

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0007.11
(264 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
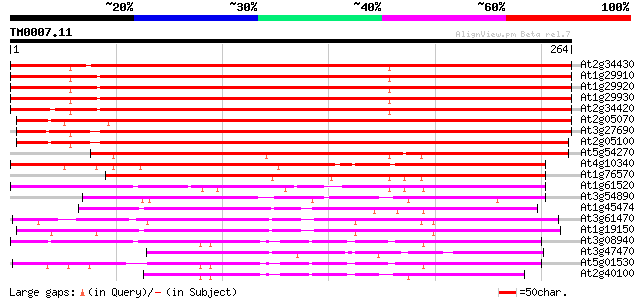
Score E
Sequences producing significant alignments: (bits) Value
At2g34430 putative photosystem II type I chlorophyll a/b binding... 476 e-135
At1g29910 chlorophyll a/b-binding protein 472 e-134
At1g29920 chlorophyll a/b-binding protein 472 e-134
At1g29930 unknown protein 470 e-133
At2g34420 photosystem II type I chlorophyll a/b binding protein 463 e-131
At2g05070 putative chlorophyll a/b binding protein 419 e-118
At3g27690 putative chlorophyll A-B binding protein 416 e-117
At2g05100 putative chlorophyll a/b binding protein 416 e-117
At5g54270 Lhcb3 chlorophyll a/b binding protein (gb|AAD28773.1) 348 1e-96
At4g10340 chlorophyll a/b-binding protein - like 215 2e-56
At1g76570 putative chlorophyll A-B binding protein 194 3e-50
At1g61520 PSI type III chlorophyll a/b-binding protein 140 9e-34
At3g54890 chlorophyll a/b-binding protein 138 3e-33
At1g45474 light-harvesting complex protein 135 2e-32
At3g61470 Lhca2 protein 135 3e-32
At1g19150 unknown protein 121 3e-28
At3g08940 Lhcb4-protein (CP29, Lhcb4.2) 120 1e-27
At3g47470 light-harvesting chlorophyll a/b-binding protein (Cab4) 118 3e-27
At5g01530 chlorophyll a/b-binding protein CP29 117 8e-27
At2g40100 putative chlorophyll a/b binding protein 97 1e-20
>At2g34430 putative photosystem II type I chlorophyll a/b binding
protein.
Length = 266
Score = 476 bits (1225), Expect = e-135
Identities = 237/268 (88%), Positives = 249/268 (92%), Gaps = 6/268 (2%)
Query: 1 MAASTMALSSPSLAGQALNLSPSSQEV---GRFSMRKAATTKKVVSSGSPWYGPDRVKYL 57
MAASTMALSSP+L G+A+ LSP++ EV GR +MRKA+ K SGSPWYG DRVKYL
Sbjct: 1 MAASTMALSSPALTGKAVKLSPAASEVFGTGRITMRKAS--KPTGPSGSPWYGSDRVKYL 58
Query: 58 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELL 117
GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFA+NRELEVIHSRWAMLGALGCVFPELL
Sbjct: 59 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 118
Query: 118 ARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAG 177
ARNGVKFGEAVWFKAGSQIFS+GGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYR+AG
Sbjct: 119 ARNGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVAG 178
Query: 178 -GPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 236
GPLGE D LYPGGSFDPLGLA DPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG
Sbjct: 179 DGPLGEAEDLLYPGGSFDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 238
Query: 237 PLENLADHLSDPVNNNAWAYATNFVPGK 264
PLENLADHL+DPVNNNAWA+ATNFVPGK
Sbjct: 239 PLENLADHLADPVNNNAWAFATNFVPGK 266
>At1g29910 chlorophyll a/b-binding protein
Length = 267
Score = 472 bits (1215), Expect = e-134
Identities = 235/268 (87%), Positives = 247/268 (91%), Gaps = 5/268 (1%)
Query: 1 MAASTMALSSPSLAGQALNLSPSSQEV---GRFSMRKAATTKKVVSSGSPWYGPDRVKYL 57
MAASTMALSSP+ AG+A+NLSP++ EV GR +MRK K SGSPWYG DRVKYL
Sbjct: 1 MAASTMALSSPAFAGKAVNLSPAASEVLGSGRVTMRKTVAKPKG-PSGSPWYGSDRVKYL 59
Query: 58 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELL 117
GPFSGE PSYLTGEFPGDYGWDTAGLSADPETFA+NRELEVIHSRWAMLGALGCVFPELL
Sbjct: 60 GPFSGESPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 119
Query: 118 ARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAG 177
ARNGVKFGEAVWFKAGSQIFS+GGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYR+AG
Sbjct: 120 ARNGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVAG 179
Query: 178 -GPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 236
GPLGE D LYPGGSFDPLGLA DPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG
Sbjct: 180 NGPLGEAEDLLYPGGSFDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 239
Query: 237 PLENLADHLSDPVNNNAWAYATNFVPGK 264
P+ENLADHL+DPVNNNAWA+ATNFVPGK
Sbjct: 240 PIENLADHLADPVNNNAWAFATNFVPGK 267
>At1g29920 chlorophyll a/b-binding protein
Length = 267
Score = 472 bits (1215), Expect = e-134
Identities = 235/268 (87%), Positives = 247/268 (91%), Gaps = 5/268 (1%)
Query: 1 MAASTMALSSPSLAGQALNLSPSSQEV---GRFSMRKAATTKKVVSSGSPWYGPDRVKYL 57
MAASTMALSSP+ AG+A+NLSP++ EV GR +MRK K SGSPWYG DRVKYL
Sbjct: 1 MAASTMALSSPAFAGKAVNLSPAASEVLGSGRVTMRKTVAKPKG-PSGSPWYGSDRVKYL 59
Query: 58 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELL 117
GPFSGE PSYLTGEFPGDYGWDTAGLSADPETFA+NRELEVIHSRWAMLGALGCVFPELL
Sbjct: 60 GPFSGESPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 119
Query: 118 ARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAG 177
ARNGVKFGEAVWFKAGSQIFS+GGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYR+AG
Sbjct: 120 ARNGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVAG 179
Query: 178 -GPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 236
GPLGE D LYPGGSFDPLGLA DPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG
Sbjct: 180 NGPLGEAEDLLYPGGSFDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 239
Query: 237 PLENLADHLSDPVNNNAWAYATNFVPGK 264
P+ENLADHL+DPVNNNAWA+ATNFVPGK
Sbjct: 240 PIENLADHLADPVNNNAWAFATNFVPGK 267
>At1g29930 unknown protein
Length = 267
Score = 470 bits (1209), Expect = e-133
Identities = 234/268 (87%), Positives = 246/268 (91%), Gaps = 5/268 (1%)
Query: 1 MAASTMALSSPSLAGQALNLSPSSQEV---GRFSMRKAATTKKVVSSGSPWYGPDRVKYL 57
MAASTMALSSP+ AG+A+ LSP++ EV GR +MRK K SGSPWYG DRVKYL
Sbjct: 1 MAASTMALSSPAFAGKAVKLSPAASEVLGSGRVTMRKTVAKPKG-PSGSPWYGSDRVKYL 59
Query: 58 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELL 117
GPFSGE PSYLTGEFPGDYGWDTAGLSADPETFA+NRELEVIHSRWAMLGALGCVFPELL
Sbjct: 60 GPFSGESPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 119
Query: 118 ARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAG 177
ARNGVKFGEAVWFKAGSQIFS+GGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYR+AG
Sbjct: 120 ARNGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVAG 179
Query: 178 -GPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 236
GPLGE D LYPGGSFDPLGLA DPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG
Sbjct: 180 NGPLGEAEDLLYPGGSFDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 239
Query: 237 PLENLADHLSDPVNNNAWAYATNFVPGK 264
P+ENLADHL+DPVNNNAWA+ATNFVPGK
Sbjct: 240 PIENLADHLADPVNNNAWAFATNFVPGK 267
>At2g34420 photosystem II type I chlorophyll a/b binding protein
Length = 265
Score = 463 bits (1192), Expect = e-131
Identities = 232/268 (86%), Positives = 245/268 (90%), Gaps = 7/268 (2%)
Query: 1 MAASTMALSSPSLAGQALNLSPSSQEV---GRFSMRKAATTKKVVSSGSPWYGPDRVKYL 57
MA+STMALSSP+ AG+A+ P++ +V GR +MRK K SGSPWYG DRVKYL
Sbjct: 1 MASSTMALSSPAFAGKAVK--PAASDVLGSGRVTMRKTVAKPKG-PSGSPWYGSDRVKYL 57
Query: 58 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELL 117
GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFA+NRELEVIHSRWAMLGALGCVFPELL
Sbjct: 58 GPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFARNRELEVIHSRWAMLGALGCVFPELL 117
Query: 118 ARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAG 177
ARNGVKFGEAVWFKAGSQIFS+GGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYR+AG
Sbjct: 118 ARNGVKFGEAVWFKAGSQIFSDGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRVAG 177
Query: 178 -GPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 236
GPLGE D LYPGGSFDPLGLA DPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG
Sbjct: 178 DGPLGEAEDLLYPGGSFDPLGLATDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKG 237
Query: 237 PLENLADHLSDPVNNNAWAYATNFVPGK 264
PLENLADHL+DPVNNNAWA+ATNFVPGK
Sbjct: 238 PLENLADHLADPVNNNAWAFATNFVPGK 265
>At2g05070 putative chlorophyll a/b binding protein
Length = 265
Score = 419 bits (1077), Expect = e-118
Identities = 206/265 (77%), Positives = 226/265 (84%), Gaps = 5/265 (1%)
Query: 4 STMALSSPSLAGQALNLSPSS---QEVGRFSMRKAATTKKVVSSG-SPWYGPDRVKYLGP 59
+T A+ S AGQ L PSS Q+VG + + V S+ S WYGPDR KYLGP
Sbjct: 2 ATSAIQQSSFAGQTA-LKPSSDLIQKVGVLGGGRVTMRRTVKSTPQSIWYGPDRPKYLGP 60
Query: 60 FSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELLAR 119
FS PSYLTGE+PGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGC FPE+L++
Sbjct: 61 FSENTPSYLTGEYPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCTFPEILSK 120
Query: 120 NGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAGGP 179
NGVKFGEAVWFKAGSQIFSEGGLDYLGNP+L+HAQSILAIWA QV+LMG +EGYRI GGP
Sbjct: 121 NGVKFGEAVWFKAGSQIFSEGGLDYLGNPNLIHAQSILAIWAVQVVLMGFIEGYRIGGGP 180
Query: 180 LGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKGPLE 239
LGE DPLYPGG+FDPL LA+DPEAF+ELKVKELKNGRLAMFSMFGFFVQAIVTGKGP+E
Sbjct: 181 LGEGLDPLYPGGAFDPLNLAEDPEAFSELKVKELKNGRLAMFSMFGFFVQAIVTGKGPIE 240
Query: 240 NLADHLSDPVNNNAWAYATNFVPGK 264
NL DHL+DPV NNAW+YATNFVPGK
Sbjct: 241 NLFDHLADPVANNAWSYATNFVPGK 265
>At3g27690 putative chlorophyll A-B binding protein
Length = 266
Score = 416 bits (1068), Expect = e-117
Identities = 204/270 (75%), Positives = 225/270 (82%), Gaps = 14/270 (5%)
Query: 4 STMALSSPSLAGQALNLSPSSQEV---------GRFSMRKAATTKKVVSSGSPWYGPDRV 54
+T A+ S AGQ L PS+ + GR MR+ + + S WYGPDR
Sbjct: 2 ATSAIQHSSFAGQT-TLKPSNDLLRKIGASNGGGRIIMRRTVKS----TPQSIWYGPDRP 56
Query: 55 KYLGPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFP 114
KYLGPFS PSYLTGE+PGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGC FP
Sbjct: 57 KYLGPFSENTPSYLTGEYPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCTFP 116
Query: 115 ELLARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYR 174
E+L++NGVKFGEAVWFKAGSQIFSEGGLDYLGNP+L+HAQSILAIWA QV+LMG +EGYR
Sbjct: 117 EILSKNGVKFGEAVWFKAGSQIFSEGGLDYLGNPNLIHAQSILAIWACQVVLMGFIEGYR 176
Query: 175 IAGGPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTG 234
I GGPLGE DPLYPGG+FDPL LA+DPEAF+ELKVKELKNGRLAMFSMFGFFVQAIVTG
Sbjct: 177 IGGGPLGEGLDPLYPGGAFDPLNLAEDPEAFSELKVKELKNGRLAMFSMFGFFVQAIVTG 236
Query: 235 KGPLENLADHLSDPVNNNAWAYATNFVPGK 264
KGP+ENL DH++DPV NNAWAYATNFVPGK
Sbjct: 237 KGPIENLFDHIADPVANNAWAYATNFVPGK 266
>At2g05100 putative chlorophyll a/b binding protein
Length = 265
Score = 416 bits (1068), Expect = e-117
Identities = 203/268 (75%), Positives = 226/268 (83%), Gaps = 13/268 (4%)
Query: 4 STMALSSPSLAGQALNLSPSSQEV--------GRFSMRKAATTKKVVSSGSPWYGPDRVK 55
+T A+ S AGQ L PS++ + GR +MR+ + + S WYGPDR K
Sbjct: 2 ATSAIQQSSFAGQTA-LKPSNELLRKVGVSGGGRVTMRRTVKS----TPQSIWYGPDRPK 56
Query: 56 YLGPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPE 115
YLGPFS PSYLTGE+PGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGC FPE
Sbjct: 57 YLGPFSENTPSYLTGEYPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCTFPE 116
Query: 116 LLARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRI 175
+L++NGVKFGEAVWFKAGSQIFSEGGLDYLGNP+L+HAQSILAIWA QV+LMG +EGYRI
Sbjct: 117 ILSKNGVKFGEAVWFKAGSQIFSEGGLDYLGNPNLIHAQSILAIWAVQVVLMGFIEGYRI 176
Query: 176 AGGPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGK 235
GGPLGE DPLYPGG+FDPL LA+DPEAF+ELKVKELKNGRLAMFSMFGFFVQAIVTGK
Sbjct: 177 GGGPLGEGLDPLYPGGAFDPLNLAEDPEAFSELKVKELKNGRLAMFSMFGFFVQAIVTGK 236
Query: 236 GPLENLADHLSDPVNNNAWAYATNFVPG 263
GP+ENL DHL+DPV NNAW+YATNFVPG
Sbjct: 237 GPIENLFDHLADPVANNAWSYATNFVPG 264
>At5g54270 Lhcb3 chlorophyll a/b binding protein (gb|AAD28773.1)
Length = 265
Score = 348 bits (894), Expect = 1e-96
Identities = 180/236 (76%), Positives = 192/236 (81%), Gaps = 12/236 (5%)
Query: 39 KKVVSSGSP--------WYGPDRVKYLGPFSGEPPSYLTGEFPGDYGWDTAGLSADPETF 90
+ VVS GSP WYGPDRVKYLGPFS + PSYLTGEFPGDYGWDTAGLSADPE F
Sbjct: 30 RDVVSLGSPKYTMGNDLWYGPDRVKYLGPFSVQTPSYLTGEFPGDYGWDTAGLSADPEAF 89
Query: 91 AKNRELEVIHSRWAMLGALGCVFPELLAR-NGVKFGEAVWFKAGSQIFSEGGLDYLGNPS 149
AKNR LEVIH RWAMLGA GC+ PE+L + V F E VWFKAGSQIFSEGGLDYLGNP+
Sbjct: 90 AKNRALEVIHGRWAMLGAFGCITPEVLQKWVRVDFKEPVWFKAGSQIFSEGGLDYLGNPN 149
Query: 150 LVHAQSILAIWATQVILMGAVEGYRIAG-GPLGEVTDPLYPGGS-FDPLGLADDPEAFAE 207
LVHAQSILA+ QVILMG VEG+RI G +GE D LYPGG FDPLGLADDP FAE
Sbjct: 150 LVHAQSILAVLGFQVILMGLVEGFRINGLDGVGEGND-LYPGGQYFDPLGLADDPVTFAE 208
Query: 208 LKVKELKNGRLAMFSMFGFFVQAIVTGKGPLENLADHLSDPVNNNAWAYATNFVPG 263
LKVKE+KNGRLAMFSMFGFFVQAIVTGKGPLENL DHL +PV NNAWA+AT F PG
Sbjct: 209 LKVKEIKNGRLAMFSMFGFFVQAIVTGKGPLENLLDHLDNPVANNAWAFATKFAPG 264
>At4g10340 chlorophyll a/b-binding protein - like
Length = 280
Score = 215 bits (548), Expect = 2e-56
Identities = 129/262 (49%), Positives = 160/262 (60%), Gaps = 15/262 (5%)
Query: 1 MAASTMALSSPSLAGQALNLSPSS-QEVGRFSMRKAATTK-KVVSSGSP----WYGPDRV 54
M + + + S + L SPS+ + V FS +K A K K VS S WYGPDR
Sbjct: 9 MLGTPLNFRAVSRSSAPLASSPSTFKTVALFSKKKPAPAKSKAVSETSDELAKWYGPDRR 68
Query: 55 KYLGPF---SGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGC 111
+L E P YL GE GDYG+D GL PE FAK + E+IH+RWAMLGA G
Sbjct: 69 IFLPDGLLDRSEIPEYLNGEVAGDYGYDPFGLGKKPENFAKYQAFELIHARWAMLGAAGF 128
Query: 112 VFPELLARNGVKFG-EAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAV 170
+ PE L + G G EAVWFK G+ + L+Y G ++ +LA+ A +V+L+G
Sbjct: 129 IIPEALNKYGANCGPEAVWFKTGALLLDGNTLNYFGKNIPINL--VLAVVA-EVVLLGGA 185
Query: 171 EGYRIAGGPLGEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQA 230
E YRI G + D L+PGG FDPLGLA DPE A LKVKE+KNGRLAMF+M GFF+QA
Sbjct: 186 EYYRITNGL--DFEDKLHPGGPFDPLGLAKDPEQGALLKVKEIKNGRLAMFAMLGFFIQA 243
Query: 231 IVTGKGPLENLADHLSDPVNNN 252
VTG+GP+ENLA HLSDP NN
Sbjct: 244 YVTGEGPVENLAKHLSDPFGNN 265
>At1g76570 putative chlorophyll A-B binding protein
Length = 327
Score = 194 bits (494), Expect = 3e-50
Identities = 106/217 (48%), Positives = 135/217 (61%), Gaps = 10/217 (4%)
Query: 46 SPWYGPDRVKYLGPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAM 105
S WYG +R ++ GP + P YLTGE PGDYG+D AGL D TF K E++H+RWAM
Sbjct: 104 SKWYGKERPRWFGPIPYDYPPYLTGELPGDYGFDIAGLGKDRLTFDKYFNFEILHARWAM 163
Query: 106 LGALGCVFPELLARNGV-KFGEAVWFKAGSQIFSEGGLDYLGNPSL--VHAQSILAIWAT 162
L ALG + PE+ G F E VW++ G L+YLG P L +Q ++ I
Sbjct: 164 LAALGALIPEVFDLTGTFHFAEPVWWRVGYSKLQGETLEYLGIPGLHVAGSQGVIVIAIC 223
Query: 163 QVILMGAVEGYRIAG----GPLGEVT--DPLYPGGS-FDPLGLADDPEAFAELKVKELKN 215
QV+LM E R G PLG D YPGG+ FDPL L++DP AF +LKVKE+KN
Sbjct: 224 QVLLMVGPEYARYCGIEALEPLGIYLPGDINYPGGTLFDPLNLSEDPVAFEDLKVKEIKN 283
Query: 216 GRLAMFSMFGFFVQAIVTGKGPLENLADHLSDPVNNN 252
GRLAM + GF+ QA TGKGP++NL DH+SDP++NN
Sbjct: 284 GRLAMVAWLGFYAQAAFTGKGPVQNLVDHVSDPLHNN 320
>At1g61520 PSI type III chlorophyll a/b-binding protein
Length = 273
Score = 140 bits (352), Expect = 9e-34
Identities = 101/277 (36%), Positives = 144/277 (51%), Gaps = 37/277 (13%)
Query: 1 MAASTMALSSPSLAGQALNLSPSSQEVGRFSMRKAATTKKVVSSGSPWYGPDRVKYLGPF 60
MAA + SS + + Q S+ V S +K++ K ++ G +R +
Sbjct: 1 MAAQALVSSSLTSSVQTARQIFGSKPVASASQKKSSFVVKAAATPPVKQGANRPLWFA-- 58
Query: 61 SGEPPSYLTGEFPGDYGWDTAGLSADPET---FAKNREL---EVIHSRWAMLGALGCVFP 114
S + SYL G PGDYG+D GLS DPE F + R L E+I+ R+AMLGA G + P
Sbjct: 59 SSQSLSYLDGSLPGDYGFDPLGLS-DPEGTGGFIEPRWLAYGEIINGRFAMLGAAGAIAP 117
Query: 115 ELLARNGVKFGEAV--WFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEG 172
E+L + G+ E WF+ G I G Y + ++ ++ LMG E
Sbjct: 118 EILGKAGLIPAETALPWFQTGV-IPPAGTYTYWADN--------YTLFVLEMALMGFAEH 168
Query: 173 YRIAG----GPLGEVT------------DPLYPGGSF-DPLGLADDPEAFAELKVKELKN 215
R+ G +G+ +P YPGG F +PLG D ++ ELK+KE+KN
Sbjct: 169 RRLQDWYNPGSMGKQYFLGLEKGLAGSGNPAYPGGPFFNPLGFGKDEKSLKELKLKEVKN 228
Query: 216 GRLAMFSMFGFFVQAIVTGKGPLENLADHLSDPVNNN 252
GRLAM ++ G+F+Q +VTG GP +NL DHL+DPVNNN
Sbjct: 229 GRLAMLAILGYFIQGLVTGVGPYQNLLDHLADPVNNN 265
>At3g54890 chlorophyll a/b-binding protein
Length = 241
Score = 138 bits (347), Expect = 3e-33
Identities = 94/228 (41%), Positives = 121/228 (52%), Gaps = 25/228 (10%)
Query: 35 AATTKKVVSSGSPWYGPDRVKYLGPFS----GEP-PSYLTGEFPGDYGWDTAGLSADPET 89
+++ K VS+G P V + + GEP P+YL G PGD+G+D GL P
Sbjct: 20 SSSKSKFVSAGVPLPNAGNVGRIRMAAHWMPGEPRPAYLDGSAPGDFGFDPLGLGEVPAN 79
Query: 90 FAKNRELEVIHSRWAMLGALGCVFPELLARNGVKFGEAVWFKAGSQIFSEGG-LDYLGNP 148
+ +E E+IH RWAML G + PE L G W KA GG YLGNP
Sbjct: 80 LERYKESELIHCRWAMLAVPGILVPEAL-------GYGNWVKAQEWAALPGGQATYLGNP 132
Query: 149 SLVHAQSILAIWATQVILMGAVEGYRIAGGPLGEVTDP---LYPGGSFDPLGLADDPEAF 205
V ++ I A + + + VE R DP YPGG+FDPLG + DP+
Sbjct: 133 --VPWGTLPTILAIEFLAIAFVEHQR------SMEKDPEKKKYPGGAFDPLGYSKDPKKL 184
Query: 206 AELKVKELKNGRLAMFSMFGFFV-QAIVTGKGPLENLADHLSDPVNNN 252
ELKVKE+KNGRLA+ + GF V Q+ G GPLENLA HL+DP +NN
Sbjct: 185 EELKVKEIKNGRLALLAFVGFCVQQSAYPGTGPLENLATHLADPWHNN 232
>At1g45474 light-harvesting complex protein
Length = 256
Score = 135 bits (340), Expect = 2e-32
Identities = 87/225 (38%), Positives = 123/225 (54%), Gaps = 17/225 (7%)
Query: 33 RKAATTKKVVSSGSPWYGPDRVKYLGPFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAK 92
R+ ++ K +P +R +L + PP YL G GDYG+D GL DPE+
Sbjct: 26 RRISSVKAAGGGINPTVAVERATWLPGLN--PPPYLDGNLAGDYGFDPLGLGEDPESLKW 83
Query: 93 NRELEVIHSRWAMLGALGCVFPELLARNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVH 152
+ E++HSR+AMLG G +F +LL G++ VW++AG+ F D+ +L+
Sbjct: 84 YVQAELVHSRFAMLGVAGILFTDLLRTTGIR-NLPVWYEAGAVKF-----DFASTKTLIV 137
Query: 153 AQSILAIWATQVILMGAV-------EGYRIAGGPLG-EVTDPLYPGGSF-DPLGLADDPE 203
Q +L +A M V EG G E +P YPGG +PLGLA D +
Sbjct: 138 VQFLLMGFAETKRYMDFVSPGSQAKEGSFFFGLEAALEGLEPGYPGGPLLNPLGLAKDVQ 197
Query: 204 AFAELKVKELKNGRLAMFSMFGFFVQAIVTGKGPLENLADHLSDP 248
+ K+KE+KNGRLAM +M GFFVQA VT GP++NL +HLS+P
Sbjct: 198 NAHDWKLKEIKNGRLAMMAMLGFFVQASVTHTGPIDNLVEHLSNP 242
>At3g61470 Lhca2 protein
Length = 257
Score = 135 bits (339), Expect = 3e-32
Identities = 95/266 (35%), Positives = 136/266 (50%), Gaps = 27/266 (10%)
Query: 2 AASTMALSSPS-LAGQALNLSPSSQEVGRFSMRKAATTKKVVSSGSPWYGPDRVKYLGPF 60
+++ A+SSPS L G+ L L + A +++ S + PDR + F
Sbjct: 8 SSAIAAISSPSFLGGKKLRLKKK--------LTVPAVSRQDASVRAVAADPDRPIW---F 56
Query: 61 SGE-PPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELLAR 119
G PP +L G PGD+G+D GLS+DP++ N + E++H RWAMLGA G PE L +
Sbjct: 57 PGSTPPEWLDGSLPGDFGFDPLGLSSDPDSLKWNVQAEIVHCRWAMLGAAGIFIPEFLTK 116
Query: 120 NGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWA-----TQVILMGAVEGYR 174
G+ W+ AG Q + + +L + IL WA +I G+V
Sbjct: 117 IGI-LNTPSWYTAGEQEY------FTDKTTLFVVELILIGWAEGRRWADIIKPGSVNTDP 169
Query: 175 IAGGPLGEVTDPLYPGGS-FDPLGL-ADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIV 232
+ TD YPGG FDPLG + P EL+ KE+KNGRLAM ++ G + Q I
Sbjct: 170 VFPNNKLTGTDVGYPGGLWFDPLGWGSGSPAKLKELRTKEIKNGRLAMLAVMGAWFQHIY 229
Query: 233 TGKGPLENLADHLSDPVNNNAWAYAT 258
TG GP++NL HL+DP + +A T
Sbjct: 230 TGTGPIDNLFAHLADPGHATIFAAFT 255
>At1g19150 unknown protein
Length = 270
Score = 121 bits (304), Expect = 3e-28
Identities = 89/268 (33%), Positives = 130/268 (48%), Gaps = 21/268 (7%)
Query: 4 STMALSSPSLAGQAL---NLSPSSQEVGRFSMRKAATTK--KVVSSGSPWYGPDRVKYLG 58
ST+ LS+ + +++ +S GR + + K VSS PDR +
Sbjct: 11 STLTLSTSRVQNPTQRRPHVASTSSTGGRLMRERLVVVRAGKEVSSVCEPLPPDRPLWFP 70
Query: 59 PFSGEPPSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELLA 118
S PP +L G PGD+G+D GL +DP+T + E+IHSRWAML G + PE L
Sbjct: 71 GSS--PPEWLDGSLPGDFGFDPLGLGSDPDTLKWFAQAELIHSRWAMLAVTGIIIPECLE 128
Query: 119 RNGVKFGEAVWFKAGSQIFSEGGLDYLGNPSLVHAQSILAIWA-----TQVILMGAVEGY 173
R G W+ AGS+ + + + +L AQ +L WA +I G+V+
Sbjct: 129 RLGF-IENFSWYDAGSREY------FADSTTLFVAQMVLMGWAEGRRWADLIKPGSVDIE 181
Query: 174 RIAGGPLGEVTDPLYPGGSFDPLGL--ADDPEAFAELKVKELKNGRLAMFSMFGFFVQAI 231
+ D YPGG + + PE L+ KE+KNGRLAM + GF QA
Sbjct: 182 PKYPHKVNPKPDVGYPGGLWFDFMMWGRGSPEPVMVLRTKEIKNGRLAMLAFLGFCFQAT 241
Query: 232 VTGKGPLENLADHLSDPVNNNAWAYATN 259
T + P+ENL HL+DP + N ++ T+
Sbjct: 242 YTSQDPIENLMAHLADPGHCNVFSAFTS 269
>At3g08940 Lhcb4-protein (CP29, Lhcb4.2)
Length = 287
Score = 120 bits (300), Expect = 1e-27
Identities = 101/294 (34%), Positives = 132/294 (44%), Gaps = 61/294 (20%)
Query: 1 MAASTMALSSPSLAGQALNLSPSSQEVGRFSMRKAATTKKVVSSGSPWYGPDRVKYLGPF 60
MAA++ A ++ S+ G + +S S RF+ R TKK + DR +
Sbjct: 1 MAATSTAAAASSIMGTRV-VSDISSNSSRFTARFGFGTKKASPKKAKTVISDRPLWFP-- 57
Query: 61 SGEPPSYLTGEFPGDYGWDTAGLSADPE-----------TFAKN---------------- 93
+ P YL G GDYG+D GL E AKN
Sbjct: 58 GAKSPEYLDGSLVGDYGFDPFGLGKPAEYLQFDLDSLDQNLAKNLYGEVIGTRTEAVDPK 117
Query: 94 ----------------RELEVIHSRWAMLGALGCVFPELLARNGVKFGEAVWFKAGSQIF 137
RE E+IH RWAML LG + E L GV W AG
Sbjct: 118 STPFQPYSEVFGLQRFRECELIHGRWAMLATLGAITVEWLT--GV-----TWQDAGKVEL 170
Query: 138 SEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAGGPLGEVTDPLYPGGSF-DPL 196
+G YLG P +++ I +V+++G +E R A + LYPGG F DPL
Sbjct: 171 VDGS-SYLGQPLPFSISTLIWI---EVLVIGYIEFQRNAEL---DSEKRLYPGGKFFDPL 223
Query: 197 GLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKGPLENLADHLSDPVN 250
GLA DP A+L++ E+K+ RLAM GF VQA TGKGPL N A HLSDP++
Sbjct: 224 GLASDPVKKAQLQLAEIKHARLAMVGFLGFAVQAAATGKGPLNNWATHLSDPLH 277
>At3g47470 light-harvesting chlorophyll a/b-binding protein (Cab4)
Length = 251
Score = 118 bits (296), Expect = 3e-27
Identities = 79/191 (41%), Positives = 100/191 (51%), Gaps = 14/191 (7%)
Query: 65 PSYLTGEFPGDYGWDTAGLSADPETFAKNRELEVIHSRWAMLGALGCVFPELLARNGVKF 124
P YLTG GD G+D GL+ DPE + E+++ RWAMLG G + PE+ + G+
Sbjct: 64 PDYLTGSLAGDNGFDPLGLAEDPENLKWFVQAELVNGRWAMLGVAGMLLPEVFTKIGI-I 122
Query: 125 GEAVWFKAGS-QIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEG---YRIAGGPL 180
W+ AG Q F+ ++ L H I W + G+V ++ P
Sbjct: 123 NVPEWYDAGKEQYFASSSTLFVIEFILFHYVEIRR-WQ-DIKNPGSVNQDPIFKQYSLPK 180
Query: 181 GEVTDPLYPGGSFDPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKGPLEN 240
GEV YPGG F+PL A EA K KEL NGRLAM + GF VQ VTGKGP EN
Sbjct: 181 GEVG---YPGGIFNPLNFAPTQEA----KEKELANGRLAMLAFLGFVVQHNVTGKGPFEN 233
Query: 241 LADHLSDPVNN 251
L HLSDP +N
Sbjct: 234 LLQHLSDPWHN 244
>At5g01530 chlorophyll a/b-binding protein CP29
Length = 290
Score = 117 bits (292), Expect = 8e-27
Identities = 103/301 (34%), Positives = 135/301 (44%), Gaps = 75/301 (24%)
Query: 2 AASTMALSSPSLAGQ--ALNLSPSSQE---VGRFSMRKAA---TTKKVVSSGSPWYGPDR 53
A S A ++ S+ G A + P S V F +KAA + KK V++ P + P
Sbjct: 3 ATSAAAAAASSIMGTRVAPGIHPGSGRFTAVFGFGKKKAAPKKSAKKTVTTDRPLWYPGA 62
Query: 54 VKYLGPFSGEPPSYLTGEFPGDYGWDTAGLSADPE-----------TFAKN--------- 93
+ P +L G GDYG+D GL E AKN
Sbjct: 63 IS---------PDWLDGSLVGDYGFDPFGLGKPAEYLQFDIDSLDQNLAKNLAGDVIGTR 113
Query: 94 -----------------------RELEVIHSRWAMLGALGCVFPELLARNGVKFGEAVWF 130
RE E+IH RWAML LG + E L GV W
Sbjct: 114 TEAADAKSTPFQPYSEVFGIQRFRECELIHGRWAMLATLGALSVEWLT--GV-----TWQ 166
Query: 131 KAGSQIFSEGGLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAGGPLGEVTDPLYPG 190
AG +G YLG P +++ I +V+++G +E R A + LYPG
Sbjct: 167 DAGKVELVDGS-SYLGQPLPFSISTLIWI---EVLVIGYIEFQRNAEL---DSEKRLYPG 219
Query: 191 GSF-DPLGLADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKGPLENLADHLSDPV 249
G F DPLGLA DPE A+L++ E+K+ RLAM + GF VQA TGKGPL N A HLSDP+
Sbjct: 220 GKFFDPLGLAADPEKTAQLQLAEIKHARLAMVAFLGFAVQAAATGKGPLNNWATHLSDPL 279
Query: 250 N 250
+
Sbjct: 280 H 280
>At2g40100 putative chlorophyll a/b binding protein
Length = 276
Score = 96.7 bits (239), Expect = 1e-20
Identities = 78/225 (34%), Positives = 98/225 (42%), Gaps = 63/225 (28%)
Query: 64 PPSYLTGEFPGDYGWDTAGLSADPE-----------TFAKN------------------- 93
PP +L G GD G+D GL E AKN
Sbjct: 65 PPEWLDGSMIGDRGFDPFGLGKPAEYLQYDFDGLDQNLAKNVAGDIIGIIQESSEIKPTP 124
Query: 94 -------------RELEVIHSRWAMLGALGCVFPELLARNGVKFGEAVWFKAGSQIFSEG 140
RE E+IH RWAMLG LG + E L G+ W AG EG
Sbjct: 125 FQPYTEVFGIQRFRECELIHGRWAMLGTLGAIAVEALT--GI-----AWQDAGKVELVEG 177
Query: 141 GLDYLGNPSLVHAQSILAIWATQVILMGAVEGYRIAGGPLGEVTDP---LYPGGSFDPLG 197
YLG P +++ I +V+++G +E R DP +YPGG FDPLG
Sbjct: 178 S-SYLGQPLPFSLTTLIWI---EVLVVGYIEFQR------NSELDPEKRIYPGGYFDPLG 227
Query: 198 LADDPEAFAELKVKELKNGRLAMFSMFGFFVQAIVTGKGPLENLA 242
LA DPE LK+ E+K+ RLAM + F +QA TGKGP+ LA
Sbjct: 228 LAADPEKLDTLKLAEIKHSRLAMVAFLIFALQAAFTGKGPVSFLA 272
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,398,999
Number of Sequences: 26719
Number of extensions: 282486
Number of successful extensions: 657
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 564
Number of HSP's gapped (non-prelim): 34
length of query: 264
length of database: 11,318,596
effective HSP length: 97
effective length of query: 167
effective length of database: 8,726,853
effective search space: 1457384451
effective search space used: 1457384451
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0007.11


