVVLYFTSLRGIRKTYEDCCNIRIILKSLGIRIDERDVSM 265
Query: 302 DSGFKEELRKLMGAE-----QVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL------- 349
SGFK+EL+KL+ + + +P VF+ + +GGV+E+ KL + +L L+
Sbjct: 266 HSGFKDELKKLLEGKFNNGVGITLPRVFLGNKYLGGVEEIKKLNENGELEKLIKDCEMVE 325
Query: 350 EGIPKAVGGCEGCGGVRFVMCVECNGSCKVLDEDKKKT--------VKCGKCNENGIMQC 401
+G P CE CG VRFV C C+GSCK+ E +++ +C CNENG+++C
Sbjct: 326 DGSPGFGNECEACGDVRFVPCETCSGSCKLYHEGEEEDEGVTEYGFQRCPYCNENGLIRC 385
Query: 402 PICC 405
+CC
Sbjct: 386 HVCC 389
Score = 35.0 bits (79), Expect = 0.074
Identities = 25/53 (47%), Positives = 27/53 (50%), Gaps = 9/53 (16%)
Query: 53 PVIAAK-----PREEPEAITTINAWELMEGLEDGVPISNQPKKSPKSSSPFLR 100
P I AK P EPE TIN WELMEGLED P+ P S F+R
Sbjct: 125 PKIVAKTPIVTPPGEPE---TINTWELMEGLEDVSPL-RSPNHLRSFSFDFVR 173
>At3g28850 hypothetical protein
Length = 428
Score = 135 bits (339), Expect = 5e-32
Identities = 127/467 (27%), Positives = 205/467 (43%), Gaps = 102/467 (21%)
Query: 1 MGCVSSKLIRK------------DIKQEHVIIDNCGGGGRYLSHVVSLTSSTYGALKL-- 46
MGC SSK ++ D+++ H + + H+V+L+SS+ G+LKL
Sbjct: 1 MGCASSKHRKRCLHCRRGYSPPVDVQRSHSV-HHASQNSEDSCHMVALSSSSLGSLKLCD 59
Query: 47 ---DKDNDQPVIAAKPREEPEAITTINAWELMEGLEDGVPISNQPKKSPKSSSPFLRGFM 103
++ ++ E++ T N + G + + KS+ M
Sbjct: 60 SSFGHNHKHLADFSEKLVSGESVKTGNGF--------GPNVVREKSDKEKSNLELQAKLM 111
Query: 104 NSDTRSPLKFLNQLGSPKTVKKST-----GKENKVQVNGVRTGGVRRLDYSNSPKGILKP 158
+ S + +N+ PK V K+ G+ + + G L SP +
Sbjct: 112 EAKVWSSM--MNEK-IPKIVPKTPIVTPPGEPETINTWEMMDGLEDVLSPLRSPNHVKSF 168
Query: 159 S-NLSPNASKNSGIPAKGSPICARRKSFGGNEKDTLQCSSRRKSQSPLFDPELVASYEKE 217
S ++ PN K++G P+ + + +D FDPE+++S+ K
Sbjct: 169 SFDVGPNGGKSNG---SVKPVWLQMEEEEEGFED--------------FDPEIISSFRKS 211
Query: 218 LSEEEEQVKRMVWATPKTRRARKSLDSLSLIQMF-------EKKCPPGGENSVVIYTTTL 270
L E P S L F E++ G+ V++Y T+L
Sbjct: 212 LQE-----------LPSDHPFHISNHDFELKPRFNFSDEEKEEEEQSVGKERVILYFTSL 260
Query: 271 RGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAE-----QVKVPVVFV 325
RGIRKT+E+ VR I++S +++ ERDVSM SGFK+EL++L+G + + +P VF+
Sbjct: 261 RGIRKTYEESCDVRVILKSLGIRVDERDVSMHSGFKDELKELLGEKFNKGVGITLPRVFL 320
Query: 326 KGRLVGGVDEVVKLEDEEKLGVLLEGIPKAVGG-------CEGCGGVRFVMCVECNGSCK 378
+ +GG +E+ KL ++ KL LL G + CE CG VRFV C C+GSCK
Sbjct: 321 GRKYIGGAEEIRKLNEDGKLEKLLGGCERVEENQNGNGLECEACGDVRFVPCETCSGSCK 380
Query: 379 VLDE-----------DKKKTVK---------CGKCNENGIMQCPICC 405
V E D ++VK C CNENG+++CP+CC
Sbjct: 381 VYYEYEDDDDDDDEGDDDESVKEEREYGFQTCPDCNENGLIRCPVCC 427
>At5g58530 unknown protein
Length = 273
Score = 128 bits (321), Expect = 6e-30
Identities = 69/158 (43%), Positives = 99/158 (61%), Gaps = 11/158 (6%)
Query: 257 PGGENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAE 316
PG E S+V+Y T+LR +R TFE C V +I+ S+ V++ ERD+SMD+ F EL+++ G +
Sbjct: 109 PGAEKSIVVYFTSLRVVRPTFEACKSVTSILHSFPVRIDERDLSMDASFSTELQRIFGKD 168
Query: 317 Q---------VKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLLEGIPKAVGG-CEGCGGVR 366
Q K+P VF+ GR +GG +EV +L + +L L++ +PK G CE CGG R
Sbjct: 169 QNQNQNQAKTPKLPRVFIGGRYIGGAEEVKQLHEIGELKKLVQELPKIEPGVCEMCGGHR 228
Query: 367 FVMCVECNGSCKVLDEDKKKTVKCGKCNENGIMQCPIC 404
FV C +C+GS KV E K C CNENG+++C C
Sbjct: 229 FVPCKDCHGSHKVHTE-KLGFRTCLTCNENGLVRCSSC 265
>At5g10660 vacuolar calcium binding protein - like
Length = 407
Score = 37.0 bits (84), Expect = 0.019
Identities = 75/305 (24%), Positives = 118/305 (38%), Gaps = 75/305 (24%)
Query: 51 DQPVIAAKPREEPEAITTINAWELMEGLEDGVPISNQPKKSP---KSSSPFLRGFMNSDT 107
D+P + PRE+P +T + + G+ S+ K P + SS + ++ ++
Sbjct: 117 DKPAV---PREKP--VTGLRSTSFHGSSRGGLRGSSTVKSPPVASRGSSGVKKSGLSGNS 171
Query: 108 RSPLKFLNQLGSPKTVKKSTGKE-------------NKVQVNGVRTGGVRRLDYSNSPKG 154
S K + GS KKS+GKE ++ ++ V T V D+ PK
Sbjct: 172 SSKSK---KEGSGNVPKKSSGKEISPDSSPLASAHEDEEEIVKVETD-VHISDHGEEPKE 227
Query: 155 ILKPSNLSPNASKNSGIPAKGSPICARRKSFGGNEKDTLQCSSRRKSQSPLFDPELVASY 214
K P+ SG + SP+ A + G D KS + +P+ +
Sbjct: 228 EDKDQFAQPD---ESGEEKETSPVAASTEEQKGELID------EDKSTEQIEEPKEPENI 278
Query: 215 EKELSEEEEQVKRMVWATPKTRRARKSLDSLSLIQMFEKKCPPGGENSVVIYTTTLRGIR 274
E+ SEEEE+VK +KS D ENS + TTT
Sbjct: 279 EENNSEEEEEVK------------KKSDDE---------------ENSETVATTT----- 306
Query: 275 KTFEDCNKVRAIIESYCVQMRERDVSMDSG----FKEELRKLMGAEQVKVPVVFVKGRLV 330
D N+ + ES + E +V + G KEE + M A+ ++P K +V
Sbjct: 307 ----DMNEAVNVEESKEEEKEEAEVKEEEGESSAAKEETTETM-AQVEELPEEGTKNEVV 361
Query: 331 GGVDE 335
G E
Sbjct: 362 QGKKE 366
>At2g34860 unknown protein
Length = 186
Score = 36.2 bits (82), Expect = 0.033
Identities = 33/136 (24%), Positives = 50/136 (36%), Gaps = 19/136 (13%)
Query: 281 NKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAE------QVKVPVVFVKGRLVGGVD 334
N+VR + +S C + + D + E K G + Q V L+
Sbjct: 25 NRVRVLAKS-CPENQSFDSNDSDSSSETTHKAQGDQKSVSRRQWMTACVCASAALISNSY 83
Query: 335 EVVKLEDEEKLGVLLEGIPKAVGGCEGCGGVRFVMCVECNGS--CKVLDEDKKKTV---- 388
V ++ L K G C C G V+C C G+ K L+ + K V
Sbjct: 84 TFVSVQSAAALD------KKPGGSCRNCQGSGAVLCDMCGGTGKWKALNRKRAKDVYEFT 137
Query: 389 KCGKCNENGIMQCPIC 404
+C C G + CP+C
Sbjct: 138 ECPNCYGRGKLVCPVC 153
>At4g08550 hypothetical protein
Length = 587
Score = 35.8 bits (81), Expect = 0.043
Identities = 28/95 (29%), Positives = 46/95 (47%), Gaps = 12/95 (12%)
Query: 263 VVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMRERDVSMDSGFKEELRKLMGAEQVKVPV 322
+++YT R E+C R + ++ E ++ + K EL K+ G + V P+
Sbjct: 226 IILYT------RLGCEECRGCRLFLHEKRLRYVEINIDIYPTRKVELEKISGGDVV--PM 277
Query: 323 VFVKGRLVGGVDEVVKLED----EEKLGVLLEGIP 353
VF +LVG E+ LE+ EEK+ L+E P
Sbjct: 278 VFFNEKLVGSYKELKVLEESGELEEKIKHLIEETP 312
>At2g36770 putative glucosyl transferase
Length = 496
Score = 34.7 bits (78), Expect = 0.096
Identities = 19/40 (47%), Positives = 28/40 (69%), Gaps = 2/40 (5%)
Query: 319 KVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL--EGIPKAV 356
K+ V +K + GV+EV+K +EEK+GVL+ EG+ KAV
Sbjct: 404 KLVVQVLKAGVSAGVEEVMKWGEEEKIGVLVDKEGVKKAV 443
>At1g69570 H-protein promoter binding factor-2b, putative
Length = 399
Score = 33.9 bits (76), Expect = 0.16
Identities = 31/128 (24%), Positives = 55/128 (42%), Gaps = 8/128 (6%)
Query: 83 PISNQPKKSPKSSSPFLRGFMNSDTRSPLKFLNQLGSPKTVKKSTGKENKVQVNGV--RT 140
PI KKS + S+ F ++ SD P K ++++ SP++ K + +++++ T
Sbjct: 58 PIRVPVKKSEQESNKFKDPYILSDLNEPPKAVSEISSPRSSKNNCDQQSEITTTTTTSTT 117
Query: 141 GGVRRLDYSNSPKGILKPSNLSPNA------SKNSGIPAKGSPICARRKSFGGNEKDTLQ 194
G + K I P S N + N P C R + GG+ ++
Sbjct: 118 SGEKSTALKKPDKLIPCPRCESANTKFCYYNNYNVNQPRYFCRNCQRYWTAGGSMRNVPV 177
Query: 195 CSSRRKSQ 202
S RRK++
Sbjct: 178 GSGRRKNK 185
>At4g28730 unknown protein
Length = 174
Score = 33.5 bits (75), Expect = 0.21
Identities = 31/94 (32%), Positives = 47/94 (49%), Gaps = 12/94 (12%)
Query: 260 ENSVVIYTTTLRGIRKTFEDCNKVRAIIESYCVQ--MRERDVSMDSG--FKEELRKLMGA 315
EN+VVIY+ T C +V+ + + VQ + E D G ++ L +L G
Sbjct: 79 ENTVVIYSKTWCSY------CTEVKTLFKRLGVQPLVVELDQLGPQGPQLQKVLERLTG- 131
Query: 316 EQVKVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL 349
Q VP VFV G+ +GG + VKL + L ++L
Sbjct: 132 -QHTVPNVFVCGKHIGGCTDTVKLNRKGDLELML 164
>At2g36780 putative glucosyl transferase
Length = 496
Score = 33.5 bits (75), Expect = 0.21
Identities = 18/40 (45%), Positives = 28/40 (70%), Gaps = 2/40 (5%)
Query: 319 KVPVVFVKGRLVGGVDEVVKLEDEEKLGVLL--EGIPKAV 356
K+ V +K + GV+EV+K +E+K+GVL+ EG+ KAV
Sbjct: 404 KLVVQVLKAGVSAGVEEVMKWGEEDKIGVLVDKEGVKKAV 443
>At5g63030 glutaredoxin-like protein
Length = 125
Score = 33.1 bits (74), Expect = 0.28
Identities = 27/90 (30%), Positives = 42/90 (46%), Gaps = 11/90 (12%)
Query: 263 VVIYTTTLRGIRKTFEDCNKVRAIIESYCVQMR--ERDVSMDSG-FKEELRKLMGAEQVK 319
VV+++ T G C +V+ ++ + E D D G + L + G Q
Sbjct: 31 VVVFSKTYCGY------CQRVKQLLTQLGATFKVLELDEMSDGGEIQSALSEWTG--QTT 82
Query: 320 VPVVFVKGRLVGGVDEVVKLEDEEKLGVLL 349
VP VF+KG +GG D V++ + KL LL
Sbjct: 83 VPNVFIKGNHIGGCDRVMETNKQGKLVPLL 112
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.134 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,091,213
Number of Sequences: 26719
Number of extensions: 481072
Number of successful extensions: 1428
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 1335
Number of HSP's gapped (non-prelim): 90
length of query: 405
length of database: 11,318,596
effective HSP length: 102
effective length of query: 303
effective length of database: 8,593,258
effective search space: 2603757174
effective search space used: 2603757174
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0010.16


