

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0007.1
(254 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
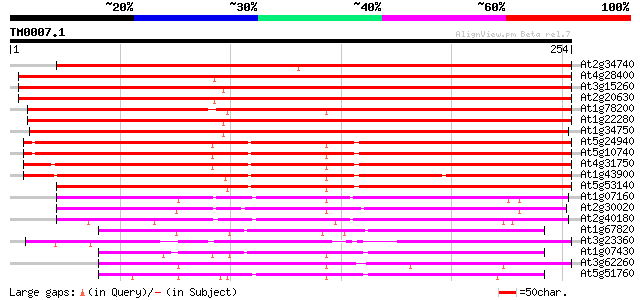
Score E
Sequences producing significant alignments: (bits) Value
At2g34740 putative protein phosphatase 2C 300 6e-82
At4g28400 protein phosphatase 2C like protein 298 2e-81
At3g15260 putative protein phosphatase type 2C 296 7e-81
At2g20630 putative protein phosphatase 2C 291 2e-79
At1g78200 putative protein phosphatase 2C 259 1e-69
At1g22280 protein phosphatase type 2C like protein 255 2e-68
At1g34750 protein phosphatase type 2C, putative 252 1e-67
At5g24940 Protein phosphatase 2C-like protein 201 3e-52
At5g10740 protein phosphatase 2C -like protein 199 2e-51
At4g31750 unknown protein 196 1e-50
At1g43900 protein phosphatase type 2C like protein 196 1e-50
At5g53140 protein phosphatase 2C-like 177 5e-45
At1g07160 unknown protein 143 1e-34
At2g30020 putative protein phosphatase 2C 138 2e-33
At2g40180 protein phosphatase 2C (AthPP2C5) 134 6e-32
At1g67820 protein phosphatase-like protein 126 1e-29
At3g23360 hypothetical protein 117 5e-27
At1g07430 protein phosphatase 2C, putative 114 4e-26
At3g62260 unknown protein 114 7e-26
At5g51760 protein phosphatase-2C; PP2C-like protein 112 1e-25
>At2g34740 putative protein phosphatase 2C
Length = 239
Score = 300 bits (767), Expect = 6e-82
Identities = 145/234 (61%), Positives = 188/234 (79%), Gaps = 1/234 (0%)
Query: 22 MEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNPVHAVKKA 81
MED++ A K + G+ LGLYAIFDGH+G DVA YLQ+HLF+NIL++PDFW+NP A+K+A
Sbjct: 1 MEDFIVADTKTVKGHNLGLYAIFDGHSGSDVADYLQNHLFDNILSQPDFWRNPKKAIKRA 60
Query: 82 CKATDEEILENIADSRGGSTAVAAILINGVKLLVFNVGDSRAISCKNG-VAKQLTEDHEP 140
K+TD+ IL+N+ RGGSTAV AI+I+G K++V NVGDSRAI C+ V KQ+T DHEP
Sbjct: 61 YKSTDDYILQNVVGPRGGSTAVTAIVIDGKKIVVANVGDSRAILCRESDVVKQITVDHEP 120
Query: 141 EKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDEDTEF 200
+KERDLV+S+GGFV +PG+VPRVDG LAMTRAFGDG LK+HI+ P++ I +I +DT+F
Sbjct: 121 DKERDLVKSKGGFVSQKPGNVPRVDGQLAMTRAFGDGGLKEHISVIPNIEIAEIHDDTKF 180
Query: 201 IILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
+ILASDGLWKVM+N E D IK +A+ AAK L+ +A ++GS DDISC+VV F
Sbjct: 181 LILASDGLWKVMSNDEVWDQIKKRGNAEEAAKMLIDKALARGSKDDISCVVVSF 234
>At4g28400 protein phosphatase 2C like protein
Length = 283
Score = 298 bits (763), Expect = 2e-81
Identities = 148/251 (58%), Positives = 194/251 (76%), Gaps = 1/251 (0%)
Query: 5 RHFSHGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENI 64
++ +HG + ++G +H MEDYV ++ KKL G+ELGL+AIFDGH G DVAKYLQ++LF+NI
Sbjct: 32 KNITHGFHCVKGKSSHPMEDYVVSEFKKLEGHELGLFAIFDGHLGHDVAKYLQTNLFDNI 91
Query: 65 LNEPDFWKNPVHAVKKACKATDEEILE-NIADSRGGSTAVAAILINGVKLLVFNVGDSRA 123
L E DFW + +A++ A ++TD IL+ ++ +GGSTAV ILI+G KL+V NVGDSRA
Sbjct: 92 LKEKDFWTDTENAIRNAYRSTDAVILQQSLKLGKGGSTAVTGILIDGKKLVVANVGDSRA 151
Query: 124 ISCKNGVAKQLTEDHEPEKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHI 183
+ KNGVA QL+ DHEP KE+ +ESRGGFV N PG VPRVDG LA+ RAFGD +LK H+
Sbjct: 152 VMSKNGVAHQLSVDHEPSKEKKEIESRGGFVSNIPGDVPRVDGQLAVARAFGDKSLKLHL 211
Query: 184 TAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGS 243
++EPD+ + ID+ TEFI+ ASDG+WKV++NQEA D IK I+D AAK L++EA S+ S
Sbjct: 212 SSEPDITHQTIDDHTEFILFASDGIWKVLSNQEAVDAIKSIKDPHAAAKHLIEEAISRKS 271
Query: 244 YDDISCIVVKF 254
DDISCIVVKF
Sbjct: 272 KDDISCIVVKF 282
>At3g15260 putative protein phosphatase type 2C
Length = 289
Score = 296 bits (758), Expect = 7e-81
Identities = 151/251 (60%), Positives = 188/251 (74%), Gaps = 1/251 (0%)
Query: 5 RHFSHGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENI 64
+ +HG +L++G H MEDYV AK K+++ ELGL+AIFDGH ++ YL SHLFENI
Sbjct: 38 KQITHGFHLVKGKAFHEMEDYVVAKFKEVDDNELGLFAIFDGHLSHEIPDYLCSHLFENI 97
Query: 65 LNEPDFWKNPVHAVKKACKATDEEILENIAD-SRGGSTAVAAILINGVKLLVFNVGDSRA 123
L EP+FW+ P A+KKA TD IL+ D +GGSTAV AILIN KL+V NVGDSRA
Sbjct: 98 LKEPNFWQEPEKAIKKAYYITDTTILDKADDLGKGGSTAVTAILINCQKLVVANVGDSRA 157
Query: 124 ISCKNGVAKQLTEDHEPEKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHI 183
+ C+NGVAK L+ DHEP E+D +E+RGGFV N PG VPRVDG LA+ RAFGD +LK H+
Sbjct: 158 VICQNGVAKPLSVDHEPNMEKDEIENRGGFVSNFPGDVPRVDGQLAVARAFGDKSLKMHL 217
Query: 184 TAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGS 243
++EP V + ID+D EF+ILASDGLWKVM+NQEA D IK I+DAK AAK L +EA ++ S
Sbjct: 218 SSEPYVTVEIIDDDAEFLILASDGLWKVMSNQEAVDSIKGIKDAKAAAKHLAEEAVARKS 277
Query: 244 YDDISCIVVKF 254
DDIS +VVKF
Sbjct: 278 SDDISVVVVKF 288
>At2g20630 putative protein phosphatase 2C
Length = 279
Score = 291 bits (746), Expect = 2e-79
Identities = 144/251 (57%), Positives = 192/251 (76%), Gaps = 1/251 (0%)
Query: 5 RHFSHGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENI 64
++ +HG ++G H MEDYV ++ KK++G++LGL+AIFDGH G DVAKYLQ++LF+NI
Sbjct: 28 KNIAHGYDFVKGKAGHPMEDYVVSEFKKVDGHDLGLFAIFDGHLGHDVAKYLQTNLFDNI 87
Query: 65 LNEPDFWKNPVHAVKKACKATDEEILE-NIADSRGGSTAVAAILINGVKLLVFNVGDSRA 123
L E DFW + +A++ A +TD ILE ++ +GGSTAV ILI+G L++ NVGDSRA
Sbjct: 88 LKEKDFWTDTKNAIRNAYISTDAVILEQSLKLGKGGSTAVTGILIDGKTLVIANVGDSRA 147
Query: 124 ISCKNGVAKQLTEDHEPEKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHI 183
+ KNGVA QL+ DHEP KE+ +ESRGGFV N PG VPRVDG LA+ RAFGD +LK H+
Sbjct: 148 VMSKNGVASQLSVDHEPSKEQKEIESRGGFVSNIPGDVPRVDGQLAVARAFGDKSLKIHL 207
Query: 184 TAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGS 243
+++PD+ ID +TEFI+ ASDG+WKVM+NQEA D IK I+D + AAK+L++EA S+ S
Sbjct: 208 SSDPDIRDENIDHETEFILFASDGVWKVMSNQEAVDLIKSIKDPQAAAKELIEEAVSKQS 267
Query: 244 YDDISCIVVKF 254
DDISCIVV+F
Sbjct: 268 TDDISCIVVRF 278
>At1g78200 putative protein phosphatase 2C
Length = 283
Score = 259 bits (662), Expect = 1e-69
Identities = 139/253 (54%), Positives = 178/253 (69%), Gaps = 10/253 (3%)
Query: 9 HGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEP 68
+G LI+G H MEDY AK NG ELGL+AIFDGH G VA YLQ HLF NIL +
Sbjct: 33 YGFSLIKGKSNHSMEDYHVAKFTNFNGNELGLFAIFDGHKGDHVAAYLQKHLFSNILKDG 92
Query: 69 DFWKNPVHAVKKACKATDEEILENIADSR-----GGSTAVAAILINGVKLLVFNVGDSRA 123
+F +P A+ KA + TD++IL AD+R GGSTAV AILING L + NVGDSRA
Sbjct: 93 EFLVDPRRAIAKAYENTDQKIL---ADNRTDLESGGSTAVTAILINGKALWIANVGDSRA 149
Query: 124 ISCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKD 181
I G AKQ++ DH+P+ ER ++ES+GGFV NRPG VPRV+G+LA++R FGD NLK
Sbjct: 150 IVSSRGKAKQMSVDHDPDDDTERSMIESKGGFVTNRPGDVPRVNGLLAVSRVFGDKNLKA 209
Query: 182 HITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQ 241
++ +EP++ ID T+F+ILASDG+ KVM+NQEA D K ++D K AA+++V EA +
Sbjct: 210 YLNSEPEIKDVTIDSHTDFLILASDGISKVMSNQEAVDVAKKLKDPKEAARQVVAEALKR 269
Query: 242 GSYDDISCIVVKF 254
S DDISCIVV+F
Sbjct: 270 NSKDDISCIVVRF 282
>At1g22280 protein phosphatase type 2C like protein
Length = 281
Score = 255 bits (651), Expect = 2e-68
Identities = 127/247 (51%), Positives = 168/247 (67%), Gaps = 1/247 (0%)
Query: 9 HGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEP 68
+G L++G H MEDY A + +ELGL+AI+DGH G V YLQ LF NIL E
Sbjct: 34 YGFSLVKGKANHPMEDYHVANFINIQDHELGLFAIYDGHMGDSVPAYLQKRLFSNILKEG 93
Query: 69 DFWKNPVHAVKKACKATDEEILENIAD-SRGGSTAVAAILINGVKLLVFNVGDSRAISCK 127
+FW +P ++ KA + TD+ IL N +D RGGSTAV AILING KL + NVGDSRA+
Sbjct: 94 EFWVDPRRSIAKAYEKTDQAILSNSSDLGRGGSTAVTAILINGRKLWIANVGDSRAVLSH 153
Query: 128 NGVAKQLTEDHEPEKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEP 187
G Q++ DHEP ER +E RGGFV N PG VPRV+G LA++RAFGD LK H+++EP
Sbjct: 154 GGAITQMSTDHEPRTERSSIEDRGGFVSNLPGDVPRVNGQLAVSRAFGDKGLKTHLSSEP 213
Query: 188 DVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGSYDDI 247
D+ +D T+ ++LASDG+WKVM N+EA + + ++D + AAK+L EA + S DDI
Sbjct: 214 DIKEATVDSQTDVLLLASDGIWKVMTNEEAMEIARRVKDPQKAAKELTAEALRRESKDDI 273
Query: 248 SCIVVKF 254
SC+VV+F
Sbjct: 274 SCVVVRF 280
>At1g34750 protein phosphatase type 2C, putative
Length = 282
Score = 252 bits (644), Expect = 1e-67
Identities = 133/245 (54%), Positives = 170/245 (69%), Gaps = 1/245 (0%)
Query: 10 GCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPD 69
G L++G H MEDY +K K++G ELGL+AI+DGH G V YLQ HLF NIL E
Sbjct: 36 GYSLVKGKANHPMEDYHVSKFVKIDGNELGLFAIYDGHLGERVPAYLQKHLFSNILKEEQ 95
Query: 70 FWKNPVHAVKKACKATDEEILENIAD-SRGGSTAVAAILINGVKLLVFNVGDSRAISCKN 128
F +P ++ A + TD+ IL + +D RGGSTAV AIL+NG +L V NVGDSRA+ +
Sbjct: 96 FRYDPQRSIIAAYEKTDQAILSHSSDLGRGGSTAVTAILMNGRRLWVANVGDSRAVLSQG 155
Query: 129 GVAKQLTEDHEPEKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPD 188
G A Q+T DHEP ER +E +GGFV N PG VPRV+G LA++RAFGD +LK H+ ++PD
Sbjct: 156 GQAIQMTIDHEPHTERLSIEGKGGFVSNMPGDVPRVNGQLAVSRAFGDKSLKTHLRSDPD 215
Query: 189 VMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGSYDDIS 248
V ID+ T+ ++LASDGLWKVMANQEA D + I+D AAK+L EA + S DDIS
Sbjct: 216 VKDSSIDDHTDVLVLASDGLWKVMANQEAIDIARRIKDPLKAAKELTTEALRRDSKDDIS 275
Query: 249 CIVVK 253
CIVV+
Sbjct: 276 CIVVR 280
>At5g24940 Protein phosphatase 2C-like protein
Length = 447
Score = 201 bits (511), Expect = 3e-52
Identities = 110/252 (43%), Positives = 161/252 (63%), Gaps = 8/252 (3%)
Query: 7 FSHGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILN 66
FS+G Y MED+ + ++G +GL+ +FDGH G A+Y++ HLF N++
Sbjct: 32 FSYG-YASSAGKRSSMEDFFETRIDGIDGEIVGLFGVFDGHGGSRAAEYVKRHLFSNLIT 90
Query: 67 EPDFWKNPVHAVKKACKATDEEIL--ENIADSRGGSTAVAAILINGVKLLVFNVGDSRAI 124
P F + A+ A TD E+L EN GSTA AIL+ G +LLV NVGDSRA+
Sbjct: 91 HPKFISDTKSAIADAYTHTDSELLKSENSHTRDAGSTASTAILV-GDRLLVANVGDSRAV 149
Query: 125 SCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDH 182
C+ G A ++ DH+P++ ER+ +E+ GGFV+ RV G+LA++RAFGD LK +
Sbjct: 150 ICRGGNAFAVSRDHKPDQSDERERIENAGGFVMW--AGTWRVGGVLAVSRAFGDRLLKQY 207
Query: 183 ITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQG 242
+ A+P++ KID+ EF+ILASDGLW V +N+EA +K++ED + + KKLV EA +G
Sbjct: 208 VVADPEIQEEKIDDSLEFLILASDGLWDVFSNEEAVAVVKEVEDPEESTKKLVGEAIKRG 267
Query: 243 SYDDISCIVVKF 254
S D+I+C+VV+F
Sbjct: 268 SADNITCVVVRF 279
>At5g10740 protein phosphatase 2C -like protein
Length = 354
Score = 199 bits (505), Expect = 2e-51
Identities = 109/252 (43%), Positives = 162/252 (64%), Gaps = 8/252 (3%)
Query: 7 FSHGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILN 66
FS+G Y MED+ + +NG +GL+ +FDGH G A+Y++ HLF N++
Sbjct: 32 FSYG-YASSAGKRSSMEDFFETRIDGINGEIVGLFGVFDGHGGARAAEYVKRHLFSNLIT 90
Query: 67 EPDFWKNPVHAVKKACKATDEEIL--ENIADSRGGSTAVAAILINGVKLLVFNVGDSRAI 124
P F + A+ A TD E+L EN + GSTA AIL+ G +L+V NVGDSRA+
Sbjct: 91 HPKFISDTKSAITDAYNHTDSELLKSENSHNRDAGSTASTAILV-GDRLVVANVGDSRAV 149
Query: 125 SCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDH 182
+ G A ++ DH+P++ ER+ +E+ GGFV+ RV G+LA++RAFGD LK +
Sbjct: 150 ISRGGKAIAVSRDHKPDQSDERERIENAGGFVMW--AGTWRVGGVLAVSRAFGDRLLKQY 207
Query: 183 ITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQG 242
+ A+P++ KID+ EF+ILASDGLW V +N+ A +K++ED +++AKKLV EA +G
Sbjct: 208 VVADPEIQEEKIDDTLEFLILASDGLWDVFSNEAAVAMVKEVEDPEDSAKKLVGEAIKRG 267
Query: 243 SYDDISCIVVKF 254
S D+I+C+VV+F
Sbjct: 268 SADNITCVVVRF 279
>At4g31750 unknown protein
Length = 311
Score = 196 bits (498), Expect = 1e-50
Identities = 109/252 (43%), Positives = 158/252 (62%), Gaps = 8/252 (3%)
Query: 7 FSHGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILN 66
FS+G G + MED+ + + G +GL+ +FDGH G A+Y++ +LF N++
Sbjct: 32 FSYGYASSPGKRS-SMEDFYETRIDGVEGEIVGLFGVFDGHGGARAAEYVKQNLFSNLIR 90
Query: 67 EPDFWKNPVHAVKKACKATDEEIL--ENIADSRGGSTAVAAILINGVKLLVFNVGDSRAI 124
P F + A+ A TD E L EN + GSTA AIL+ G +LLV NVGDSRA+
Sbjct: 91 HPKFISDTTAAIADAYNQTDSEFLKSENSQNRDAGSTASTAILV-GDRLLVANVGDSRAV 149
Query: 125 SCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDH 182
C+ G A ++ DH+P++ ER +E GGFV+ RV G+LA++RAFGD LK +
Sbjct: 150 ICRGGNAIAVSRDHKPDQSDERQRIEDAGGFVMW--AGTWRVGGVLAVSRAFGDRLLKQY 207
Query: 183 ITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQG 242
+ A+P++ K+D EF+ILASDGLW V++N+EA IK IED + AK+L+ EA +G
Sbjct: 208 VVADPEIQEEKVDSSLEFLILASDGLWDVVSNEEAVGMIKAIEDPEEGAKRLMMEAYQRG 267
Query: 243 SYDDISCIVVKF 254
S D+I+C+VV+F
Sbjct: 268 SADNITCVVVRF 279
>At1g43900 protein phosphatase type 2C like protein
Length = 371
Score = 196 bits (498), Expect = 1e-50
Identities = 110/252 (43%), Positives = 163/252 (64%), Gaps = 9/252 (3%)
Query: 7 FSHGCYLIQGMMAHGMEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILN 66
FS+G ++G A MEDY + +NG + + +FDGH G A+YL+++LF+N+++
Sbjct: 122 FSYGYSSLKGKRAT-MEDYFETRISDVNGQMVAFFGVFDGHGGARTAEYLKNNLFKNLVS 180
Query: 67 EPDFWKNPVHAVKKACKATDEEILENIADS--RGGSTAVAAILINGVKLLVFNVGDSRAI 124
DF + A+ + K TDEE L A GSTA A LI G KL+V NVGDSR +
Sbjct: 181 HDDFISDTKKAIVEVFKQTDEEYLIEEAGQPKNAGSTAATAFLI-GDKLIVANVGDSRVV 239
Query: 125 SCKNGVAKQLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDH 182
+ +NG A L++DH+P++ ER +E GGF++ RV GILA++RAFGD LK +
Sbjct: 240 ASRNGSAVPLSDDHKPDRSDERQRIEDAGGFIIW--AGTWRVGGILAVSRAFGDKQLKPY 297
Query: 183 ITAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQG 242
+ AEP++ I EFI++ASDGLW V++N++A ++DI DA+ AA+KLV+E ++G
Sbjct: 298 VIAEPEIQEEDIST-LEFIVVASDGLWNVLSNKDAVAIVRDISDAETAARKLVQEGYARG 356
Query: 243 SYDDISCIVVKF 254
S D+I+CIVV+F
Sbjct: 357 SCDNITCIVVRF 368
>At5g53140 protein phosphatase 2C-like
Length = 420
Score = 177 bits (449), Expect = 5e-45
Identities = 102/237 (43%), Positives = 145/237 (61%), Gaps = 7/237 (2%)
Query: 22 MEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNPVHAVKKA 81
MED+ K + G + ++ IFDGH G A+YL+ HLF N++ P F + A+ +
Sbjct: 114 MEDFYDIKASTIEGQAVCMFGIFDGHGGSRAAEYLKEHLFNNLMKHPQFLTDTKLALNET 173
Query: 82 CKATDEEILENIADSR--GGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHE 139
K TD LE+ D+ GSTA AA+L+ G L V NVGDSR I K G A L++DH+
Sbjct: 174 YKQTDVAFLESEKDTYRDDGSTASAAVLV-GNHLYVANVGDSRTIVSKAGKAIALSDDHK 232
Query: 140 PEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDED 197
P + ER +ES GG ++ RV G+LAM+RAFG+ LK + AEP++ +ID +
Sbjct: 233 PNRSDERKRIESAGGVIMW--AGTWRVGGVLAMSRAFGNRMLKQFVVAEPEIQDLEIDHE 290
Query: 198 TEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
E ++LASDGLW V+ N++A + E+ + AA+KL A S+GS D+I+CIVVKF
Sbjct: 291 AELLVLASDGLWDVVPNEDAVALAQSEEEPEAAARKLTDTAFSRGSADNITCIVVKF 347
>At1g07160 unknown protein
Length = 380
Score = 143 bits (360), Expect = 1e-34
Identities = 95/244 (38%), Positives = 138/244 (55%), Gaps = 15/244 (6%)
Query: 22 MEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNPV---HAV 78
MED A + ++ ++DGH G A++ +L NIL E +N AV
Sbjct: 135 MEDRFSAITNLQGDPKQAIFGVYDGHGGPTAAEFAAKNLCSNILGEIVGGRNESKIEEAV 194
Query: 79 KKACKATDEEILENIADSRGGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDH 138
K+ ATD E L+ + +GGS V A++ +G L+V N GD RA+ G A+ LT DH
Sbjct: 195 KRGYLATDSEFLKE-KNVKGGSCCVTALISDG-NLVVANAGDCRAVLSVGGFAEALTSDH 252
Query: 139 EPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDE 196
P + ER+ +ES GG+V + SV R+ G LA++R GD +LK I +EP++ I +I+
Sbjct: 253 RPSRDDERNRIESSGGYV-DTFNSVWRIQGSLAVSRGIGDAHLKQWIISEPEINILRINP 311
Query: 197 DTEFIILASDGLWKVMANQEACDCIKDI---EDAKN----AAKKLVKEAKSQGSYDDISC 249
EF+ILASDGLW ++NQEA D + D K A KKLV + S+GS DDIS
Sbjct: 312 QHEFLILASDGLWDKVSNQEAVDIARPFCKGTDQKRKPLLACKKLVDLSVSRGSLDDISV 371
Query: 250 IVVK 253
++++
Sbjct: 372 MLIQ 375
>At2g30020 putative protein phosphatase 2C
Length = 396
Score = 138 bits (348), Expect = 2e-33
Identities = 93/242 (38%), Positives = 132/242 (54%), Gaps = 14/242 (5%)
Query: 22 MEDYVFAKHKKLNGYELGLYAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNP---VHAV 78
MED A + ++ ++DGH G A++ +L +NI+ E ++ AV
Sbjct: 152 MEDRFSAITNLHGDRKQAIFGVYDGHGGVKAAEFAAKNLDKNIVEEVVGKRDESEIAEAV 211
Query: 79 KKACKATDEEILENIADSRGGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDH 138
K ATD L+ D +GGS V A L+N L+V N GD RA+ GVAK L+ DH
Sbjct: 212 KHGYLATDASFLKE-EDVKGGSCCVTA-LVNEGNLVVSNAGDCRAVMSVGGVAKALSSDH 269
Query: 139 EPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDE 196
P + ER +E+ GG+V G V R+ G LA++R GD LK + AEP+ I +I+
Sbjct: 270 RPSRDDERKRIETTGGYVDTFHG-VWRIQGSLAVSRGIGDAQLKKWVIAEPETKISRIEH 328
Query: 197 DTEFIILASDGLWKVMANQEACDCIKDIEDAKN------AAKKLVKEAKSQGSYDDISCI 250
D EF+ILASDGLW ++NQEA D + + A KKLV + S+GS DDIS +
Sbjct: 329 DHEFLILASDGLWDKVSNQEAVDIARPLCLGTEKPLLLAACKKLVDLSASRGSSDDISVM 388
Query: 251 VV 252
++
Sbjct: 389 LI 390
>At2g40180 protein phosphatase 2C (AthPP2C5)
Length = 390
Score = 134 bits (336), Expect = 6e-32
Identities = 88/248 (35%), Positives = 137/248 (54%), Gaps = 20/248 (8%)
Query: 22 MEDYVFAKHKKLN--GYELGLYAIFDGHAGRDVAKYLQSHLFENI------LNEPDFWKN 73
MED FA + + GY+ + +FDGH G A++ +L NI + +
Sbjct: 141 MEDRYFAAVDRNDDGGYKNAFFGVFDGHGGSKAAEFAAMNLGNNIEAAMASARSGEDGCS 200
Query: 74 PVHAVKKACKATDEEILENIADSRGGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQ 133
A+++ TDE+ L+ SRGG+ V A++ G +L V N GD RA+ + G A+
Sbjct: 201 MESAIREGYIKTDEDFLKE--GSRGGACCVTALISKG-ELAVSNAGDCRAVMSRGGTAEA 257
Query: 134 LTEDHEPEKERDL--VESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMI 191
LT DH P + +L +E+ GG+V + V R+ G LA++R GD LK+ + AEP+
Sbjct: 258 LTSDHNPSQANELKRIEALGGYV-DCCNGVWRIQGTLAVSRGIGDRYLKEWVIAEPETRT 316
Query: 192 RKIDEDTEFIILASDGLWKVMANQEACDCIK----DIED--AKNAAKKLVKEAKSQGSYD 245
+I + EF+ILASDGLW + NQEA D ++ +E+ +A KKL + + +GS D
Sbjct: 317 LRIKPEFEFLILASDGLWDKVTNQEAVDVVRPYCVGVENPMTLSACKKLAELSVKRGSLD 376
Query: 246 DISCIVVK 253
DIS I+++
Sbjct: 377 DISLIIIQ 384
>At1g67820 protein phosphatase-like protein
Length = 447
Score = 126 bits (316), Expect = 1e-29
Identities = 75/224 (33%), Positives = 127/224 (56%), Gaps = 24/224 (10%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNP---VHAVKKACKATDEEILENIADSR 97
+ ++DGH G A+++ +L + ++ + K V A K A TD + LE + +
Sbjct: 135 FGVYDGHGGAKAAEFVAENLHKYVVEMMENCKGKEEKVEAFKAAFLRTDRDFLEKVIKEQ 194
Query: 98 G------GSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEP--EKERDLVES 149
G+ V A+ I +++V N+GD RA+ C+ GVA+ LT+DH+P + E++ +ES
Sbjct: 195 SLKGVVSGACCVTAV-IQDQEMIVSNLGDCRAVLCRAGVAEALTDDHKPGRDDEKERIES 253
Query: 150 R-----------GGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDEDT 198
+ GG+V N G+ RV GILA++R+ GD +LK + AEP+ + ++++D
Sbjct: 254 QSLIPFMTFGLQGGYVDNHQGAW-RVQGILAVSRSIGDAHLKKWVVAEPETRVLELEQDM 312
Query: 199 EFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQG 242
EF++LASDGLW V++NQEA + + + K+ +E QG
Sbjct: 313 EFLVLASDGLWDVVSNQEAVYTVLHVLAQRKTPKESEEENLVQG 356
>At3g23360 hypothetical protein
Length = 256
Score = 117 bits (294), Expect = 5e-27
Identities = 77/251 (30%), Positives = 128/251 (50%), Gaps = 37/251 (14%)
Query: 8 SHGCYLIQGMMA---HGMEDYVFAKHKKLNG-YELGLYAIFDGHAGRDVAKYLQSHLFEN 63
SHG Y + + +D VF + ++ + E+ L+ + + G+++ KY+Q+HLF+
Sbjct: 38 SHGYYTVDRLSYADNSSNDDSVFVQREQQSDELEIWLFGVSNAGTGKEIVKYMQNHLFDK 97
Query: 64 ILNEPDFWKNPVHAVKKACKATDEEILENIADSRGGSTAVAAILINGVKLLVFNVGDSRA 123
+ NE + + CK T + + R G +A + +++NG KL + ++GD R
Sbjct: 98 LPNEL--------GIMRKCKETMRRAY--VEEERTGGSAASVMVVNGEKLAIASIGDHRV 147
Query: 124 ISCKNGVAKQLTEDHEPEKERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHI 183
+ CK+G A Q+ + K F+ PG + + D
Sbjct: 148 VVCKDGEAHQIRDRKASTKHWSQ------FIF--PGE---------------EEDESDPR 184
Query: 184 TAEPDVMIRKIDEDTEFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQGS 243
+E V+ KI+ DTEFII+ S G+W+VM +QEA + I+ IED K AAK L KEA ++ S
Sbjct: 185 NSELVVITEKINSDTEFIIIGSPGIWEVMKSQEAINLIRHIEDPKEAAKCLAKEALNRIS 244
Query: 244 YDDISCIVVKF 254
ISC+V++F
Sbjct: 245 KSSISCVVIRF 255
>At1g07430 protein phosphatase 2C, putative
Length = 442
Score = 114 bits (286), Expect = 4e-26
Identities = 74/224 (33%), Positives = 122/224 (54%), Gaps = 28/224 (12%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEP-----DFWKNPVHAVKKACKATDEEIL---EN 92
+ ++DGH VA + L E + E + WK ++++ D+E++ E
Sbjct: 158 FGVYDGHGCSHVAARCKERLHELVQEEALSDKKEEWKK---MMERSFTRMDKEVVRWGET 214
Query: 93 IADSRG------------GSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEP 140
+ + GSTAV ++ I K++V N GDSRA+ C+NG A L+ DH+P
Sbjct: 215 VMSANCRCELQTPDCDAVGSTAVVSV-ITPEKIIVANCGDSRAVLCRNGKAVPLSTDHKP 273
Query: 141 EK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVMIRKIDEDT 198
++ E D ++ GG V+ G+ RV G+LAM+RA GD LK ++T+EP+V + E+
Sbjct: 274 DRPDELDRIQEAGGRVIYWDGA--RVLGVLAMSRAIGDNYLKPYVTSEPEVTVTDRTEED 331
Query: 199 EFIILASDGLWKVMANQEACDCIKDIEDAKNAAKKLVKEAKSQG 242
EF+ILA+DGLW V+ N+ AC ++ + K+ + E ++ G
Sbjct: 332 EFLILATDGLWDVVTNEAACTMVRMCLNRKSGRGRRRGETQTPG 375
>At3g62260 unknown protein
Length = 383
Score = 114 bits (284), Expect = 7e-26
Identities = 73/236 (30%), Positives = 119/236 (49%), Gaps = 26/236 (11%)
Query: 41 YAIFDGHAGRDVAKYLQSHLFENILNEPDFWKNPV----------HAVKKACKATDEEIL 90
YA+FDGH G + A Y++ + + F + +++ A D +
Sbjct: 117 YAVFDGHGGPEAAAYVRENAIRFFFEDEQFPQTSEVSSVYVEEVETSLRNAFLQADLALA 176
Query: 91 ENIADSRGGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAKQLTEDHEPEK--ERDLVE 148
E+ + S T LI G L+V N GD RA+ C+ G A ++EDH+P ER VE
Sbjct: 177 EDCSISDSCGTTALTALICGRLLMVANAGDCRAVLCRKGRAIDMSEDHKPINLLERRRVE 236
Query: 149 SRGGFVLNRPGSVPRVDGILAMTRAFGDGNLK------DHITAEPDVMIRKIDEDTEFII 202
GGF+ N ++ +LA+TRA GD +LK + +EP++ + ED EF++
Sbjct: 237 ESGGFITND----GYLNEVLAVTRALGDWDLKLPHGSQSPLISEPEIKQITLTEDDEFLV 292
Query: 203 LASDGLWKVMANQEACDCIK----DIEDAKNAAKKLVKEAKSQGSYDDISCIVVKF 254
+ DG+W V+ +QEA ++ D A++LV EA + S+D+++ +VV F
Sbjct: 293 IGCDGIWDVLTSQEAVSIVRRGLNRHNDPTRCARELVMEALGRNSFDNLTAVVVCF 348
>At5g51760 protein phosphatase-2C; PP2C-like protein
Length = 416
Score = 112 bits (281), Expect = 1e-25
Identities = 78/236 (33%), Positives = 125/236 (52%), Gaps = 37/236 (15%)
Query: 41 YAIFDGHAGRDVAK--------YLQSHLFENILNEPDFWKNPV------HAVKKACKATD 86
+A++DGH G V+ +++ L +N+ E + +N V +K++ K D
Sbjct: 145 FAVYDGHGGSQVSTLCSTTMHTFVKEELEQNLEEEEEGSENDVVERKWRGVMKRSFKRMD 204
Query: 87 EEILENIA----------DSR----GGSTAVAAILINGVKLLVFNVGDSRAISCKNGVAK 132
E D R GSTAV A+L + ++V N GDSRA+ C+NG+A
Sbjct: 205 EMATSTCVCGTSVPLCNCDPREAAISGSTAVTAVLTHD-HIIVANTGDSRAVLCRNGMAI 263
Query: 133 QLTEDHEPEK--ERDLVESRGGFVLNRPGSVPRVDGILAMTRAFGDGNLKDHITAEPDVM 190
L+ DH+P++ ER +E+ GG VL G+ RV+GILA +RA GD LK + EP+V
Sbjct: 264 PLSNDHKPDRPDERARIEAAGGRVLVVDGA--RVEGILATSRAIGDRYLKPMVAWEPEVT 321
Query: 191 IRKIDEDTEFIILASDGLWKVMANQEACD----CIKDIEDAKNAAKKLVKEAKSQG 242
+ + E ++LASDGLW V+++Q ACD C+++ + ++ +E + G
Sbjct: 322 FMRRESGDECLVLASDGLWDVLSSQLACDIARFCLREETPSSLDLNRMAQEDDNDG 377
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,793,984
Number of Sequences: 26719
Number of extensions: 238965
Number of successful extensions: 873
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 627
Number of HSP's gapped (non-prelim): 106
length of query: 254
length of database: 11,318,596
effective HSP length: 97
effective length of query: 157
effective length of database: 8,726,853
effective search space: 1370115921
effective search space used: 1370115921
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0007.1


