

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0009.6
(309 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
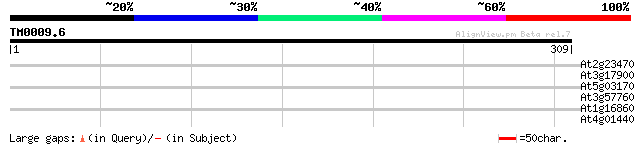
At2g23470 hypothetical protein 30 2.2
At3g17900 unknown protein 29 2.8
At5g03170 fasciclin-like arabinogalactan protein FLA11 28 6.3
At3g57760 unknown protein 28 6.3
At1g16860 unknown protein 28 6.3
At4g01440 predicted protein of unknown function 28 8.2
>At2g23470 hypothetical protein
Length = 405
Score = 29.6 bits (65), Expect = 2.2
Identities = 20/76 (26%), Positives = 34/76 (44%), Gaps = 1/76 (1%)
Query: 92 LPPGCQSSRSGTVLLRSRCRVDPYLVPLKHTGFAFRRRSTTSAEPFCCKEGARVKNVNRT 151
L P Q + ++S C D Y + ++ + FRR S C +EGA +V +
Sbjct: 267 LDPKAQIPTLSMMAMQSLCSDDGYFITMELSSQGFRR-IPKSGIVICLREGANSVDVITS 325
Query: 152 TINHALITRSVFINQT 167
+ I +S+ N+T
Sbjct: 326 LLQTCYIRKSLGANRT 341
>At3g17900 unknown protein
Length = 838
Score = 29.3 bits (64), Expect = 2.8
Identities = 53/186 (28%), Positives = 69/186 (36%), Gaps = 39/186 (20%)
Query: 103 TVLLRSRCRVDPYLVPLKHTGFAFRRRSTTSAEPFCCKEG--ARVKNVNRTTINHALITR 160
TVL + P +V L T TT PF RV+N R T
Sbjct: 686 TVLAPASFTSPPTVVSLNST-------PTTPISPFLGFSDFTERVQNEKRNT-------- 730
Query: 161 SVFINQTLVPVSLELK-DTVVRAYRTDPVQRFLYQRVPKNESHIFRPWIYKREEGKDDTV 219
+V Q+L P+ LE + + ++P VPK+ W+ R
Sbjct: 731 TVRKQQSLPPIPLETRTENNTNGESSNPSDV-----VPKSGLGCTHLWLQSRV------- 778
Query: 220 ANPCLCVQCKSFEVIKLELLILKCYTHSLIPIQDCSIHGPPK---QFPENFLLKQPIKST 276
P CV KS IKLELL L T +I + IH K PE LK S+
Sbjct: 779 --PLGCVPSKSTATIKLELLPL---TDGIITLDTLQIHAKEKGRRYIPEQ-SLKINATSS 832
Query: 277 ESEGMF 282
S G+F
Sbjct: 833 ISSGIF 838
>At5g03170 fasciclin-like arabinogalactan protein FLA11
Length = 246
Score = 28.1 bits (61), Expect = 6.3
Identities = 20/63 (31%), Positives = 29/63 (45%), Gaps = 4/63 (6%)
Query: 1 MVKSTPRIDQRMKKIHNSKDKSNPTILGRFLPTSGGFSSTLGGSFTQISGFVLVFLLGHH 60
++KST DQ I+ + S+ L F PT F+S G+ +S V L+ H
Sbjct: 53 LLKSTQASDQ----INTQLNSSSSNGLTVFAPTDNAFNSLKSGTLNSLSDQQKVQLVQFH 108
Query: 61 HLP 63
LP
Sbjct: 109 VLP 111
>At3g57760 unknown protein
Length = 378
Score = 28.1 bits (61), Expect = 6.3
Identities = 13/32 (40%), Positives = 17/32 (52%)
Query: 265 ENFLLKQPIKSTESEGMFREVCRLDAYYRLYK 296
+NF Q IK+T + R+D YYR YK
Sbjct: 41 KNFSYDQIIKATNNFCQSNRASRIDVYYRCYK 72
>At1g16860 unknown protein
Length = 474
Score = 28.1 bits (61), Expect = 6.3
Identities = 18/62 (29%), Positives = 28/62 (45%), Gaps = 5/62 (8%)
Query: 32 PTSGGFSSTLGGSFTQISGFVLVFLLGHHHLPLTAATLSSTAIPVAAAATLASSSLFRRH 91
P SGG + GS + L+ P+T+ L+S+ P + L SS L + H
Sbjct: 127 PQSGGVTRQNSGSIPILPATGLIT-----SGPITSGPLNSSGAPRKVSGPLDSSGLMKSH 181
Query: 92 LP 93
+P
Sbjct: 182 MP 183
>At4g01440 predicted protein of unknown function
Length = 365
Score = 27.7 bits (60), Expect = 8.2
Identities = 20/53 (37%), Positives = 27/53 (50%), Gaps = 8/53 (15%)
Query: 37 FSSTLGGSFTQISGFVLVFLLGHHHLPLTAATLSSTAIPVAAAATLASSSLFR 89
FS+ +G S TQ FLLG L T+ATL+ I + A T + +FR
Sbjct: 78 FSALVGASLTQY-----FFLLG---LSYTSATLACAFISMTPAITFVMALIFR 122
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,993,505
Number of Sequences: 26719
Number of extensions: 289049
Number of successful extensions: 642
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 640
Number of HSP's gapped (non-prelim): 6
length of query: 309
length of database: 11,318,596
effective HSP length: 99
effective length of query: 210
effective length of database: 8,673,415
effective search space: 1821417150
effective search space used: 1821417150
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0009.6


